

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.10
(265 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
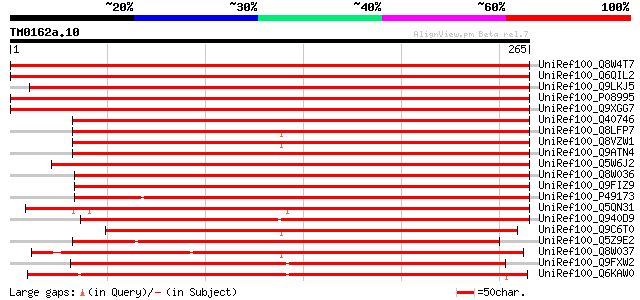
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8W4T7 Multifunctional aquaporin [Medicago truncatula] 392 e-108
UniRef100_Q6QIL2 NIP2 [Medicago truncatula] 375 e-103
UniRef100_Q9LKJ5 Multifunctional transport intrinsic membrane pr... 374 e-102
UniRef100_P08995 Nodulin-26 [Glycine max] 370 e-101
UniRef100_Q9XGG7 Nodulin26-like major intrinsic protein [Pisum s... 368 e-100
UniRef100_Q40746 Major intrinsic protein [Oryza sativa] 319 5e-86
UniRef100_Q8LFP7 Aquaporin NIP1.2 [Arabidopsis thaliana] 319 5e-86
UniRef100_Q8VZW1 Aquaporin NIP1.1 [Arabidopsis thaliana] 314 1e-84
UniRef100_Q9ATN4 NOD26-like membrane integral protein ZmNIP1-1 [... 312 5e-84
UniRef100_Q5W6J2 Hypothetical protein OSJNBb0115F21.2 [Oryza sat... 298 1e-79
UniRef100_Q8W036 Probable aquaporin NIP4.2 [Arabidopsis thaliana] 273 4e-72
UniRef100_Q9FIZ9 Putative aquaporin NIP4.1 [Arabidopsis thaliana] 271 1e-71
UniRef100_P49173 Probable aquaporin NIP-type [Nicotiana alata] 269 5e-71
UniRef100_Q5QN31 Putative membrane integral protein ZmNIP1-1 [Or... 264 2e-69
UniRef100_Q940D9 Early embryogenesis aquaglyceroporin [Pinus taeda] 254 2e-66
UniRef100_Q9C6T0 Putative aquaporin NIP3.1 [Arabidopsis thaliana] 247 3e-64
UniRef100_Q5Z9E2 Putative major intrinsic protein [Oryza sativa] 238 2e-61
UniRef100_Q8W037 Aquaporin NIP2.1 [Arabidopsis thaliana] 231 1e-59
UniRef100_Q9FXW2 MIP [Adiantum capillus-veneris] 221 1e-56
UniRef100_Q6KAW0 Nod26-like major intrinsic protein [Cicer ariet... 221 1e-56
>UniRef100_Q8W4T7 Multifunctional aquaporin [Medicago truncatula]
Length = 276
Score = 392 bits (1008), Expect = e-108
Identities = 192/266 (72%), Positives = 228/266 (85%), Gaps = 1/266 (0%)
Query: 1 MAN-NSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIV 59
MAN NSA ETH+VVL NKD+S T + S ++ SV F+QKL+AE VGT+FLIF GCASIV
Sbjct: 1 MANDNSARIETHEVVLDTNKDSSDTCKGSGSFVSVPFLQKLIAEMVGTYFLIFAGCASIV 60
Query: 60 VNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYI 119
VNK+NDNVVTLPGIA+VWGL L+VLIYS+GHISGAHFNPAVT AFATT+RFP +QV YI
Sbjct: 61 VNKDNDNVVTLPGIAIVWGLTLLVLIYSLGHISGAHFNPAVTIAFATTRRFPLLQVPAYI 120
Query: 120 ASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR 179
++QLLGA LASG LK++FSG HD FSGT+PSG+NLQAFV+EFITTF LMF IS VATD R
Sbjct: 121 SAQLLGATLASGTLKLIFSGAHDHFSGTLPSGSNLQAFVLEFITTFYLMFTISGVATDTR 180
Query: 180 AIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVA 239
AIGE+AGIAIGSTLLLN++I+GP+TGASMNP RTLGPA H++YR I +Y +S I GA+A
Sbjct: 181 AIGELAGIAIGSTLLLNVMIAGPVTGASMNPVRTLGPAFVHNEYRGIWIYLLSPILGAIA 240
Query: 240 GAWVFNILRYTDKPLHEITKGSSILK 265
GAWV+N +RYT+KPL EIT+ +S LK
Sbjct: 241 GAWVYNTVRYTNKPLREITQSASFLK 266
>UniRef100_Q6QIL2 NIP2 [Medicago truncatula]
Length = 269
Score = 375 bits (962), Expect = e-103
Identities = 177/265 (66%), Positives = 225/265 (84%)
Query: 1 MANNSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVV 60
