

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0159.10
(353 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
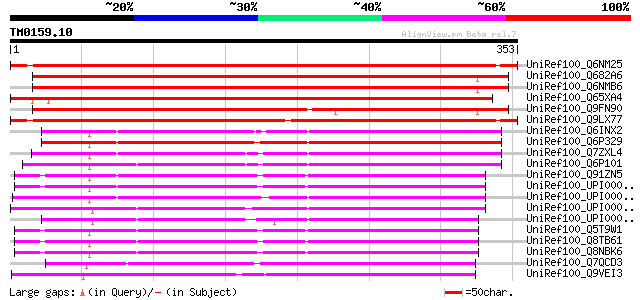
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6NM25 At3g46180 [Arabidopsis thaliana] 559 e-158
UniRef100_Q682A6 MRNA, complete cds, clone: RAFL21-88-P21 [Arabi... 551 e-156
UniRef100_Q6NMB6 At5g59740 [Arabidopsis thaliana] 551 e-155
UniRef100_Q65XA4 Hypothetical protein OJ1654_B10.14 [Oryza sativa] 513 e-144
UniRef100_Q9FN90 UDP-galactose transporter related protein-like ... 498 e-139
UniRef100_Q9LX77 Hypothetical protein F12M12_150 [Arabidopsis th... 491 e-137
UniRef100_Q6INX2 LOC432208 protein [Xenopus laevis] 230 5e-59
UniRef100_Q6P329 LOC394918 protein [Xenopus tropicalis] 229 1e-58
UniRef100_Q7ZXL4 Slc35b1-prov protein [Xenopus laevis] 226 1e-57
UniRef100_Q6P101 Hypothetical protein zgc:77349 [Brachydanio rerio] 224 3e-57
UniRef100_Q91ZN5 Embryonic seven-span transmembrane protein-like... 216 8e-55
UniRef100_UPI0000181102 UPI0000181102 UniRef100 entry 215 1e-54
UniRef100_UPI00003ACCF3 UPI00003ACCF3 UniRef100 entry 214 4e-54
UniRef100_UPI000027CE36 UPI000027CE36 UniRef100 entry 211 3e-53
UniRef100_UPI0000362925 UPI0000362925 UniRef100 entry 210 5e-53
UniRef100_Q5T9W1 OTTHUMP00000016513 [Homo sapiens] 208 2e-52
UniRef100_Q8TB61 Solute carrier family 35, member B2 (Adenosine ... 208 2e-52
UniRef100_Q8NBK6 Hypothetical protein PSEC0149 [Homo sapiens] 207 4e-52
UniRef100_Q7QCD3 ENSANGP00000012787 [Anopheles gambiae str. PEST] 199 1e-49
UniRef100_Q9VEI3 CG7623-PA [Drosophila melanogaster] 194 3e-48
>UniRef100_Q6NM25 At3g46180 [Arabidopsis thaliana]
Length = 347
Score = 559 bits (1441), Expect = e-158
Identities = 270/353 (76%), Positives = 310/353 (87%), Gaps = 6/353 (1%)
Query: 1 MAESPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHS 60
MAE S + + + KLWK +FA+SGIM+TLV YG+LQEKIMRVPYG +KEYFKHS
Sbjct: 1 MAEPDSVNEAKE----KKKKLWKAVFAISGIMLTLVIYGLLQEKIMRVPYGLKKEYFKHS 56
Query: 61 LFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPV 120
LFLVFCNR+TTSAVSA +LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPV
Sbjct: 57 LFLVFCNRLTTSAVSAAALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPV 116
Query: 121 QTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGREN 180
QTLAKCAKMIPVM+WGT+IMQK+Y+G DY++AFLVTLGCSVFIL+PAG+DISPY++GREN
Sbjct: 117 QTLAKCAKMIPVMVWGTLIMQKKYRGFDYLVAFLVTLGCSVFILFPAGDDISPYNKGREN 176
Query: 181 TVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIP 240
TVWGV LM GYLG DGFTSTFQDKLF+GY+MEIHNQIFYTT+CS ILS TGLILQGHL+P
Sbjct: 177 TVWGVSLMVGYLGFDGFTSTFQDKLFKGYNMEIHNQIFYTTICSSILSFTGLILQGHLLP 236
Query: 241 AIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWF 300
A++FV RH DC FDIALLSTVATASQFFISYTIRTFGALTFA IMTTRQL SIMLSC+WF
Sbjct: 237 AVDFVSRHRDCLFDIALLSTVATASQFFISYTIRTFGALTFAAIMTTRQLASIMLSCIWF 296
Query: 301 SHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDSIPLVQSGDSNNLKDNP 353
SHPLSWEQ IG+VIVFGSLYAK+F +K +K ++ +P + ++ LK NP
Sbjct: 297 SHPLSWEQCIGSVIVFGSLYAKTFVKKKSEKPPAAQELP--RDEEAQPLKGNP 347
>UniRef100_Q682A6 MRNA, complete cds, clone: RAFL21-88-P21 [Arabidopsis thaliana]
Length = 344
Score = 551 bits (1421), Expect = e-156
Identities = 264/333 (79%), Positives = 299/333 (89%), Gaps = 2/333 (0%)
Query: 17 RDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSA 76
++NKLWKG+FAVSGIM TLV YGVLQEKIMRVPYG KE+FKHSLFLVFCNR+TTSAVSA
Sbjct: 12 KENKLWKGVFAVSGIMSTLVIYGVLQEKIMRVPYGVNKEFFKHSLFLVFCNRLTTSAVSA 71
Query: 77 GSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWG 136
G+LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPVQTLAKCAKMIPVM+WG
Sbjct: 72 GALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMVWG 131
