

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0142.11
(270 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
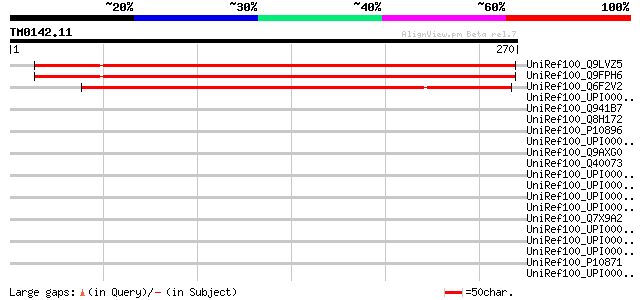
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LVZ5 Arabidopsis thaliana genomic DNA, chromosome 3,... 370 e-101
UniRef100_Q9FPH6 AT3g15840 [Arabidopsis thaliana] 370 e-101
UniRef100_Q6F2V2 Expressed protein [Oryza sativa] 318 7e-86
UniRef100_UPI000029057B UPI000029057B UniRef100 entry 43 0.010
UniRef100_Q941B7 At2g39730/T5I7.3 [Arabidopsis thaliana] 40 0.049
UniRef100_Q8H172 At2g39730/T5I7.3 [Arabidopsis thaliana] 40 0.049
UniRef100_P10896 Ribulose bisphosphate carboxylase/oxygenase act... 40 0.049
UniRef100_UPI00002B7355 UPI00002B7355 UniRef100 entry 40 0.064
UniRef100_Q9AXG0 Ribulose-1,5-bisphosphate carboxylase/oxygenase... 40 0.064
UniRef100_Q40073 Ribulose bisphosphate carboxylase/oxygenase act... 40 0.064
UniRef100_UPI0000325A5B UPI0000325A5B UniRef100 entry 40 0.083
UniRef100_UPI00002BD7F4 UPI00002BD7F4 UniRef100 entry 40 0.083
UniRef100_UPI0000295B6E UPI0000295B6E UniRef100 entry 40 0.083
UniRef100_UPI000028A292 UPI000028A292 UniRef100 entry 39 0.11
UniRef100_Q7X9A2 Rubisco activase alpha form precursor [Deschamp... 39 0.11
UniRef100_UPI000030BF31 UPI000030BF31 UniRef100 entry 39 0.19
UniRef100_UPI000029647E UPI000029647E UniRef100 entry 39 0.19
UniRef100_UPI00002688A8 UPI00002688A8 UniRef100 entry 38 0.24
UniRef100_P10871 Ribulose bisphosphate carboxylase/oxygenase act... 38 0.24
UniRef100_UPI00003254BB UPI00003254BB UniRef100 entry 37 0.41
>UniRef100_Q9LVZ5 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone: MSJ11
[Arabidopsis thaliana]
Length = 268
Score = 370 bits (951), Expect = e-101
Identities = 174/256 (67%), Positives = 208/256 (80%), Gaps = 1/256 (0%)
Query: 14 PSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFEL 73
PS P++S+ +FL S L + K+ + A VA P +EI+EYK+PSWA FE+
Sbjct: 14 PSQPLSSKRSFLYSSRIGPILRRFPRKKLDLQVKA-VATTLAPLEEIKEYKLPSWAMFEM 72
Query: 74 GKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAML 133
G A VYWKTMNGLPPTSGEKLKLFYNP +++L NE++G+AFNGGFNQPIMCGGEPRAML
Sbjct: 73 GTAPVYWKTMNGLPPTSGEKLKLFYNPAASKLTLNEDYGVAFNGGFNQPIMCGGEPRAML 132
Query: 134 RKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEG 193
+KDRGKAD+PIY++QIC+PKHA+NLIFSFTNGVDWDGPYRLQFQVPK QNKPIEFFNEG
Sbjct: 133 KKDRGKADSPIYTMQICIPKHAVNLIFSFTNGVDWDGPYRLQFQVPKRWQNKPIEFFNEG 192
Query: 194 LAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLA 253
LA EL ++GACE+AIFPDSN V T+C M+ NLTVEGGDRC+L+LV GC D +S +NP A
Sbjct: 193 LANELSQDGACERAIFPDSNVVPTRCTMIANLTVEGGDRCNLDLVPGCMDTNSEHFNPYA 252
Query: 254 NVDDGTCPLDLDSDSE 269
NVDDG+CPL+L E
Sbjct: 253 NVDDGSCPLELSDSDE 268
>UniRef100_Q9FPH6 AT3g15840 [Arabidopsis thaliana]
Length = 268
Score = 370 bits (950), Expect = e-101
Identities = 174/256 (67%), Positives = 208/256 (80%), Gaps = 1/256 (0%)
Query: 14 PSNPITSRHTFLNVSLPRCYLVKERNVKVTRTINAAVAVATTPAQEIEEYKIPSWANFEL 73