MA NSAS T+++VL+VNKD S E S ++ + F+QK VAE +GT+FLIF GCAS++V
Sbjct: 1 MAENSASNATNEIVLNVNKDVSNKSEDSTSHATASFLQKSVAEVIGTYFLIFAGCASVLV 60
Query: 61 NKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIA 120
NKNN+NVVTLPGI++VWGL +MVL+YS+GHISGAHFNPAVT AFA+TKRFP QV Y+A
Sbjct: 61 NKNNENVVTLPGISIVWGLAVMVLVYSLGHISGAHFNPAVTIAFASTKRFPLKQVPAYVA 120
Query: 121 SQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA 180
+Q+ G+ LASG L+++F+G H+QF GT+P+G++LQAFVIEFI TF MF+IS VATDNRA
Sbjct: 121 AQVFGSTLASGTLRLIFTGKHNQFVGTLPAGSDLQAFVIEFIITFYPMFIISGVATDNRA 180
Query: 181 IGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAG 240
IGE+AGIA+GST+LLN++ +GPITGASMNPAR++GPA+ HS+YR I +Y VS I GAVAG
Sbjct: 181 IGELAGIAVGSTVLLNVMFAGPITGASMNPARSIGPALLHSEYRGIWIYLVSPILGAVAG 240
Query: 241 AWVFNILRYTDKPLHEITKGSSILK 265
AWV+N++RYTDKP+ EITK SS LK
Sbjct: 241 AWVYNVIRYTDKPVREITKSSSFLK 265
>UniRef100_Q9LKJ5 Multifunctional transport intrinsic membrane protein 2 [Lotus
japonicus]
Length = 270
Score = 374 bits (959), Expect = e-102
Identities = 177/255 (69%), Positives = 219/255 (85%)
Query: 11 HDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTL 70
H+VV++VNKDASKTIE SDT +V F+QK++AE VGT+F IF GCASIVVNKNNDNVVTL
Sbjct: 14 HEVVVNVNKDASKTIEVSDTNFTVSFLQKVIAELVGTYFFIFAGCASIVVNKNNDNVVTL 73
Query: 71 PGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLAS 130
PGIALVWGL +MVL+YS+GHISGAHFNPA T AFA+TKRFPW QV Y+++Q+LG+ LAS
Sbjct: 74 PGIALVWGLAVMVLVYSLGHISGAHFNPAATIAFASTKRFPWKQVPAYVSAQVLGSTLAS 133
Query: 131 GILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIG 190
G L+++FSG H+QF+G +P+G+NLQAFVIEFI TF L+F++ VATD+RAIGE+AGI +G
Sbjct: 134 GTLRLIFSGKHNQFAGALPTGSNLQAFVIEFIITFFLIFILFGVATDDRAIGEVAGIVVG 193
Query: 191 STLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYT 250
ST+LLN+L +GPITGASMNPAR++G A H++YR I +Y +S GAVAGAWV+NI+RYT
Sbjct: 194 STVLLNVLFAGPITGASMNPARSIGSAFVHNEYRGIWIYLLSPTLGAVAGAWVYNIVRYT 253
Query: 251 DKPLHEITKGSSILK 265
DKPL EITK S LK
Sbjct: 254 DKPLREITKNVSFLK 268
>UniRef100_P08995 Nodulin-26 [Glycine max]
Length = 271
Score = 370 bits (951), Expect = e-101
Identities = 178/265 (67%), Positives = 220/265 (82%)
Query: 1 MANNSASFETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVV 60
MA+ SA E+ +VV++V K+ S+TI+ SD+ SV F+QKLVAE VGT+FLIF GCAS+VV
Sbjct: 1 MADYSAGTESQEVVVNVTKNTSETIQRSDSLVSVPFLQKLVAEAVGTYFLIFAGCASLVV 60
Query: 61 NKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIA 120
N+N N++T PGIA+VWGLVL VL+Y+VGHISG HFNPAVT AFA+T+RFP IQV Y+
Sbjct: 61 NENYYNMITFPGIAIVWGLVLTVLVYTVGHISGGHFNPAVTIAFASTRRFPLIQVPAYVV 120
Query: 121 SQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRA 180
+QLLG++LASG L++LF G HDQFSGT+P+GTNLQAFV EFI TF LMFVI VATDNRA
Sbjct: 121 AQLLGSILASGTLRLLFMGNHDQFSGTVPNGTNLQAFVFEFIMTFFLMFVICGVATDNRA 180
Query: 181 IGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAG 240
+GE AGIAIGSTLLLN++I GP+TGASMNPAR+LGPA H +Y I +Y ++ + GA+AG
Sbjct: 181 VGEFAGIAIGSTLLLNVIIGGPVTGASMNPARSLGPAFVHGEYEGIWIYLLAPVVGAIAG 240
Query: 241 AWVFNILRYTDKPLHEITKGSSILK 265
AWV+NI+RYTDKPL E TK +S LK
