

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0162a.1
(118 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
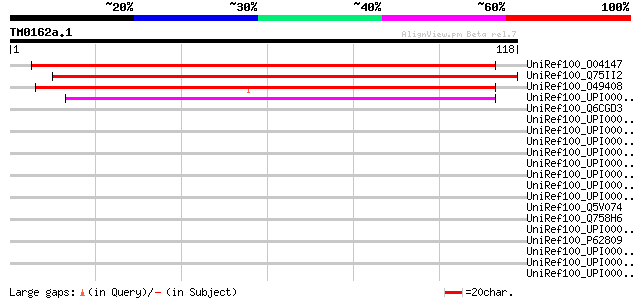
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O04147 Cyclic phosphodiesterase [Arabidopsis thaliana] 131 3e-30
UniRef100_Q75II2 Hypothetical protein B1130G10.20 [Oryza sativa] 125 3e-28
UniRef100_O49408 Hypothetical protein AT4g18940 [Arabidopsis tha... 105 2e-22
UniRef100_UPI000029E4A6 UPI000029E4A6 UniRef100 entry 45 5e-04
UniRef100_Q6CGD3 Similar to sp|P53314 Saccharomyces cerevisiae Y... 44 0.001
UniRef100_UPI00002F1A13 UPI00002F1A13 UniRef100 entry 42 0.003
UniRef100_UPI0000312722 UPI0000312722 UniRef100 entry 38 0.047
UniRef100_UPI00002B3C08 UPI00002B3C08 UniRef100 entry 38 0.047
UniRef100_UPI000030A849 UPI000030A849 UniRef100 entry 37 0.14
UniRef100_UPI000031C135 UPI000031C135 UniRef100 entry 35 0.30
UniRef100_UPI00002F151A UPI00002F151A UniRef100 entry 35 0.40
UniRef100_UPI00002D0634 UPI00002D0634 UniRef100 entry 35 0.40
UniRef100_UPI0000338466 UPI0000338466 UniRef100 entry 35 0.52
UniRef100_Q5V074 Probable cysteine desulfurase [Haloarcula maris... 34 0.68
UniRef100_Q758H6 AEL214Cp [Ashbya gossypii] 33 1.5
UniRef100_UPI000029218F UPI000029218F UniRef100 entry 32 3.4
UniRef100_P62809 Cyclic phosphodiesterase [Triticum aestivum] 32 3.4
UniRef100_UPI000033A6AC UPI000033A6AC UniRef100 entry 32 4.4
UniRef100_UPI00002E3BF0 UPI00002E3BF0 UniRef100 entry 32 4.4
UniRef100_UPI00002C3112 UPI00002C3112 UniRef100 entry 32 4.4
>UniRef100_O04147 Cyclic phosphodiesterase [Arabidopsis thaliana]
Length = 181
Score = 131 bits (330), Expect = 3e-30
Identities = 61/108 (56%), Positives = 75/108 (68%)
Query: 6 IETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKL 65
+E KK+VYSVWA+P + PR KLM LRSEF GP F PH+TV + LTAD+A
Sbjct: 1 MEEVKKDVYSVWALPDEESEPRFKKLMEALRSEFTGPRFVPHVTVAVSAYLTADEAKKMF 60
Query: 66 RSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
SAC+ LKA+ TVDRV+ GTFF+QCV+LLL +P V+E HC NHF
Sbjct: 61 ESACDGLKAYTATVDRVSTGTFFFQCVFLLLQTTPEVMEAGEHCKNHF 108
>UniRef100_Q75II2 Hypothetical protein B1130G10.20 [Oryza sativa]
Length = 185
Score = 125 bits (313), Expect = 3e-28
Identities = 60/108 (55%), Positives = 73/108 (67%)
Query: 11 KEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACE 70
KEVYSVWA+PP+ VR R+ +M LR+ GGP FEPH TVVGAI L A+ LR+A
Sbjct: 9 KEVYSVWALPPEPVRARLRGVMAGLRAAHGGPAFEPHATVVGAIRLRRSAAVEALRAAAA 68
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGYKSS 118
++ + V VA G FFYQC+YLLL P+P VVE S HC HFGY+ S
Sbjct: 69 GVRPYTARVVGVARGDFFYQCIYLLLEPTPEVVEASDHCCGHFGYERS 116
>UniRef100_O49408 Hypothetical protein AT4g18940 [Arabidopsis thaliana]
Length = 196
Score = 105 bits (263), Expect = 2e-22
Identities = 48/109 (44%), Positives = 72/109 (66%), Gaps = 2/109 (1%)
Query: 7 ETQKKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAI--TLTADDALNK 64
+ ++K +Y++WA+P D V R+ +LM LRSEFGGP F+PH+T+VG LTA +A