Query: 137 TIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDG 196
T+IMQK+Y+G DY++AFLVTLGCSVFIL+PAG+D+SPY++GRENTVWGV LM GYLG DG
Sbjct: 132 TLIMQKKYKGFDYLVAFLVTLGCSVFILFPAGDDVSPYNKGRENTVWGVSLMAGYLGFDG 191
Query: 197 FTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIA 256
FTSTFQDKLF+GY+MEIHNQIFYTTLCSC+LS TGLILQGHL+PA++FV H DC DIA
Sbjct: 192 FTSTFQDKLFKGYNMEIHNQIFYTTLCSCVLSFTGLILQGHLLPAVDFVSLHRDCLLDIA 251
Query: 257 LLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVF 316
LLSTVATASQFFISYTIRTFGALTFA IMTTRQL SIMLSC+WFSHPLSWEQ IG+VIVF
Sbjct: 252 LLSTVATASQFFISYTIRTFGALTFAAIMTTRQLASIMLSCIWFSHPLSWEQCIGSVIVF 311
Query: 317 GSLYAKSF--TRKAPQKTTSSDSIPLVQSGDSN 347
GSLYAK+ ++K Q +P + +S+
Sbjct: 312 GSLYAKNLLNSKKNSQTQPPPPELPQYEKVESS 344
>UniRef100_Q6NMB6 At5g59740 [Arabidopsis thaliana]
Length = 344
Score = 551 bits (1420), Expect = e-155
Identities = 264/333 (79%), Positives = 298/333 (89%), Gaps = 2/333 (0%)
Query: 17 RDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSA 76
++NKLWKG+FAVSGIM TLV YGVLQEKIMRVPYG KE+FKHSLFLVFCNR+TTSAVSA
Sbjct: 12 KENKLWKGVFAVSGIMSTLVIYGVLQEKIMRVPYGVNKEFFKHSLFLVFCNRLTTSAVSA 71
Query: 77 GSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWG 136
G+LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPVQTLAKCAKMIPVM+WG
Sbjct: 72 GALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMVWG 131
Query: 137 TIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDG 196
T+IMQK+Y+G DY++AFLVTLGCSVFIL+PAG+D+SPY++GRENTVWGV LM GYLG DG
Sbjct: 132 TLIMQKKYKGFDYLVAFLVTLGCSVFILFPAGDDVSPYNKGRENTVWGVSLMAGYLGFDG 191
Query: 197 FTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIA 256
FTSTFQDKLF+GY+MEIHNQIFYTTLCSC+LS TGLILQGHL+PA++FV H DC DIA
Sbjct: 192 FTSTFQDKLFKGYNMEIHNQIFYTTLCSCVLSFTGLILQGHLLPAVDFVSLHRDCLLDIA 251
Query: 257 LLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVF 316
LLSTVATASQFFISYTIRTFGALTFA IMTTRQL SIMLSC+WFSHPLSWEQ IG+VIVF
Sbjct: 252 LLSTVATASQFFISYTIRTFGALTFAAIMTTRQLASIMLSCIWFSHPLSWEQCIGSVIVF 311
Query: 317 GSLYAKSF--TRKAPQKTTSSDSIPLVQSGDSN 347
GSLYAK+ +K Q +P + +S+
Sbjct: 312 GSLYAKNLLNNKKNSQTQPPPPELPQYEKVESS 344
>UniRef100_Q65XA4 Hypothetical protein OJ1654_B10.14 [Oryza sativa]
Length = 360
Score = 513 bits (1322), Expect = e-144
Identities = 252/352 (71%), Positives = 294/352 (82%), Gaps = 16/352 (4%)
Query: 1 MAESPSTSSSSSPP--------------DSRDNKLWKGI--FAVSGIMVTLVTYGVLQEK 44
MA++ S+S+S+S D D + G FAV GIM TL+ YG+LQEK
Sbjct: 1 MADAASSSASASAAVAGTLPVAAAATGKDKEDRRRLVGRCGFAVVGIMSTLLIYGLLQEK 60
Query: 45 IMRVPYGAQKEYFKHSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNIL 104
IMRVPYGA+KE+F++SLFLVFCNRITTS VSA L ASKK+LDPVAP+ KYC+VSVSNIL
Sbjct: 61 IMRVPYGAEKEFFRYSLFLVFCNRITTSTVSALVLTASKKSLDPVAPLQKYCVVSVSNIL 120
Query: 105 TTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFIL 164
TTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIM+K+Y G DY A +VT+GCS+FIL
Sbjct: 121 TTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMRKKYGGKDYFFAVVVTVGCSLFIL 180
Query: 165 YPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCS 224
YPA D SP++RGRENT+WGV LM GYLG DGFTSTFQDKLF+GYDMEIHNQIFYTT+CS
Sbjct: 181 YPASMDASPFNRGRENTIWGVSLMLGYLGFDGFTSTFQDKLFKGYDMEIHNQIFYTTMCS 240
Query: 225 CILSLTGLILQGHLIPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATI 284
C+LSL+GLILQ +IPA++F++RH DCF+D+ +LS+VATASQFFISYTIRTFGALTFATI
Sbjct: 241 CVLSLSGLILQNQMIPAVDFMFRHPDCFYDVIILSSVATASQFFISYTIRTFGALTFATI 300
Query: 285 MTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSD 336
MTTRQLVSI+LSCVWF HPLSW QW+GA IVFG+LY KSF R PQK +++
Sbjct: 301 MTTRQLVSILLSCVWFVHPLSWMQWVGAAIVFGALYTKSFLRSKPQKPAAAN 352