Sbjct: 241 AWVYNIVRYTDKPLSETTKSASFLK 265
>UniRef100_Q9XGG7 Nodulin26-like major intrinsic protein [Pisum sativum]
Length = 270
Score = 368 bits (944), Expect = e-100
Identities = 176/266 (66%), Positives = 221/266 (82%), Gaps = 1/266 (0%)
Query: 1 MANNSASFETHDVVLSVNKDASK-TIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIV 59
M +NS ET+++V++VNKD S T E S + + +QKLVAE VGT+FLIF GCA++
Sbjct: 1 MGDNSGCNETNEIVVNVNKDVSNITQEDSTAHATASLLQKLVAEVVGTYFLIFAGCAAVA 60
Query: 60 VNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYI 119
VNKNNDNVVTLPGI++VWGL +MVL+YS+GHISGAHFNPAVT AFATT+RFP QV YI
Sbjct: 61 VNKNNDNVVTLPGISIVWGLAVMVLVYSLGHISGAHFNPAVTIAFATTRRFPLKQVPAYI 120
Query: 120 ASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNR 179
A+Q+ G+ LASG L++LFSG HDQF GT+ +G+NLQAFV+EFI TF LMF+IS VATDNR
Sbjct: 121 AAQVFGSTLASGTLRLLFSGKHDQFVGTLAAGSNLQAFVMEFIITFYLMFIISGVATDNR 180
Query: 180 AIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVA 239
AIGE+AGIA+GST+LLN++ +GPITGASMNPAR++GPA H++YR I +Y +S I GAV+
Sbjct: 181 AIGELAGIAVGSTVLLNVMFAGPITGASMNPARSIGPAFVHNEYRGIWIYMISPIVGAVS 240
Query: 240 GAWVFNILRYTDKPLHEITKGSSILK 265
GAWV+N++RYTDKP+ EITK S LK
Sbjct: 241 GAWVYNVIRYTDKPVREITKSGSFLK 266
>UniRef100_Q40746 Major intrinsic protein [Oryza sativa]
Length = 284
Score = 319 bits (817), Expect = 5e-86
Identities = 144/233 (61%), Positives = 193/233 (82%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV F+QK++AE GT+FLIF GC ++ +N++ + +T PG+A+VWGL +MV++Y+VGHIS
Sbjct: 43 SVPFIQKIIAEIFGTYFLIFAGCGAVTINQSKNGQITFPGVAIVWGLAVMVMVYAVGHIS 102
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGT 152
GAHFNPAVT AFAT +RFPW QV Y A+Q+LGA LA+G L+++F G H+ F GT+P+G+
Sbjct: 103 GAHFNPAVTLAFATCRRFPWRQVPAYAAAQMLGATLAAGTLRLMFGGRHEHFPGTLPAGS 162
Query: 153 NLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPAR 212
++Q+ V+EFI TF LMFVIS VATDNRAIGE+AG+A+G+T+LLN+LI+GPI+GASMNPAR
Sbjct: 163 DVQSLVLEFIITFYLMFVISGVATDNRAIGELAGLAVGATILLNVLIAGPISGASMNPAR 222
Query: 213 TLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
+LGPA+ +YR+I VY V + GAVAGAW +NI+R+T+KPL EITK S LK
Sbjct: 223 SLGPAMIGGEYRSIWVYIVGPVAGAVAGAWAYNIIRFTNKPLREITKSGSFLK 275
>UniRef100_Q8LFP7 Aquaporin NIP1.2 [Arabidopsis thaliana]
Length = 294
Score = 319 bits (817), Expect = 5e-86
Identities = 155/240 (64%), Positives = 195/240 (80%), Gaps = 7/240 (2%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV F+QKL+AE +GT+FLIF GCA++ VN +D VTLPGIA+VWGL +MVL+YS+GHIS
Sbjct: 47 SVPFLQKLMAEVLGTYFLIFAGCAAVAVNTQHDKAVTLPGIAIVWGLTVMVLVYSLGHIS 106
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF-------SGTHDQFS 145
GAHFNPAVT AFA+ RFP QV Y+ SQ++G+ LA+ L++LF SG HD F
Sbjct: 107 GAHFNPAVTIAFASCGRFPLKQVPAYVISQVIGSTLAAATLRLLFGLDQDVCSGKHDVFV 166
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITG 205
GT+PSG+NLQ+FVIEFI TF LMFVIS VATDNRAIGE+AG+A+GST+LLN++I+GP++G
Sbjct: 167 GTLPSGSNLQSFVIEFIITFYLMFVISGVATDNRAIGELAGLAVGSTVLLNVIIAGPVSG 226
Query: 206 ASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
ASMNP R+LGPA+ +S YR + +Y VS I GAV+GAWV+N++RYTDKPL EITK S LK