Sbjct: 18 DEEEKAMYAIWAVPEDDVEDRLQRLMEGLRSEFGGPAFDPHLTLVGPFPYKLTASEAKRM 77
Query: 65 LRSACEELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
+SACE K + TVD+V+AGT ++QC+Y+ L + V+ + H HF
Sbjct: 78 FKSACEGFKVYPATVDQVSAGTSYFQCLYVSLRHTVEVMNAAGHFMAHF 126
>UniRef100_UPI000029E4A6 UPI000029E4A6 UniRef100 entry
Length = 167
Score = 44.7 bits (104), Expect = 5e-04
Identities = 21/100 (21%), Positives = 43/100 (43%)
Query: 14 YSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELK 73
YS+W IP R R +++ L + P F PH T+ G + + + + + +
Sbjct: 4 YSIWLIPERDSRSRFREIILRLSKKLRSPIFHPHCTIYGRLNIPLGKIIPAVDHLSKSID 63
Query: 74 AFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHF 113
F V ++ G ++ +YL + + ++ +C F
Sbjct: 64 QFSTIVKKLKTGKTKWKSLYLTIEKKEEMEKSYNYCKKLF 103
>UniRef100_Q6CGD3 Similar to sp|P53314 Saccharomyces cerevisiae YGR247w hypothetical
protein [Yarrowia lipolytica]
Length = 219
Score = 43.5 bits (101), Expect = 0.001
Identities = 30/109 (27%), Positives = 57/109 (51%), Gaps = 8/109 (7%)
Query: 15 SVWAIPPDH--VRPRIAKLMTDLRSEFG-GPHFEPHITVVGAITLTA----DDALNKLRS 67
S+W PP + + ++A + L+ F +FEPHIT+ I++ D L++ +
Sbjct: 7 SLWLQPPRNSPIWQQLATTIQGLKPIFSDSENFEPHITLTSNISVNTQGQVDFVLDRAVA 66
Query: 68 ACEEL-KAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFGY 115
A + + + FQ+ + V G+ F++ VYL + P+P ++ + C F Y
Sbjct: 67 AAKCVPQGFQIHLSSVKYGSRFFKKVYLQVEPTPELLSLARICREDFVY 115
>UniRef100_UPI00002F1A13 UPI00002F1A13 UniRef100 entry
Length = 156
Score = 42.0 bits (97), Expect = 0.003
Identities = 25/84 (29%), Positives = 42/84 (49%), Gaps = 7/84 (8%)
Query: 13 VYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAIT--LTADDALNKLRSACE 70
+YS+W + I K++ L E +FEPH+T+V + +A A NK+
Sbjct: 1 MYSIWVESSSNQLTEIKKVIKILSEEEDKDYFEPHVTLVTNLNSERSAMSAFNKIPK--- 57
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYL 94
K F +T D + GT ++Q ++L
Sbjct: 58 --KRFDITFDTIQEGTTYFQRLFL 79
>UniRef100_UPI0000312722 UPI0000312722 UniRef100 entry
Length = 162
Score = 38.1 bits (87), Expect = 0.047
Identities = 18/86 (20%), Positives = 46/86 (52%), Gaps = 7/86 (8%)
Query: 11 KEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAIT--LTADDALNKLRSA 68
K+++S+W +P D ++K++ L + P F+PH T+ +I A+ ++++
Sbjct: 3 KKIFSIWLLPIDSDNHYLSKIVQSLSEVYNAPFFQPHCTLFSSINNLKNAEKIIDQI--- 59
Query: 69 CEELKAFQVTVDRVAAGTFFYQCVYL 94
++ +F+V V+ + ++ V++
Sbjct: 60 --DIDSFKVDVNGINHSNDVWKTVFI 83
>UniRef100_UPI00002B3C08 UPI00002B3C08 UniRef100 entry
Length = 156
Score = 38.1 bits (87), Expect = 0.047
Identities = 24/84 (28%), Positives = 41/84 (48%), Gaps = 7/84 (8%)
Query: 13 VYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITL--TADDALNKLRSACE 70
+YS+W + I ++ L E +FEPH+T+V + +A D NK+
Sbjct: 1 MYSIWVESSLNQLSDIKGIIKTLSLEEDKEYFEPHVTLVTNLKTERSAMDVFNKIPK--- 57
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYL 94
K F +T D + GT ++Q ++L
Sbjct: 58 --KRFDITFDTIQEGTTYFQRLFL 79
>UniRef100_UPI000030A849 UPI000030A849 UniRef100 entry
Length = 111
Score = 36.6 bits (83), Expect = 0.14
