

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0153b.6
(156 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
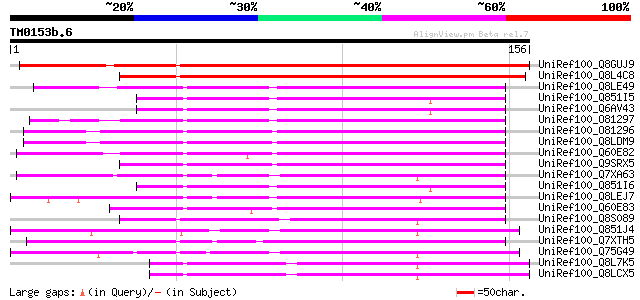
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GUJ9 Hypothetical protein At5g19590 [Arabidopsis tha... 196 2e-49
UniRef100_Q8L4C8 Hypothetical protein B1103C09.17 [Oryza sativa] 129 2e-29
UniRef100_Q8LE49 Hypothetical protein [Arabidopsis thaliana] 80 2e-14
UniRef100_Q851I5 Hypothetical protein OSJNBb0021O11.27 [Oryza sa... 78 6e-14
UniRef100_Q6AV43 Hypothetical protein OSJNBb0101N11.18 [Oryza sa... 78 6e-14
UniRef100_O81297 T14P8.17 protein [Arabidopsis thaliana] 77 1e-13
UniRef100_O81296 T14P8.18 protein [Arabidopsis thaliana] 76 3e-13
UniRef100_Q8LDM9 Hypothetical protein [Arabidopsis thaliana] 76 3e-13
UniRef100_Q60E82 Hypothetical protein P0692D12.13 [Oryza sativa] 75 4e-13
UniRef100_Q9SRX5 Hypothetical protein F22D16.19 [Arabidopsis tha... 75 5e-13
UniRef100_Q7XA63 At5g19860 [Arabidopsis thaliana] 73 2e-12
UniRef100_Q851I6 Hypothetical protein OSJNBb0021O11.26 [Oryza sa... 70 1e-11
UniRef100_Q8LEJ7 Hypothetical protein [Arabidopsis thaliana] 70 1e-11
UniRef100_Q60E83 Hypothetical protein P0692D12.12 [Oryza sativa] 68 6e-11
UniRef100_Q8S089 P0470A12.30 protein [Oryza sativa] 63 2e-09
UniRef100_Q851J4 Hypothetical protein OSJNBb0021O11.18 [Oryza sa... 62 3e-09
UniRef100_Q7XTH5 P0041A24.4 protein [Oryza sativa] 60 2e-08
UniRef100_Q75G49 Hypothetical protein B1003C08.8 [Oryza sativa] 57 1e-07
UniRef100_Q8L7K5 Hypothetical protein At3g07460 [Arabidopsis tha... 56 2e-07
UniRef100_Q8LCX5 Hypothetical protein [Arabidopsis thaliana] 55 6e-07
>UniRef100_Q8GUJ9 Hypothetical protein At5g19590 [Arabidopsis thaliana]
Length = 151
Score = 196 bits (498), Expect = 2e-49
Identities = 100/153 (65%), Positives = 120/153 (78%), Gaps = 3/153 (1%)
Query: 4 KILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTS 63
KI + L L S Q+P P+ K P+ AH ELTN+GFP+GLLP +V + +NQTS
Sbjct: 2 KISLFLLSILILLPISLQDPTPEVKK--PTRAHAELTNHGFPIGLLP-LSVKDYFLNQTS 58
Query: 64 GDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSS 123
GDFS+ L GACKITLPPDNY+ATYSN +TG+I +G+IAEL GIRVRAFF+ WSITGIRSS
Sbjct: 59 GDFSLFLNGACKITLPPDNYIATYSNKVTGRISQGKIAELQGIRVRAFFKSWSITGIRSS 118
Query: 124 GDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
GDN+VFEV +TAKYPSKNFD+S CEG+RSSS
Sbjct: 119 GDNLVFEVAGITAKYPSKNFDESLDCEGKRSSS 151
>UniRef100_Q8L4C8 Hypothetical protein B1103C09.17 [Oryza sativa]
Length = 160
Score = 129 bits (325), Expect = 2e-29