Sbjct: 227 ASMNPGRSLGPAMVYSCYRGLWIYIVSPIVGAVSGAWVYNMVRYTDKPLREITKSGSFLK 286
>UniRef100_Q8VZW1 Aquaporin NIP1.1 [Arabidopsis thaliana]
Length = 296
Score = 314 bits (805), Expect = 1e-84
Identities = 153/240 (63%), Positives = 191/240 (78%), Gaps = 7/240 (2%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV F+QKL+AEF+GT+FL+FTGCAS+VVN NDNVVTLPGIA+VWGL +MVLIYS+GHIS
Sbjct: 50 SVPFLQKLIAEFLGTYFLVFTGCASVVVNMQNDNVVTLPGIAIVWGLTIMVLIYSLGHIS 109
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLF-------SGTHDQFS 145
GAH NPAVT AFA+ RFP QV Y+ SQ++G+ LA+ L++LF SG HD F
Sbjct: 110 GAHINPAVTIAFASCGRFPLKQVPAYVISQVIGSTLAAATLRLLFGLDHDVCSGKHDVFI 169
Query: 146 GTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITG 205
G+ P G++LQAF +EFI TF LMF+IS VATDNRAIGE+AG+AIGST+LLN+LI+ P++
Sbjct: 170 GSSPVGSDLQAFTMEFIVTFYLMFIISGVATDNRAIGELAGLAIGSTVLLNVLIAAPVSS 229
Query: 206 ASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
ASMNP R+LGPA+ + Y+ I +Y V+ GA+AGAWV+N +RYTDKPL EITK S LK
Sbjct: 230 ASMNPGRSLGPALVYGCYKGIWIYLVAPTLGAIAGAWVYNTVRYTDKPLREITKSGSFLK 289
>UniRef100_Q9ATN4 NOD26-like membrane integral protein ZmNIP1-1 [Zea mays]
Length = 282
Score = 312 bits (800), Expect = 5e-84
Identities = 139/233 (59%), Positives = 191/233 (81%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
SV F+QK++AE GT+FL+F GC ++ +N + + +T PG+A+VWGL +MV++Y+VGHIS
Sbjct: 39 SVPFIQKIIAEIFGTYFLMFAGCGAVTINASKNGQITFPGVAIVWGLAVMVMVYAVGHIS 98
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGT 152
GAHFNPAVT AFAT+ RFPW Q+ Y+ +Q+LGA LASG L+++F G H+ F GT+P+G+
Sbjct: 99 GAHFNPAVTLAFATSGRFPWRQLPAYVLAQMLGATLASGTLRLMFGGRHEHFPGTLPTGS 158
Query: 153 NLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPAR 212
+Q+ VIE ITTF LMFVIS VATDNRAIGE+AG+A+G+T+LLN+LI+GP++GASMNPAR
Sbjct: 159 EVQSLVIEIITTFYLMFVISGVATDNRAIGELAGLAVGATILLNVLIAGPVSGASMNPAR 218
Query: 213 TLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
++GPA+ +Y +I VY V + GAVAGAW +N++R+T+KPL EITK +S LK
Sbjct: 219 SVGPALVSGEYTSIWVYVVGPVVGAVAGAWAYNLIRFTNKPLREITKSTSFLK 271
>UniRef100_Q5W6J2 Hypothetical protein OSJNBb0115F21.2 [Oryza sativa]
Length = 286
Score = 298 bits (763), Expect = 1e-79
Identities = 139/244 (56%), Positives = 183/244 (74%)
Query: 22 SKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVL 81
SK+ ++ SV F QK++AE +GTFFLIF GCA++ VNK VT PGI + WGL +
Sbjct: 38 SKSSSNNHPMFSVQFAQKVIAEILGTFFLIFAGCAAVAVNKRTGGTVTFPGICITWGLAV 97
Query: 82 MVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTH 141
MV++YSVGHISGAH NPAVT AFAT RFPW +V Y A+Q+ G+ AS L+ LF G
Sbjct: 98 MVMVYSVGHISGAHLNPAVTLAFATCGRFPWRRVPAYAAAQVAGSAAASAALRALFGGAP 157
Query: 142 DQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISG 201
+ F GT P+G+++Q+ +EFI TF LMFV+S VATDNRAIGE+AG+A+G+T+L+N+L +G
Sbjct: 158 EHFFGTAPAGSDVQSLAMEFIITFYLMFVVSGVATDNRAIGELAGLAVGATVLVNVLFAG 217
Query: 202 PITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGS 261
PI+GASMNPART+GPAI +Y I VY +FGAVAGAW +N++R+TDKPL EIT +
Sbjct: 218 PISGASMNPARTIGPAIILGRYTGIWVYIAGPVFGAVAGAWAYNLIRFTDKPLREITMTA 277
