

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0154.6
(227 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
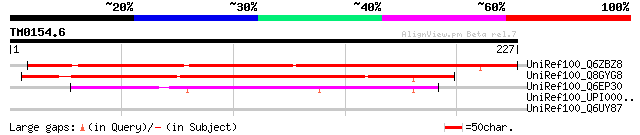
UniRef100_Q6ZBZ8 Hypothetical protein P0493A04.32 [Oryza sativa] 234 1e-60
UniRef100_Q8GYG8 Hypothetical protein At1g02180/T6A9_2 [Arabidop... 141 1e-32
UniRef100_Q6EP30 Ferredoxin-like [Oryza sativa] 124 2e-27
UniRef100_UPI00002D7802 UPI00002D7802 UniRef100 entry 35 1.9
UniRef100_Q6UY87 Bestrophin [Macaca fascicularis] 32 9.6
>UniRef100_Q6ZBZ8 Hypothetical protein P0493A04.32 [Oryza sativa]
Length = 226
Score = 234 bits (597), Expect = 1e-60
Identities = 119/220 (54%), Positives = 153/220 (69%), Gaps = 6/220 (2%)
Query: 9 ILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIK 68
+L+ + +A SAA+ K NNPAD+LVA IN NRTA K S L DN GL CIALQYIK
Sbjct: 12 LLAAVLAALLLSAASAADSK--NNPADQLVALINSNRTASKASTLDDNQGLGCIALQYIK 69
Query: 69 AYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSE 128
AY+G C VG ++KKPPE+ FAE FAPNCGV+A++L ITGR L CQ+ Y +AF+
Sbjct: 70 AYEGQCNQVG--ESKKPPETSFAETFAPNCGVQAATLTKITGRLLACQSNYATPDQAFN- 126
Query: 129 VLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFEGGVAKLTKP 188
L+ + +S+++LHSKNHT+VGAAV+GT GG PYFWCVLFS GKP ++F +GGV K +P
Sbjct: 127 FLVNDAKSIQVLHSKNHTEVGAAVSGTSGGGPYFWCVLFSSGKPTTSFKVDGGVPKSVRP 186
Query: 189 GCFSGANDECSGAHDWSPLSV-MWLFAASVLIALGFAFPL 227
GCFSG ND+C GA+ + W A++L + F L
Sbjct: 187 GCFSGNNDDCMGANAAVSIGAGTWRLVAALLFSAACVFAL 226
>UniRef100_Q8GYG8 Hypothetical protein At1g02180/T6A9_2 [Arabidopsis thaliana]
Length = 226
Score = 141 bits (356), Expect = 1e-32
Identities = 75/195 (38%), Positives = 119/195 (60%), Gaps = 8/195 (4%)
Query: 6 LVVILSFLHSASTGSAATNTQIKVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQ 65
L +IL FL +S +++ K+ N A ++V+ +N+NRTA K+ +L ++ GL C+ALQ
Sbjct: 8 LELILLFLSLSSVLASS-----KLHGNSAHEMVSILNQNRTARKLGKLNESPGLGCMALQ 62
Query: 66 YIKAYQGDCGAVGGPDAKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEA 125
Y++ +G+C V + + PE F +VFAPNCGV+ + ITG LGC +KY A
Sbjct: 63 YVELCEGNCN-VNNTLSCEHPEDDFTQVFAPNCGVELPTFGTITGHILGCSSKYAAPEVA 121
Query: 126 FSEVLIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFE-GGVAK 184
FS++L ++ +L +L +++HT+VG + G+ +FWC+LFS G NS+F E G
Sbjct: 122 FSDILFRDSSALSVLRNRSHTEVGVGMARLHKGT-FFWCLLFSDGVKNSSFVLEDNGRGI 180
Query: 185 LTKPGCFSGANDECS 199
+ GC+SG+ CS
Sbjct: 181 KQRTGCYSGSAFPCS 195
>UniRef100_Q6EP30 Ferredoxin-like [Oryza sativa]
Length = 230
Score = 124 bits (311), Expect = 2e-27
Identities = 74/175 (42%), Positives = 93/175 (52%), Gaps = 14/175 (8%)
Query: 28 KVTNNPADKLVAAINENRTAYKVSELYDNAGLACIALQYIKAYQGDCGAVG------GPD 81
K+ NPA+ LVA +N NRTA K+ L +AGL C+ALQYI DC +G
Sbjct: 24 KIHGNPANDLVALVNANRTATKLPHLRTSAGLGCMALQYI----SDCIGIGIGCAGDNTV 79
Query: 82 AKKPPESQFAEVFAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSL---E 138
A +PPE+ EV+A NCGV+ ++ ITGR LGC + A A VL + S
Sbjct: 80 ACQPPEAHITEVYAANCGVELPTVDVITGRLLGCHRQRSDAEAALEAVLSGSGNSTAARA 139
Query: 139 ILHSKNHTQVGAAVTGTDGGSPYFWCVLFSGGKPNSTFAFE-GGVAKLTKPGCFS 192
++ K HTQVGA P+FWC+LFS G NSTF E G GCFS
Sbjct: 140 VIRGKEHTQVGAGFDRAHRRGPFFWCLLFSSGSANSTFLLEAAGKGVHQSHGCFS 194
>UniRef100_UPI00002D7802 UPI00002D7802 UniRef100 entry
Length = 175
Score = 34.7 bits (78), Expect = 1.9
Identities = 23/82 (28%), Positives = 36/82 (43%), Gaps = 7/82 (8%)
Query: 130 LIQNQRSLEILHSKNHTQVGAAVTGTDGGSPYFWCV----LFSGGKPNSTFAFEG-GVAK 184
LI N LE+L+ +++ A G FW V ++ G + F G G+ +
Sbjct: 73 LISNTNILELLNIVRKSKLVVANDSAVGHISAFWGVPTLTIYGGPSNENIFKPSGEGITQ 132
Query: 185 LTKPGCF--SGANDECSGAHDW 204
+ +P C S D CS +DW
Sbjct: 133 IIRPNCICKSSEQDRCSNPNDW 154
>UniRef100_Q6UY87 Bestrophin [Macaca fascicularis]
Length = 585
Score = 32.3 bits (72), Expect = 9.6
Identities = 23/66 (34%), Positives = 29/66 (43%), Gaps = 3/66 (4%)
Query: 94 FAPNCGVKASSLAPITGRFLGCQTKYVHAPEAFSEVLIQNQRSLEILHS---KNHTQVGA 150
F PN K + I GRFLG Q+ H P A S + + +LH KNH V
Sbjct: 375 FQPNQEDKEDAHTGIIGRFLGLQSHDHHPPGANSRTKLLWPKRESLLHEGLPKNHKAVKQ 434
Query: 151 AVTGTD 156
V G +
Sbjct: 435 NVRGLE 440
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 391,388,229
Number of Sequences: 2790947
Number of extensions: 16391045
Number of successful extensions: 32729
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 32720
Number of HSP's gapped (non-prelim): 5
length of query: 227
length of database: 848,049,833
effective HSP length: 123
effective length of query: 104
effective length of database: 504,763,352
effective search space: 52495388608
effective search space used: 52495388608
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0154.6