>UniRef100_Q9FN90 UDP-galactose transporter related protein-like [Arabidopsis
thaliana]
Length = 345
Score = 498 bits (1281), Expect = e-139
Identities = 247/337 (73%), Positives = 283/337 (83%), Gaps = 9/337 (2%)
Query: 17 RDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHSLFLVFCNRITTSAVSA 76
++NKLWKG+FAVSGIM TLV YGVLQEKIMRVPYG KE+FKHSLFLVFCNR+TTSAVSA
Sbjct: 12 KENKLWKGVFAVSGIMSTLVIYGVLQEKIMRVPYGVNKEFFKHSLFLVFCNRLTTSAVSA 71
Query: 77 GSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWG 136
G+LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPVQTLAKCAKMIPVM+WG
Sbjct: 72 GALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMVWG 131
Query: 137 TIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDG 196
T+IMQK+Y+G DY++AFLVTLGCSVFIL+PAG+D+SPY++GRENTVWGV LM GYLG DG
Sbjct: 132 TLIMQKKYKGFDYLVAFLVTLGCSVFILFPAGDDVSPYNKGRENTVWGVSLMAGYLGFDG 191
Query: 197 FTSTFQDKLFRGYDMEIHNQIFYTTLCSC----ILSLTGLILQGHLIPAIEFVYRHHDCF 252
FTSTFQDKLF+ ++ LCS ++ GLILQGHL+PA++FV H DC
Sbjct: 192 FTSTFQDKLFK---PSKRRSMWSIILCSYHYVNLMVDVGLILQGHLLPAVDFVSLHRDCL 248
Query: 253 FDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGA 312
DIALLSTVATASQFFISYTIRTFGALTFA IMTTRQL SIMLSC+WFSHPLSWEQ IG+
Sbjct: 249 LDIALLSTVATASQFFISYTIRTFGALTFAAIMTTRQLASIMLSCIWFSHPLSWEQCIGS 308
Query: 313 VIVFGSLYAKSF--TRKAPQKTTSSDSIPLVQSGDSN 347
VIVFGSLYAK+ +K Q +P + +S+
Sbjct: 309 VIVFGSLYAKNLLNNKKNSQTQPPPPELPQYEKVESS 345
>UniRef100_Q9LX77 Hypothetical protein F12M12_150 [Arabidopsis thaliana]
Length = 344
Score = 491 bits (1263), Expect = e-137
Identities = 245/353 (69%), Positives = 287/353 (80%), Gaps = 9/353 (2%)
Query: 1 MAESPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEYFKHS 60
MAE S + + + KLWK +FA+SGIM+TLV YG+LQEKIMRVPYG +KEYFKHS
Sbjct: 1 MAEPDSVNEAKE----KKKKLWKAVFAISGIMLTLVIYGLLQEKIMRVPYGLKKEYFKHS 56
Query: 61 LFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPV 120
LFLVFCNR+TTSAVSA +LLASKK LDPVAP+YKYCL+SV+NILTTTCQYEALKYVSFPV
Sbjct: 57 LFLVFCNRLTTSAVSAAALLASKKVLDPVAPVYKYCLISVTNILTTTCQYEALKYVSFPV 116
Query: 121 QTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGREN 180
QTLAKCAKMIPVM+WGT+IMQK+Y+G DY++AFLVTLGCSVFIL+PAG+DISPY++GREN
Sbjct: 117 QTLAKCAKMIPVMVWGTLIMQKKYRGFDYLVAFLVTLGCSVFILFPAGDDISPYNKGREN 176
Query: 181 TVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIP 240
TVWGV LM GYL G+ S L G+ + + Y + LTGLILQGHL+P
Sbjct: 177 TVWGVSLMVGYL---GYISVLLVSLLFGFGIGTLILVCYVHNWVLVKWLTGLILQGHLLP 233
Query: 241 AIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWF 300
A++FV RH DC FDIALLSTVATASQFFISYTIRTFGALTFA IMTTRQL SIMLSC+WF
Sbjct: 234 AVDFVSRHRDCLFDIALLSTVATASQFFISYTIRTFGALTFAAIMTTRQLASIMLSCIWF 293
Query: 301 SHPLSWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDSIPLVQSGDSNNLKDNP 353
SHPLSWEQ IG+VIVFGSLYAK+F +K +K ++ +P + ++ LK NP
Sbjct: 294 SHPLSWEQCIGSVIVFGSLYAKTFVKKKSEKPPAAQELP--RDEEAQPLKGNP 344
>UniRef100_Q6INX2 LOC432208 protein [Xenopus laevis]
Length = 458
Score = 230 bits (586), Expect = 5e-59
Identities = 133/325 (40%), Positives = 196/325 (59%), Gaps = 11/325 (3%)
Query: 23 KGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKHSLFLVFCNRITTSAVSAGS 78
K +F +G+ V+ +T+GVLQE++M YG+++ E F+ S FLVF NRI V AG
Sbjct: 138 KLLFCAAGLQVSYLTWGVLQERVMTRTYGSEEGGPGERFRDSQFLVFMNRILALTV-AGL 196
Query: 79 LLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTI 138
+ K AP+YKY S+SNIL++ CQYEALK++SFP Q LAK +K+IPVM+ G +
Sbjct: 197 YCSVTKQPRHGAPMYKYSFASLSNILSSWCQYEALKFISFPTQVLAKASKVIPVMLMGKL 256
Query: 139 IMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFT 198
+ K Y+ +Y+ A L+++G S+F+L G D P+ T GV+++ GY+ D FT
Sbjct: 257 VSHKSYEYWEYLTAVLISVGVSMFLLSNGGGD-RPWG---VTTFSGVVILAGYIVFDSFT 312
Query: 199 STFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALL 258
