

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0141.1
(705 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
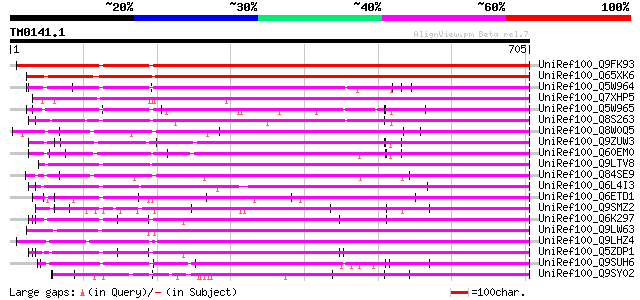
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FK93 Selenium-binding protein-like [Arabidopsis thal... 688 0.0
UniRef100_Q65XK6 Hypothetical protein OJ1735_C10.12 [Oryza sativa] 646 0.0
UniRef100_Q5W964 PPR986-12 [Physcomitrella patens] 489 e-136
UniRef100_Q7XHP5 Putative pentatricopeptide (PPR) repeat-contain... 456 e-127
UniRef100_Q5W965 PPR868-14 [Physcomitrella patens] 456 e-126
UniRef100_Q8S263 Putative pentatricopeptide (PPR) repeat-contain... 455 e-126
UniRef100_Q8W0Q5 Putative vegetative storage protein [Sorghum bi... 449 e-124
UniRef100_Q9ZUW3 Putative selenium-binding protein [Arabidopsis ... 444 e-123
UniRef100_Q60EM0 Unknow protein [Oryza sativa] 441 e-122
UniRef100_Q9LTV8 Selenium-binding protein-like [Arabidopsis thal... 440 e-122
UniRef100_Q84SE9 P0020E09.15 protein [Oryza sativa] 440 e-122
UniRef100_Q6L4I3 Hypothetical protein OSJNBa0074P11.10 [Oryza sa... 440 e-122
UniRef100_Q6ETD1 Putative pentatricopeptide (PPR) repeat-contain... 440 e-122
UniRef100_Q9SMZ2 Hypothetical protein F4I10.100 [Arabidopsis tha... 439 e-121
UniRef100_Q6K297 Pentatricopeptide (PPR) repeat-containing prote... 437 e-121
UniRef100_Q9LW63 Emb|CAB37460.1 [Arabidopsis thaliana] 436 e-121
UniRef100_Q9LHZ4 Pentatricopeptide (PPR) repeat-containing prote... 436 e-120
UniRef100_Q5ZDP1 Pentatricopeptide (PPR) repeat-containing prote... 434 e-120
UniRef100_Q9SUH6 Hypothetical protein T10C21.50 [Arabidopsis tha... 428 e-118
UniRef100_Q9SY02 Hypothetical protein T5J8.5 [Arabidopsis thaliana] 427 e-118
>UniRef100_Q9FK93 Selenium-binding protein-like [Arabidopsis thaliana]
Length = 710
Score = 688 bits (1775), Expect = 0.0
Identities = 330/697 (47%), Positives = 481/697 (68%), Gaps = 7/697 (1%)
Query: 10 VVVNSRYLP-PLEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLY 68
+V S+ P P++ + +LLK+ A++ L G++IHA L+V +Q+S+ D Q+NSLI+LY
Sbjct: 20 LVPKSKKTPFPIDRLNELLKVCANSSYLRIGESIHAHLIVTNQSSRAEDAYQINSLINLY 79
Query: 69 VKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNE 128
VKC AR +FD MP R+V SW +M GY +SG EVL LFK++ S ++ PNE
Sbjct: 80 VKCRETVRARKLFDLMPERNVVSWCAMMKGYQNSGFDFEVLKLFKSMFFSGESR---PNE 136
Query: 129 YVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
+V T V SCS+SGR+ EG Q HG KYGL+S++FV++ L MY+ C A++VL+
Sbjct: 137 FVATVVFKSCSNSGRIEEGKQFHGCFLKYGLISHEFVRNTLVYMYSLCSGNGEAIRVLDD 196
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
D+ +++S L+ +E G ++E + VLR+ NE VW+N TY+ + L + + DL
Sbjct: 197 LP---YCDLSVFSSALSGYLECGAFKEGLDVLRKTANEDFVWNNLTYLSSLRLFSNLRDL 253
Query: 249 QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAY 308
L L+VH+R++R G + LI+MYGKC VL A++VFD +N+ + T++M AY
Sbjct: 254 NLALQVHSRMVRFGFNAEVEACGALINMYGKCGKVLYAQRVFDDTHAQNIFLNTTIMDAY 313
Query: 309 LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYV 368
Q++ FEE L+L + MD ++ PNE TF +LL + A ++LLK GDLLH + K G++N+V
Sbjct: 314 FQDKSFEEALNLFSKMDTKEVPPNEYTFAILLNSIAELSLLKQGDLLHGLVLKSGYRNHV 373
Query: 369 IVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSA 428
+V NAL+NMY+KSG I+ + FS RD+ TWN+MI G SHHGLG+EAL F M
Sbjct: 374 MVGNALVNMYAKSGSIEDARKAFSGMTFRDIVTWNTMISGCSHHGLGREALEAFDRMIFT 433
Query: 429 GESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEK 488
GE PN +TF+GVL AC+H+ V++GL Y MK ++P ++HYTC+V L +AG+ +
Sbjct: 434 GEIPNRITFIGVLQACSHIGFVEQGLHYFNQLMKKFDVQPDIQHYTCIVGLLSKAGMFKD 493
Query: 489 AENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHAT 548
AE+FM+T ++WD+VAWRTLLN+C V RNY LGK++AE ++ P D G Y LLSN+HA
Sbjct: 494 AEDFMRTAPIEWDVVAWRTLLNACYVRRNYRLGKKVAEYAIEKYPNDSGVYVLLSNIHAK 553
Query: 549 ANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAM 608
+ W+GV ++R +M R +KKEPG SW+ IRN HVF++E +HPE I KV+++++
Sbjct: 554 SREWEGVAKVRSLMNNRGVKKEPGVSWIGIRNQTHVFLAEDNQHPEITLIYAKVKEVMSK 613
Query: 609 IKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDC 668
IKPLGY P+++ HDV++EQ+E L+ HSEKLA+AYGL+K P +P+ + KN+R+CDDC
Sbjct: 614 IKPLGYSPDVAGAFHDVDEEQREDNLSYHSEKLAVAYGLIKTPEKSPLYVTKNVRICDDC 673
Query: 669 HTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
H+A+KLISK++ R+I++RD+NRFHHF DG C+C D+W
Sbjct: 674 HSAIKLISKISKRYIVIRDSNRFHHFLDGQCSCCDYW 710
>UniRef100_Q65XK6 Hypothetical protein OJ1735_C10.12 [Oryza sativa]
Length = 687
Score = 646 bits (1666), Expect = 0.0
Identities = 325/682 (47%), Positives = 454/682 (65%), Gaps = 14/682 (2%)
Query: 24 VKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDK 83
V +L+ +A A L+ GKA+HA+++ + R DVVQ N+LI LYVKCGRL LAR +FD
Sbjct: 20 VAVLRAAAAAGELSLGKAVHARVV----RAARFDVVQYNNLIALYVKCGRLGLARQVFDA 75
Query: 84 MPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGR 143
MP R+ S N LM+GY SG + + L L + NEYV ++ +++ +H
Sbjct: 76 MPSRNPVSGNLLMSGYASSGRHRDALALLR-------VADFGLNEYVLSSAVAATAHVRS 128
Query: 144 VREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSL 203
G QCHGY K GL + +V SA+ +MY +C H++ A++V + S + +F +NS+
Sbjct: 129 YDMGRQCHGYAIKAGLAEHPYVCSAVLHMYCQCAHMDEAVKVFDNVSSFN---VFAFNSM 185
Query: 204 LNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGL 263
+N ++ G+ + + ++R M WD+ +YV V+G CA ++ LG +VH + L+ L
Sbjct: 186 INGFLDRGQMDGSTSIVRSMVRNVGQWDHVSYVAVLGHCASTKEVVLGSQVHTQALKRRL 245
Query: 264 LFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTC 323
+ +VGS L+DMYGKC A +VF+ L +N+V WT++M+AY QN FE+ L L
Sbjct: 246 ELNVYVGSALVDMYGKCDFPHEANRVFEVLPEKNIVSWTAIMTAYTQNELFEDALQLFLD 305
Query: 324 MDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGD 383
M+ E PNE T+ V L +CAG+A LK+G+ L A K G + V NAL+NMYSKSG
Sbjct: 306 MEMEGVRPNEFTYAVALNSCAGLATLKNGNALGACTMKTGHWGLLPVCNALMNMYSKSGS 365
Query: 384 IDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSA 443
++ + VF CRDV +WNS+I GY+HHG +EA+ F DM A E P+ VTF+GVLSA
Sbjct: 366 VEDARRVFLSMPCRDVVSWNSIIIGYAHHGRAREAMEAFHDMLFAEEVPSYVTFIGVLSA 425
Query: 444 CAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV 503
CA L LVD+G YL MK ++PG EHYTCMV L CR G L++AE F+++ + D+V
Sbjct: 426 CAQLGLVDEGFYYLNIMMKEVGVKPGKEHYTCMVGLLCRVGRLDEAERFIESNCIGTDVV 485
Query: 504 AWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMK 563
AWR+LL+SC+V+RNY LG R+AE I Q+ P+D+GTY LLSNM+A AN WDGVV++R++M+
Sbjct: 486 AWRSLLSSCQVYRNYGLGHRVAEQIFQLKPKDVGTYVLLSNMYAKANRWDGVVKVRRLMR 545
Query: 564 ERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLH 623
E ++KEPG SW+++ + HVF SE KHP +I KK+Q+L+ IK +GYVPNI+ LH
Sbjct: 546 ELGVRKEPGVSWIQVGSEVHVFTSEDKKHPYMEQITKKLQELIDKIKVIGYVPNIAVALH 605
Query: 624 DVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFI 683
DVEDEQKE +L HSEKLA+A+GL++ P IRI+KN+R+CDDCH A+KLIS T R I
Sbjct: 606 DVEDEQKEEHLMYHSEKLALAFGLIRTPKGEAIRIMKNVRICDDCHVAIKLISLATGRRI 665
Query: 684 IVRDANRFHHFRDGSCTCADHW 705
+VRD RFH DG C+C D+W
Sbjct: 666 VVRDTVRFHCIEDGVCSCDDYW 687
>UniRef100_Q5W964 PPR986-12 [Physcomitrella patens]
Length = 986
Score = 489 bits (1258), Expect = e-136
Identities = 247/682 (36%), Positives = 400/682 (58%), Gaps = 11/682 (1%)
Query: 24 VKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDK 83
V LL+ + L GK +HA++ + +++ +++ +Y KCG + A +FD
Sbjct: 316 VSLLRACNHPEALEQGKKVHARM---KEVGWDTEIYVGTAILSMYTKCGSMEDALEVFDL 372
Query: 84 MPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGR 143
+ R+V SW ++ G+ G E + F ++ S PN F ++L +CS
Sbjct: 373 VKGRNVVSWTAMIAGFAQHGRIDEAFLFFNKMIESGIE----PNRVTFMSILGACSSPSA 428
Query: 144 VREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSL 203
++ G Q ++ + G S V++AL +MYA+C + A +V S + ++ +N++
Sbjct: 429 LKRGQQIQDHIIEAGYGSDDRVRTALLSMYAKCGSLKDAHRVFEKIS---KQNVVAWNAM 485
Query: 204 LNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGL 263
+ V+ +++ A+ + + E + +++T+ ++ +C L+LG VH +++ GL
Sbjct: 486 ITAYVQHEQYDNALATFQALLKEGIKPNSSTFTSILNVCKSSDSLELGKWVHFLIMKAGL 545
Query: 264 LFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTC 323
D V + L+ M+ C +++A+ +F+ + R++V W ++++ ++Q+ + D
Sbjct: 546 ESDLHVSNALVSMFVNCGDLMSAKNLFNDMPKRDLVSWNTIIAGFVQHGKNQVAFDYFKM 605
Query: 324 MDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGD 383
M + P++ TF LL ACA L G LHA + + F V+V LI+MY+K G
Sbjct: 606 MQESGIKPDKITFTGLLNACASPEALTEGRRLHALITEAAFDCDVLVGTGLISMYTKCGS 665
Query: 384 IDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSA 443
I+ ++ VF ++V +W SMI GY+ HG G+EAL +F M+ G P+ +TFVG LSA
Sbjct: 666 IEDAHQVFHKLPKKNVYSWTSMIAGYAQHGRGKEALELFYQMQQEGVKPDWITFVGALSA 725
Query: 444 CAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV 503
CAH L+++GL + + MK IEP +EHY CMV L+ RAGLL +A F+ +V+ D
Sbjct: 726 CAHAGLIEEGLHH-FQSMKEFNIEPRMEHYGCMVDLFGRAGLLNEAVEFIIKMQVEPDSR 784
Query: 504 AWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMK 563
W LL +C+VH N +L ++ A+ L++DP D G + +LSN++A A MW V ++RK+M
Sbjct: 785 VWGALLGACQVHLNVELAEKAAQKKLELDPNDNGVFVILSNIYAAAGMWKEVAKMRKVML 844
Query: 564 ERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLH 623
+R + K+PG SW+E+ H F S+ HP++ EI+ ++++L ++ LGYVP+ VLH
Sbjct: 845 DRGVVKKPGQSWIEVDGKVHTFYSDDKTHPQTEEIHAELERLHMEMRQLGYVPDTRYVLH 904
Query: 624 DVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFI 683
DVED +KE L HSE+LAI YGL+K P PI I KNLR+C DCHTA K ISK+T R I
Sbjct: 905 DVEDNEKEQALFYHSERLAITYGLLKTPPLTPIVISKNLRVCGDCHTATKFISKITKRQI 964
Query: 684 IVRDANRFHHFRDGSCTCADHW 705
I RD+NRFHHF+DG C+C D W
Sbjct: 965 IARDSNRFHHFKDGVCSCGDFW 986
Score = 233 bits (594), Expect = 1e-59
Identities = 146/511 (28%), Positives = 258/511 (49%), Gaps = 16/511 (3%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