PS P++S+ +FL S L + K+ + A VA P +EI+EYK+PSWA FE+
Sbjct: 14 PSQPLSSKRSFLYSSRIGPILRRFPRKKLDLQVKA-VATTLAPLEEIKEYKLPSWAMFEM 72
Query: 74 GKASVYWKTMNGLPPTSGEKLKLFYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAML 133
G A VYWKTMNGLPPTSGEKLKLFYNP +++L NE++G+AFNGGFNQPIMCGGEPRAML
Sbjct: 73 GTAPVYWKTMNGLPPTSGEKLKLFYNPAASKLTLNEDYGVAFNGGFNQPIMCGGEPRAML 132
Query: 134 RKDRGKADAPIYSIQICVPKHALNLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEG 193
+KDRGKAD+PIY++QIC+PKHA+NLIFSFTNGVDWDGPYRLQFQVPK QNKPIEFFNEG
Sbjct: 133 KKDRGKADSPIYTMQICIPKHAVNLIFSFTNGVDWDGPYRLQFQVPKRRQNKPIEFFNEG 192
Query: 194 LAEELGKEGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLA 253
LA EL ++GACE+AIFPDSN V T+C M+ NLTVEGGDRC+L+LV GC D +S +NP A
Sbjct: 193 LANELSQDGACERAIFPDSNVVPTRCKMIANLTVEGGDRCNLDLVPGCMDTNSEHFNPYA 252
Query: 254 NVDDGTCPLDLDSDSE 269
NVDDG+CPL+L E
Sbjct: 253 NVDDGSCPLELSDSDE 268
>UniRef100_Q6F2V2 Expressed protein [Oryza sativa]
Length = 271
Score = 318 bits (816), Expect = 7e-86
Identities = 149/231 (64%), Positives = 184/231 (79%), Gaps = 3/231 (1%)
Query: 39 NVKVTRT-INAAVAVATTPA-QEIEEYKIPSWANFELGKASVYWKTMNGLPPTSGEKLKL 96
+ +V RT AA A AT PA + E +P+WA FELGKA VYWKTMNGLPP++GE L L
Sbjct: 41 SARVVRTGAAAAPAAATAPAVPQTNECSLPTWAEFELGKAPVYWKTMNGLPPSAGEGLIL 100
Query: 97 FYNPTSTQLAPNEEFGIAFNGGFNQPIMCGGEPRAMLRKDRGKADAPIYSIQICVPKHAL 156
FYNP +T++ PN +FGIAFNGGFNQPIMCGGEPR M ++RG AD PIY+I+I VP+HA+
Sbjct: 101 FYNPAATKMTPNAQFGIAFNGGFNQPIMCGGEPRQMTLQERGSADPPIYTIRIRVPQHAM 160
Query: 157 NLIFSFTNGVDWDGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGACEQAIFPDSNKVI 216
L+FSFTNGVDWDGPY L+F+VPK NKP+ FFNEGLA+EL +EGAC++AIFPD N VI
Sbjct: 161 TLVFSFTNGVDWDGPYTLKFRVPKPWLNKPLSFFNEGLADELNREGACDRAIFPDENVVI 220
Query: 217 TKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDLDSD 267
T C M G+ EGGDRC L++V GC DP+SH+++PLA VDDG+CP+D DS+
Sbjct: 221 TSCEM-GSYYEEGGDRCKLDIVSGCMDPNSHMFDPLATVDDGSCPMDSDSE 270
>UniRef100_UPI000029057B UPI000029057B UniRef100 entry
Length = 321
Score = 42.7 bits (99), Expect = 0.010
Identities = 18/37 (48%), Positives = 23/37 (61%)
Query: 224 NLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
N D + ++ GCTD +S LYNPLAN DDG+C
Sbjct: 131 NANANTPDGSCIPVINGCTDSTSPLYNPLANTDDGSC 167
Score = 39.3 bits (90), Expect = 0.11
Identities = 22/60 (36%), Positives = 32/60 (52%), Gaps = 4/60 (6%)
Query: 201 EGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+G+C I +N + L N+ D + ++ GCTD S+ YNPLANVD+ TC
Sbjct: 8 DGSCIATINGCTNPIAFNYDSLANVN----DGSCIAVIGGCTDLSAINYNPLANVDNNTC 63
>UniRef100_Q941B7 At2g39730/T5I7.3 [Arabidopsis thaliana]
Length = 474
Score = 40.4 bits (93), Expect = 0.049
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>UniRef100_Q8H172 At2g39730/T5I7.3 [Arabidopsis thaliana]
Length = 474
Score = 40.4 bits (93), Expect = 0.049
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>UniRef100_P10896 Ribulose bisphosphate carboxylase/oxygenase activase, chloroplast
precursor [Arabidopsis thaliana]
Length = 474