S +QD LF+ Y M +F L SC+ ++ L+ QG L+ I F+ RH D F ALL
Sbjct: 313 SNWQDSLFK-YKMSSVQMMFGVNLFSCLFTVGSLLEQGALLEDIHFMNRHPDFAFHAALL 371
Query: 259 STVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGS 318
S + Q FI YTI FGA F IMT RQ ++I+LSC+ + HP++ +G IVF +
Sbjct: 372 SVCSAFGQLFIFYTINKFGAAIFTIIMTLRQALAILLSCLLYGHPVTAVGGVGVAIVFLA 431
Query: 319 LYAKSFTR-KAPQKTTSSDSIPLVQ 342
L+ + + R + +K + P+VQ
Sbjct: 432 LFLRVYARGRMRKKVRKTADTPVVQ 456
>UniRef100_Q6P329 LOC394918 protein [Xenopus tropicalis]
Length = 453
Score = 229 bits (583), Expect = 1e-58
Identities = 130/325 (40%), Positives = 196/325 (60%), Gaps = 11/325 (3%)
Query: 23 KGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKHSLFLVFCNRITTSAVSAGS 78
K +F +G+ V+ +T+GVLQE++M YG+++ E F+ S FLVF NRI V AG
Sbjct: 133 KLLFCAAGLQVSYLTWGVLQERVMTRTYGSEEGSPGERFRDSQFLVFMNRILALTV-AGV 191
Query: 79 LLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTI 138
+ K AP+YKY S+SNIL++ CQYEALK++SFP Q LAK +K+IPVM+ G +
Sbjct: 192 YCSISKQPRHGAPMYKYSFASLSNILSSWCQYEALKFISFPTQVLAKASKVIPVMLMGKL 251
Query: 139 IMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFT 198
+ K Y+ +Y+ A L+++G S+F+L G + T GV++++GY+ D FT
Sbjct: 252 VSHKTYEYWEYLTAVLISVGVSMFLLSNGGGN----RPSGVTTFSGVVILSGYIVFDSFT 307
Query: 199 STFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALL 258
S +QD LF+ Y M +F L SC+ ++ L+ QG L+ A+ F+ RH D F ALL
Sbjct: 308 SNWQDSLFK-YKMSSVQMMFGVNLFSCLFTVGSLLEQGALLEAVHFMSRHPDFAFHAALL 366
Query: 259 STVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGS 318
S + Q FI YTI FGA F IMT RQ ++I+LSC+ + HP++ +G +VF +
Sbjct: 367 SVCSAFGQLFIFYTINKFGAAIFTIIMTLRQALAILLSCLLYGHPVTAVGGVGVAVVFLA 426
Query: 319 LYAKSFTR-KAPQKTTSSDSIPLVQ 342
L+ + + R + +K + P+VQ
Sbjct: 427 LFLRVYARGRMRKKGRKAADTPVVQ 451
>UniRef100_Q7ZXL4 Slc35b1-prov protein [Xenopus laevis]
Length = 439
Score = 226 bits (575), Expect = 1e-57
Identities = 131/332 (39%), Positives = 197/332 (58%), Gaps = 11/332 (3%)
Query: 16 SRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKHSLFLVFCNRITT 71
S + K +F +G+ V+ +T+GVLQE++M YG+++ E F+ S FLVF NRI
Sbjct: 112 SNTQQALKLLFCTAGLQVSYLTWGVLQERVMTRTYGSEEGGPGERFRDSQFLVFMNRILA 171
Query: 72 SAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIP 131
V AG + K AP+YKY S+SNIL++ CQYEALK++SFP Q LAK +K+IP
Sbjct: 172 LTV-AGLYCSVTKQPRHGAPMYKYSFASLSNILSSWCQYEALKFISFPTQVLAKASKVIP 230
Query: 132 VMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGY 191
VM+ G ++ K Y+ +Y+ A L+++G S+F+L GE P T G++++ GY
Sbjct: 231 VMLMGKLVSHKSYEYWEYLTAVLISVGVSMFLL-SNGEGNRP---SGVTTFSGLVILAGY 286
Query: 192 LGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDC 251
+ D FTS +QD LF+ Y M +F + SC+ ++ L+ QG ++ AI F+ RH D
Sbjct: 287 IVFDSFTSNWQDSLFK-YKMSSVQMMFGVNMFSCLFTVGSLLEQGAMLEAIHFMTRHPDF 345
Query: 252 FFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIG 311
F LLS + Q FI YTI FGA F IMT RQ ++I+LSC+ + HP++ +G
Sbjct: 346 AFHACLLSVCSAFGQLFIFYTINKFGAAVFTIIMTLRQALAILLSCLLYGHPVTAVGGVG 405
Query: 312 AVIVFGSLYAKSFTR-KAPQKTTSSDSIPLVQ 342
VF +L+ + + R + +K + IP+VQ
Sbjct: 406 VATVFLALFLRVYARGRMRKKGRKAADIPVVQ 437
>UniRef100_Q6P101 Hypothetical protein zgc:77349 [Brachydanio rerio]
Length = 435
Score = 224 bits (571), Expect = 3e-57
Identities = 132/338 (39%), Positives = 196/338 (57%), Gaps = 11/338 (3%)
Query: 10 SSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK-----EYFKHSLFLV 64
+ S S + +K IF +G+ V+ +T+GVLQE++M YG+ + E F+ S FLV
Sbjct: 102 NESESSSSARQAFKLIFCAAGLQVSYLTWGVLQERVMTRSYGSSEAEGSGERFRDSQFLV 161
Query: 65 FCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLA 124
F NRI VS + K+ AP+YKY S+SNIL++ CQYEALK++SFP Q LA
Sbjct: 162 FMNRILALTVSGLWCVLFKQPRHG-APMYKYSFASLSNILSSWCQYEALKFISFPTQVLA 220