Query: 262 SILK 265
S ++
Sbjct: 278 SFIR 281
>UniRef100_Q8W036 Probable aquaporin NIP4.2 [Arabidopsis thaliana]
Length = 283
Score = 273 bits (697), Expect = 4e-72
Identities = 130/232 (56%), Positives = 172/232 (74%)
Query: 34 VFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISG 93
V QKL+AE +GT+F+IF+GC +VVN +T PGI + WGL++MV+IYS GHISG
Sbjct: 39 VCLTQKLIAEMIGTYFIIFSGCGVVVVNVLYGGTITFPGICVTWGLIVMVMIYSTGHISG 98
Query: 94 AHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTN 153
AHFNPAVT FA +RFPW QV YI +QL G++LAS L+++F+ T F GT P+ ++
Sbjct: 99 AHFNPAVTVTFAVFRRFPWYQVPLYIGAQLTGSLLASLTLRLMFNVTPKAFFGTTPTDSS 158
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
QA V E I +FLLMFVIS VATD+RA GE+AGIA+G T++LN+ ++GPI+GASMNPAR+
Sbjct: 159 GQALVAEIIISFLLMFVISGVATDSRATGELAGIAVGMTIILNVFVAGPISGASMNPARS 218
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
LGPAI +Y+ I VY V G AG +V+N +R+TDKPL E+TK +S L+
Sbjct: 219 LGPAIVMGRYKGIWVYIVGPFVGIFAGGFVYNFMRFTDKPLRELTKSASFLR 270
>UniRef100_Q9FIZ9 Putative aquaporin NIP4.1 [Arabidopsis thaliana]
Length = 283
Score = 271 bits (693), Expect = 1e-71
Identities = 125/232 (53%), Positives = 173/232 (73%)
Query: 34 VFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISG 93
V QKL+AE +GT+F++F+GC +VVN +T PGI + WGL++MV+IYS GHISG
Sbjct: 39 VCLTQKLIAEMIGTYFIVFSGCGVVVVNVLYGGTITFPGICVTWGLIVMVMIYSTGHISG 98
Query: 94 AHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTN 153
AHFNPAVT FA +RFPW QV YI +Q G++LAS L+++F T + F GT P+ +
Sbjct: 99 AHFNPAVTVTFAIFRRFPWHQVPLYIGAQFAGSLLASLTLRLMFKVTPEAFFGTTPADSP 158
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
+A V E I +FLLMFVIS VATDNRA+GE+AGIA+G T+++N+ ++GPI+GASMNPAR+
Sbjct: 159 ARALVAEIIISFLLMFVISGVATDNRAVGELAGIAVGMTIMVNVFVAGPISGASMNPARS 218
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
LGPA+ Y+ I VY V + G ++G +V+N++R+TDKPL E+TK +S L+
Sbjct: 219 LGPALVMGVYKHIWVYIVGPVLGVISGGFVYNLIRFTDKPLRELTKSASFLR 270
>UniRef100_P49173 Probable aquaporin NIP-type [Nicotiana alata]
Length = 270
Score = 269 bits (688), Expect = 5e-71
Identities = 130/232 (56%), Positives = 175/232 (75%), Gaps = 1/232 (0%)
Query: 34 VFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISG 93
V +QKL+AE +GT+F+IF GC S+ VNK +V T PGI + WGL++MV++Y+VG+ISG
Sbjct: 39 VVILQKLIAEAIGTYFVIFAGCGSVAVNKIYGSV-TFPGICVTWGLIVMVMVYTVGYISG 97
Query: 94 AHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTN 153
AHFNPAVT F+ RFPW QV YI +QL+G++LASG L +LF T + GT+P G+N
Sbjct: 98 AHFNPAVTITFSIFGRFPWKQVPLYIIAQLMGSILASGTLALLFDVTPQAYFGTVPVGSN 157
Query: 154 LQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPART 213
Q+ IE I +FLLMFVIS VATD+RAIG++AGIA+G T+ LN+ ++GPI+GASMNPAR+
Sbjct: 158 GQSLAIEIIISFLLMFVISGVATDDRAIGQVAGIAVGMTITLNVFVAGPISGASMNPARS 217
Query: 214 LGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
+GPAI Y + VY V I G +AGA+V+N++R TDKPL E+ K +S L+
Sbjct: 218 IGPAIVKHVYTGLWVYVVGPIIGTLAGAFVYNLIRSTDKPLRELAKSASSLR 269
>UniRef100_Q5QN31 Putative membrane integral protein ZmNIP1-1 [Oryza sativa]
Length = 303