Score = 40.4 bits (93), Expect = 0.049
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 9/91 (9%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEELG------KEGACEQAIFPDSNKVITKCAMLGNLTVEG 229
F+ P++ K +E+ N + E+ E QA D+N G +G
Sbjct: 383 FEQPEMTYEKLMEYGNMLVMEQENVKRVQLAETYLSQAALGDAN---ADAIGRGTFYGKG 439
Query: 230 GDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ +L + EGCTDP + ++P A DDGTC
Sbjct: 440 AQQVNLPVPEGCTDPVAENFDPTARSDDGTC 470
>UniRef100_UPI00002B7355 UPI00002B7355 UniRef100 entry
Length = 136
Score = 40.0 bits (92), Expect = 0.064
Identities = 16/28 (57%), Positives = 20/28 (71%)
Query: 233 CDLNLVEGCTDPSSHLYNPLANVDDGTC 260
C +V GCTDP + ++PLA VDDGTC
Sbjct: 36 CLYQMVPGCTDPQAINFDPLATVDDGTC 63
Score = 37.0 bits (84), Expect = 0.54
Identities = 18/36 (50%), Positives = 24/36 (66%), Gaps = 1/36 (2%)
Query: 226 TVEGGDRCDLNL-VEGCTDPSSHLYNPLANVDDGTC 260
TV+ G +N V+GCTDP + Y+PLA DDG+C
Sbjct: 57 TVDDGTCIYINQPVQGCTDPLATNYDPLATQDDGSC 92
Score = 36.2 bits (82), Expect = 0.92
Identities = 16/36 (44%), Positives = 21/36 (57%)
Query: 225 LTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
L + C V GCTDP ++ Y+PLA DDG+C
Sbjct: 84 LATQDDGSCVYPPVLGCTDPLANNYDPLATQDDGSC 119
Score = 34.7 bits (78), Expect = 2.7
Identities = 18/37 (48%), Positives = 24/37 (64%), Gaps = 2/37 (5%)
Query: 224 NLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
N TV+ G C + ++ GCTDPS+ Y AN DDG+C
Sbjct: 2 NATVDDGS-C-IPIIYGCTDPSAINYFAGANTDDGSC 36
>UniRef100_Q9AXG0 Ribulose-1,5-bisphosphate carboxylase/oxygenase activase 2
[Gossypium hirsutum]
Length = 435
Score = 40.0 bits (92), Expect = 0.064
Identities = 27/92 (29%), Positives = 44/92 (47%), Gaps = 11/92 (11%)
Query: 176 FQVPKVLQNKPIEFFNEGLAEE-------LGKEGACEQAIFPDSNKVITKCAMLGNLTVE 228
F+ PK+ K +E+ N +AE+ L + E A+ + I + G +
Sbjct: 344 FEQPKMTIEKLLEYGNMLVAEQENVKRVQLADKYLSEAALGEANEDSINRGTFYGKAAQQ 403
Query: 229 GGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G + + EGCTDP++ ++P A DDGTC
Sbjct: 404 VG----VPVPEGCTDPNADNFDPTARSDDGTC 431
>UniRef100_Q40073 Ribulose bisphosphate carboxylase/oxygenase activase A, chloroplast
precursor [Hordeum vulgare]
Length = 464
Score = 40.0 bits (92), Expect = 0.064
Identities = 29/98 (29%), Positives = 48/98 (48%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+N+ K
Sbjct: 368 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANQDAMKT--- 422
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+ +G + L + EGCTD ++ Y+P A DDG+C
Sbjct: 423 GSFYGKGAQQGTLPVPEGCTDQNAKNYDPTARSDDGSC 460
>UniRef100_UPI0000325A5B UPI0000325A5B UniRef100 entry
Length = 265
Score = 39.7 bits (91), Expect = 0.083
Identities = 18/43 (41%), Positives = 28/43 (64%), Gaps = 1/43 (2%)
Query: 221 MLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLD 263
ML + E D C + ++EGCTD +++ +NP A +D+GTC D
Sbjct: 223 MLSAVCWESCDACPV-VIEGCTDSTANNFNPEATLDNGTCEYD 264
>UniRef100_UPI00002BD7F4 UPI00002BD7F4 UniRef100 entry
Length = 236
Score = 39.7 bits (91), Expect = 0.083
