

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.8
(247 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
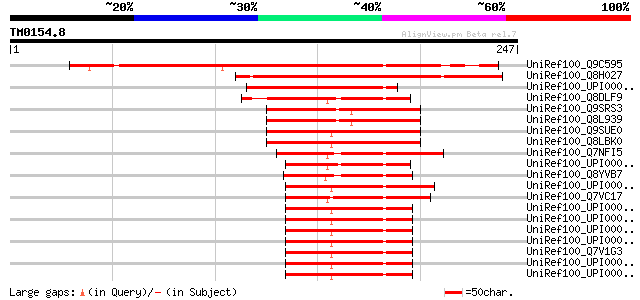
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C595 Hypothetical protein At5g21920 [Arabidopsis tha... 220 2e-56
UniRef100_Q8H027 Hypothetical protein OSJNBa0050H14.3 [Oryza sat... 181 2e-44
UniRef100_UPI000029D3BB UPI000029D3BB UniRef100 entry 103 3e-21
UniRef100_Q8DLF9 Ycf19 protein [Synechococcus elongatus] 80 5e-14
UniRef100_Q9SRS3 F21O3.14 protein [Arabidopsis thaliana] 80 5e-14
UniRef100_Q8L939 Hypothetical protein [Arabidopsis thaliana] 80 5e-14
UniRef100_Q9SUE0 Hypothetical protein T13J8.100 [Arabidopsis tha... 79 1e-13
UniRef100_Q8LBK0 Hypothetical protein [Arabidopsis thaliana] 79 1e-13
UniRef100_Q7NFI5 Ycf19 protein [Gloeobacter violaceus] 79 1e-13
UniRef100_UPI00002D1B67 UPI00002D1B67 UniRef100 entry 77 4e-13
UniRef100_Q8YVB7 Asl2061 protein [Anabaena sp.] 77 5e-13
UniRef100_UPI00002DD7D2 UPI00002DD7D2 UniRef100 entry 76 9e-13
UniRef100_Q7VC17 Uncharacterized YGGT family conserved membrane ... 76 9e-13
UniRef100_UPI000027E6C3 UPI000027E6C3 UniRef100 entry 74 3e-12
UniRef100_UPI000026ECE5 UPI000026ECE5 UniRef100 entry 73 8e-12
UniRef100_UPI000033D170 UPI000033D170 UniRef100 entry 72 1e-11
UniRef100_UPI00002FE8C7 UPI00002FE8C7 UniRef100 entry 72 1e-11
UniRef100_Q7V1G3 Conserved hypothetical membrane protein [Prochl... 72 1e-11
UniRef100_UPI000031B03B UPI000031B03B UniRef100 entry 72 1e-11
UniRef100_UPI0000306656 UPI0000306656 UniRef100 entry 72 1e-11
>UniRef100_Q9C595 Hypothetical protein At5g21920 [Arabidopsis thaliana]
Length = 251
Score = 220 bits (561), Expect = 2e-56
Identities = 123/215 (57%), Positives = 150/215 (69%), Gaps = 19/215 (8%)
Query: 30 TQTPFALP-----ILKPPISNLTSFLSPPELHKSFTTATDNCFRFLHSLASQNPFLNKIL 84
TQTP +L PP+++ + + H+S +AT+ FLHSLAS+NP + +
Sbjct: 33 TQTPLVRSNKPNLLLLPPVADSVKLIQ--DFHQSLISATEKFKGFLHSLASKNPLFQEAV 90
Query: 85 SLRSEFDSLCFQIRCSNY-RSRRWLSSHNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLV 143
L SEF LC +IR N R R +S+H FAAVLPGDSVAGLVV NG+ NFLNIYNT+LV
Sbjct: 91 RLSSEFHILCDEIRLRNTTRVRFAMSNHGFAAVLPGDSVAGLVVANGLINFLNIYNTILV 150
Query: 144 VRLVLTWFPNSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAFLVLNAFTSASA 203
VRLVLTWFP++PPAIV PLST+CDPYLN+FRG IPPLGG LDLSPILAFLVLNAFTS++
Sbjct: 151 VRLVLTWFPSAPPAIVNPLSTLCDPYLNIFRGFIPPLGG-LDLSPILAFLVLNAFTSSAM 209
Query: 204 ALPAELPVTQQSEQGVAATLQSTDITSSQKKWMKR 238
ALP ELP S G + SS+ KW++R
Sbjct: 210 ALPCELP----SADGAVSP------ASSETKWVRR 234
>UniRef100_Q8H027 Hypothetical protein OSJNBa0050H14.3 [Oryza sativa]
Length = 157
Score = 181 bits (458), Expect = 2e-44
Identities = 91/130 (70%), Positives = 110/130 (84%), Gaps = 2/130 (1%)
Query: 111 HNFAAVLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYL 170
H FAAVL GDSVAG+VV +GI NFL++YNT+LVVRLVLTWFPN+PPAIV PLSTICDPYL
Sbjct: 19 HCFAAVL-GDSVAGVVVSSGINNFLSLYNTVLVVRLVLTWFPNTPPAIVAPLSTICDPYL 77