Query: 125 KCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWG 184
K +K+IPVM+ G I+ +K Y+ +Y+ A L++LG S+F+L + D P T G
Sbjct: 221 KASKVIPVMLMGKIVSRKSYEYWEYLTAVLISLGVSMFLL-SSSTDKHP---STVTTFSG 276
Query: 185 VLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEF 244
VL++ GY+ D FTS +QD LF+ Y M +F L SC+ ++ L+ QG ++ F
Sbjct: 277 VLILAGYIVFDSFTSNWQDNLFK-YKMSSVQMMFGVNLFSCLFTVGSLLEQGAFFNSLAF 335
Query: 245 VYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPL 304
+ RH + F LLS + Q FI +TI FGA F IMT RQ ++I+LSC + HP+
Sbjct: 336 MTRHSEFAFHAVLLSVCSAFGQLFIFFTIAQFGAAVFTIIMTLRQALAILLSCFLYGHPV 395
Query: 305 SWEQWIGAVIVFGSLYAKSFTRKAPQKTTSSDSIPLVQ 342
S +G +VF +L+ + + R +K+T + VQ
Sbjct: 396 SLTGGLGVGVVFLALFLRIYARSRVKKSTKRPAQTPVQ 433
>UniRef100_Q91ZN5 Embryonic seven-span transmembrane protein-like protein [Mus
musculus]
Length = 431
Score = 216 bits (550), Expect = 8e-55
Identities = 125/332 (37%), Positives = 190/332 (56%), Gaps = 13/332 (3%)
Query: 4 SPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKH 59
+P T ++ S P + KL +F SG+ V+ +T+G+LQE++M YGA E+F
Sbjct: 95 APRTETAESTPSWQVLKL---VFCASGLQVSYLTWGILQERVMTGSYGATATSPGEHFTD 151
Query: 60 SLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFP 119
S FLV NR+ V AG +K AP+Y+Y S+SN+L++ CQYEALK+VSFP
Sbjct: 152 SQFLVLMNRVLALVV-AGLYCVLRKQPRHGAPMYRYSFASLSNVLSSWCQYEALKFVSFP 210
Query: 120 VQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRE 179
Q LAK +K+IPVM+ G ++ ++ Y+ +Y+ A L+++G S+F+L E S
Sbjct: 211 TQVLAKASKVIPVMMMGKLVSRRSYEHWEYLTAGLISIGVSMFLLSSGPEPRS----SPA 266
Query: 180 NTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLI 239
T+ G++L+ GY+ D FTS +QD LF Y M +F L SC+ ++ L+ QG L+
Sbjct: 267 TTLSGLVLLAGYIAFDSFTSNWQDALF-AYKMSSVQMMFGVNLFSCLFTVGSLLEQGALL 325
Query: 240 PAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVW 299
F+ RH + LLS + Q FI YTI FGA F IMT RQ ++I+LSC+
Sbjct: 326 EGARFMGRHSEFALHALLLSICSAFGQLFIFYTIGQFGAAVFTIIMTLRQAIAILLSCLL 385
Query: 300 FSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQK 331
+ H ++ +G +VF +L + + R Q+
Sbjct: 386 YGHTVTVVGGLGVAVVFTALLLRVYARGRKQR 417
>UniRef100_UPI0000181102 UPI0000181102 UniRef100 entry
Length = 429
Score = 215 bits (548), Expect = 1e-54
Identities = 125/332 (37%), Positives = 190/332 (56%), Gaps = 13/332 (3%)
Query: 4 SPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKH 59
+P T ++ S P + KL +F +G+ V+ +T+GVLQE++M YGA E+F
Sbjct: 93 APRTETAESTPSWQVLKL---VFCAAGLQVSYLTWGVLQERVMTGSYGATATSPGEHFTD 149
Query: 60 SLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFP 119
S FLV NR+ V AG +K AP+Y+Y S+SN+L++ CQYEALK+VSFP
Sbjct: 150 SQFLVLMNRVLALVV-AGLYCILRKQPRHGAPMYRYSFASLSNVLSSWCQYEALKFVSFP 208
Query: 120 VQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRE 179
Q LAK +K+IPVM+ G ++ ++ Y+ +Y+ A L+++G S+F+L E S
Sbjct: 209 TQVLAKASKVIPVMMMGKLVSRRSYEHWEYLTAGLISIGVSMFLLSSGPEPRS----SPA 264
Query: 180 NTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLI 239
T+ G++L+ GY+ D FTS +QD LF Y M +F L SC+ ++ L+ QG L+
Sbjct: 265 TTLSGLVLLAGYIAFDSFTSNWQDALF-AYKMSSVQMMFGVNLFSCLFTVGSLLEQGALL 323
Query: 240 PAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVW 299
F+ RH + LLS + Q FI YTI FGA F IMT RQ ++I+LSC+
Sbjct: 324 EGARFMGRHSEFALHALLLSVCSAFGQLFIFYTIGQFGAAVFTIIMTLRQAIAILLSCLL 383
Query: 300 FSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQK 331
+ H ++ +G +VF +L + + R Q+
Sbjct: 384 YGHTVTVVGGLGVAVVFTALLLRVYARGRKQR 415
>UniRef100_UPI00003ACCF3 UPI00003ACCF3 UniRef100 entry
Length = 432
Score = 214 bits (544), Expect = 4e-54
Identities = 129/335 (38%), Positives = 192/335 (56%), Gaps = 13/335 (3%)
Query: 3 ESPSTSSSSSPPDSRD-NKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYF 57