LL+L K L G+ I+ + ++ + D+ N+LI++Y KCG A+ +FD M
Sbjct: 116 LLQLCIKFKNLGDGERIYNHI---KKSGVQPDIFMRNTLINMYAKCGNTISAKQIFDDMR 172
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
+DV+SWN L+ GY+ G Y E L + +V S P++ F ++L++C+ + V
Sbjct: 173 EKDVYSWNLLLGGYVQHGLYEEAFKLHEQMVQDSVK----PDKRTFVSMLNACADARNVD 228
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLN 205
+G + + + K G + FV +AL NM+ +C + A +V + D+ + S++
Sbjct: 229 KGRELYNLILKAGWDTDLFVGTALINMHIKCGDIGDATKVFDNLP---TRDLVTWTSMIT 285
Query: 206 VLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLF 265
L GR+++A + +RM E + D +V ++ C L+ G +VHAR+ G
Sbjct: 286 GLARHGRFKQACNLFQRMEEEGVQPDKVAFVSLLRACNHPEALEQGKKVHARMKEVGWDT 345
Query: 266 DEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMD 325
+ +VG+ ++ MY KC + +A +VFD ++ RNVV WT++++ + Q+ +E M
Sbjct: 346 EIYVGTAILSMYTKCGSMEDALEVFDLVKGRNVVSWTAMIAGFAQHGRIDEAFLFFNKMI 405
Query: 326 QEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDID 385
+ PN TF +LGAC+ + LK G + + + G+ + V AL++MY+K G +
Sbjct: 406 ESGIEPNRVTFMSILGACSSPSALKRGQQIQDHIIEAGYGSDDRVRTALLSMYAKCGSLK 465
Query: 386 SSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACA 445
++ VF ++V WN+MI Y H AL FQ + G PN+ TF +L+ C
Sbjct: 466 DAHRVFEKISKQNVVAWNAMITAYVQHEQYDNALATFQALLKEGIKPNSSTFTSILNVCK 525
Query: 446 HLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAW 505
++ G +++ + +E L +V ++ G L A+N K D+V+W
Sbjct: 526 SSDSLELG-KWVHFLIMKAGLESDLHVSNALVSMFVNCGDLMSAKNLFNDMP-KRDLVSW 583
Query: 506 RTLLNSCRVHRN----YDLGKRIAESILQMD 532
T++ H +D K + ES ++ D
Sbjct: 584 NTIIAGFVQHGKNQVAFDYFKMMQESGIKPD 614
Score = 208 bits (529), Expect = 5e-52
Identities = 126/465 (27%), Positives = 240/465 (51%), Gaps = 18/465 (3%)
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
++D N ++ +G + E + + + + SS H ++ +L C +
Sbjct: 72 IKDTQKANAVLNRLSKAGQFNEAMQVLERVDSS----HIQIYRQTYSALLQLCIKFKNLG 127
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLN 205
+G + + ++ K G+ F+++ L NMYA+C + A Q+ + E D++ +N LL
Sbjct: 128 DGERIYNHIKKSGVQPDIFMRNTLINMYAKCGNTISAKQIFDDMR---EKDVYSWNLLLG 184
Query: 206 VLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLF 265
V+ G +EEA + +M + + D T+V ++ CA ++ G ++ +L+ G
Sbjct: 185 GYVQHGLYEEAFKLHEQMVQDSVKPDKRTFVSMLNACADARNVDKGRELYNLILKAGWDT 244
Query: 266 DEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMD 325
D FVG+ LI+M+ KC + +A KVFD L R++V WTS+++ ++ F++ +L M+
Sbjct: 245 DLFVGTALINMHIKCGDIGDATKVFDNLPTRDLVTWTSMITGLARHGRFKQACNLFQRME 304
Query: 326 QEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDID 385
+E P++ F LL AC L+ G +HAR++++G+ + V A+++MY+K G ++
Sbjct: 305 EEGVQPDKVAFVSLLRACNHPEALEQGKKVHARMKEVGWDTEIYVGTAILSMYTKCGSME 364
Query: 386 SSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACA 445
+ VF R+V +W +MI G++ HG EA F M +G PN VTF+ +L AC+
Sbjct: 365 DALEVFDLVKGRNVVSWTAMIAGFAQHGRIDEAFLFFNKMIESGIEPNRVTFMSILGACS 424
Query: 446 HLALVDKGLDYLYNEMKNHKIEPGL----EHYTCMVVLYCRAGLLEKAENFMKTTKVKWD 501
+ + +G ++++H IE G T ++ +Y + G L+ A + K +
Sbjct: 425 SPSALKRG-----QQIQDHIIEAGYGSDDRVRTALLSMYAKCGSLKDAHRVFEKIS-KQN 478
Query: 502 IVAWRTLLNSCRVHRNYDLGKRIAESILQMD-PQDMGTYTLLSNM 545
+VAW ++ + H YD +++L+ + T+T + N+
Sbjct: 479 VVAWNAMITAYVQHEQYDNALATFQALLKEGIKPNSSTFTSILNV 523
Score = 171 bits (432), Expect = 9e-41
Identities = 100/327 (30%), Positives = 174/327 (52%), Gaps = 2/327 (0%)
Query: 193 DENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGL 252
D D N++LN L ++G++ EA+ VL R+ + + TY ++ LC + +L G
Sbjct: 71 DIKDTQKANAVLNRLSKAGQFNEAMQVLERVDSSHIQIYRQTYSALLQLCIKFKNLGDGE 130
Query: 253 RVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNR 312
R++ + + G+ D F+ + LI+MY KC + ++A+++FD ++ ++V W L+ Y+Q+
Sbjct: 131 RIYNHIKKSGVQPDIFMRNTLINMYAKCGNTISAKQIFDDMREKDVYSWNLLLGGYVQHG 190
Query: 313 YFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNN 372
+EE L M Q+ P++ TF +L ACA + G L+ + K G+ + V
Sbjct: 191 LYEEAFKLHEQMVQDSVKPDKRTFVSMLNACADARNVDKGRELYNLILKAGWDTDLFVGT 250
Query: 373 ALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESP 432
ALINM+ K GDI + VF + RD+ TW SMI G + HG ++A +FQ M+ G P
Sbjct: 251 ALINMHIKCGDIGDATKVFDNLPTRDLVTWTSMITGLARHGRFKQACNLFQRMEEEGVQP 310
Query: 433 NNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENF 492
+ V FV +L AC H +++G ++ MK + + T ++ +Y + G +E A
Sbjct: 311 DKVAFVSLLRACNHPEALEQG-KKVHARMKEVGWDTEIYVGTAILSMYTKCGSMEDALEV 369
Query: 493 MKTTKVKWDIVAWRTLLNSCRVHRNYD 519
K + ++V+W ++ H D
Sbjct: 370 FDLVKGR-NVVSWTAMIAGFAQHGRID 395
Score = 43.1 bits (100), Expect = 0.028
Identities = 57/283 (20%), Positives = 102/283 (35%), Gaps = 65/283 (22%)
Query: 390 VFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLAL 449
VF+D +D N+++ S G EA++V + + S+ T+ +L C
Sbjct: 68 VFADI--KDTQKANAVLNRLSKAGQFNEAMQVLERVDSSHIQIYRQTYSALLQLCIKFKN 125
Query: 450 VDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
+ G + +YN +K ++P + ++ +Y + G A+ + K D+ +W LL
Sbjct: 126 LGDG-ERIYNHIKKSGVQPDIFMRNTLINMYAKCGNTISAKQIFDDMREK-DVYSWNLLL 183
Query: 510 -----------------------------------NSCRVHRNYDLGKRIAESILQMD-P 533
N+C RN D G+ + IL+
Sbjct: 184 GGYVQHGLYEEAFKLHEQMVQDSVKPDKRTFVSMLNACADARNVDKGRELYNLILKAGWD 243
Query: 534 QDMGTYTLLSNMHATA----------------------NMWDGVVEIRKMMKERNIKKEP 571
D+ T L NMH +M G+ + + N+ +
Sbjct: 244 TDLFVGTALINMHIKCGDIGDATKVFDNLPTRDLVTWTSMITGLARHGRFKQACNLFQRM 303
Query: 572 GASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGY 614
++ V V + C HPE+ E KKV A +K +G+
Sbjct: 304 EEEGVQPDKVAFVSLLRACNHPEALEQGKKVH---ARMKEVGW 343
>UniRef100_Q7XHP5 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 720
Score = 456 bits (1174), Expect = e-127
Identities = 247/717 (34%), Positives = 402/717 (55%), Gaps = 45/717 (6%)
Query: 31 ADAKCLTSGKAIH---AQLLVRDQASKRSDVV--------QLNSLIHLYVKCGRLRLARS 79
A A+ L+ + H A LL+R +A++ + + S++ +V+ R AR
Sbjct: 7 AAARLLSPRASTHPFSAALLLRGRAARGGSLEARLATVPHERASVLRFWVRRRRFHDARG 66
Query: 80 MFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCS 139
+FD+ P R W ++G G Y + + F +++ A PN +V V+ C+
Sbjct: 67 VFDERPTRTAPVWTLTISGCARRGRYADGMRAFAEMLAEG---EATPNAFVLAAVVRCCA 123
Query: 140 HSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT----------- 188
G V G + HG++ + G+ + +A+ +MYA+C A +V
Sbjct: 124 GMGDVESGKRVHGWMLRNGVHLDVVLCNAVLDMYAKCGQFERARRVFGAMAERDAVSWNI 183
Query: 189 ------ESGG--------DEN---DIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWD 231
+SG DE+ D +N++++ L+ SG +A+ LRRM +V++
Sbjct: 184 AIGACIQSGDILGSMQLFDESPLRDTTSWNTIISGLMRSGHAADALSHLRRMAQAGVVFN 243
Query: 232 NATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFD 291
+ TY L + LG ++H R+L L D FV S L+DMY KC + A VFD
Sbjct: 244 HYTYSTAFVLAGMLLLPDLGRQLHGRVLIAALEGDAFVRSSLMDMYCKCGLLEAAASVFD 303
Query: 292 G---LQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIAL 348
L W+++++ Y+QN EE LDL M +E + T + ACA + +
Sbjct: 304 HWSPLTRDMNFAWSTMVAGYVQNGREEEALDLFRRMLREGVAADRFTLTSVAAACANVGM 363
Query: 349 LKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICG 408
++ G +H +EKL +K + +A+++MY+K G+++ + ++F +++ W SM+C
Sbjct: 364 VEQGRQVHGCVEKLWYKLDAPLASAIVDMYAKCGNLEDARSIFDRACTKNIAVWTSMLCS 423
Query: 409 YSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEP 468
Y+ HG G+ A+ +F+ M + +PN +T VGVLSAC+H+ LV +G Y + + I P
Sbjct: 424 YASHGQGRIAIELFERMTAEKMTPNEITLVGVLSACSHVGLVSEGELYFKQMQEEYGIVP 483
Query: 469 GLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESI 528
+EHY C+V LY R+GLL+KA+NF++ + + + W+TLL++CR+H++ + K +E +
Sbjct: 484 SIEHYNCIVDLYGRSGLLDKAKNFIEENNINHEAIVWKTLLSACRLHQHNEYAKLASEKL 543
Query: 529 LQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSE 588
+Q++ D G+Y +LSN++AT N W E+R M+ER ++K+PG SW+ ++N H FV+
Sbjct: 544 VQLEQCDAGSYVMLSNIYATNNKWHDTFELRVSMQERKVRKQPGRSWIHLKNTVHTFVAG 603
Query: 589 GCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLM 648
HP+S EI +++L+ +K +GY V+HDVEDEQ+E L HSEKLAIA+G++
Sbjct: 604 DASHPQSAEIYAYLEKLVERLKEIGYTSRTDLVVHDVEDEQRETALKFHSEKLAIAFGII 663
Query: 649 KMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
PS P+RI KNLR+C+DCH A+K IS T R I+VRD RFHHF+D SC+C D W
Sbjct: 664 STPSGTPLRIFKNLRVCEDCHEAIKYISLATGREIVVRDLYRFHHFKDASCSCEDFW 720
>UniRef100_Q5W965 PPR868-14 [Physcomitrella patens]
Length = 868
Score = 456 bits (1172), Expect = e-126
Identities = 249/718 (34%), Positives = 383/718 (52%), Gaps = 46/718 (6%)
Query: 24 VKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDK 83
+ +LK + L G+ IH + +DV +LI +Y KCG + +A +F K
Sbjct: 161 LSILKACNNYSILEKGRKIHT---IVKAMGMETDVAVATALITMYSKCGEISVACEVFHK 217
Query: 84 MPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGR 143
M R+V SW ++ E L++ ++ + + PN F ++L+SC+
Sbjct: 218 MTERNVVSWTAIIQANAQHRKLNEAFELYEQMLQAGIS----PNAVTFVSLLNSCNTPEA 273
Query: 144 VREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSL 203
+ G + H ++ + GL + V +AL MY +C V A ++ + S + D+ ++++
Sbjct: 274 LNRGRRIHSHISERGLETDMIVANALITMYCKCNSVQEAREIFDRMS---KRDVISWSAM 330
Query: 204 LNVLVESG-----RWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARL 258
+ +SG +E +L RM E + + T++ ++ C G L+ G ++HA L
Sbjct: 331 IAGYAQSGYKDKESIDEVFQLLERMRREGVFPNKVTFMSILRACTAHGALEQGRQIHAEL 390
Query: 259 LRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYL--------- 309
+ G D + + + +MY KC + A +VF + N+NVV WTS +S Y+
Sbjct: 391 SKVGFELDRSLQTAIFNMYAKCGSIYEAEQVFSKMANKNVVAWTSFLSMYIKCGDLSSAE 450
Query: 310 ----------------------QNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIA 347
QN + +LL+ M E P+ T +L AC +A
Sbjct: 451 KVFSEMPTRNVVSWNLMIAGYAQNGDIVKVFELLSSMKAEGFQPDRVTVITILEACGALA 510