Query: 171 NLFRGLIPPLGGTLDLSPILAFLVLNAFTSASAALPAELPVTQQSEQGVAATLQSTDITS 230
N+FRG+IPPLGGTLDLSPILAFLVLNA +S +AALPAELP AT S+ +T+
Sbjct: 78 NIFRGIIPPLGGTLDLSPILAFLVLNALSSTAAALPAELP-DPAPPTSRGATSSSSVLTA 136
Query: 231 SQKKWMKRLQ 240
+++KWM+R++
Sbjct: 137 NRRKWMRRIR 146
>UniRef100_UPI000029D3BB UPI000029D3BB UniRef100 entry
Length = 81
Score = 103 bits (258), Expect = 3e-21
Identities = 48/74 (64%), Positives = 61/74 (81%), Gaps = 1/74 (1%)
Query: 116 VLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTICDPYLNLFRG 175
++PGDS V+ +GI + LNIYNTL++ RL+LTWFPN P I+ PL+T+CDPYLNLFRG
Sbjct: 1 MIPGDSAVESVITSGIFSTLNIYNTLIIGRLILTWFPNPPRQIMYPLATLCDPYLNLFRG 60
Query: 176 LIPPLGGTLDLSPI 189
+IPPLGG +DLSPI
Sbjct: 61 IIPPLGG-IDLSPI 73
>UniRef100_Q8DLF9 Ycf19 protein [Synechococcus elongatus]
Length = 96
Score = 80.1 bits (196), Expect = 5e-14
Identities = 45/86 (52%), Positives = 60/86 (69%), Gaps = 14/86 (16%)
Query: 114 AAVLPGDSVAGLVVGNGIQNFLNIYNTLLVVRLVLTWFPN----SPPAIVGPLSTICDPY 169
AAVLP ++ GI NFL IY +L++R++L+WFPN +PP + LS + DPY
Sbjct: 4 AAVLP-------ILLMGIANFLQIYLIILLIRVLLSWFPNINWYNPPFSI--LSQLTDPY 54
Query: 170 LNLFRGLIPPLGGTLDLSPILAFLVL 195
LN+FRGLIPP+GG LD SPI+AF +L
Sbjct: 55 LNIFRGLIPPIGG-LDFSPIIAFFLL 79
>UniRef100_Q9SRS3 F21O3.14 protein [Arabidopsis thaliana]
Length = 232
Score = 80.1 bits (196), Expect = 5e-14
Identities = 43/78 (55%), Positives = 56/78 (71%), Gaps = 4/78 (5%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV GI+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 149 VVAVGIKKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPIFD 207
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 208 TLDVSPLLAFAVLGTLGS 225
>UniRef100_Q8L939 Hypothetical protein [Arabidopsis thaliana]
Length = 234
Score = 80.1 bits (196), Expect = 5e-14
Identities = 43/78 (55%), Positives = 56/78 (71%), Gaps = 4/78 (5%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSPPAIVGPLSTI---CDPYLNLFRGLIPPLGG 182
VV GI+ +L+IY+ +L+VR++L+WFPN P PLS I CDPYLNLFR +IPP+
Sbjct: 151 VVAVGIKKWLDIYSGVLMVRVLLSWFPNIPWERQ-PLSAIRDLCDPYLNLFRNIIPPIFD 209
Query: 183 TLDLSPILAFLVLNAFTS 200
TLD+SP+LAF VL S
Sbjct: 210 TLDVSPLLAFAVLGTLGS 227
>UniRef100_Q9SUE0 Hypothetical protein T13J8.100 [Arabidopsis thaliana]
Length = 218
Score = 79.0 bits (193), Expect = 1e-13
Identities = 38/77 (49%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGT 183
VV G+ +L+IY+ +L+VR++L+WFPN P + + +CDPYLNLFR +IPP+ T
Sbjct: 135 VVAAGLSKWLDIYSGVLMVRVLLSWFPNIPWDRQPLSAIRDLCDPYLNLFRNIIPPVFDT 194
Query: 184 LDLSPILAFLVLNAFTS 200
LD+SP+LAF VL S
Sbjct: 195 LDVSPLLAFAVLGTLGS 211
>UniRef100_Q8LBK0 Hypothetical protein [Arabidopsis thaliana]
Length = 218
Score = 79.0 bits (193), Expect = 1e-13
Identities = 38/77 (49%), Positives = 54/77 (69%), Gaps = 2/77 (2%)
Query: 126 VVGNGIQNFLNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGT 183
VV G+ +L+IY+ +L+VR++L+WFPN P + + +CDPYLNLFR +IPP+ T