Identities = 17/30 (56%), Positives = 22/30 (72%), Gaps = 1/30 (3%)
Query: 231 DRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
D C + V GCTDP++ YNPLAN D+G+C
Sbjct: 3 DTC-IAYVYGCTDPTAFNYNPLANTDNGSC 31
Score = 35.8 bits (81), Expect = 1.2
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 4/59 (6%)
Query: 202 GACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+CE +F ++ + L N + C + + GCTDPS YNP AN +D +C
Sbjct: 29 GSCEAFVFGCTDSTMFNYNPLANAD---NNTC-VPFIYGCTDPSMLNYNPQANTEDFSC 83
>UniRef100_UPI0000295B6E UPI0000295B6E UniRef100 entry
Length = 293
Score = 39.7 bits (91), Expect = 0.083
Identities = 21/55 (38%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Query: 208 IFPDSNKVIT--KCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+ PD N +T C + +G + CD ++GCTD + YNP A DDGTC
Sbjct: 195 LHPDLNIPLTYRNCDGVCYYDEDGDNVCDEEEIQGCTDETKCNYNPDATDDDGTC 249
>UniRef100_UPI000028A292 UPI000028A292 UniRef100 entry
Length = 310
Score = 39.3 bits (90), Expect = 0.11
Identities = 22/57 (38%), Positives = 28/57 (48%), Gaps = 4/57 (7%)
Query: 204 CEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
C IF +N L N D + +V GCT+P + YNP ANVDDG+C
Sbjct: 166 CIPVIFGCTNPTALNYDSLANTD----DGSCIAMVFGCTNPQAFNYNPQANVDDGSC 218
Score = 38.9 bits (89), Expect = 0.14
Identities = 22/60 (36%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Query: 201 EGACEQAIFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
+G+C +F +N A N D + +V GCTDP+ YNP ANVD+G+C
Sbjct: 189 DGSCIAMVFGCTNPQ----AFNYNPQANVDDGSCIAIVYGCTDPTMFNYNPNANVDNGSC 244
Score = 33.9 bits (76), Expect = 4.6
Identities = 15/30 (50%), Positives = 18/30 (60%)
Query: 231 DRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
D C + GCTD S Y+ LANVD+G C
Sbjct: 112 DSCGVYATFGCTDSLSFNYDSLANVDNGGC 141
Score = 33.1 bits (74), Expect = 7.8
Identities = 12/23 (52%), Positives = 17/23 (73%)
Query: 238 VEGCTDPSSHLYNPLANVDDGTC 260
+ GCTDPS+ Y+P AN +D +C
Sbjct: 276 IYGCTDPSAFNYDPTANTEDFSC 298
>UniRef100_Q7X9A2 Rubisco activase alpha form precursor [Deschampsia antarctica]
Length = 465
Score = 39.3 bits (90), Expect = 0.11
Identities = 29/98 (29%), Positives = 47/98 (47%), Gaps = 11/98 (11%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNKVITKCAML 222
DGP + F+ PK+ K +E+ + + E+ + QA D+NK K
Sbjct: 369 DGP--VTFEQPKMTVEKLLEYGHMLVQEQDNVKRVQLADTYMSQAALGDANKDAMKT--- 423
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTC 260
G+ +G + L + EGCTD + ++P A DDG+C
Sbjct: 424 GSFYGKGAQQGTLPVPEGCTDRDAKNFDPTARSDDGSC 461
>UniRef100_UPI000030BF31 UPI000030BF31 UniRef100 entry
Length = 306
Score = 38.5 bits (88), Expect = 0.19
Identities = 17/29 (58%), Positives = 21/29 (71%)
Query: 238 VEGCTDPSSHLYNPLANVDDGTCPLDLDS 266
V GCTDPS++ Y+ A VDDG+C D DS
Sbjct: 264 VFGCTDPSAYNYDETATVDDGSCYYDGDS 292
>UniRef100_UPI000029647E UPI000029647E UniRef100 entry
Length = 292
Score = 38.5 bits (88), Expect = 0.19
Identities = 19/48 (39%), Positives = 31/48 (64%), Gaps = 2/48 (4%)