Query: 348 LLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMIC 407
L+ G L+HA KLG ++ +V +LI MYSK G + + VF RD WN+M+
Sbjct: 511 GLERGKLVHAEAVKLGLESDTVVATSLIGMYSKCGQVAEARTVFDKMSNRDTVAWNAMLA 570
Query: 408 GYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIE 467
GY HG G EA+ +F+ M SPN +T V+SAC+ LV +G + ++ K+
Sbjct: 571 GYGQHGDGLEAVDLFKRMLKERVSPNEITLTAVISACSRAGLVQEGREIFRMMQEDFKMT 630
Query: 468 PGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAES 527
P +HY CMV L RAG L++AE F+++ + DI W LL +C+ H N L +R A
Sbjct: 631 PRKQHYGCMVDLLGRAGRLQEAEEFIQSMPCEPDISVWHALLGACKSHNNVQLAERAAHH 690
Query: 528 ILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVS 587
IL+++P Y LSN++A A WD ++R++M +R +KK+ G S +EI H FV+
Sbjct: 691 ILELEPSYASVYITLSNIYAQAGRWDDSTKVRRVMDDRGLKKDRGESSIEIDGRIHTFVA 750
Query: 588 EGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGL 647
E C HPE + I+ +++ L +K GY P++ VLHDV+D QKE L HSEKLAIAYGL
Sbjct: 751 EDCAHPEIDAIHAELETLTKEMKEAGYTPDMRFVLHDVDDVQKEKALCHHSEKLAIAYGL 810
Query: 648 MKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+K PS PIRI+KNLR+C DCHTA K ISK+ R I+ RDANRFH+F +G+C+C D W
Sbjct: 811 LKTPSGTPIRIMKNLRVCGDCHTATKFISKIRKREIVARDANRFHYFNNGTCSCGDFW 868
Score = 212 bits (539), Expect = 3e-53
Identities = 149/574 (25%), Positives = 270/574 (46%), Gaps = 55/574 (9%)
Query: 31 ADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVF 90
A A+ GK +H QL D+ D+ NSLI+ Y K + A +F +M LRDV
Sbjct: 67 AKARRFEDGKMVHKQL---DELGVEIDIYLGNSLINFYSKFEDVASAEQVFRRMTLRDVV 123
Query: 91 SWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQC 150
+W+ ++ Y + + + F+ + ++ PN F ++L +C++ + +G +
Sbjct: 124 TWSSMIAAYAGNNHPAKAFDTFERMTDANIE----PNRITFLSILKACNNYSILEKGRKI 179
Query: 151 HGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVES 210
H + G+ + V +AL MY++C +++A +V + + E ++ + +++ +
Sbjct: 180 HTIVKAMGMETDVAVATALITMYSKCGEISVACEVFHKMT---ERNVVSWTAIIQANAQH 236
Query: 211 GRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVG 270
+ EA + +M + + T+V ++ C L G R+H+ + GL D V
Sbjct: 237 RKLNEAFELYEQMLQAGISPNAVTFVSLLNSCNTPEALNRGRRIHSHISERGLETDMIVA 296
Query: 271 SKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRY-----FEETLDLLTCMD 325
+ LI MY KC V AR++FD + R+V+ W+++++ Y Q+ Y +E LL M
Sbjct: 297 NALITMYCKCNSVQEAREIFDRMSKRDVISWSAMIAGYAQSGYKDKESIDEVFQLLERMR 356
Query: 326 QEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFK-------------------- 365
+E PN+ TF +L AC L+ G +HA L K+GF+
Sbjct: 357 REGVFPNKVTFMSILRACTAHGALEQGRQIHAELSKVGFELDRSLQTAIFNMYAKCGSIY 416
Query: 366 -----------NYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGL 414
V+ + ++MY K GD+ S+ VFS+ R+V +WN MI GY+ +G
Sbjct: 417 EAEQVFSKMANKNVVAWTSFLSMYIKCGDLSSAEKVFSEMPTRNVVSWNLMIAGYAQNGD 476
Query: 415 GQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYT 474
+ + MK+ G P+ VT + +L AC LA +++G ++ E +E T
Sbjct: 477 IVKVFELLSSMKAEGFQPDRVTVITILEACGALAGLERG-KLVHAEAVKLGLESDTVVAT 535
Query: 475 CMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRN----YDLGKRIAESILQ 530
++ +Y + G + +A + D VAW +L H + DL KR+ + +
Sbjct: 536 SLIGMYSKCGQVAEARTVFDKMSNR-DTVAWNAMLAGYGQHGDGLEAVDLFKRMLKE--R 592
Query: 531 MDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKE 564
+ P ++ T T + + + A + EI +MM+E
Sbjct: 593 VSPNEI-TLTAVISACSRAGLVQEGREIFRMMQE 625
Score = 159 bits (402), Expect = 3e-37
Identities = 104/389 (26%), Positives = 181/389 (45%), Gaps = 10/389 (2%)
Query: 127 NEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVL 186
N + V+ C+ + R +G H L + G+ ++ ++L N Y++ V A QV
Sbjct: 55 NSNTYGCVIEHCAKARRFEDGKMVHKQLDELGVEIDIYLGNSLINFYSKFEDVASAEQVF 114
Query: 187 NTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIG 246
+ D+ ++S++ + +A RM + + + T++ ++ C
Sbjct: 115 RRMT---LRDVVTWSSMIAAYAGNNHPAKAFDTFERMTDANIEPNRITFLSILKACNNYS 171
Query: 247 DLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMS 306
L+ G ++H + G+ D V + LI MY KC + A +VF + RNVV WT+++
Sbjct: 172 ILEKGRKIHTIVKAMGMETDVAVATALITMYSKCGEISVACEVFHKMTERNVVSWTAIIQ 231
Query: 307 AYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKN 366
A Q+R E +L M Q PN TF LL +C L G +H+ + + G +
Sbjct: 232 ANAQHRKLNEAFELYEQMLQAGISPNAVTFVSLLNSCNTPEALNRGRRIHSHISERGLET 291
Query: 367 YVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLG-----QEALRV 421
+IV NALI MY K + + +F RDV +W++MI GY+ G E ++
Sbjct: 292 DMIVANALITMYCKCNSVQEAREIFDRMSKRDVISWSAMIAGYAQSGYKDKESIDEVFQL 351
Query: 422 FQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYC 481
+ M+ G PN VTF+ +L AC +++G ++ E+ E T + +Y
Sbjct: 352 LERMRREGVFPNKVTFMSILRACTAHGALEQGRQ-IHAELSKVGFELDRSLQTAIFNMYA 410
Query: 482 RAGLLEKAENFMKTTKVKWDIVAWRTLLN 510
+ G + +AE K ++VAW + L+
Sbjct: 411 KCGSIYEAEQVFSKMANK-NVVAWTSFLS 438
Score = 148 bits (373), Expect = 6e-34
Identities = 88/303 (29%), Positives = 159/303 (52%), Gaps = 2/303 (0%)
Query: 207 LVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFD 266
L ++GR EAI +L + ++ ++ TY V+ CA+ + G VH +L G+ D
Sbjct: 31 LCKAGRLREAIQLLGIIKQRGLLVNSNTYGCVIEHCAKARRFEDGKMVHKQLDELGVEID 90
Query: 267 EFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQ 326
++G+ LI+ Y K V +A +VF + R+VV W+S+++AY N + + D M
Sbjct: 91 IYLGNSLINFYSKFEDVASAEQVFRRMTLRDVVTWSSMIAAYAGNNHPAKAFDTFERMTD 150
Query: 327 EDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDS 386
+ PN TF +L AC ++L+ G +H ++ +G + V V ALI MYSK G+I
Sbjct: 151 ANIEPNRITFLSILKACNNYSILEKGRKIHTIVKAMGMETDVAVATALITMYSKCGEISV 210
Query: 387 SYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAH 446
+ VF R+V +W ++I + H EA +++ M AG SPN VTFV +L++C
Sbjct: 211 ACEVFHKMTERNVVSWTAIIQANAQHRKLNEAFELYEQMLQAGISPNAVTFVSLLNSCNT 270
Query: 447 LALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWR 506
+++G +++ + +E + ++ +YC+ +++A K D+++W
Sbjct: 271 PEALNRG-RRIHSHISERGLETDMIVANALITMYCKCNSVQEAREIFDRMS-KRDVISWS 328
Query: 507 TLL 509
++
Sbjct: 329 AMI 331
>UniRef100_Q8S263 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 1062
Score = 455 bits (1170), Expect = e-126
Identities = 243/685 (35%), Positives = 397/685 (57%), Gaps = 15/685 (2%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
+ + S + L G+ +HA +L ++ + N L+++Y KCG + A +F M
Sbjct: 388 IAEFSTAEQGLRKGREVHAHVLRAGHIYRK--IAVSNGLVNMYAKCGAIDKACRVFQLME 445
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
RD SWN ++T +G Y E ++ L+ ++ P+ + + LSSC+ G +
Sbjct: 446 ARDRISWNTIITALDQNG-YCEAAMMNYCLMRQNSIG---PSNFAAISGLSSCAGLGLLA 501
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLN 205
G Q H K+GL V +AL MY C ++ ++ N+ S +D+ +NS++
Sbjct: 502 AGQQLHCDAVKWGLYLDTSVSNALVKMYGECGRMSECWEIFNSMSA---HDVVSWNSIMG 558
Query: 206 VLVES-GRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLL 264
V+ S E++ V M +V + T+V + + L+LG ++H+ +L+ G+
Sbjct: 559 VMASSQAPITESVQVFSNMMKSGLVPNKVTFVNFLAALTPLSVLELGKQIHSVMLKHGVT 618
Query: 265 FDEFVGSKLIDMYGKCCHVLNARKVFDGLQNR-NVVVWTSLMSAYLQNRYFEETLDLLTC 323
D V + L+ Y K V + ++F + R + + W S++S Y+ N + +E +D +
Sbjct: 619 EDNAVDNALMSCYAKSGDVDSCERLFSRMSGRRDAISWNSMISGYIYNGHLQEAMDCVCL 678
Query: 324 MDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGD 383
M + + + CTF ++L ACA +A L+ G +HA + ++ V+V +AL++MYSK G
Sbjct: 679 MMHSEQMMDHCTFSIVLNACASVAALERGMEMHAFGLRSHLESDVVVESALVDMYSKCGR 738
Query: 384 IDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSA 443
ID + VF ++ +WNSMI GY+ HGLG++AL +F++M+ +GESP++VTFV VLSA
Sbjct: 739 IDYASKVFHSMSQKNEFSWNSMISGYARHGLGRKALEIFEEMQESGESPDHVTFVSVLSA 798
Query: 444 CAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV 503
C+H LV++GLDY + M+++ I P +EHY+C++ L RAG L+K + +MK +K + +
Sbjct: 799 CSHAGLVERGLDY-FELMEDYGILPRIEHYSCVIDLLGRAGELDKIQEYMKRMPMKPNTL 857
Query: 504 AWRTLLNSCRVHRN---YDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRK 560
WRT+L +C+ ++ DLG + +L+++PQ+ Y L S HA W+ + R
Sbjct: 858 IWRTVLVACQQSKHRAKIDLGTEASRMLLELEPQNPVNYVLSSKFHAAIGRWEDTAKARA 917
Query: 561 MMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISA 620
MK +KKE G SW+ + + H F++ HP + EI +K+ L+ I+ GYVP
Sbjct: 918 AMKGAAVKKEAGRSWVTLTDGVHTFIAGDRSHPNTKEIYEKLNFLIQKIRNAGYVPLTEY 977
Query: 621 VLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTN 680
VLHD+E+E KE L HSEKLA+A+ L + S PIRI+KNLR+C DCHTA + IS++
Sbjct: 978 VLHDLEEENKEELLRYHSEKLAVAFVLTRSSSGGPIRIMKNLRVCGDCHTAFRYISQIVG 1037
Query: 681 RFIIVRDANRFHHFRDGSCTCADHW 705
R II+RD+ RFHHF+DG C+C D+W
Sbjct: 1038 RQIILRDSIRFHHFKDGKCSCGDYW 1062
Score = 145 bits (367), Expect = 3e-33
Identities = 128/492 (26%), Positives = 226/492 (45%), Gaps = 31/492 (6%)
Query: 35 CLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWND 94
C S +++H +++ R D+ N L++ Y K RL AR +FD MP R+ SW
Sbjct: 80 CDASPESLHLEVVKRGLTH---DLFLANHLVNSYAKGARLDAARRVFDGMPGRNAVSWTC 136
Query: 95 LMTGYMHSGNYLEVLVLFKNLVSSSTTQHAC-PNEYVFTTVLSSCSHSGRVREG--MQCH 151
L++G++ SG + LF+ ++ C P + F +VL +C SG R G +Q H
Sbjct: 137 LISGHVLSGLPEDAFPLFRAMLREGP---GCRPTSFTFGSVLRACQDSGPDRLGFAVQVH 193
Query: 152 GYLFKYGLVSYQFVKSALFNMYARCL--HVNLALQVLNTESGGDENDIFLYNSLLNVLVE 209
G + K S V +AL +MY C LA +V +T D+ +N+L++V +
Sbjct: 194 GLVSKTEFTSNTTVCNALISMYGSCSVGPPILAQRVFDTT---PVRDLITWNALMSVYAK 250
Query: 210 SGRWEEAIGVLRRM-----GNECMVWDNATYVGVMGLCAQIGDLQLGL--RVHARLLRGG 262
G + R M G E ++ G + + LGL ++ R+L+ G
Sbjct: 251 RGDAICTFTLFRAMQYDDSGIELRPTEHT--FGSLITATYLSSCSLGLLDQLFVRVLKSG 308
Query: 263 LLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLT 322
D +VGS L+ + + + A+ ++ GL+ RN V L++ ++ ++ E ++
Sbjct: 309 CSSDLYVGSALVSAFARHGMLDEAKDIYLGLKERNAVTLNGLIAGLVKQQHGEAAAEIFM 368