Identities = 61/122 (50%), Positives = 91/122 (74%), Gaps = 1/122 (0%)
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
+A+ EL + GFP+GLLP V G+ ++ SGDF+V L +C+I LP +Y+A++S+ +TG
Sbjct: 40 TAYDELRHRGFPLGLLP-ANVRGYTLDSGSGDFAVDLASSCRIVLPAGSYLASFSDRLTG 98
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPACEGQR 153
++ RI+ L+GIRVRAFF+WWSITGIR+ GD +VFEVG V+AK+P+++F+ S C +
Sbjct: 99 RLDDRRISGLSGIRVRAFFRWWSITGIRADGDELVFEVGSVSAKFPARHFNASLECPAKA 158
Query: 154 SS 155
S
Sbjct: 159 DS 160
>UniRef100_Q8LE49 Hypothetical protein [Arabidopsis thaliana]
Length = 149
Score = 80.1 bits (196), Expect = 2e-14
Identities = 48/142 (33%), Positives = 74/142 (51%), Gaps = 8/142 (5%)
Query: 8 LFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFS 67
+F +FL L S S K S + L NY P G+LP+ V + +N+ +G F
Sbjct: 3 IFIIFLLLLSTSVSVSGQKK-----RSVYQVLENYTLPRGILPEG-VHDYDLNRRTGVFK 56
Query: 68 VRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNI 127
VR C+ ++ D+Y Y I+G I +GR+ L G+ V+ F W +I+ + GD++
Sbjct: 57 VRFNTTCQFSI--DSYKVKYKPVISGIITRGRVIRLIGVSVKVLFFWINISEVSRDGDDV 114
Query: 128 VFEVGMVTAKYPSKNFDDSPAC 149
F VG + ++ SK F DSP C
Sbjct: 115 EFFVGAASEEFSSKYFVDSPKC 136
>UniRef100_Q851I5 Hypothetical protein OSJNBb0021O11.27 [Oryza sativa]
Length = 171
Score = 78.2 bits (191), Expect = 6e-14
Identities = 38/113 (33%), Positives = 66/113 (57%), Gaps = 5/113 (4%)
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L YG P GL+PD+ V ++ ++ +G+F + L G C + +++ Y ++ G++ G
Sbjct: 38 LPKYGLPRGLIPDS-VASYSFDEATGEFEIHLAGTCYVWF--GSHLVYYERSVRGRLSYG 94
Query: 99 RIAELNGIRVRAFFQWWSITGIRSSGD--NIVFEVGMVTAKYPSKNFDDSPAC 149
I++L+GI+ + F W S+TGI + D + F+VG V+ P+ FD PAC
Sbjct: 95 AISDLSGIQAKKLFLWVSVTGIVAHPDQGTVEFQVGFVSEALPASQFDAVPAC 147
>UniRef100_Q6AV43 Hypothetical protein OSJNBb0101N11.18 [Oryza sativa]
Length = 204
Score = 78.2 bits (191), Expect = 6e-14
Identities = 38/113 (33%), Positives = 66/113 (57%), Gaps = 5/113 (4%)
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L YG P GL+PD+ V ++ ++ +G+F + L G C + +++ Y ++ G++ G
Sbjct: 38 LPKYGLPRGLIPDS-VASYSFDEATGEFEIHLAGTCYVWF--GSHLVYYERSVRGRLSYG 94
Query: 99 RIAELNGIRVRAFFQWWSITGIRSSGD--NIVFEVGMVTAKYPSKNFDDSPAC 149
I++L+GI+ + F W S+TGI + D + F+VG V+ P+ FD PAC
Sbjct: 95 AISDLSGIQAKKLFLWVSVTGIVAHPDQGTVEFQVGFVSEALPASQFDAVPAC 147
>UniRef100_O81297 T14P8.17 protein [Arabidopsis thaliana]
Length = 154
Score = 77.4 bits (189), Expect = 1e-13
Identities = 45/143 (31%), Positives = 74/143 (51%), Gaps = 12/143 (8%)
Query: 7 ILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDF 66
I+F LF + SF KP +A+ + Y P G+LP V+ + +N +G+F