Score = 264 bits (675), Expect = 2e-69
Identities = 133/280 (47%), Positives = 188/280 (66%), Gaps = 23/280 (8%)
Query: 9 ETHDVVLSVNKDASKTIESSDTY----TSVFFVQK-------------LVAEFVGTFFLI 51
++ +V ++D S + SS + SV F+QK ++AE +GT+F+I
Sbjct: 18 DSKEVKCESSEDGSSSSSSSRCHGNDVISVQFMQKVHPWCMCMNKNLLILAEILGTYFMI 77
Query: 52 FTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFP 111
F GC ++VVN++ VT PGI VWGLV+MVL+Y+V HISGAHFNPAVT AFAT RF
Sbjct: 78 FAGCGAVVVNQSTGGAVTFPGICAVWGLVVMVLVYTVSHISGAHFNPAVTVAFATCGRFR 137
Query: 112 WIQVAPYIASQLLGAVLASGILKMLFSGT------HDQFSGTIPSGTNLQAFVIEFITTF 165
W QV Y+ +Q+LG+ +AS L+++F G F GT P+G+ QA +EF+ +F
Sbjct: 138 WKQVPSYVVAQVLGSTMASLTLRVVFGGGGGGARGEHLFFGTTPAGSMAQAAALEFVISF 197
Query: 166 LLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRA 225
LMFV+S VATDNRAIGE+AG+A+G+T+ +N+L +GP+TGASMNPAR+LGPA+ +Y
Sbjct: 198 FLMFVVSGVATDNRAIGELAGLAVGATVAVNVLFAGPVTGASMNPARSLGPAMVAGRYGG 257
Query: 226 IVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
+ VY + + G V GAW +N+LR+TDKPL +I +S L+
Sbjct: 258 VWVYVAAPVSGTVCGAWAYNLLRFTDKPLRDIANTASFLR 297
>UniRef100_Q940D9 Early embryogenesis aquaglyceroporin [Pinus taeda]
Length = 264
Score = 254 bits (649), Expect = 2e-66
Identities = 121/229 (52%), Positives = 169/229 (72%), Gaps = 1/229 (0%)
Query: 37 VQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHF 96
V+K+VAEF+GTFFLIF GC S+VV+K ++ +T G++LVWG+ M++IYS+GHISGAH
Sbjct: 28 VRKVVAEFIGTFFLIFVGCGSVVVDKISNGSITHLGVSLVWGMAAMIVIYSIGHISGAHL 87
Query: 97 NPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGTNLQA 156
NPAVT A A KRFPW+QV YI +Q+ G++ A +L+ +F G T+PSG+ +Q+
Sbjct: 88 NPAVTLALAAVKRFPWVQVPGYIVAQVFGSISAGFLLRFMF-GEVAFMGATVPSGSEMQS 146
Query: 157 FVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGP 216
F +E ITT LL+FV+SAVATD +A+GE+ G+AIG+T+ +N+ ISGPI+GASMNPART+G
Sbjct: 147 FALEIITTSLLVFVVSAVATDTKAVGELGGLAIGATIAMNVAISGPISGASMNPARTIGS 206
Query: 217 AIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITKGSSILK 265
A+ +KY +I VY V + GA+ GA +N++R T EI K S +K
Sbjct: 207 AVAGNKYTSIWVYMVGPVIGALMGAMSYNMIRETKMSEREIMKSGSFVK 255
Score = 42.0 bits (97), Expect = 0.016
Identities = 28/101 (27%), Positives = 52/101 (50%), Gaps = 6/101 (5%)
Query: 153 NLQAFVIEFITTFLLMFV-ISAVATDNRAIGEMAGIAI----GSTLLLNILISGPITGAS 207
+++ V EFI TF L+FV +V D + G + + + G ++ I G I+GA
Sbjct: 27 SVRKVVAEFIGTFFLIFVGCGSVVVDKISNGSITHLGVSLVWGMAAMIVIYSIGHISGAH 86
Query: 208 MNPARTLG-PAIFHSKYRAIVVYFVSTIFGAVAGAWVFNIL 247
+NPA TL A+ + + Y V+ +FG+++ ++ +
Sbjct: 87 LNPAVTLALAAVKRFPWVQVPGYIVAQVFGSISAGFLLRFM 127
>UniRef100_Q9C6T0 Putative aquaporin NIP3.1 [Arabidopsis thaliana]
Length = 269
Score = 247 bits (630), Expect = 3e-64
Identities = 124/217 (57%), Positives = 164/217 (75%), Gaps = 7/217 (3%)
Query: 50 LIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKR 109
+IF GC++IVVN+ VTLPGIALVWGLV+ V+IYS+GH+SGAHFNPAV+ AFA++K+
Sbjct: 1 MIFAGCSAIVVNETYGKPVTLPGIALVWGLVVTVMIYSIGHVSGAHFNPAVSIAFASSKK 60
Query: 110 FPWIQVAPYIASQLLGAVLASGILKMLF-------SGTHDQFSGTIPSGTNLQAFVIEFI 162
FP+ QV YIA+QLLG+ LA+ +L+++F S D + GT PS +N +FV+EFI