Query: 323 CMDQEDTLPNECTFKVLLGACAGIAL----LKHGDLLHARLEKLG-FKNYVIVNNALINM 377
++ N T+ VLL A A + L+ G +HA + + G + V+N L+NM
Sbjct: 369 GA-RDSAAVNVDTYVVLLSAIAEFSTAEQGLRKGREVHAHVLRAGHIYRKIAVSNGLVNM 427
Query: 378 YSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTF 437
Y+K G ID + VF RD +WN++I +G + A+ + M+ P+N
Sbjct: 428 YAKCGAIDKACRVFQLMEARDRISWNTIITALDQNGYCEAAMMNYCLMRQNSIGPSNFAA 487
Query: 438 VGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTK 497
+ LS+CA L L+ G L+ + + +V +Y G + + +
Sbjct: 488 ISGLSSCAGLGLLAAG-QQLHCDAVKWGLYLDTSVSNALVKMYGECGRMSECWEIFNSMS 546
Query: 498 VKWDIVAWRTLL 509
D+V+W +++
Sbjct: 547 AH-DVVSWNSIM 557
>UniRef100_Q8W0Q5 Putative vegetative storage protein [Sorghum bicolor]
Length = 779
Score = 449 bits (1156), Expect = e-124
Identities = 252/713 (35%), Positives = 392/713 (54%), Gaps = 21/713 (2%)
Query: 5 WQGPMVVVNSRY-------LPPLEYIVK-LLKLSADAKCLTSGKAIHAQLLVRDQASKRS 56
W+GP Y +PP +Y +LK + L +G+ IHA +
Sbjct: 76 WRGPFHAAIDLYRSMLYFRVPPNKYTFPFVLKACSALADLCAGRTIHAHAAA---VGLHT 132
Query: 57 DVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLV 116
D+ +LI LY++C R A ++F KMP+RDV +WN ++ GY + G Y + +L+
Sbjct: 133 DLFVSTALIDLYIRCARFGPAANVFAKMPMRDVVAWNAMLAGYANHGMYHHAIA---HLL 189
Query: 117 SSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQ---FVKSALFNMY 173
PN ++L + G + +G H Y + L + + +AL +MY
Sbjct: 190 DMQDRGGLRPNASTLVSLLPLLAQHGALFQGTSVHAYCLRAYLDQNEEQVLIGTALLDMY 249
Query: 174 ARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA 233
A+C H+ A +V + + +E +++L+ V R EA + + M E M + +A
Sbjct: 250 AKCKHLVYACRVFHGMTVRNE---VTWSALIGGFVLCDRMTEAFNLFKDMLVEGMCFLSA 306
Query: 234 TYVG-VMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDG 292
T V + +CA + DL++G ++HA L + G+ D G+ L+ MY K + A +FD
Sbjct: 307 TSVASALRVCASLADLRMGTQLHALLAKSGIHADLTAGNSLLSMYAKAGLINEATMLFDE 366
Query: 293 LQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHG 352
+ ++ + + +L+S Y+QN EE + M + P+ T L+ AC+ +A L+HG
Sbjct: 367 IAIKDTISYGALLSGYVQNGKAEEAFLVFKKMQACNVQPDIATMVSLIPACSHLAALQHG 426
Query: 353 DLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHH 412
H + G + N+LI+MY+K G ID S VF RD+ +WN+MI GY H
Sbjct: 427 RCSHGSVIIRGLALETSICNSLIDMYAKCGRIDLSRQVFDKMPARDIVSWNTMIAGYGIH 486
Query: 413 GLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEH 472
GLG+EA +F MK+ G P++VTF+ +++AC+H LV +G + + I P +EH
Sbjct: 487 GLGKEATTLFLSMKNQGFEPDDVTFICLIAACSHSGLVTEGKHWFDTMTHKYGILPRMEH 546
Query: 473 YTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMD 532
Y CMV L R G L++A F+++ +K D+ W LL +CR+H+N DLGK+++ I ++
Sbjct: 547 YICMVDLLARGGFLDEAYQFIQSMPLKADVRVWGALLGACRIHKNIDLGKQVSRMIQKLG 606
Query: 533 PQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKH 592
P+ G + LLSN+ + A +D E+R + K + KK PG SW+EI H FV H
Sbjct: 607 PEGTGNFVLLSNIFSAAGRFDEAAEVRIIQKVKGFKKSPGCSWIEINGSLHAFVGGDQSH 666
Query: 593 PESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPS 652
P S +I ++ +L IK LGY + S VL D+E+E+KE L HSEKLAIA+G++ +
Sbjct: 667 PCSPDIYHELDNILIDIKKLGYQADTSFVLQDLEEEEKEKALLYHSEKLAIAFGVLSLNE 726
Query: 653 PAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
I + KNLR+C DCHTA+K ++ V NR IIVRDANRFHHF++G C+C D W
Sbjct: 727 DKTIFVTKNLRVCGDCHTAIKYMTLVRNRTIIVRDANRFHHFKNGQCSCGDFW 779
Score = 186 bits (472), Expect = 2e-45
Identities = 134/473 (28%), Positives = 229/473 (48%), Gaps = 19/473 (4%)
Query: 68 YVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPN 127
++ G+L LAR +FD++P D ++N L+ Y G + + L+++++ PN
Sbjct: 43 HIARGQLALARQVFDRIPAPDARAYNALIRAYSWRGPFHAAIDLYRSMLYFRVP----PN 98
Query: 128 EYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLN 187
+Y F VL +CS + G H + GL + FV +AL ++Y RC A V
Sbjct: 99 KYTFPFVLKACSALADLCAGRTIHAHAAAVGLHTDLFVSTALIDLYIRCARFGPAANVF- 157
Query: 188 TESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA-TYVGVMGLCAQIG 246
+ D+ +N++L G + AI L M + + NA T V ++ L AQ G
Sbjct: 158 --AKMPMRDVVAWNAMLAGYANHGMYHHAIAHLLDMQDRGGLRPNASTLVSLLPLLAQHG 215
Query: 247 DLQLGLRVHARLLRGGLLFDE---FVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTS 303
L G VHA LR L +E +G+ L+DMY KC H++ A +VF G+ RN V W++
Sbjct: 216 ALFQGTSVHAYCLRAYLDQNEEQVLIGTALLDMYAKCKHLVYACRVFHGMTVRNEVTWSA 275
Query: 304 LMSAYLQNRYFEETLDLLTCMDQED-TLPNECTFKVLLGACAGIALLKHGDLLHARLEKL 362
L+ ++ E +L M E + + L CA +A L+ G LHA L K
Sbjct: 276 LIGGFVLCDRMTEAFNLFKDMLVEGMCFLSATSVASALRVCASLADLRMGTQLHALLAKS 335
Query: 363 GFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVF 422
G + N+L++MY+K+G I+ + +F + +D ++ +++ GY +G +EA VF
Sbjct: 336 GIHADLTAGNSLLSMYAKAGLINEATMLFDEIAIKDTISYGALLSGYVQNGKAEEAFLVF 395
Query: 423 QDMKSAGESPNNVTFVGVLSACAHLALVDKG-LDYLYNEMKNHKIEPGLEHYTCMVVLYC 481
+ M++ P+ T V ++ AC+HLA + G + ++ +E + ++ +Y
Sbjct: 396 KKMQACNVQPDIATMVSLIPACSHLAALQHGRCSHGSVIIRGLALETSI--CNSLIDMYA 453
Query: 482 RAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQ 534
+ G ++ + + DIV+W T++ +H LGK L M Q
Sbjct: 454 KCGRIDLSRQVFDKMPAR-DIVSWNTMIAGYGIH---GLGKEATTLFLSMKNQ 502
Score = 89.7 bits (221), Expect = 3e-16
Identities = 74/275 (26%), Positives = 118/275 (42%), Gaps = 30/275 (10%)
Query: 286 ARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAG 345
AR+VFD + + + +L+ AY F +DL M PN+ TF +L AC+
Sbjct: 52 ARQVFDRIPAPDARAYNALIRAYSWRGPFHAAIDLYRSMLYFRVPPNKYTFPFVLKACSA 111
Query: 346 IALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSM 405
+A L G +HA +G + V+ ALI++Y + + NVF+ RDV WN+M
Sbjct: 112 LADLCAGRTIHAHAAAVGLHTDLFVSTALIDLYIRCARFGPAANVFAKMPMRDVVAWNAM 171
Query: 406 ICGYSHHGLGQEALRVFQDMKS-AGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNH 464
+ GY++HG+ A+ DM+ G PN T V +L A + +G
Sbjct: 172 LAGYANHGMYHHAIAHLLDMQDRGGLRPNASTLVSLLPLLAQHGALFQGTS--------- 222
Query: 465 KIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVA-WRTLLNSCRVHRNYDLGKR 523
V YC L++ E + D+ A + L+ +CRV +
Sbjct: 223 ------------VHAYCLRAYLDQNEEQVLIGTALLDMYAKCKHLVYACRVFHGMTVRNE 270
Query: 524 IAESILQMDPQDMGTYTLLSNMHATANMW-DGVVE 557
+ S L +G + L M N++ D +VE
Sbjct: 271 VTWSAL------IGGFVLCDRMTEAFNLFKDMLVE 299
>UniRef100_Q9ZUW3 Putative selenium-binding protein [Arabidopsis thaliana]
Length = 868
Score = 444 bits (1142), Expect = e-123
Identities = 239/646 (36%), Positives = 377/646 (57%), Gaps = 12/646 (1%)
Query: 62 NSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTT 121
NSLI+LY+KCG +R AR +FDK ++ V +WN +++GY +G LE L +F ++
Sbjct: 233 NSLINLYLKCGNVRKARILFDKTEVKSVVTWNSMISGYAANGLDLEALGMFYSM----RL 288
Query: 122 QHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNL 181
+ +E F +V+ C++ +R Q H + KYG + Q +++AL Y++C +
Sbjct: 289 NYVRLSESSFASVIKLCANLKELRFTEQLHCSVVKYGFLFDQNIRTALMVAYSKCTAMLD 348
Query: 182 ALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGL 241
AL++ + G ++ + ++++ +++ EEA+ + M + + + TY ++
Sbjct: 349 ALRLF--KEIGCVGNVVSWTAMISGFLQNDGKEEAVDLFSEMKRKGVRPNEFTYSVILTA 406
Query: 242 CAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
I + VHA++++ VG+ L+D Y K V A KVF G+ ++++V W
Sbjct: 407 LPVISPSE----VHAQVVKTNYERSSTVGTALLDAYVKLGKVEEAAKVFSGIDDKDIVAW 462
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGI-ALLKHGDLLHARLE 360
+++++ Y Q E + + + + PNE TF +L CA A + G H
Sbjct: 463 SAMLAGYAQTGETEAAIKMFGELTKGGIKPNEFTFSSILNVCAATNASMGQGKQFHGFAI 522
Query: 361 KLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALR 420
K + + V++AL+ MY+K G+I+S+ VF +D+ +WNSMI GY+ HG +AL
Sbjct: 523 KSRLDSSLCVSSALLTMYAKKGNIESAEEVFKRQREKDLVSWNSMISGYAQHGQAMKALD 582
Query: 421 VFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLY 480
VF++MK + VTF+GV +AC H LV++G Y +++ KI P EH +CMV LY
Sbjct: 583 VFKEMKKRKVKMDGVTFIGVFAACTHAGLVEEGEKYFDIMVRDCKIAPTKEHNSCMVDLY 642
Query: 481 CRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYT 540
RAG LEKA ++ WRT+L +CRVH+ +LG+ AE I+ M P+D Y
Sbjct: 643 SRAGQLEKAMKVIENMPNPAGSTIWRTILAACRVHKKTELGRLAAEKIIAMKPEDSAAYV 702
Query: 541 LLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINK 600
LLSNM+A + W ++RK+M ERN+KKEPG SW+E++N + F++ HP ++I
Sbjct: 703 LLSNMYAESGDWQERAKVRKLMNERNVKKEPGYSWIEVKNKTYSFLAGDRSHPLKDQIYM 762
Query: 601 KVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIK 660
K++ L +K LGY P+ S VL D++DE KE L HSE+LAIA+GL+ P +P+ IIK
Sbjct: 763 KLEDLSTRLKDLGYEPDTSYVLQDIDDEHKEAVLAQHSERLAIAFGLIATPKGSPLLIIK 822
Query: 661 NLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHF-RDGSCTCADHW 705
NLR+C DCH +KLI+K+ R I+VRD+NRFHHF DG C+C D W
Sbjct: 823 NLRVCGDCHLVIKLIAKIEEREIVVRDSNRFHHFSSDGVCSCGDFW 868
Score = 209 bits (531), Expect = 3e-52
Identities = 140/513 (27%), Positives = 263/513 (50%), Gaps = 22/513 (4%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
+LK+SA G+ +H Q + + DV SL+ Y+K + R +FD+M
Sbjct: 99 VLKVSATLCDELFGRQLHCQCI---KFGFLDDVSVGTSLVDTYMKGSNFKDGRKVFDEMK 155
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
R+V +W L++GY + EVL LF + + T PN + F L + G
Sbjct: 156 ERNVVTWTTLISGYARNSMNDEVLTLFMRMQNEGTQ----PNSFTFAAALGVLAEEGVGG 211
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLN 205
G+Q H + K GL V ++L N+Y +C +V A + + + + +NS+++
Sbjct: 212 RGLQVHTVVVKNGLDKTIPVSNSLINLYLKCGNVRKARILFDKT---EVKSVVTWNSMIS 268
Query: 206 VLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLF 265
+G EA+G+ M + +++ V+ LCA + +L+ ++H +++ G LF
Sbjct: 269 GYAANGLDLEALGMFYSMRLNYVRLSESSFASVIKLCANLKELRFTEQLHCSVVKYGFLF 328
Query: 266 DEFVGSKLIDMYGKCCHVLNARKVFDGLQ-NRNVVVWTSLMSAYLQNRYFEETLDLLTCM 324
D+ + + L+ Y KC +L+A ++F + NVV WT+++S +LQN EE +DL + M
Sbjct: 329 DQNIRTALMVAYSKCTAMLDALRLFKEIGCVGNVVSWTAMISGFLQNDGKEEAVDLFSEM 388
Query: 325 DQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