Sbjct: 10 IVFFLFFTV---SFAVSGQKP------TAYDAVKLYNLPPGILPKG-VVDYELNPKTGNF 59
Query: 67 SVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDN 126
V C+ T+ +Y Y +TI+G I G + L G+ V+ F W +I + G +
Sbjct: 60 KVYFNDTCEFTI--QSYQLKYKSTISGVISPGHVKNLKGVSVKVLFFWVNIAEVSLDGAD 117
Query: 127 IVFEVGMVTAKYPSKNFDDSPAC 149
+ F VG+ +A +P+ NF++SP C
Sbjct: 118 LDFSVGIASASFPAANFEESPQC 140
>UniRef100_O81296 T14P8.18 protein [Arabidopsis thaliana]
Length = 167
Score = 75.9 bits (185), Expect = 3e-13
Identities = 44/145 (30%), Positives = 77/145 (52%), Gaps = 6/145 (4%)
Query: 5 ILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
ILI LFL+ + + +S +A+ L +Y FPVG+LP V+ + ++ T+G
Sbjct: 6 ILIASCLFLSSLTAAVVTA----AESDTPTAYSLLQSYNFPVGILPKG-VVAYDLDTTTG 60
Query: 65 DFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSG 124
F +C L +Y Y +TI+G I + ++ +L G++V+ F W +I + +G
Sbjct: 61 KFHAYFNDSCSFNLV-GSYQLNYKSTISGYISENKLKKLTGVKVKVLFLWLNIVEVIRNG 119
Query: 125 DNIVFEVGMVTAKYPSKNFDDSPAC 149
D + F VG+ +A + + F +SP C
Sbjct: 120 DEMEFSVGITSANFAIQEFLESPQC 144
>UniRef100_Q8LDM9 Hypothetical protein [Arabidopsis thaliana]
Length = 167
Score = 75.9 bits (185), Expect = 3e-13
Identities = 44/145 (30%), Positives = 77/145 (52%), Gaps = 6/145 (4%)
Query: 5 ILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSG 64
ILI LFL+ + + +S +A+ L +Y FPVG+LP V+ + ++ T+G
Sbjct: 6 ILIASCLFLSSLTAAVVTA----AESDTPTAYSLLQSYNFPVGILPKG-VVAYDLDTTTG 60
Query: 65 DFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSG 124
F +C L +Y Y +TI+G I + ++ +L G++V+ F W +I + +G
Sbjct: 61 KFHAYFNDSCSFNLV-GSYQLNYKSTISGYISENKLKKLTGVKVKVLFLWLNIVEVIRNG 119
Query: 125 DNIVFEVGMVTAKYPSKNFDDSPAC 149
D + F VG+ +A + + F +SP C
Sbjct: 120 DEMEFSVGITSANFAIQEFLESPQC 144
>UniRef100_Q60E82 Hypothetical protein P0692D12.13 [Oryza sativa]
Length = 182
Score = 75.5 bits (184), Expect = 4e-13
Identities = 45/156 (28%), Positives = 75/156 (47%), Gaps = 16/156 (10%)
Query: 3 PKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQT 62
P++ +L L + + + + K P +A+ L +Y FPVG+LP V + + T
Sbjct: 2 PRLHLLLLAVLAVAAAAAEAAAEKKP-----TAYEVLESYDFPVGILPKG-VTSYTLEAT 55
Query: 63 SGDFSVRL---------GGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQ 113
+GDF+ L C+ + +Y Y ITG+I G + +L G+ V+ F
Sbjct: 56 TGDFTATLDTGDDDDSSSSTCEFAIE-GSYSLRYQRAITGRIATGHLTDLRGVAVKVLFF 114
Query: 114 WWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
W +I + GD + F VG+ +A + NF +SP C
Sbjct: 115 WLNIVEVTRRGDRLEFSVGIASADFTVDNFLESPQC 150
>UniRef100_Q9SRX5 Hypothetical protein F22D16.19 [Arabidopsis thaliana]
Length = 166