Sbjct: 135 VVAAGLSKWLDIYSGVLMVRVLLSWFPNIPWDGQPLSAIRDLCDPYLNLFRNIIPPVFDT 194
Query: 184 LDLSPILAFLVLNAFTS 200
LD+SP+LAF VL S
Sbjct: 195 LDVSPLLAFAVLGTLGS 211
>UniRef100_Q7NFI5 Ycf19 protein [Gloeobacter violaceus]
Length = 95
Score = 78.6 bits (192), Expect = 1e-13
Identities = 46/86 (53%), Positives = 56/86 (64%), Gaps = 9/86 (10%)
Query: 131 IQNFLNIYNTLLVVRLVLTWFPN-----SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLD 185
+ NF+ IY LL VR++L+WFPN +P AI LS + DPYLNLFR +IPPLGG +D
Sbjct: 14 LSNFIQIYVVLLFVRVLLSWFPNIDWSSNPWAI---LSQLTDPYLNLFRSIIPPLGG-ID 69
Query: 186 LSPILAFLVLNAFTSASAALPAELPV 211
LSPILAFL L + A LPV
Sbjct: 70 LSPILAFLALQVVGGLLVSGLASLPV 95
>UniRef100_UPI00002D1B67 UPI00002D1B67 UniRef100 entry
Length = 100
Score = 77.0 bits (188), Expect = 4e-13
Identities = 40/64 (62%), Positives = 50/64 (77%), Gaps = 5/64 (7%)
Query: 135 LNIYNTLLVVRLVLTWFPN---SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILA 191
L IY+T+L++R++LTWFPN S P +V L I DPYLN FRG+IPPL G LDLSPILA
Sbjct: 17 LGIYSTILIIRVLLTWFPNLDMSNPILVN-LCAITDPYLNFFRGIIPPLAG-LDLSPILA 74
Query: 192 FLVL 195
F+V+
Sbjct: 75 FVVI 78
>UniRef100_Q8YVB7 Asl2061 protein [Anabaena sp.]
Length = 92
Score = 76.6 bits (187), Expect = 5e-13
Identities = 42/68 (61%), Positives = 50/68 (72%), Gaps = 9/68 (13%)
Query: 134 FLNIYNTLLVVRLVLTWFP-----NSPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSP 188
F+ IY+ LL++R++LTWFP N P A LS I DPYLNLFR +IPPLGG +D SP
Sbjct: 11 FVTIYSYLLIIRVLLTWFPQIDWYNQPFAA---LSQITDPYLNLFRSIIPPLGG-MDFSP 66
Query: 189 ILAFLVLN 196
ILAFLVLN
Sbjct: 67 ILAFLVLN 74
>UniRef100_UPI00002DD7D2 UPI00002DD7D2 UniRef100 entry
Length = 96
Score = 75.9 bits (185), Expect = 9e-13
Identities = 39/75 (52%), Positives = 54/75 (72%), Gaps = 3/75 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY +L+VR++L+WFPN ++ +S+I DPYLN+FRGLIPPLGG LDLS I+AF
Sbjct: 14 LSIYTVVLIVRVLLSWFPNLDWGNPVLSAVSSITDPYLNVFRGLIPPLGG-LDLSAIIAF 72
Query: 193 LVLNAFTSASAALPA 207
+ LN S ++ A
Sbjct: 73 IALNLLQSLLSSASA 87
>UniRef100_Q7VC17 Uncharacterized YGGT family conserved membrane protein
[Prochlorococcus marinus]
Length = 101
Score = 75.9 bits (185), Expect = 9e-13
Identities = 42/74 (56%), Positives = 55/74 (73%), Gaps = 5/74 (6%)
Query: 135 LNIYNTLLVVRLVLTWFPN---SPPAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILA 191
L IY+ +L++R++LTWFPN S P I+ +S I DPYLNLFRG+IP +GG LD+SPILA
Sbjct: 16 LFIYSYILILRVLLTWFPNLDWSNP-ILSNISAITDPYLNLFRGIIPAIGG-LDISPILA 73
Query: 192 FLVLNAFTSASAAL 205
F+VLN S + L
Sbjct: 74 FIVLNLAESVLSNL 87
>UniRef100_UPI000027E6C3 UPI000027E6C3 UniRef100 entry
Length = 95
Score = 74.3 bits (181), Expect = 3e-12
Identities = 37/64 (57%), Positives = 50/64 (77%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY +L+VR++L+WFPN ++ +S+I DPYLN+FRGLIPPLGG LDLS I+AF
Sbjct: 14 LSIYTVVLIVRVLLSWFPNLDWGNPVLSAVSSITDPYLNVFRGLIPPLGG-LDLSAIIAF 72
Query: 193 LVLN 196
+ LN
Sbjct: 73 VALN 76
>UniRef100_UPI000026ECE5 UPI000026ECE5 UniRef100 entry