E S + P +S ++ K +F +G+ V+ +T+GV+QE++M YGA + E F
Sbjct: 90 EDGSLPPRAEPTESSTARQVLKLLFCAAGLQVSYLTWGVVQERVMTRTYGATETDPGEKF 149
Query: 58 KHSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVS 117
K S FLVF NRI V AG A K A +YKY S+SNIL++ CQYEALKY+S
Sbjct: 150 KDSQFLVFMNRILAFTV-AGLYCALTKQPRHGAAMYKYSFASLSNILSSWCQYEALKYIS 208
Query: 118 FPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPA-GEDISPYSR 176
FP Q LAK +K+IPVM+ G ++ K Y+ +Y+ A L+++G S+F+L A +S +
Sbjct: 209 FPTQVLAKASKVIPVMMMGKLVSHKSYEYWEYLTAALISVGVSMFLLSSAPNRHVSTVT- 267
Query: 177 GRENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQG 236
T GV+L+ GY+ D FTS +QD LF Y M +F + SC+ ++ L+ QG
Sbjct: 268 ----TFSGVVLLAGYIVFDSFTSNWQDALFT-YKMSPVQMMFGVNVFSCLFTVGSLLEQG 322
Query: 237 HLIPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLS 296
L+ ++ F+ RH + LLS + Q FI YTI FGA F IMT RQ +I+LS
Sbjct: 323 ALLESVHFMARHSEFTAHAVLLSVCSACGQLFIFYTINQFGAAVFTIIMTLRQAFAILLS 382
Query: 297 CVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQK 331
C+ + H ++ +G +VF +L+ + + R +K
Sbjct: 383 CLIYGHTVTVVGGLGIAVVFMALFLRVYARSRMKK 417
>UniRef100_UPI000027CE36 UPI000027CE36 UniRef100 entry
Length = 429
Score = 211 bits (536), Expect = 3e-53
Identities = 123/335 (36%), Positives = 184/335 (54%), Gaps = 10/335 (2%)
Query: 1 MAESPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEY---- 56
+ + P+ S + S + K +F +G+ V+ +T+GVLQE++M Y A E
Sbjct: 89 LVDDPALSRNEGDSGSSVKQAVKLLFCAAGLQVSYLTWGVLQERVMTRSYAASPEAAGEK 148
Query: 57 FKHSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYV 116
F S FLVF NRI VS + + AP+YKY S+SNIL++ CQYEALKY+
Sbjct: 149 FTDSQFLVFMNRILALTVSGLWCVVFHQPRHG-APMYKYSFASLSNILSSWCQYEALKYI 207
Query: 117 SFPVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSR 176
SFP Q LAK +K+IPVM+ G II K Y+ +Y A L+++G S+F+L +
Sbjct: 208 SFPAQVLAKASKVIPVMLMGKIISHKSYEYWEYFTAALISVGVSMFLL----SSHNTKHL 263
Query: 177 GRENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQG 236
T G+++++GY+ D FTS +QD LF+ Y M +F + SC+ ++ L+ QG
Sbjct: 264 STATTFSGLIILSGYIVFDSFTSNWQDNLFK-YKMSSVQMMFGVNMFSCLFTVGSLLEQG 322
Query: 237 HLIPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLS 296
++ F+ RH + F LLS + Q FI YTI FGA F IMT RQ +I+LS
Sbjct: 323 AFFDSLAFMTRHSEFAFHAVLLSVCSACGQLFIFYTINQFGAAVFTIIMTLRQAFAILLS 382
Query: 297 CVWFSHPLSWEQWIGAVIVFGSLYAKSFTRKAPQK 331
C + H ++ G +VF +L+ + + R +K
Sbjct: 383 CFLYGHAITVVGGTGVAVVFLALFLRVYARSRMKK 417
>UniRef100_UPI0000362925 UPI0000362925 UniRef100 entry
Length = 421
Score = 210 bits (534), Expect = 5e-53
Identities = 124/311 (39%), Positives = 178/311 (56%), Gaps = 16/311 (5%)
Query: 23 KGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQKEY----FKHSLFLVFCNRITTSAVSAGS 78
K IF +G+ V+ +T+GVLQE++M Y A E F S FLVF NRI VS
Sbjct: 111 KLIFCAAGLQVSYLTWGVLQERVMTRSYAATPEEAGEKFTDSQFLVFMNRILALTVSGLW 170
Query: 79 LLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTI 138
L + AP+YKY S+SNIL++ CQYEALKY+SFP Q LAK +K+IPVM+ G I
Sbjct: 171 CLMFHQPRHG-APMYKYSFASLSNILSSWCQYEALKYISFPTQVLAKASKVIPVMLMGKI 229
Query: 139 IMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVW---GVLLMTGYLGCD 195
I K Y+ +Y A L+++G S+F+L S ++ +TV G++++ GY+ D
Sbjct: 230 ISHKSYEYWEYFTAVLISMGVSMFLL-------SSHASKHPSTVTTFSGLVILVGYIVFD 282
Query: 196 GFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDI 255
FTS +QD LF+ Y M +F + SC+ ++ L+ QG ++ F+ RH + F
Sbjct: 283 SFTSNWQDNLFK-YKMSSVQMMFGVNMFSCLFTVGSLLEQGAFFDSLAFMTRHSEFAFHA 341
Query: 256 ALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIV 315
LLS + Q FI YTI FGA F IMT RQ +I+LSC + H ++ IG +V
Sbjct: 342 VLLSVCSACGQLFIFYTINQFGAAVFTIIMTLRQAFAILLSCFLYGHAITAVGGIGVAVV 401
Query: 316 FGSLYAKSFTR 326