Score = 75.1 bits (183), Expect = 5e-13
Identities = 37/116 (31%), Positives = 66/116 (56%), Gaps = 2/116 (1%)
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
+A+ L +Y FPVG+LP V+ + +++++G F +C L +Y Y +TI+G
Sbjct: 31 TAYTLLQSYNFPVGILPKG-VVSYDLDKSTGQFHAYFNKSCSFALQ-GSYQLDYKSTISG 88
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I + +I +L G++V+ F W +I + +GD + F VG+ +A + F +SP C
Sbjct: 89 YISENKITKLTGVKVKVLFLWLNIVEVIRNGDELEFSVGITSANFEIDEFYESPQC 144
>UniRef100_Q7XA63 At5g19860 [Arabidopsis thaliana]
Length = 181
Score = 73.2 bits (178), Expect = 2e-12
Identities = 51/150 (34%), Positives = 74/150 (49%), Gaps = 9/150 (6%)
Query: 3 PKILILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQT 62
P + +F L L + + + N P S S+ + L YG P GLLPDT V F ++
Sbjct: 5 PFFISIFIFSLTLFTTTTHSLNEPDPDSI-STVYELLPKYGLPSGLLPDT-VTDFTLSD- 61
Query: 63 SGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIR- 121
G F V L +C+I +Y+ Y TI+G+I G I EL GI+V+ FF W + I+
Sbjct: 62 DGRFVVHLPNSCEIEF---DYLVHYDKTISGRIGYGSITELKGIQVKKFFIWLDVDEIKV 118
Query: 122 --SSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
D+I F+VG + K F +C
Sbjct: 119 DLPPSDSIYFKVGFINKKLDIDQFKTIHSC 148
>UniRef100_Q851I6 Hypothetical protein OSJNBb0021O11.26 [Oryza sativa]
Length = 174
Score = 70.5 bits (171), Expect = 1e-11
Identities = 38/113 (33%), Positives = 60/113 (52%), Gaps = 5/113 (4%)
Query: 39 LTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKG 98
L YG P GL+P+T + + + +GDF +RL C I +++A + + I G+I G
Sbjct: 39 LPEYGLPRGLIPET-IASYTFDNDTGDFEIRLTSTCYIWF--GSHLAYFEDAIRGRIAYG 95
Query: 99 RIAELNGIRVRAFFQWWSITGIRSSGD--NIVFEVGMVTAKYPSKNFDDSPAC 149
I L+GI+ + FF W SIT I + D + F G ++ P +F + P C
Sbjct: 96 TITGLSGIQAQKFFVWVSITTIVAHPDQGTVEFRAGFISEALPESDFAEVPVC 148
>UniRef100_Q8LEJ7 Hypothetical protein [Arabidopsis thaliana]
Length = 175
Score = 70.5 bits (171), Expect = 1e-11
Identities = 52/170 (30%), Positives = 78/170 (45%), Gaps = 25/170 (14%)
Query: 1 MNPKILILFA-LFLNLTSKS------------------FQNPNPKPPKSTPSSAHIELTN 41
M +L LF+ LFL+L S S F + N P H L
Sbjct: 1 MASSLLALFSCLFLSLLSLSSSLNLRRPIFSQSNDLDLFSSLNLDRPSLAADDIHDLLPR 60
Query: 42 YGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIA 101
YGFP GLLP+ V + ++ GDF+V L +C + + + Y I GK+ G +
Sbjct: 61 YGFPKGLLPNN-VKSYTISD-DGDFTVDLISSCYVKF--SDQLVFYGKNIAGKLSYGSVK 116
Query: 102 ELNGIRVRAFFQWWSITGIRS--SGDNIVFEVGMVTAKYPSKNFDDSPAC 149
++ GI+ + F W IT + S S +VF VG V+ P+ F++ P+C
Sbjct: 117 DVRGIQAKEAFLWLPITAMESDPSSATVVFSVGFVSKTLPASMFENVPSC 166
>UniRef100_Q60E83 Hypothetical protein P0692D12.12 [Oryza sativa]