Query: 223 GNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDLDSDSEA 270
G V+ G C+ ++V+GCTDP ++ Y+ LAN D+GTC ++ +A
Sbjct: 212 GEAIVDDGS-CE-DVVQGCTDPDAYNYDNLANNDNGTCEAIVEGCKDA 257
Score = 37.4 bits (85), Expect = 0.41
Identities = 14/23 (60%), Positives = 18/23 (77%)
Query: 238 VEGCTDPSSHLYNPLANVDDGTC 260
+ GCT+PS+ YNP AN+ DGTC
Sbjct: 65 LSGCTNPSAFNYNPEANIPDGTC 87
Score = 35.4 bits (80), Expect = 1.6
Identities = 13/24 (54%), Positives = 17/24 (70%)
Query: 237 LVEGCTDPSSHLYNPLANVDDGTC 260
++ GCTD +S YN AN DDG+C
Sbjct: 90 VINGCTDEASPSYNSFANTDDGSC 113
Score = 35.4 bits (80), Expect = 1.6
Identities = 22/69 (31%), Positives = 32/69 (45%), Gaps = 12/69 (17%)
Query: 196 EELGKEGACEQAIF----PDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNP 251
E + +G+CE + PD+ NL C+ +VEGC D ++ YN
Sbjct: 213 EAIVDDGSCEDVVQGCTDPDAYNY-------DNLANNDNGTCEA-IVEGCKDANAFNYNS 264
Query: 252 LANVDDGTC 260
AN DDG+C
Sbjct: 265 SANTDDGSC 273
Score = 33.9 bits (76), Expect = 4.6
Identities = 11/24 (45%), Positives = 19/24 (78%)
Query: 237 LVEGCTDPSSHLYNPLANVDDGTC 260
++ GCTD ++ YNP++N D+G+C
Sbjct: 38 VIYGCTDELANNYNPISNTDNGSC 61
>UniRef100_UPI00002688A8 UPI00002688A8 UniRef100 entry
Length = 288
Score = 38.1 bits (87), Expect = 0.24
Identities = 16/25 (64%), Positives = 19/25 (76%)
Query: 236 NLVEGCTDPSSHLYNPLANVDDGTC 260
+ V GCTDP+S YN ANVDDG+C
Sbjct: 36 DFVYGCTDPNSSNYNADANVDDGSC 60
>UniRef100_P10871 Ribulose bisphosphate carboxylase/oxygenase activase, chloroplast
precursor [Spinacia oleracea]
Length = 472
Score = 38.1 bits (87), Expect = 0.24
Identities = 29/103 (28%), Positives = 47/103 (45%), Gaps = 13/103 (12%)
Query: 169 DGPYRLQFQVPKVLQNKPIEFFNEGLAEELGKEGA------CEQAIFPDSNK-VITKCAM 221
DGP F+ P++ K +E+ N + E+ + A D+NK I +
Sbjct: 376 DGPP--VFEQPEMTLQKLMEYGNMLVQEQENVKRVQLADQYMSSAALGDANKDAIDRGTF 433
Query: 222 LGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLDL 264
G + + L + +GCTDP + Y+P A DDG+C +L
Sbjct: 434 FG----KAAQQVSLPVAQGCTDPEAKNYDPTARSDDGSCTYNL 472
>UniRef100_UPI00003254BB UPI00003254BB UniRef100 entry
Length = 150
Score = 37.4 bits (85), Expect = 0.41
Identities = 22/56 (39%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Query: 208 IFPDSNKVITKCAMLGNLTVEGGDRCDLNLVEGCTDPSSHLYNPLANVDDGTCPLD 263
+F D + V T G + G + +LV GCTDP + YNP A DDG+C D
Sbjct: 85 VFIDGSLVDT----FGEDLMVGSTFINESLVSGCTDPMAFNYNPDALWDDGSCDYD 136
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 495,457,710
Number of Sequences: 2790947
Number of extensions: 21561982
Number of successful extensions: 36076
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 56
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 35930
Number of HSP's gapped (non-prelim): 154
length of query: 270
length of database: 848,049,833
effective HSP length: 125
effective length of query: 145
effective length of database: 499,181,458
effective search space: 72381311410
effective search space used: 72381311410
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0142.11