++ PNE T+ V+L A I+ + +HA++ K ++ V AL++ Y K G +
Sbjct: 389 KRKGVRPNEFTYSVILTALPVISPSE----VHAQVVKTNYERSSTVGTALLDAYVKLGKV 444
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
+ + VFS +D+ W++M+ GY+ G + A+++F ++ G PN TF +L+ C
Sbjct: 445 EEAAKVFSGIDDKDIVAWSAMLAGYAQTGETEAAIKMFGELTKGGIKPNEFTFSSILNVC 504
Query: 445 AHL-ALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIV 503
A A + +G + +K+ +++ L + ++ +Y + G +E AE K + K D+V
Sbjct: 505 AATNASMGQGKQFHGFAIKS-RLDSSLCVSSALLTMYAKKGNIESAEEVFKRQREK-DLV 562
Query: 504 AWRTLLNSCRVH----RNYDLGKRIAESILQMD 532
+W ++++ H + D+ K + + ++MD
Sbjct: 563 SWNSMISGYAQHGQAMKALDVFKEMKKRKVKMD 595
Score = 146 bits (369), Expect = 2e-33
Identities = 95/311 (30%), Positives = 154/311 (48%), Gaps = 1/311 (0%)
Query: 200 YNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLL 259
Y SLL GR +EA + + M D + + V+ + A + D G ++H + +
Sbjct: 61 YISLLFGFSRDGRTQEAKRLFLNIHRLGMEMDCSIFSSVLKVSATLCDELFGRQLHCQCI 120
Query: 260 RGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLD 319
+ G L D VG+ L+D Y K + + RKVFD ++ RNVV WT+L+S Y +N +E L
Sbjct: 121 KFGFLDDVSVGTSLVDTYMKGSNFKDGRKVFDEMKERNVVTWTTLISGYARNSMNDEVLT 180
Query: 320 LLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYS 379
L M E T PN TF LG A + G +H + K G + V+N+LIN+Y
Sbjct: 181 LFMRMQNEGTQPNSFTFAAALGVLAEEGVGGRGLQVHTVVVKNGLDKTIPVSNSLINLYL 240
Query: 380 KSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVG 439
K G++ + +F T + V TWNSMI GY+ +GL EAL +F M+ + +F
Sbjct: 241 KCGNVRKARILFDKTEVKSVVTWNSMISGYAANGLDLEALGMFYSMRLNYVRLSESSFAS 300
Query: 440 VLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVK 499
V+ CA+L + + + L+ + + T ++V Y + + A K
Sbjct: 301 VIKLCANLKEL-RFTEQLHCSVVKYGFLFDQNIRTALMVAYSKCTAMLDALRLFKEIGCV 359
Query: 500 WDIVAWRTLLN 510
++V+W +++
Sbjct: 360 GNVVSWTAMIS 370
Score = 140 bits (353), Expect = 1e-31
Identities = 114/442 (25%), Positives = 203/442 (45%), Gaps = 14/442 (3%)
Query: 69 VKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNE 128
V RL A ++FDK P RD S+ L+ G+ G E LF N+
Sbjct: 38 VSSSRLYNAHNLFDKSPGRDRESYISLLFGFSRDGRTQEAKRLFLNIHRLGMEMDCS--- 94
Query: 129 YVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
+F++VL + G Q H K+G + V ++L + Y + + +V +
Sbjct: 95 -IFSSVLKVSATLCDELFGRQLHCQCIKFGFLDDVSVGTSLVDTYMKGSNFKDGRKVFDE 153
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
E ++ + +L++ + +E + + RM NE ++ T+ +G+ A+ G
Sbjct: 154 MK---ERNVVTWTTLISGYARNSMNDEVLTLFMRMQNEGTQPNSFTFAAALGVLAEEGVG 210
Query: 249 QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAY 308
GL+VH +++ GL V + LI++Y KC +V AR +FD + ++VV W S++S Y
Sbjct: 211 GRGLQVHTVVVKNGLDKTIPVSNSLINLYLKCGNVRKARILFDKTEVKSVVTWNSMISGY 270
Query: 309 LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYV 368
N E L + M +E +F ++ CA + L+ + LH + K GF
Sbjct: 271 AANGLDLEALGMFYSMRLNYVRLSESSFASVIKLCANLKELRFTEQLHCSVVKYGFLFDQ 330
Query: 369 IVNNALINMYSKSGDIDSSYNVFSDTMC-RDVTTWNSMICGYSHHGLGQEALRVFQDMKS 427
+ AL+ YSK + + +F + C +V +W +MI G+ + +EA+ +F +MK
Sbjct: 331 NIRTALMVAYSKCTAMLDALRLFKEIGCVGNVVSWTAMISGFLQNDGKEEAVDLFSEMKR 390
Query: 428 AGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLE 487
G PN T+ +L+A ++ + ++ ++ E T ++ Y + G +E
Sbjct: 391 KGVRPNEFTYSVILTALPVISPSE-----VHAQVVKTNYERSSTVGTALLDAYVKLGKVE 445
Query: 488 KAENFMKTTKVKWDIVAWRTLL 509
+A K DIVAW +L
Sbjct: 446 EAAKVFSGIDDK-DIVAWSAML 466
>UniRef100_Q60EM0 Unknow protein [Oryza sativa]
Length = 874
Score = 441 bits (1134), Expect = e-122
Identities = 238/681 (34%), Positives = 388/681 (56%), Gaps = 14/681 (2%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
+L A L G+ +HAQ + + RS V NSL+++Y KCG + A+S+F+ M
Sbjct: 207 VLSAVASQGALDLGQRVHAQSV---KFGCRSSVFVCNSLMNMYAKCGLVEDAKSVFNWME 263
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
RD+ SWN LM G + LE L LF S + + TV+ C++ ++
Sbjct: 264 TRDMVSWNTLMAGLQLNECELEALQLFHE----SRATMGKMTQSTYATVIKLCANLKQLA 319
Query: 146 EGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLN 205
Q H + K+G V +AL + Y++C + AL + + +G ++ + ++++
Sbjct: 320 LARQLHSCVLKHGFHLTGNVMTALADAYSKCGELADALNIFSMTTGS--RNVVSWTAIIS 377
Query: 206 VLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLF 265
+++G A+ + RM + ++ + TY ++ I L ++HA++++
Sbjct: 378 GCIQNGDIPLAVVLFSRMREDRVMPNEFTYSAMLKASLSI----LPPQIHAQVIKTNYQH 433
Query: 266 DEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMD 325
FVG+ L+ Y K +A +F ++ ++VV W++++S + Q E L M
Sbjct: 434 IPFVGTALLASYSKFGSTEDALSIFKMIEQKDVVAWSAMLSCHAQAGDCEGATYLFNKMA 493
Query: 326 QEDTLPNECTFKVLLGACA-GIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
+ PNE T ++ ACA A + G HA K + + + V++AL++MYS+ G+I
Sbjct: 494 IQGIKPNEFTISSVIDACACPSAGVDQGRQFHAISIKYRYHDAICVSSALVSMYSRKGNI 553
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
DS+ VF RD+ +WNSMI GY+ HG +A+ F+ M+++G + VTF+ V+ C
Sbjct: 554 DSAQIVFERQTDRDLVSWNSMISGYAQHGYSMKAIETFRQMEASGIQMDGVTFLAVIMGC 613
Query: 445 AHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVA 504
H LV +G Y + +++HKI P +EHY CMV LY RAG L++ + ++ +
Sbjct: 614 THNGLVVEGQQYFDSMVRDHKINPTMEHYACMVDLYSRAGKLDETMSLIRDMPFPAGAMV 673
Query: 505 WRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKE 564
WRTLL +CRVH+N +LGK A+ +L ++P D TY LLSN++A A W E+RK+M
Sbjct: 674 WRTLLGACRVHKNVELGKFSADKLLSLEPHDSSTYVLLSNIYAAAGKWKERDEVRKLMDY 733
Query: 565 RNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHD 624
R +KKE G SW++I+N H F++ HP S++I KK++ ++ +K GY PN S VLHD
Sbjct: 734 RKVKKEAGCSWIQIKNKVHSFIAFDKSHPMSDQIYKKLKVIITRLKQDGYSPNTSFVLHD 793
Query: 625 VEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFII 684
+ ++QKE L HSE+LA+A+GL+ P P++I+KNLR+C DCH +K++S + +R II
Sbjct: 794 IAEDQKEAMLVAHSERLALAFGLIATPPGTPLQIVKNLRVCGDCHMVMKMVSMIEDREII 853
Query: 685 VRDANRFHHFRDGSCTCADHW 705
+RD +RFHHF G+C+C D W
Sbjct: 854 MRDCSRFHHFNGGACSCGDFW 874
Score = 184 bits (468), Expect = 6e-45
Identities = 133/484 (27%), Positives = 237/484 (48%), Gaps = 19/484 (3%)
Query: 55 RSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKN 114
R +V SL+ +Y+KCG + +F+ MP ++V +W L+TG H+ + EV+ LF
Sbjct: 132 RGEVSAGTSLVDMYMKCGSVCEGIEVFEGMPKKNVVTWTSLLTGCAHAQMHSEVMALFFR 191
Query: 115 LVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYA 174
+ + PN + F +VLS+ + G + G + H K+G S FV ++L NMYA
Sbjct: 192 M----RAEGIWPNPFTFASVLSAVASQGALDLGQRVHAQSVKFGCRSSVFVCNSLMNMYA 247
Query: 175 RCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNAT 234
+C V A V N + D+ +N+L+ L + EA+ + +T
Sbjct: 248 KCGLVEDAKSVFNWM---ETRDMVSWNTLMAGLQLNECELEALQLFHESRATMGKMTQST 304
Query: 235 YVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFD-GL 293
Y V+ LCA + L L ++H+ +L+ G V + L D Y KC + +A +F
Sbjct: 305 YATVIKLCANLKQLALARQLHSCVLKHGFHLTGNVMTALADAYSKCGELADALNIFSMTT 364
Query: 294 QNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGD 353
+RNVV WT+++S +QN + L + M ++ +PNE T+ +L A I
Sbjct: 365 GSRNVVSWTAIISGCIQNGDIPLAVVLFSRMREDRVMPNEFTYSAMLKASLSIL----PP 420
Query: 354 LLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHG 413
+HA++ K +++ V AL+ YSK G + + ++F +DV W++M+ ++ G
Sbjct: 421 QIHAQVIKTNYQHIPFVGTALLASYSKFGSTEDALSIFKMIEQKDVVAWSAMLSCHAQAG 480
Query: 414 LGQEALRVFQDMKSAGESPNNVTFVGVLSACA-HLALVDKGLDYLYNEMKNHKIEPGLEH 472
+ A +F M G PN T V+ ACA A VD+G + +K ++ +
Sbjct: 481 DCEGATYLFNKMAIQGIKPNEFTISSVIDACACPSAGVDQGRQFHAISIK-YRYHDAICV 539
Query: 473 YTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH----RNYDLGKRIAESI 528
+ +V +Y R G ++ A+ + + D+V+W ++++ H + + +++ S
Sbjct: 540 SSALVSMYSRKGNIDSAQ-IVFERQTDRDLVSWNSMISGYAQHGYSMKAIETFRQMEASG 598
Query: 529 LQMD 532
+QMD
Sbjct: 599 IQMD 602
Score = 132 bits (333), Expect = 3e-29
Identities = 87/299 (29%), Positives = 146/299 (48%), Gaps = 5/299 (1%)
Query: 215 EAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEF-VGSKL 273
+ V RR G ++ D+AT V+ C + D LG ++H ++ G E G+ L
Sbjct: 85 DQFSVARRGG---VLVDSATLSCVLKACRSVPDRVLGEQLHCLCVKCGHDRGEVSAGTSL 141
Query: 274 IDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNE 333
+DMY KC V +VF+G+ +NVV WTSL++ + E + L M E PN
Sbjct: 142 VDMYMKCGSVCEGIEVFEGMPKKNVVTWTSLLTGCAHAQMHSEVMALFFRMRAEGIWPNP 201
Query: 334 CTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSD 393
TF +L A A L G +HA+ K G ++ V V N+L+NMY+K G ++ + +VF+
Sbjct: 202 FTFASVLSAVASQGALDLGQRVHAQSVKFGCRSSVFVCNSLMNMYAKCGLVEDAKSVFNW 261
Query: 394 TMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKG 453
RD+ +WN+++ G + EAL++F + ++ T+ V+ CA+L +
Sbjct: 262 METRDMVSWNTLMAGLQLNECELEALQLFHESRATMGKMTQSTYATVIKLCANLKQLALA 321
Query: 454 LDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSC 512
L++ + H T + Y + G L A N T ++V+W +++ C
Sbjct: 322 RQ-LHSCVLKHGFHLTGNVMTALADAYSKCGELADALNIFSMTTGSRNVVSWTAIISGC 379
Score = 127 bits (320), Expect = 9e-28
Identities = 121/442 (27%), Positives = 195/442 (43%), Gaps = 26/442 (5%)
Query: 77 ARSMFDKMPLRDV-FSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVL 135
AR D++P RD N ++ Y G LEVL F S + + + VL
Sbjct: 51 ARYPLDEIPRRDAAVGANRVLFDYARRGMVLEVLDQF----SVARRGGVLVDSATLSCVL 106
Query: 136 SSCSHSGRVREGMQCHGYLFKYGLVSYQF-VKSALFNMYARCLHVNLALQVLNTESGGDE 194
+C G Q H K G + ++L +MY +C V ++V G +
Sbjct: 107 KACRSVPDRVLGEQLHCLCVKCGHDRGEVSAGTSLVDMYMKCGSVCEGIEVFE---GMPK 163
Query: 195 NDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA-TYVGVMGLCAQIGDLQLGLR 253
++ + SLL + E + + RM E +W N T+ V+ A G L LG R
Sbjct: 164 KNVVTWTSLLTGCAHAQMHSEVMALFFRMRAEG-IWPNPFTFASVLSAVASQGALDLGQR 222
Query: 254 VHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRY 313
VHA+ ++ G FV + L++MY KC V +A+ VF+ ++ R++V W +LM+ N