F +L+ + + R
Sbjct: 402 FLALFLRVYAR 412
>UniRef100_Q5T9W1 OTTHUMP00000016513 [Homo sapiens]
Length = 480
Score = 208 bits (530), Expect = 2e-52
Identities = 121/327 (37%), Positives = 186/327 (56%), Gaps = 13/327 (3%)
Query: 4 SPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKH 59
+P T ++ + P + KL +F +G+ V+ +T+GVLQE++M YGA E F
Sbjct: 143 APRTEAAETTPMWQALKL---LFCATGLQVSYLTWGVLQERVMTRSYGATATSPGERFTD 199
Query: 60 SLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFP 119
S FLV NR+ V+ S + K+ AP+Y+Y S+SN+L++ CQYEALK+VSFP
Sbjct: 200 SQFLVLMNRVLALIVAGLSCVLCKQPRHG-APMYRYSFASLSNVLSSWCQYEALKFVSFP 258
Query: 120 VQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRE 179
Q LAK +K+IPVM+ G ++ ++ Y+ +Y+ A L+++G S+F+L E S
Sbjct: 259 TQVLAKASKVIPVMLMGKLVSRRSYEHWEYLTATLISIGVSMFLLSSGPEPRS----SPA 314
Query: 180 NTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLI 239
T+ G++L+ GY+ D FTS +QD LF Y M +F SC+ ++ L+ QG L+
Sbjct: 315 TTLSGLILLAGYIAFDSFTSNWQDALF-AYKMSSVQMMFGVNFFSCLFTVGSLLEQGALL 373
Query: 240 PAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVW 299
F+ RH + LLS + Q FI YTI FGA F IMT RQ +I+LSC+
Sbjct: 374 EGTRFMGRHSEFAAHALLLSICSACGQLFIFYTIGQFGAAVFTIIMTLRQAFAILLSCLL 433
Query: 300 FSHPLSWEQWIGAVIVFGSLYAKSFTR 326
+ H ++ +G +VF +L + + R
Sbjct: 434 YGHTVTVVGGLGVAVVFAALLLRVYAR 460
>UniRef100_Q8TB61 Solute carrier family 35, member B2 (Adenosine 3'-phospho 5'-
phosphosulfate (PAPS) transporter) [Homo sapiens]
Length = 432
Score = 208 bits (530), Expect = 2e-52
Identities = 121/327 (37%), Positives = 186/327 (56%), Gaps = 13/327 (3%)
Query: 4 SPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKH 59
+P T ++ + P + KL +F +G+ V+ +T+GVLQE++M YGA E F
Sbjct: 95 APRTEAAETTPMWQALKL---LFCATGLQVSYLTWGVLQERVMTRSYGATATSPGERFTD 151
Query: 60 SLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFP 119
S FLV NR+ V+ S + K+ AP+Y+Y S+SN+L++ CQYEALK+VSFP
Sbjct: 152 SQFLVLMNRVLALIVAGLSCVLCKQPRHG-APMYRYSFASLSNVLSSWCQYEALKFVSFP 210
Query: 120 VQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRE 179
Q LAK +K+IPVM+ G ++ ++ Y+ +Y+ A L+++G S+F+L E S
Sbjct: 211 TQVLAKASKVIPVMLMGKLVSRRSYEHWEYLTATLISIGVSMFLLSSGPEPRS----SPA 266
Query: 180 NTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLI 239
T+ G++L+ GY+ D FTS +QD LF Y M +F SC+ ++ L+ QG L+
Sbjct: 267 TTLSGLILLAGYIAFDSFTSNWQDALF-AYKMSSVQMMFGVNFFSCLFTVGSLLEQGALL 325
Query: 240 PAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVW 299
F+ RH + LLS + Q FI YTI FGA F IMT RQ +I+LSC+
Sbjct: 326 EGTRFMGRHSEFAAHALLLSICSACGQLFIFYTIGQFGAAVFTIIMTLRQAFAILLSCLL 385
Query: 300 FSHPLSWEQWIGAVIVFGSLYAKSFTR 326
+ H ++ +G +VF +L + + R
Sbjct: 386 YGHTVTVVGGLGVAVVFAALLLRVYAR 412
>UniRef100_Q8NBK6 Hypothetical protein PSEC0149 [Homo sapiens]
Length = 432
Score = 207 bits (527), Expect = 4e-52
Identities = 121/327 (37%), Positives = 185/327 (56%), Gaps = 13/327 (3%)
Query: 4 SPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPYGAQK----EYFKH 59
+P T ++ + P + KL +F +G+ V+ +T+GVLQE++M YGA E F
Sbjct: 95 APRTEAAETTPMWQALKL---LFCATGLQVSYLTWGVLQERVMTRSYGATATSPGERFTD 151
Query: 60 SLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFP 119
S FLV NR+ V+ S + K+ AP+Y+Y S+SN+L++ CQYEALK+VSFP
Sbjct: 152 SQFLVLMNRVLALIVAGLSCVLCKQPRHG-APMYRYSFASLSNVLSSWCQYEALKFVSFP 210
Query: 120 VQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRE 179
Q LAK +K+IPVM+ G ++ ++ Y+ +Y+ A L+++G S+F+L E S
Sbjct: 211 TQVLAKASKVIPVMLMGKLVSRRSYEHWEYLTATLISIGVSMFLLSSGPEPRS----SPA 266
Query: 180 NTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLI 239
T+ G++L+ GY+ D FTS +QD LF Y M +F SC+ ++ L+ QG L+
Sbjct: 267 TTLSGLILLAGYIAFDSFTSNWQDALF-AYKMSSVQMMFGVNFFSCLFTVGSLLEQGALL 325