Length = 92
Score = 72.8 bits (177), Expect = 8e-12
Identities = 34/64 (53%), Positives = 50/64 (78%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP ++ L++I DPYLN+FRG+IPPLGG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGVLSALTSITDPYLNIFRGIIPPLGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
>UniRef100_UPI000033D170 UPI000033D170 UniRef100 entry
Length = 92
Score = 72.4 bits (176), Expect = 1e-11
Identities = 34/64 (53%), Positives = 50/64 (78%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP I+ L++I DPYLN+FRG+IPP+GG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGILSALTSITDPYLNIFRGIIPPIGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
>UniRef100_UPI00002FE8C7 UPI00002FE8C7 UniRef100 entry
Length = 92
Score = 72.4 bits (176), Expect = 1e-11
Identities = 34/64 (53%), Positives = 50/64 (78%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP I+ L++I DPYLN+FRG+IPP+GG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGILSALASITDPYLNIFRGIIPPIGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
>UniRef100_Q7V1G3 Conserved hypothetical membrane protein [Prochlorococcus marinus
subsp. pastoris]
Length = 92
Score = 72.4 bits (176), Expect = 1e-11
Identities = 34/64 (53%), Positives = 50/64 (78%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP I+ L++I DPYLN+FRG+IPP+GG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGILSALTSITDPYLNIFRGIIPPIGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
>UniRef100_UPI000031B03B UPI000031B03B UniRef100 entry
Length = 92
Score = 72.0 bits (175), Expect = 1e-11
Identities = 33/64 (51%), Positives = 50/64 (77%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP ++ L++I DPYLN+FRG+IPP+GG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGVLSALTSITDPYLNIFRGIIPPIGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
>UniRef100_UPI0000306656 UPI0000306656 UniRef100 entry
Length = 92
Score = 72.0 bits (175), Expect = 1e-11
Identities = 33/64 (51%), Positives = 50/64 (77%), Gaps = 3/64 (4%)
Query: 135 LNIYNTLLVVRLVLTWFPNSP--PAIVGPLSTICDPYLNLFRGLIPPLGGTLDLSPILAF 192
L+IY+ +L++R++LTWFP ++ L++I DPYLN+FRG+IPP+GG D+S +LAF
Sbjct: 13 LSIYSFILIIRILLTWFPGIDWSNGVLSALTSITDPYLNIFRGIIPPIGG-FDISSLLAF 71
Query: 193 LVLN 196
L+LN
Sbjct: 72 LLLN 75
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 413,341,007
Number of Sequences: 2790947
Number of extensions: 16454287
Number of successful extensions: 40871
Number of sequences better than 10.0: 233
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 187
Number of HSP's that attempted gapping in prelim test: 40650
Number of HSP's gapped (non-prelim): 241
length of query: 247
length of database: 848,049,833
effective HSP length: 124
effective length of query: 123
effective length of database: 501,972,405
effective search space: 61742605815
effective search space used: 61742605815
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0154.8