Length = 173
Score = 68.2 bits (165), Expect = 6e-11
Identities = 36/121 (29%), Positives = 62/121 (50%), Gaps = 4/121 (3%)
Query: 31 TPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLG--GACKITLPPDNYVATYS 88
T +A+ L + FP G+LP V+ + ++ +GDF+ L C ++ +Y Y
Sbjct: 26 TKPTAYEALATFDFPPGILPKG-VVSYTLDDATGDFTATLNTTSTCAFSIQ-GSYSLRYQ 83
Query: 89 NTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGDNIVFEVGMVTAKYPSKNFDDSPA 148
++G+I R+ L G+ V+ F W +I + GD + F VG+ +A + NF +SP
Sbjct: 84 RRLSGRIAADRLTNLQGVSVKILFLWVNIVEVTRHGDELGFSVGIASADFGIDNFLESPQ 143
Query: 149 C 149
C
Sbjct: 144 C 144
>UniRef100_Q8S089 P0470A12.30 protein [Oryza sativa]
Length = 161
Score = 63.2 bits (152), Expect = 2e-09
Identities = 41/119 (34%), Positives = 58/119 (48%), Gaps = 7/119 (5%)
Query: 34 SAHIELTNYGFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITG 93
+AH L +G P GLLP + F ++ SG F LG +C Y+ T+ G
Sbjct: 22 AAHEVLRAHGLPRGLLP-AGIADFRHDEGSGRFEAALGESCTAQFEVG---LRYNATVAG 77
Query: 94 KIVKGRIAELNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPAC 149
I GRIA L+G+ + F W+ + GIR S I F+VG+V +P F+ P C
Sbjct: 78 VISYGRIASLSGVSAQDLFLWFPVRGIRVDVPSSGVIYFDVGVVFKHFPLAVFEAPPPC 136
>UniRef100_Q851J4 Hypothetical protein OSJNBb0021O11.18 [Oryza sativa]
Length = 172
Score = 62.4 bits (150), Expect = 3e-09
Identities = 42/161 (26%), Positives = 70/161 (43%), Gaps = 14/161 (8%)
Query: 1 MNPKILILFALFLNLTSKSFQNP---NPKPPKSTPSSAHIELTNYGFPVGLLP-DTTVLG 56
M P +L+L L L + + + P + PP ++ L YG P GL P T
Sbjct: 1 MPPPLLLLLLLLLGGAAAAEEPPAAPSSSPPPHKNATLSEILPRYGLPPGLFPASVTAFS 60
Query: 57 FAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWS 116
A N G +V LGG C Y+ + +TG + G + L+G++VR F W+
Sbjct: 61 LAAN---GSLAVDLGGPCYAHY---EYLTYFEPRVTGVLRYGSLTGLSGVKVRRFLVWFD 114
Query: 117 ITGIR----SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQR 153
+ ++ + ++G +T K P+ F+ CE +
Sbjct: 115 VVRVKVDLPPPPRYVYLDIGWITRKLPADEFESPHECEDSK 155
>UniRef100_Q7XTH5 P0041A24.4 protein [Oryza sativa]
Length = 162
Score = 59.7 bits (143), Expect = 2e-08
Identities = 37/144 (25%), Positives = 59/144 (40%), Gaps = 4/144 (2%)
Query: 6 LILFALFLNLTSKSFQNPNPKPPKSTPSSAHIELTNYGFPVGLLPDTTVLGFAVNQTSGD 65
L++ AL + S S P + + +G P LLP T + ++ G
Sbjct: 10 LLIAALVVPTASASASAAESAAGPYDPPTVPELMDRFGLPRALLP-VTARRYLLHD-DGS 67
Query: 66 FSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSITGIRSSGD 125
F + L G C Y Y ++G + GR L G+RVR F W +T + +G
Sbjct: 68 FQLFLDGGC--VAEAGGYRVGYGVKLSGAVAPGRATGLGGVRVRVLFAWVPVTAVEVAGG 125
Query: 126 NIVFEVGMVTAKYPSKNFDDSPAC 149
+ +G + +P+ F SP C