Identities = 23/84 (27%), Positives = 41/84 (48%), Gaps = 7/84 (8%)
Query: 13 VYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITL--TADDALNKLRSACE 70
+YS+W + I ++ L E +F+PH+T+V + +A D NK+
Sbjct: 1 MYSIWVESSLNQLSDIKGIIKTLSLEEDKEYFDPHVTLVTNLKTERSAMDVFNKIPK--- 57
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYL 94
K F +T D + GT ++Q ++L
Sbjct: 58 --KRFDITFDTIQEGTNYFQRLFL 79
>UniRef100_UPI000031C135 UPI000031C135 UniRef100 entry
Length = 161
Score = 35.4 bits (80), Expect = 0.30
Identities = 23/85 (27%), Positives = 42/85 (49%), Gaps = 4/85 (4%)
Query: 10 KKEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSAC 69
+K YS+W H + + L+ E F PH+T+V I+ + ++A+N L
Sbjct: 4 EKTTYSIWLQNSSHNQD-LLNLIDKFAKEESKDSFFPHVTLVTHIS-SEEEAINILSKL- 60
Query: 70 EELKAFQVTVDRVAAGTFFYQCVYL 94
+ +V D+++ G F+Q +YL
Sbjct: 61 -RINKDEVVFDKISTGDTFFQKLYL 84
>UniRef100_UPI00002F151A UPI00002F151A UniRef100 entry
Length = 159
Score = 35.0 bits (79), Expect = 0.40
Identities = 28/86 (32%), Positives = 40/86 (45%), Gaps = 8/86 (9%)
Query: 11 KEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLT--ADDALNKLRSA 68
KE YS+W H ++ +L+ E F PH+T+V I A LNKL
Sbjct: 3 KETYSIWLQNSIH-NEKVYELIKRYAKEEDKDSFFPHVTLVSHIDSEEKALKILNKLL-- 59
Query: 69 CEELKAFQVTVDRVAAGTFFYQCVYL 94
+K V DRV+ G ++Q +YL
Sbjct: 60 ---VKKSTVIFDRVSTGDTYFQKLYL 82
>UniRef100_UPI00002D0634 UPI00002D0634 UniRef100 entry
Length = 188
Score = 35.0 bits (79), Expect = 0.40
Identities = 23/90 (25%), Positives = 39/90 (42%), Gaps = 9/90 (10%)
Query: 16 VWAIPPDHVRPRIAKLMTDLRSEFG-GPHFEPHITVVGAITLTADDALNKLRSACEEL-- 72
+W P AK + D+ P+F PH+T+VGA + D + S+
Sbjct: 1 MWLEPTTSTYAPCAKFILDVAHRHPEAPYFPPHVTLVGAFHASDDRRATSIFSSAARAFS 60
Query: 73 ------KAFQVTVDRVAAGTFFYQCVYLLL 96
+ ++ ++RV G +QCVY+ L
Sbjct: 61 SVDGGRRGPRIGLERVECGERRHQCVYIRL 90
>UniRef100_UPI0000338466 UPI0000338466 UniRef100 entry
Length = 165
Score = 34.7 bits (78), Expect = 0.52
Identities = 17/82 (20%), Positives = 36/82 (43%)
Query: 15 SVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKA 74
S+W P + K++ L + P F PH T++G I L D N + + + +
Sbjct: 3 SIWLCPTHEDELYLQKIINKLSENYNSPIFLPHCTLLGGIKLNEFDLENIIDISIDNIGL 62
Query: 75 FQVTVDRVAAGTFFYQCVYLLL 96
++ + + ++ V++ L
Sbjct: 63 IKIKSNGIDYTNIIWKTVFIKL 84
>UniRef100_Q5V074 Probable cysteine desulfurase [Haloarcula marismortui]
Length = 367
Score = 34.3 bits (77), Expect = 0.68
Identities = 26/99 (26%), Positives = 42/99 (42%), Gaps = 19/99 (19%)
Query: 13 VYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEEL 72
+ +V AI D ++PR+ +L L+ G P G +T TADD
Sbjct: 276 IETVEAIGFDTIQPRVERLTDRLKDGLGDRLLSPRDYESGLVTFTADDP----------- 324
Query: 73 KAFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSN 111
+ TV+R+AA + + + P+P + S H N
Sbjct: 325 ---EATVERLAA-----EGIIIRSLPNPDALRASVHAFN 355
>UniRef100_Q758H6 AEL214Cp [Ashbya gossypii]
Length = 216
Score = 33.1 bits (74), Expect = 1.5
Identities = 27/94 (28%), Positives = 44/94 (46%), Gaps = 14/94 (14%)