Query: 240 PAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVW 299
F+ RH LLS + Q FI YTI FGA F IMT RQ +I+LSC+
Sbjct: 326 EGTRFMGRHSGFAAHALLLSICSACGQLFIFYTIGQFGAAVFTIIMTLRQAFAILLSCLL 385
Query: 300 FSHPLSWEQWIGAVIVFGSLYAKSFTR 326
+ H ++ +G +VF +L + + R
Sbjct: 386 YGHTVTVVGGLGVAVVFAALLLRVYAR 412
>UniRef100_Q7QCD3 ENSANGP00000012787 [Anopheles gambiae str. PEST]
Length = 419
Score = 199 bits (505), Expect = 1e-49
Identities = 116/302 (38%), Positives = 173/302 (56%), Gaps = 8/302 (2%)
Query: 26 FAVSGIMVTLVTYGVLQEKIMRVPYGA---QKEYFKHSLFLVFCNRITTSAVSAGSLLAS 82
+ + G+M + +T+GVLQEKIM Y +K +FK S FLVF NR+ ++A L+A
Sbjct: 102 YCLVGLMGSYLTWGVLQEKIMTQEYEGPEKRKSHFKDSQFLVFSNRVLGFLITAVYLVA- 160
Query: 83 KKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSFPVQTLAKCAKMIPVMIWGTIIMQK 142
K+ AP+YKY S SNI++ QYEALK+V+FP Q LAK K+IPVMI G II +
Sbjct: 161 KRQFRHRAPLYKYSFASFSNIMSAWFQYEALKFVNFPTQVLAKSCKIIPVMIMGKIISRN 220
Query: 143 RYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGRENTVWGVLLMTGYLGCDGFTSTFQ 202
+Y+ +Y+ A ++++G F+ E T+ GVLL+T Y+ D FTS +Q
Sbjct: 221 KYEFYEYLTAVMISVGMIFFLTGSTDES----KASAMTTLTGVLLLTFYMIFDSFTSNWQ 276
Query: 203 DKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHLIPAIEFVYRHHDCFFDIALLSTVA 262
+LF+ Y M + L S + + L +QG ++ F H D +LS +
Sbjct: 277 GELFKSYSMSSIQMMCGVNLFSTLFTGASLAMQGGFYSSLVFAVDHPKFVVDCVVLSISS 336
Query: 263 TASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCVWFSHPLSWEQWIGAVIVFGSLYAK 322
Q FI YTI TFGA+ F IMT RQ V+I+LSC+ + H +S+ +G +IVF +++ +
Sbjct: 337 AIGQLFIFYTIATFGAVVFTIIMTLRQAVAILLSCLIYQHRISFLGVVGVLIVFLAIFLR 396
Query: 323 SF 324
+
Sbjct: 397 VY 398
>UniRef100_Q9VEI3 CG7623-PA [Drosophila melanogaster]
Length = 465
Score = 194 bits (493), Expect = 3e-48
Identities = 114/326 (34%), Positives = 179/326 (53%), Gaps = 7/326 (2%)
Query: 2 AESPSTSSSSSPPDSRDNKLWKGIFAVSGIMVTLVTYGVLQEKIMRVPY---GAQKEYFK 58
A+ ++S++P + + + ++ G+M++ +T+GVLQEKIM Y + FK
Sbjct: 123 ADKDRPAASTAPKRTSSQEAVQLLWCFGGLMISYLTWGVLQEKIMTQNYLNFTGESAKFK 182
Query: 59 HSLFLVFCNRITTSAVSAGSLLASKKALDPVAPIYKYCLVSVSNILTTTCQYEALKYVSF 118
S FLVF NR+ V+ L + AP+YKY S SNI++ QYEALK+V+F
Sbjct: 183 DSQFLVFSNRLLAFLVALAYLQWQPSPVRHRAPLYKYSYASFSNIMSAWFQYEALKFVNF 242
Query: 119 PVQTLAKCAKMIPVMIWGTIIMQKRYQGPDYMLAFLVTLGCSVFILYPAGEDISPYSRGR 178
P Q LAK K+IPVM+ G I+ + +Y+ +Y+ A L++LG I + +G S + G
Sbjct: 243 PTQVLAKSCKIIPVMLMGKIMSKAKYESYEYVTALLISLG---MIFFMSGSSDSSKASG- 298
Query: 179 ENTVWGVLLMTGYLGCDGFTSTFQDKLFRGYDMEIHNQIFYTTLCSCILSLTGLILQGHL 238
T+ G+ L++ Y+ D FT+ +Q LF+ Y M + L S I + L +QG
Sbjct: 299 VTTLTGIFLLSMYMVFDSFTANWQGSLFKSYGMTPLQMMCGVNLFSSIFTGASLSMQGGF 358
Query: 239 IPAIEFVYRHHDCFFDIALLSTVATASQFFISYTIRTFGALTFATIMTTRQLVSIMLSCV 298
+ ++ F H FD+ +LS + Q FI +TI FG + F IMT RQ V+IMLSC
Sbjct: 359 MDSLAFATEHPKFVFDMVVLSVCSAVGQLFIYHTIDVFGPVVFTIIMTLRQAVAIMLSCF 418
Query: 299 WFSHPLSWEQWIGAVIVFGSLYAKSF 324
+ H +S G +IVF +++ + +
Sbjct: 419 IYQHSISLLGIFGVLIVFVAIFLRVY 444
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.137 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 560,264,459
Number of Sequences: 2790947
Number of extensions: 21335766
Number of successful extensions: 61341
Number of sequences better than 10.0: 184
Number of HSP's better than 10.0 without gapping: 127
Number of HSP's successfully gapped in prelim test: 57
Number of HSP's that attempted gapping in prelim test: 60932
Number of HSP's gapped (non-prelim): 206
length of query: 353
length of database: 848,049,833
effective HSP length: 128
effective length of query: 225
effective length of database: 490,808,617
effective search space: 110431938825
effective search space used: 110431938825
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0159.10