Sbjct: 61 FPFNQVPGYIAAQLLGSTLAAAVLRLVFHLDDDVCSLKGDVYVGTYPSNSNTTSFVMEFI 120
Query: 163 TTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSK 222
TF LMFVISAVATD RA G AGIAIG+T++L+IL SGPI+GASMNPAR+LGPA+
Sbjct: 121 ATFNLMFVISAVATDKRATGSFAGIAIGATIVLDILFSGPISGASMNPARSLGPALIWGC 180
Query: 223 YRAIVVYFVSTIFGAVAGAWVFNILRYTDKPLHEITK 259
Y+ + +Y VS + GA++GAW + +LR T K EI +
Sbjct: 181 YKDLWLYIVSPVIGALSGAWTYGLLRSTKKSYSEIIR 217
Score = 32.7 bits (73), Expect = 9.8
Identities = 42/149 (28%), Positives = 66/149 (44%), Gaps = 18/149 (12%)
Query: 8 FETHDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNV 67
F D V S+ D S++ T+ F V EF+ TF L+F +++ +K
Sbjct: 88 FHLDDDVCSLKGDVYVGTYPSNSNTTSF-----VMEFIATFNLMFV-ISAVATDKRATG- 140
Query: 68 VTLPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFAT---TKRFPWIQVAPYIASQLL 124
+ GIA+ G +++ I G ISGA NPA + A + W+ YI S ++
Sbjct: 141 -SFAGIAI--GATIVLDILFSGPISGASMNPARSLGPALIWGCYKDLWL----YIVSPVI 193
Query: 125 GAVLASGILKMLFSGTHDQFSGTIPSGTN 153
GA+ + +L S T +S I N
Sbjct: 194 GALSGAWTYGLLRS-TKKSYSEIIRPNCN 221
>UniRef100_Q5Z9E2 Putative major intrinsic protein [Oryza sativa]
Length = 273
Score = 238 bits (606), Expect = 2e-61
Identities = 109/218 (50%), Positives = 152/218 (69%), Gaps = 1/218 (0%)
Query: 33 SVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHIS 92
S F+Q L+AEF+ TFFL+F G +I V + VT PG+A+ WG +M ++Y+VGH+S
Sbjct: 52 SFTFLQMLLAEFLATFFLMFAGLGAITVEEKK-GAVTFPGVAVAWGAAVMAMVYAVGHVS 110
Query: 93 GAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSGT 152
GAH NPAVT FA RFPW + Y +Q A AS +L+++F G H T+P G
Sbjct: 111 GAHLNPAVTLGFAVAGRFPWRRAPAYALAQTAAATAASVVLRLMFGGRHAPVPATLPGGA 170
Query: 153 NLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPAR 212
+ Q+ VIEF+ TF LMFVI AVATD++A+G MAG+A+G T++LN+L +GP++GASMNPAR
Sbjct: 171 HAQSLVIEFVITFYLMFVIMAVATDDQAVGHMAGVAVGGTIMLNVLFAGPVSGASMNPAR 230
Query: 213 TLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYT 250
++GPA+ SKY A+ VY + GA AGAW ++++R T
Sbjct: 231 SIGPALVGSKYTALWVYILGPFAGAAAGAWAYSLIRLT 268
>UniRef100_Q8W037 Aquaporin NIP2.1 [Arabidopsis thaliana]
Length = 288
Score = 231 bits (589), Expect = 1e-59
Identities = 125/258 (48%), Positives = 172/258 (66%), Gaps = 12/258 (4%)
Query: 12 DVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLP 71
D L NK S SS SV F+QKL+AE VGT++LIF GCA+I VN +++VVTL
Sbjct: 26 DTSLPSNKHES----SSPPLLSVHFLQKLLAELVGTYYLIFAGCAAIAVNAQHNHVVTLV 81
Query: 72 GIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASG 131
GIA+VWG+V+MVL+Y +GH+S AHFNPAVT A A+++RFP QV YI Q++G+ LAS
Sbjct: 82 GIAVVWGIVIMVLVYCLGHLS-AHFNPAVTLALASSQRFPLNQVPAYITVQVIGSTLASA 140
Query: 132 ILKMLF-------SGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEM 184
L++LF S HD F G+ PSG++LQAFV+EFI T LM V+ AV T R E+
Sbjct: 141 TLRLLFDLNNDVCSKKHDVFLGSSPSGSDLQAFVMEFIITGFLMLVVCAVTTTKRTTEEL 200
Query: 185 AGIAIGSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVF 244
G+ IG+T+ LN++ +G ++GASMNPAR++GPA+ Y+ I +Y ++ GAV+GA +
Sbjct: 201 EGLIIGATVTLNVIFAGEVSGASMNPARSIGPALVWGCYKGIWIYLLAPTLGAVSGALIH 260