Sbjct: 223 VHAQSVKFGCRSSVFVCNSLMNMYAKCGLVEDAKSVFNWMETRDMVSWNTLMAGLQLNEC 282
Query: 314 FEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNA 373
E L L + T+ ++ CA + L LH+ + K GF V A
Sbjct: 283 ELEALQLFHESRATMGKMTQSTYATVIKLCANLKQLALARQLHSCVLKHGFHLTGNVMTA 342
Query: 374 LINMYSKSGDIDSSYNVFS-DTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESP 432
L + YSK G++ + N+FS T R+V +W ++I G +G A+ +F M+ P
Sbjct: 343 LADAYSKCGELADALNIFSMTTGSRNVVSWTAIISGCIQNGDIPLAVVLFSRMREDRVMP 402
Query: 433 NNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY----TCMVVLYCRAGLLEK 488
N T+ +L A L L ++ I+ +H T ++ Y + G E
Sbjct: 403 NEFTYSAMLKA---------SLSILPPQIHAQVIKTNYQHIPFVGTALLASYSKFGSTED 453
Query: 489 AENFMKTTKVKWDIVAWRTLLN 510
A + K + K D+VAW +L+
Sbjct: 454 ALSIFKMIEQK-DVVAWSAMLS 474
>UniRef100_Q9LTV8 Selenium-binding protein-like [Arabidopsis thaliana]
Length = 694
Score = 440 bits (1132), Expect = e-122
Identities = 226/666 (33%), Positives = 379/666 (55%), Gaps = 9/666 (1%)
Query: 40 KAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGY 99
K IHA+LLV + + LIH G + AR +FD +P +F WN ++ GY
Sbjct: 38 KQIHARLLV---LGLQFSGFLITKLIHASSSFGDITFARQVFDDLPRPQIFPWNAIIRGY 94
Query: 100 MHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGL 159
+ ++ + L+++ N+ + + P+ + F +L +CS ++ G H +F+ G
Sbjct: 95 SRNNHFQDALLMYSNMQLARVS----PDSFTFPHLLKACSGLSHLQMGRFVHAQVFRLGF 150
Query: 160 VSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGV 219
+ FV++ L +YA+C + A V E I + ++++ ++G EA+ +
Sbjct: 151 DADVFVQNGLIALYAKCRRLGSARTVFEGLPL-PERTIVSWTAIVSAYAQNGEPMEALEI 209
Query: 220 LRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGK 279
+M + D V V+ + DL+ G +HA +++ GL + + L MY K
Sbjct: 210 FSQMRKMDVKPDWVALVSVLNAFTCLQDLKQGRSIHASVVKMGLEIEPDLLISLNTMYAK 269
Query: 280 CCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVL 339
C V A+ +FD +++ N+++W +++S Y +N Y E +D+ M +D P+ +
Sbjct: 270 CGQVATAKILFDKMKSPNLILWNAMISGYAKNGYAREAIDMFHEMINKDVRPDTISITSA 329
Query: 340 LGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDV 399
+ ACA + L+ ++ + + +++ V +++ALI+M++K G ++ + VF T+ RDV
Sbjct: 330 ISACAQVGSLEQARSMYEYVGRSDYRDDVFISSALIDMFAKCGSVEGARLVFDRTLDRDV 389
Query: 400 TTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYN 459
W++MI GY HG +EA+ +++ M+ G PN+VTF+G+L AC H +V +G + +N
Sbjct: 390 VVWSAMIVGYGLHGRAREAISLYRAMERGGVHPNDVTFLGLLMACNHSGMVREGW-WFFN 448
Query: 460 EMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYD 519
M +HKI P +HY C++ L RAG L++A +K V+ + W LL++C+ HR+ +
Sbjct: 449 RMADHKINPQQQHYACVIDLLGRAGHLDQAYEVIKCMPVQPGVTVWGALLSACKKHRHVE 508
Query: 520 LGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIR 579
LG+ A+ + +DP + G Y LSN++A A +WD V E+R MKE+ + K+ G SW+E+R
Sbjct: 509 LGEYAAQQLFSIDPSNTGHYVQLSNLYAAARLWDRVAEVRVRMKEKGLNKDVGCSWVEVR 568
Query: 580 NVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSE 639
F HP EI ++V+ + + +K G+V N A LHD+ DE+ E L HSE
Sbjct: 569 GRLEAFRVGDKSHPRYEEIERQVEWIESRLKEGGFVANKDASLHDLNDEEAEETLCSHSE 628
Query: 640 KLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSC 699
++AIAYGL+ P P+RI KNLR C +CH A KLISK+ +R I+VRD NRFHHF+DG C
Sbjct: 629 RIAIAYGLISTPQGTPLRITKNLRACVNCHAATKLISKLVDREIVVRDTNRFHHFKDGVC 688
Query: 700 TCADHW 705
+C D+W
Sbjct: 689 SCGDYW 694
>UniRef100_Q84SE9 P0020E09.15 protein [Oryza sativa]
Length = 813
Score = 440 bits (1131), Expect = e-122
Identities = 248/696 (35%), Positives = 384/696 (54%), Gaps = 24/696 (3%)
Query: 22 YIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMF 81
+ +K AD C G+AIH + A ++D+ +L+ +YVKC L A +F
Sbjct: 130 FALKACSALADHHC---GRAIHRHAI---HAGLQADLFVSTALLDMYVKCACLPDAAHIF 183
Query: 82 DKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHAC-PNEYVFTTVLSSCSH 140
MP RD+ +WN ++ GY H G Y + +L+S H PN +L +
Sbjct: 184 ATMPARDLVAWNAMLAGYAHHGMYHHAVA---HLLSMQMQMHRLRPNASTLVALLPLLAQ 240
Query: 141 SGRVREGMQCHGYLFKYGLVSYQFVKS----------ALFNMYARCLHVNLALQVLNTES 190
G + +G H Y + L + KS AL +MYA+C + A +V +
Sbjct: 241 QGALAQGTSVHAYCIRACLHPNRNSKSKLTDGVLLGTALLDMYAKCGSLLYARRVFDAMP 300
Query: 191 GGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVG-VMGLCAQIGDLQ 249
+E +++L+ V R +A + + M + + + + T + + CA + L+
Sbjct: 301 ARNE---VTWSALIGGFVLCSRMTQAFLLFKAMLAQGLCFLSPTSIASALRACASLDHLR 357
Query: 250 LGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYL 309
+G ++HA L + G+ D G+ L+ MY K + A +FD + ++ V +++L+S Y+
Sbjct: 358 MGEQLHALLAKSGVHADLTAGNSLLSMYAKAGLIDQAIALFDEMAVKDTVSYSALVSGYV 417
Query: 310 QNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVI 369
QN EE + M + P+ T L+ AC+ +A L+HG H + G +
Sbjct: 418 QNGRAEEAFLVFKKMQACNVEPDAATMVSLIPACSHLAALQHGRCSHGSVIIRGLASETS 477
Query: 370 VNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAG 429
+ NALI+MY+K G ID S VF+ RD+ +WN+MI GY HGLG+EA +F +M + G
Sbjct: 478 ICNALIDMYAKCGRIDLSRQVFNMMPSRDIVSWNTMIAGYGIHGLGKEATALFLEMNNLG 537
Query: 430 ESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKA 489
P+ VTF+ +LSAC+H LV +G + + + + P +EHY CMV L R G L++A
Sbjct: 538 FPPDGVTFICLLSACSHSGLVIEGKHWFHVMGHGYGLTPRMEHYICMVDLLSRGGFLDEA 597
Query: 490 ENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATA 549
F+++ ++ D+ W LL +CRV++N DLGK+++ I ++ P+ G + LLSN+++ A
Sbjct: 598 YEFIQSMPLRADVRVWVALLGACRVYKNIDLGKKVSRMIQELGPEGTGNFVLLSNIYSAA 657
Query: 550 NMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMI 609
+D E+R + K + KK PG SW+EI H FV HP+S EI +++ +L I
Sbjct: 658 GRFDEAAEVRIIQKVQGFKKSPGCSWIEINGSLHAFVGGDQSHPQSPEIYRELDNILVGI 717
Query: 610 KPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCH 669
K LGY P+ S VL D+E+E+KE L CHSEKLAIAYG++ + I + KNLR+C DCH
Sbjct: 718 KKLGYQPDTSFVLQDLEEEEKEKALICHSEKLAIAYGILSLSEDKTIFVTKNLRVCGDCH 777
Query: 670 TAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
T +K IS V R IIVRDANRFHHF++G C+C D W
Sbjct: 778 TVIKHISLVKRRAIIVRDANRFHHFKNGQCSCGDFW 813
Score = 178 bits (451), Expect = 6e-43
Identities = 139/498 (27%), Positives = 231/498 (45%), Gaps = 39/498 (7%)
Query: 68 YVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEV--LVLFKNLVSSSTTQHAC 125
++ G L A +FD++P DV ++NDL+ Y S L L++ ++
Sbjct: 67 HIASGHLSRAHHLFDQIPSPDVRTYNDLIRAYSSSSPTAAADGLHLYRRMLR----HRVA 122
Query: 126 PNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQV 185
PN Y F L +CS G H + GL + FV +AL +MY +C + A +
Sbjct: 123 PNNYTFPFALKACSALADHHCGRAIHRHAIHAGLQADLFVSTALLDMYVKCACLPDAAHI 182
Query: 186 LNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNEC--MVWDNATYVGVMGLCA 243
T D+ +N++L G + A+ L M + + + +T V ++ L A
Sbjct: 183 FATMPA---RDLVAWNAMLAGYAHHGMYHHAVAHLLSMQMQMHRLRPNASTLVALLPLLA 239
Query: 244 QIGDLQLGLRVHARLLRGGL---------LFD-EFVGSKLIDMYGKCCHVLNARKVFDGL 293
Q G L G VHA +R L L D +G+ L+DMY KC +L AR+VFD +
Sbjct: 240 QQGALAQGTSVHAYCIRACLHPNRNSKSKLTDGVLLGTALLDMYAKCGSLLYARRVFDAM 299
Query: 294 QNRNVVVWTSLMSAY-LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHG 352
RN V W++L+ + L +R + L + Q + + L ACA + L+ G
Sbjct: 300 PARNEVTWSALIGGFVLCSRMTQAFLLFKAMLAQGLCFLSPTSIASALRACASLDHLRMG 359
Query: 353 DLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHH 412
+ LHA L K G + N+L++MY+K+G ID + +F + +D ++++++ GY +
Sbjct: 360 EQLHALLAKSGVHADLTAGNSLLSMYAKAGLIDQAIALFDEMAVKDTVSYSALVSGYVQN 419
Query: 413 GLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEH 472
G +EA VF+ M++ P+ T V ++ AC+HLA + G I GL
Sbjct: 420 GRAEEAFLVFKKMQACNVEPDAATMVSLIPACSHLAALQHG-----RCSHGSVIIRGLAS 474
Query: 473 YT----CMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESI 528
T ++ +Y + G ++ + + DIV+W T++ +H LGK
Sbjct: 475 ETSICNALIDMYAKCGRIDLSRQVFNMMPSR-DIVSWNTMIAGYGIH---GLGKEATALF 530
Query: 529 LQMD----PQDMGTYTLL 542
L+M+ P D T+ L
Sbjct: 531 LEMNNLGFPPDGVTFICL 548
>UniRef100_Q6L4I3 Hypothetical protein OSJNBa0074P11.10 [Oryza sativa]
Length = 822
Score = 440 bits (1131), Expect = e-122
Identities = 240/674 (35%), Positives = 383/674 (56%), Gaps = 22/674 (3%)
Query: 35 CLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWND 94
CL G + LV +D+ ++LI + + G L AR +FD + + V W
Sbjct: 168 CLVGGVVLG---LVHKMGLWGTDIAVGSALIDMLARNGDLASARKVFDGLIEKTVVVWTL 224
Query: 95 LMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYL 154
L++ Y+ E + +F + + P+ Y ++++S+C+ G VR G+Q H
Sbjct: 225 LISRYVQGECAEEAVEIFLDFLEDGFE----PDRYTMSSMISACTELGSVRLGLQLHSLA 280
Query: 155 FKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTE-SGGDENDIFLYNSLLNVLVESGRW 213
+ G S V L +MYA+ ++ A+ N +ND+ + +L++ V+SG
Sbjct: 281 LRMGFASDACVSCGLVDMYAKS-NIEQAMDYANKVFERMRKNDVISWTALISGYVQSGVQ 339
Query: 214 EEAIGVL-RRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSK 272
E + VL M NE + ++ TY ++ CA I D G +VHA +++ VG+
Sbjct: 340 ENKVMVLFGEMLNESIKPNHITYSSILKACANISDHDSGRQVHAHVIKSNQAAAHTVGNA 399
Query: 273 LIDMYGKCCHVLNARKVFDGLQNRNVVVW-TSLMSAYLQNRYFEETLDLLTCMDQEDTLP 331
L+ MY + + AR+VF+ L R+++ T A L +R + + D
Sbjct: 400 LVSMYAESGCMEEARRVFNQLYERSMISCITEGRDAPLDHR-----------IGRMDMGI 448
Query: 332 NECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVF 391
+ TF L+ A A + +L G LHA K GF + V+N+L++MYS+ G ++ + F
Sbjct: 449 SSSTFASLISAAASVGMLTKGQQLHAMTLKAGFGSDRFVSNSLVSMYSRCGYLEDACRSF 508
Query: 392 SDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVD 451
++ R+V +W SMI G + HG + AL +F DM G PN+VT++ VLSAC+H+ LV
Sbjct: 509 NELKDRNVISWTSMISGLAKHGYAERALSLFHDMILTGVKPNDVTYIAVLSACSHVGLVR 568
Query: 452 KGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNS 511
+G +Y + ++H + P +EHY CMV L R+GL+++A F+ +K D + W+TLL +
Sbjct: 569 EGKEYFRSMQRDHGLIPRMEHYACMVDLLARSGLVKEALEFINEMPLKADALVWKTLLGA 628