Query: 15 SVWAIPPDH--VRPRIAKLMTDLRSEF-GGPHFEPHITVVGAITL-TADDALNKLRSACE 70
++W PP + + L+ L++ F FEPH+T+ + +ADDA L +AC
Sbjct: 4 ALWYCPPANSPCYDTVNSLILSLQTLFPDAVLFEPHVTITSQLRCDSADDAAAVLAAACA 63
Query: 71 ELKAFQ----------VTVDRVAAGTFFYQCVYL 94
L+A + V D VA G ++ V+L
Sbjct: 64 ALRACRPQLDARGSPVVRFDAVAVGRRYFDKVHL 97
>UniRef100_UPI000029218F UPI000029218F UniRef100 entry
Length = 159
Score = 32.0 bits (71), Expect = 3.4
Identities = 17/101 (16%), Positives = 42/101 (40%)
Query: 14 YSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELK 73
+SVW IP + + +L+ ++ F+PH+T+ G I + + +
Sbjct: 5 WSVWLIPSIQEKKKYKQLIRLYSKKYSFSLFDPHVTLFGRIDIEPGSTFSFFEECVKGQD 64
Query: 74 AFQVTVDRVAAGTFFYQCVYLLLHPSPPVVETSAHCSNHFG 114
++ V G ++ +Y+ + +++ + +N G
Sbjct: 65 RIRLDTLDVVRGAPPWKSLYIQFKLNSTIIDLQSKINNKLG 105
>UniRef100_P62809 Cyclic phosphodiesterase [Triticum aestivum]
Length = 51
Score = 32.0 bits (71), Expect = 3.4
Identities = 13/19 (68%), Positives = 14/19 (73%)
Query: 40 GGPHFEPHITVVGAITLTA 58
GGP FEPH TVVGA+ A
Sbjct: 4 GGPPFEPHATVVGAVAAAA 22
>UniRef100_UPI000033A6AC UPI000033A6AC UniRef100 entry
Length = 165
Score = 31.6 bits (70), Expect = 4.4
Identities = 17/82 (20%), Positives = 35/82 (41%)
Query: 15 SVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEELKA 74
S+W P + I ++ DL F PH T++ + +D N + + + +
Sbjct: 3 SIWLCPNNDDELYIQNIINDLSDIHNCARFYPHCTLLSGVKENNNDLENIIDKSIKNIGP 62
Query: 75 FQVTVDRVAAGTFFYQCVYLLL 96
V R++ F++ V++ L
Sbjct: 63 IMVKSKRISFKDIFWKTVFIEL 84
>UniRef100_UPI00002E3BF0 UPI00002E3BF0 UniRef100 entry
Length = 158
Score = 31.6 bits (70), Expect = 4.4
Identities = 17/83 (20%), Positives = 40/83 (47%), Gaps = 3/83 (3%)
Query: 12 EVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACEE 71
++YS+WA ++ + +++ ++ + F PH+T+V + + + K+ +
Sbjct: 2 DIYSIWAKSKNNSLNKAKEIINEISQKEESIPFNPHVTLV--TNIYSQEKAEKILEKINK 59
Query: 72 LKAFQVTVDRVAAGTFFYQCVYL 94
K V D V G ++Q ++L
Sbjct: 60 SKII-VNFDSVGTGNTYFQRLFL 81
>UniRef100_UPI00002C3112 UPI00002C3112 UniRef100 entry
Length = 184
Score = 31.6 bits (70), Expect = 4.4
Identities = 19/93 (20%), Positives = 37/93 (39%)
Query: 11 KEVYSVWAIPPDHVRPRIAKLMTDLRSEFGGPHFEPHITVVGAITLTADDALNKLRSACE 70
K+ YS+W + KL + GP F+ H+T+ G + ++ L L +
Sbjct: 6 KKTYSIWGEFSKIDFASLNKLQHMVNHSLNGPLFKVHLTLSGPFQILDENILGGLENLAM 65
Query: 71 ELKAFQVTVDRVAAGTFFYQCVYLLLHPSPPVV 103
++ D +Q Y+ + SP ++
Sbjct: 66 TNSIIEMRTDGYGTEDNVFQSFYVQIQVSPELL 98
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 196,961,389
Number of Sequences: 2790947
Number of extensions: 6596597
Number of successful extensions: 14338
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 14323
Number of HSP's gapped (non-prelim): 30
length of query: 118
length of database: 848,049,833
effective HSP length: 94
effective length of query: 24
effective length of database: 585,700,815
effective search space: 14056819560
effective search space used: 14056819560
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0162a.1