Query: 245 NILRYTDKPLHEITKGSS 262
+L E +K S
Sbjct: 261 KMLPSIQNAEPEFSKTGS 278
>UniRef100_Q9FXW2 MIP [Adiantum capillus-veneris]
Length = 282
Score = 221 bits (563), Expect = 1e-56
Identities = 100/222 (45%), Positives = 159/222 (71%), Gaps = 1/222 (0%)
Query: 32 TSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVTLPGIALVWGLVLMVLIYSVGHI 91
T+ +QK+ AE + T+ L+F GC + +V++ + +T G++ +GLV+M++IYSVGHI
Sbjct: 44 TNAGILQKVGAELISTYILVFAGCGAAMVDEKSGGAITHFGVSAAFGLVVMIMIYSVGHI 103
Query: 92 SGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLASGILKMLFSGTHDQFSGTIPSG 151
SGAH NPAVT AFAT + FPW QV YI +Q++ A+ A+ L+++ G + T+P G
Sbjct: 104 SGAHMNPAVTLAFATVRHFPWAQVPAYIGAQVVAAISAAFSLRLILGGAA-KIGATLPVG 162
Query: 152 TNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAIGSTLLLNILISGPITGASMNPA 211
+++Q+ +E IT+++LMFV+SAVATD RAIGE+AG+A+GS + L+ + +GPI GASMNPA
Sbjct: 163 SDVQSLALEVITSYILMFVVSAVATDTRAIGELAGLAVGSAVALDAIFAGPICGASMNPA 222
Query: 212 RTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRYTDKP 253
R++GPA+ ++++ VY V I G + GAW + +++ ++P
Sbjct: 223 RSIGPAVASYDFKSLWVYIVGPILGCLLGAWSYTMIKLPEQP 264
>UniRef100_Q6KAW0 Nod26-like major intrinsic protein [Cicer arietinum]
Length = 273
Score = 221 bits (563), Expect = 1e-56
Identities = 114/262 (43%), Positives = 170/262 (64%), Gaps = 9/262 (3%)
Query: 10 THDVVLSVNKDASKTIESSDTYTSVFFVQKLVAEFVGTFFLIFTGCASIVVNKNNDNVVT 69
TH +V + N + + Y S F +K++AE +GT+ L+F G S +N ++N V+
Sbjct: 5 THSLVNATNDFQNHITQKQSLYPSGF-PRKVLAEVIGTYLLVFVGSGSAAMNAIDENKVS 63
Query: 70 LPGIALVWGLVLMVLIYSVGHISGAHFNPAVTFAFATTKRFPWIQVAPYIASQLLGAVLA 129
G ++ G ++ V+IY++GHISGAH NPAV+ AFAT FPW QV YIA+QL GA+ A
Sbjct: 64 KLGASMAGGFIVTVMIYAIGHISGAHMNPAVSLAFATVSHFPWKQVPFYIAAQLTGAISA 123
Query: 130 SGILKMLFSGTHDQFSGTIPSGTNLQAFVIEFITTFLLMFVISAVATDNRAIGEMAGIAI 189
S LK+L + Q T PSG+N+QA +IE +TTF ++ + +AV+TD +AIGE++G+A+
Sbjct: 124 SYTLKVLLEPSK-QLGATSPSGSNIQALIIEIVTTFTMVLISTAVSTDPKAIGELSGVAV 182
Query: 190 GSTLLLNILISGPITGASMNPARTLGPAIFHSKYRAIVVYFVSTIFGAVAGAWVFNILRY 249
GS++ + +++GPI+G SMNPARTLGPAI S Y+ I VY V I GA+ G W + +++
Sbjct: 183 GSSVCIASIVAGPISGGSMNPARTLGPAIATSSYKGIWVYMVGPITGALLGTWSYVVIQE 242
Query: 250 TDK-------PLHEITKGSSIL 264
T+K LH KG ++
Sbjct: 243 TNKQALTTSLKLHHEMKGIELV 264
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.138 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 396,230,083
Number of Sequences: 2790947
Number of extensions: 15086932
Number of successful extensions: 56432
Number of sequences better than 10.0: 1236
Number of HSP's better than 10.0 without gapping: 1101
Number of HSP's successfully gapped in prelim test: 135
Number of HSP's that attempted gapping in prelim test: 52267
Number of HSP's gapped (non-prelim): 1809
length of query: 265
length of database: 848,049,833
effective HSP length: 125
effective length of query: 140
effective length of database: 499,181,458
effective search space: 69885404120
effective search space used: 69885404120
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0162a.10