Sbjct: 126 EVTVSLGPIKKSFPAAGFKSSPRC 149
>UniRef100_Q75G49 Hypothetical protein B1003C08.8 [Oryza sativa]
Length = 171
Score = 57.0 bits (136), Expect = 1e-07
Identities = 38/160 (23%), Positives = 72/160 (44%), Gaps = 13/160 (8%)
Query: 1 MNPKILILFALFLNLTSKSFQNPNP---KPPKSTPSSAHIELTNYGFPVGLLPDTTVLGF 57
M P +L+LF L + + + P PP + + I L YG P G+ P T++ F
Sbjct: 1 MPPPLLLLFFLLAGVGATTTVEEPPAALSPPHKNATLSEI-LLRYGLPPGVFP-TSITAF 58
Query: 58 AVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAELNGIRVRAFFQWWSI 117
+ +G +V L G C Y+ + + G + G + +L+G++VR F W+ +
Sbjct: 59 TL-AANGSLAVDLQGPCYSHY---EYLTYFEARVVGLLRYGSLTDLSGVKVRRFLVWFDV 114
Query: 118 TGIR----SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQR 153
++ + ++G +T K P+ F+ C+ +
Sbjct: 115 IRVKVDLPPPPHYVYLDIGWITRKLPADEFESPHKCDDSK 154
>UniRef100_Q8L7K5 Hypothetical protein At3g07460 [Arabidopsis thaliana]
Length = 177
Score = 56.2 bits (134), Expect = 2e-07
Identities = 37/117 (31%), Positives = 56/117 (47%), Gaps = 7/117 (5%)
Query: 43 GFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAE 102
G P+GL P V GF VN +G FSV L +C+ + + Y ++G I +I +
Sbjct: 38 GLPLGLFPKG-VKGFTVNGETGRFSVYLNQSCQAKYETELH---YDEIVSGTIGYAQIRD 93
Query: 103 LNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
L+GI + F W + GIR S I F+VG++ +Y F+ C R +
Sbjct: 94 LSGISAQELFLWLQVKGIRVDVPSSGLIFFDVGVLRKQYSLSLFETPRDCVAVRGDA 150
>UniRef100_Q8LCX5 Hypothetical protein [Arabidopsis thaliana]
Length = 177
Score = 55.1 bits (131), Expect = 6e-07
Identities = 37/117 (31%), Positives = 55/117 (46%), Gaps = 7/117 (5%)
Query: 43 GFPVGLLPDTTVLGFAVNQTSGDFSVRLGGACKITLPPDNYVATYSNTITGKIVKGRIAE 102
G P+GL P V GF VN +G FSV L +C+ + + Y +G I +I +
Sbjct: 38 GLPLGLFPKG-VKGFTVNGETGRFSVYLNQSCQAKYETELH---YDEIFSGTIGYAQIRD 93
Query: 103 LNGIRVRAFFQWWSITGIR---SSGDNIVFEVGMVTAKYPSKNFDDSPACEGQRSSS 156
L+GI + F W + GIR S I F+VG++ +Y F+ C R +
Sbjct: 94 LSGISAQELFLWLQVKGIRVDVPSSGLIFFDVGVLRKQYSLSLFETPRDCVAVRGDA 150
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 277,799,269
Number of Sequences: 2790947
Number of extensions: 11188459
Number of successful extensions: 25368
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 41
Number of HSP's that attempted gapping in prelim test: 25310
Number of HSP's gapped (non-prelim): 60
length of query: 156
length of database: 848,049,833
effective HSP length: 116
effective length of query: 40
effective length of database: 524,299,981
effective search space: 20971999240
effective search space used: 20971999240
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0153b.6