Query: 512 CRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEP 571
CR H N ++G+ A+++++++P+D Y LLSN++A A +WD V IR M++ N+ KE
Sbjct: 629 CRSHDNIEVGEIAAKNVIELEPRDPAPYVLLSNLYADAGLWDEVARIRSAMRDNNLNKET 688
Query: 572 GASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKE 631
G SW+E+ N H F + HP + +I K+ L+ IK +GYVP+ S VLHD+ DE KE
Sbjct: 689 GLSWMEVENTTHEFRAGDTSHPRAQDIYGKLDTLVGEIKGMGYVPDTSIVLHDMSDELKE 748
Query: 632 GYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRF 691
YL HSEK+A+A+GL+ +P PIRI KNLR+C DCH+A+K +SK T R II+RD+NRF
Sbjct: 749 QYLLQHSEKIAVAFGLITTSAPKPIRIFKNLRVCADCHSAIKYMSKATRREIILRDSNRF 808
Query: 692 HHFRDGSCTCADHW 705
H +DG C+C ++W
Sbjct: 809 HRMKDGECSCGEYW 822
Score = 155 bits (391), Expect = 5e-36
Identities = 149/553 (26%), Positives = 250/553 (44%), Gaps = 34/553 (6%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKM- 84
LL +A A L G+A+H +LL D + D V NSL+ LY +CG + AR++FD M
Sbjct: 54 LLAAAARAGDLRLGRALHRRLLRGDLLDR--DAVVANSLLTLYSRCGAVASARNVFDGMR 111
Query: 85 PLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSC-SHSGR 143
LRD+ SW + + +G E L+L ++ S PN Y V +C H
Sbjct: 112 GLRDIVSWTAMASCLARNGAERESLLLIGEMLESG----LLPNAYTLCAVAHACFPHELY 167
Query: 144 VREGMQCHGYLFKYGLVSYQF-VKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNS 202
G G + K GL V SAL +M AR + A +V + G E + ++
Sbjct: 168 CLVGGVVLGLVHKMGLWGTDIAVGSALIDMLARNGDLASARKVFD---GLIEKTVVVWTL 224
Query: 203 LLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGG 262
L++ V+ EEA+ + + D T ++ C ++G ++LGL++H+ LR G
Sbjct: 225 LISRYVQGECAEEAVEIFLDFLEDGFEPDRYTMSSMISACTELGSVRLGLQLHSLALRMG 284
Query: 263 LLFDEFVGSKLIDMYGKC---CHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFE-ETL 318
D V L+DMY K + A KVF+ ++ +V+ WT+L+S Y+Q+ E + +
Sbjct: 285 FASDACVSCGLVDMYAKSNIEQAMDYANKVFERMRKNDVISWTALISGYVQSGVQENKVM 344
Query: 319 DLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMY 378
L M E PN T+ +L ACA I+ G +HA + K V NAL++MY
Sbjct: 345 VLFGEMLNESIKPNHITYSSILKACANISDHDSGRQVHAHVIKSNQAAAHTVGNALVSMY 404
Query: 379 SKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFV 438
++SG ++ + VF+ R + S I L R+ + S+ TF
Sbjct: 405 AESGCMEEARRVFNQLYERSMI---SCITEGRDAPLDHRIGRMDMGISSS-------TFA 454
Query: 439 GVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY--TCMVVLYCRAGLLEKAENFMKTT 496
++SA A + ++ KG K G + + +V +Y R G LE A
Sbjct: 455 SLISAAASVGMLTKGQQL---HAMTLKAGFGSDRFVSNSLVSMYSRCGYLEDACRSFNEL 511
Query: 497 KVKWDIVAWRTLLNSCRVHRNYDLGKRIAESIL--QMDPQDMGTYTLLSNMHATANMWDG 554
K + ++++W ++++ H + + ++ + P D+ +LS + +G
Sbjct: 512 KDR-NVISWTSMISGLAKHGYAERALSLFHDMILTGVKPNDVTYIAVLSACSHVGLVREG 570
Query: 555 VVEIRKMMKERNI 567
R M ++ +
Sbjct: 571 KEYFRSMQRDHGL 583
>UniRef100_Q6ETD1 Putative pentatricopeptide (PPR) repeat-containing protein-like
protein [Oryza sativa]
Length = 751
Score = 440 bits (1131), Expect = e-122
Identities = 244/704 (34%), Positives = 376/704 (52%), Gaps = 62/704 (8%)
Query: 61 LNSLIHLYVKCGRLRLARSMFDKMP-------------------------------LRDV 89
LN L+ Y K GRL AR +FD+MP RD
Sbjct: 51 LNHLLTAYAKSGRLARARRVFDEMPDPNLFTRNALLSALAHSRLVPDMERLFASMPERDA 110
Query: 90 FSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQ 149
S+N L+TG+ +G+ + L++ L+ + + P + ++ S G
Sbjct: 111 VSYNALITGFSSTGSPARSVQLYRALLREESVR---PTRITLSAMIMVASALSDRALGHS 167
Query: 150 CHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVL-----------NTESGG------ 192
H + + G +Y FV S L +MYA+ + A +V NT G
Sbjct: 168 VHCQVLRLGFGAYAFVGSPLVDMYAKMGLIRDARRVFQEMEAKTVVMYNTLITGLLRCKM 227
Query: 193 -----------DENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGL 241
+ D + +++ L ++G EA+ V RRM E + D T+ ++
Sbjct: 228 IEDAKGLFQLMVDRDSITWTTMVTGLTQNGLQLEALDVFRRMRAEGVGIDQYTFGSILTA 287
Query: 242 CAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVW 301
C + L+ G ++HA + R + FVGS L+DMY KC + A VF + RN++ W
Sbjct: 288 CGALAALEEGKQIHAYITRTWYEDNVFVGSALVDMYSKCRSIRLAEAVFRRMTCRNIISW 347
Query: 302 TSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEK 361
T+++ Y QN EE + + M + P++ T ++ +CA +A L+ G H
Sbjct: 348 TAMIVGYGQNACSEEAVRAFSEMQMDGIKPDDFTLGSVISSCANLASLEEGAQFHCLALV 407
Query: 362 LGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRV 421
G Y+ V+NAL+ +Y K G I+ ++ +F + D +W +++ GY+ G +E + +
Sbjct: 408 SGLMRYITVSNALVTLYGKCGSIEDAHRLFDEMSFHDQVSWTALVTGYAQFGKAKETIDL 467
Query: 422 FQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYC 481
F+ M + G P+ VTF+GVLSAC+ LV+KG DY + K+H I P +HYTCM+ LY
Sbjct: 468 FEKMLANGLKPDGVTFIGVLSACSRAGLVEKGCDYFDSMQKDHGIVPIDDHYTCMIDLYS 527
Query: 482 RAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTL 541
R+G ++AE F+K D W TLL+SCR+ N ++GK AE++L+ DPQ+ +Y L
Sbjct: 528 RSGRFKEAEEFIKQMPHSPDAFGWATLLSSCRLRGNMEIGKWAAENLLETDPQNPASYVL 587
Query: 542 LSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKK 601
L +MHA W V +R+ M++R +KKEPG SW++ +N H+F ++ HP S+ I +K
Sbjct: 588 LCSMHAAKGQWTEVAHLRRGMRDRQVKKEPGCSWIKYKNKVHIFSADDQSHPFSSRIYEK 647
Query: 602 VQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKN 661
++ L + + GY P++S+VLHDV D K ++ HSEKLAIA+GL+ +P PIRI+KN
Sbjct: 648 LEWLNSKMAEEGYKPDVSSVLHDVADADKVHMISHHSEKLAIAFGLIFVPQEMPIRIVKN 707
Query: 662 LRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
LR+C DCH A K ISK+T R I+VRDA RFH F DG+C+C D W
Sbjct: 708 LRVCVDCHNATKFISKITGRDILVRDAVRFHKFSDGTCSCGDFW 751
Score = 134 bits (337), Expect = 9e-30
Identities = 94/342 (27%), Positives = 165/342 (47%), Gaps = 13/342 (3%)
Query: 47 LVRD-----QASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMH 101
L+RD Q + VV N+LI ++C + A+ +F M RD +W ++TG
Sbjct: 196 LIRDARRVFQEMEAKTVVMYNTLITGLLRCKMIEDAKGLFQLMVDRDSITWTTMVTGLTQ 255
Query: 102 SGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVS 161
+G LE L +F+ + + ++Y F ++L++C + EG Q H Y+ +
Sbjct: 256 NGLQLEALDVFRRM----RAEGVGIDQYTFGSILTACGALAALEEGKQIHAYITRTWYED 311
Query: 162 YQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLR 221
FV SAL +MY++C + LA V + +I + +++ ++ EEA+
Sbjct: 312 NVFVGSALVDMYSKCRSIRLAEAVFRRMTC---RNIISWTAMIVGYGQNACSEEAVRAFS 368
Query: 222 RMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCC 281
M + + D+ T V+ CA + L+ G + H L GL+ V + L+ +YGKC
Sbjct: 369 EMQMDGIKPDDFTLGSVISSCANLASLEEGAQFHCLALVSGLMRYITVSNALVTLYGKCG 428
Query: 282 HVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLG 341
+ +A ++FD + + V WT+L++ Y Q +ET+DL M P+ TF +L
Sbjct: 429 SIEDAHRLFDEMSFHDQVSWTALVTGYAQFGKAKETIDLFEKMLANGLKPDGVTFIGVLS 488
Query: 342 ACAGIALLKHG-DLLHARLEKLGFKNYVIVNNALINMYSKSG 382
AC+ L++ G D + + G +I++YS+SG
Sbjct: 489 ACSRAGLVEKGCDYFDSMQKDHGIVPIDDHYTCMIDLYSRSG 530
Score = 78.6 bits (192), Expect = 6e-13
Identities = 75/347 (21%), Positives = 141/347 (40%), Gaps = 66/347 (19%)
Query: 268 FVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQE 327
F+ + L+ Y K + AR+VFD + + N+ +L+SA +R + L M +
Sbjct: 49 FLLNHLLTAYAKSGRLARARRVFDEMPDPNLFTRNALLSALAHSRLVPDMERLFASMPER 108
Query: 328 DTL--------------------------------PNECTFKVLLGACAGIALLKHGDLL 355
D + P T ++ + ++ G +
Sbjct: 109 DAVSYNALITGFSSTGSPARSVQLYRALLREESVRPTRITLSAMIMVASALSDRALGHSV 168
Query: 356 HARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSD---------------------- 393
H ++ +LGF Y V + L++MY+K G I + VF +
Sbjct: 169 HCQVLRLGFGAYAFVGSPLVDMYAKMGLIRDARRVFQEMEAKTVVMYNTLITGLLRCKMI 228
Query: 394 ---------TMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
+ RD TW +M+ G + +GL EAL VF+ M++ G + TF +L+AC
Sbjct: 229 EDAKGLFQLMVDRDSITWTTMVTGLTQNGLQLEALDVFRRMRAEGVGIDQYTFGSILTAC 288
Query: 445 AHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVA 504
LA +++G ++ + E + + +V +Y + + AE + + +I++
Sbjct: 289 GALAALEEG-KQIHAYITRTWYEDNVFVGSALVDMYSKCRSIRLAEAVFRRMTCR-NIIS 346
Query: 505 WRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANM 551
W ++ + + R A S +QMD +TL S + + AN+
Sbjct: 347 WTAMIVGYGQNACSEEAVR-AFSEMQMDGIKPDDFTLGSVISSCANL 392
Score = 65.9 bits (159), Expect = 4e-09
Identities = 43/146 (29%), Positives = 70/146 (47%), Gaps = 8/146 (5%)
Query: 31 ADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVF 90
A+ L G H LV + + N+L+ LY KCG + A +FD+M D
Sbjct: 390 ANLASLEEGAQFHCLALV---SGLMRYITVSNALVTLYGKCGSIEDAHRLFDEMSFHDQV 446
Query: 91 SWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQC 150
SW L+TGY G E + LF+ ++++ P+ F VLS+CS +G V +G
Sbjct: 447 SWTALVTGYAQFGKAKETIDLFEKMLANGLK----PDGVTFIGVLSACSRAGLVEKGCDY 502
Query: 151 HGYLFK-YGLVSYQFVKSALFNMYAR 175
+ K +G+V + + ++Y+R
Sbjct: 503 FDSMQKDHGIVPIDDHYTCMIDLYSR 528
>UniRef100_Q9SMZ2 Hypothetical protein F4I10.100 [Arabidopsis thaliana]
Length = 990
Score = 439 bits (1128), Expect = e-121
Identities = 237/648 (36%), Positives = 372/648 (56%), Gaps = 15/648 (2%)
Query: 62 NSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTT 121
NSLI++Y K + AR++FD M RD+ SWN ++ G +G +E + LF L+
Sbjct: 354 NSLINMYCKLRKFGFARTVFDNMSERDLISWNSVIAGIAQNGLEVEAVCLFMQLLRCGLK 413
Query: 122 QHACPNEYVFTTVLSSCSHSGRVREGM----QCHGYLFKYGLVSYQFVKSALFNMYARCL 177
P++Y T+VL + S + EG+ Q H + K VS FV +AL + Y+R
Sbjct: 414 ----PDQYTMTSVLKAASS---LPEGLSLSKQVHVHAIKINNVSDSFVSTALIDAYSR-- 464
Query: 178 HVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVG 237
N ++ D+ +N+++ +S + + + M + D+ T
Sbjct: 465 --NRCMKEAEILFERHNFDLVAWNAMMAGYTQSHDGHKTLKLFALMHKQGERSDDFTLAT 522
Query: 238 VMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRN 297
V C + + G +VHA ++ G D +V S ++DMY KC + A+ FD + +
Sbjct: 523 VFKTCGFLFAINQGKQVHAYAIKSGYDLDLWVSSGILDMYVKCGDMSAAQFAFDSIPVPD 582
Query: 298 VVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHA 357
V WT+++S ++N E + + M LP+E T L A + + L+ G +HA
Sbjct: 583 DVAWTTMISGCIENGEEERAFHVFSQMRLMGVLPDEFTIATLAKASSCLTALEQGRQIHA 642
Query: 358 RLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQE 417
KL N V +L++MY+K G ID +Y +F ++T WN+M+ G + HG G+E
Sbjct: 643 NALKLNCTNDPFVGTSLVDMYAKCGSIDDAYCLFKRIEMMNITAWNAMLVGLAQHGEGKE 702
Query: 418 ALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMV 477
L++F+ MKS G P+ VTF+GVLSAC+H LV + ++ + ++ I+P +EHY+C+
Sbjct: 703 TLQLFKQMKSLGIKPDKVTFIGVLSACSHSGLVSEAYKHMRSMHGDYGIKPEIEHYSCLA 762
Query: 478 VLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMG 537
RAGL+++AEN +++ ++ +RTLL +CRV + + GKR+A +L+++P D
Sbjct: 763 DALGRAGLVKQAENLIESMSMEASASMYRTLLAACRVQGDTETGKRVATKLLELEPLDSS 822
Query: 538 TYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNE 597
Y LLSNM+A A+ WD + R MMK +KK+PG SW+E++N H+FV + + ++
Sbjct: 823 AYVLLSNMYAAASKWDEMKLARTMMKGHKVKKDPGFSWIEVKNKIHIFVVDDRSNRQTEL 882
Query: 598 INKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIR 657
I +KV+ ++ IK GYVP L DVE+E+KE L HSEKLA+A+GL+ P PIR
Sbjct: 883 IYRKVKDMIRDIKQEGYVPETDFTLVDVEEEEKERALYYHSEKLAVAFGLLSTPPSTPIR 942
Query: 658 IIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+IKNLR+C DCH A+K I+KV NR I++RDANRFH F+DG C+C D+W
Sbjct: 943 VIKNLRVCGDCHNAMKYIAKVYNREIVLRDANRFHRFKDGICSCGDYW 990
Score = 138 bits (347), Expect = 6e-31
Identities = 118/496 (23%), Positives = 217/496 (42%), Gaps = 21/496 (4%)
Query: 82 DKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHS 141
D + ++ N ++ Y+HSG Y +L F ++V S C ++ F +L++
Sbjct: 273 DASSVSEIIFRNKGLSEYLHSGQYSALLKCFADMVESDVE---C-DQVTFILMLATAVKV 328
Query: 142 GRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYN 201
+ G Q H K GL V ++L NMY + A V + S E D+ +N
Sbjct: 329 DSLALGQQVHCMALKLGLDLMLTVSNSLINMYCKLRKFGFARTVFDNMS---ERDLISWN 385
Query: 202 SLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGD-LQLGLRVHARLLR 260
S++ + ++G EA+ + ++ + D T V+ + + + L L +VH ++
Sbjct: 386 SVIAGIAQNGLEVEAVCLFMQLLRCGLKPDQYTMTSVLKAASSLPEGLSLSKQVHVHAIK 445
Query: 261 GGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDL 320
+ D FV + LID Y + + A +F+ N ++V W ++M+ Y Q+ +TL L
Sbjct: 446 INNVSDSFVSTALIDAYSRNRCMKEAEILFE-RHNFDLVAWNAMMAGYTQSHDGHKTLKL 504
Query: 321 LTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSK 380
M ++ ++ T + C + + G +HA K G+ + V++ +++MY K
Sbjct: 505 FALMHKQGERSDDFTLATVFKTCGFLFAINQGKQVHAYAIKSGYDLDLWVSSGILDMYVK 564
Query: 381 SGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGV 440
GD+ ++ F D W +MI G +G + A VF M+ G P+ T +
Sbjct: 565 CGDMSAAQFAFDSIPVPDDVAWTTMISGCIENGEEERAFHVFSQMRLMGVLPDEFTIATL 624
Query: 441 LSACAHLALVDKGLDYLYNEMK-NHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVK 499
A + L +++G N +K N +P + T +V +Y + G ++ A K ++
Sbjct: 625 AKASSCLTALEQGRQIHANALKLNCTNDPFVG--TSLVDMYAKCGSIDDAYCLFKRIEM- 681
Query: 500 WDIVAWRTLLNSCRVHRNYDLGKRIAESILQM-----DPQDMGTYTLLSNMHATANMWDG 554
+I AW +L H GK + QM P + +LS + + +
Sbjct: 682 MNITAWNAMLVGLAQHGE---GKETLQLFKQMKSLGIKPDKVTFIGVLSACSHSGLVSEA 738
Query: 555 VVEIRKMMKERNIKKE 570
+R M + IK E
Sbjct: 739 YKHMRSMHGDYGIKPE 754
Score = 135 bits (339), Expect = 5e-30
Identities = 134/552 (24%), Positives = 233/552 (41%), Gaps = 88/552 (15%)
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
L GK HA++L ++ +R +N+LI +Y KCG L AR +FDKMP RD+ SWN +
Sbjct: 55 LMLGKCTHARILTFEENPER---FLINNLISMYSKCGSLTYARRVFDKMPDRDLVSWNSI 111
Query: 96 MTGYMHSG-----NYLEVLVLFKNL--------------------------VSSSTTQHA 124
+ Y S N + +LF+ L S S +A
Sbjct: 112 LAAYAQSSECVVENIQQAFLLFRILRQDVVYTSRMTLSPMLKLCLHSGYVWASESFHGYA 171
Query: 125 CP-----NEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNM------- 172
C +E+V +++ G+V+EG + +V + + A M
Sbjct: 172 CKIGLDGDEFVAGALVNIYLKFGKVKEGKVLFEEMPYRDVVLWNLMLKAYLEMGFKEEAI 231
Query: 173 ------YARCLHVN-LALQVLNTESGGDEN-----------------DIFLYNSLLNVLV 208
++ L+ N + L++L SG D + +I N L+ +
Sbjct: 232 DLSSAFHSSGLNPNEITLRLLARISGDDSDAGQVKSFANGNDASSVSEIIFRNKGLSEYL 291
Query: 209 ESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEF 268
SG++ + M + D T++ ++ ++ L LG +VH L+ GL
Sbjct: 292 HSGQYSALLKCFADMVESDVECDQVTFILMLATAVKVDSLALGQQVHCMALKLGLDLMLT 351
Query: 269 VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEET----LDLLTCM 324
V + LI+MY K AR VFD + R+++ W S+++ QN E + LL C
Sbjct: 352 VSNSLINMYCKLRKFGFARTVFDNMSERDLISWNSVIAGIAQNGLEVEAVCLFMQLLRCG 411
Query: 325 DQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
+ D K G++L K +H K+ + V+ ALI+ YS++ +
Sbjct: 412 LKPDQYTMTSVLKAASSLPEGLSLSKQ---VHVHAIKINNVSDSFVSTALIDAYSRNRCM 468
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
+ +F + D+ WN+M+ GY+ G + L++F M GE ++ T V C
Sbjct: 469 KEAEILF-ERHNFDLVAWNAMMAGYTQSHDGHKTLKLFALMHKQGERSDDFTLATVFKTC 527
Query: 445 AHLALVDKGLDYLYNEMKNHKIEPG--LEHYTCMVVL--YCRAGLLEKAENFMKTTKVKW 500
L +++G ++ + I+ G L+ + +L Y + G + A+ + V
Sbjct: 528 GFLFAINQG-----KQVHAYAIKSGYDLDLWVSSGILDMYVKCGDMSAAQFAFDSIPVP- 581
Query: 501 DIVAWRTLLNSC 512
D VAW T+++ C
Sbjct: 582 DDVAWTTMISGC 593
Score = 96.3 bits (238), Expect = 3e-18
Identities = 58/195 (29%), Positives = 93/195 (46%), Gaps = 5/195 (2%)
Query: 247 DLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMS 306
DL LG HAR+L + F+ + LI MY KC + AR+VFD + +R++V W S+++
Sbjct: 54 DLMLGKCTHARILTFEENPERFLINNLISMYSKCGSLTYARRVFDKMPDRDLVSWNSILA 113
Query: 307 AYLQN-----RYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEK 361
AY Q+ ++ L + Q+ + T +L C + + H K
Sbjct: 114 AYAQSSECVVENIQQAFLLFRILRQDVVYTSRMTLSPMLKLCLHSGYVWASESFHGYACK 173
Query: 362 LGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRV 421
+G V AL+N+Y K G + +F + RDV WN M+ Y G +EA+ +
Sbjct: 174 IGLDGDEFVAGALVNIYLKFGKVKEGKVLFEEMPYRDVVLWNLMLKAYLEMGFKEEAIDL 233
Query: 422 FQDMKSAGESPNNVT 436
S+G +PN +T
Sbjct: 234 SSAFHSSGLNPNEIT 248
>UniRef100_Q6K297 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 877
Score = 437 bits (1124), Expect = e-121
Identities = 234/670 (34%), Positives = 381/670 (55%), Gaps = 10/670 (1%)
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
+ +G+ +HA ++ + DV N+L+ +YVK GR+ +A +F+KMP DV SWN L
Sbjct: 218 IDAGRQVHAMVV---RMGYEKDVFTANALVDMYVKMGRVDIASVIFEKMPDSDVVSWNAL 274
Query: 96 MTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLF 155
++G + +G+ + L + SS PN ++ +++L +C+ +G G Q HG++
Sbjct: 275 ISGCVLNGHDHRAIELLLQMKSSGLV----PNVFMLSSILKACAGAGAFDLGRQIHGFMI 330
Query: 156 KYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEE 215
K S ++ L +MYA+ ++ A++V + S D+ L+N+L++ GR +E
Sbjct: 331 KANADSDDYIGVGLVDMYAKNHFLDDAMKVFDWMS---HRDLILWNALISGCSHGGRHDE 387
Query: 216 AIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLID 275
A + + E + + T V+ A + +VHA + G +FD V + LID
Sbjct: 388 AFSIFYGLRKEGLGVNRTTLAAVLKSTASLEAASATRQVHALAEKIGFIFDAHVVNGLID 447
Query: 276 MYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECT 335
Y KC + +A +VF+ + +++ TS+++A Q + E + L M ++ P+
Sbjct: 448 SYWKCSCLSDAIRVFEECSSGDIIAVTSMITALSQCDHGEGAIKLFMEMLRKGLEPDPFV 507
Query: 336 FKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTM 395
LL ACA ++ + G +HA L K F + NAL+ Y+K G I+ + FS
Sbjct: 508 LSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGNALVYTYAKCGSIEDAELAFSSLP 567
Query: 396 CRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLD 455
R V +W++MI G + HG G+ AL +F M G +PN++T VL AC H LVD+
Sbjct: 568 ERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGINPNHITMTSVLCACNHAGLVDEAKR 627
Query: 456 YLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
Y + + I+ EHY+CM+ L RAG L+ A + + + + W LL + RVH
Sbjct: 628 YFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVNSMPFQANASVWGALLGASRVH 687
Query: 516 RNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASW 575
++ +LGK AE + ++P+ GT+ LL+N +A++ MW+ V ++RK+MK+ NIKKEP SW
Sbjct: 688 KDPELGKLAAEKLFILEPEKSGTHVLLANTYASSGMWNEVAKVRKLMKDSNIKKEPAMSW 747
Query: 576 LEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLN 635
+E+++ H F+ HP + EI K+ +L ++ GY+PN+ LHD++ +KE L+
Sbjct: 748 VEVKDKVHTFIVGDKSHPMTKEIYSKLDELGDLMSKAGYIPNVDVDLHDLDRSEKELLLS 807
Query: 636 CHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFR 695
HSE+LA+A+ L+ P APIR+ KNLR+C DCH A K IS + +R II+RD NRFHHFR
Sbjct: 808 HHSERLAVAFALLSTPPGAPIRVKKNLRICRDCHMAFKFISNIVSREIIIRDINRFHHFR 867
Query: 696 DGSCTCADHW 705
DG+C+C D+W
Sbjct: 868 DGTCSCGDYW 877
Score = 157 bits (397), Expect = 1e-36
Identities = 122/423 (28%), Positives = 190/423 (44%), Gaps = 26/423 (6%)
Query: 31 ADAKCLTSGKAIHAQLLVRDQASKRSDVVQL-NSLIHLYVKCGRLRLARSMFDKMPLRDV 89
A A+ L G +HA LL K + L N LI Y KC R AR +FD++P
Sbjct: 15 AAAQALLPGAHLHANLL------KSGFLASLRNHLISFYSKCRRPCCARRVFDEIPDPCH 68
Query: 90 FSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQ 149
SW+ L+T Y ++G + F + + C NE+ VL C ++ G Q
Sbjct: 69 VSWSSLVTAYSNNGLPRSAIQAFHGM----RAEGVCCNEFALPVVLK-CVPDAQL--GAQ 121
Query: 150 CHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVE 209
H G S FV +AL MY ++ A +V + G E + +N L++ V+
Sbjct: 122 VHAMAMATGFGSDVFVANALVAMYGGFGFMDDARRVF--DEAGSERNAVSWNGLMSAYVK 179
Query: 210 SGRWEEAIGVLRRMGNECMVWDNAT-----YVGVMGLCAQIGDLQLGLRVHARLLRGGLL 264
+ + +AI V M VW + V+ C ++ G +VHA ++R G
Sbjct: 180 NDQCGDAIQVFGEM-----VWSGIQPTEFGFSCVVNACTGSRNIDAGRQVHAMVVRMGYE 234
Query: 265 FDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCM 324
D F + L+DMY K V A +F+ + + +VV W +L+S + N + ++LL M
Sbjct: 235 KDVFTANALVDMYVKMGRVDIASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQM 294
Query: 325 DQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
+PN +L ACAG G +H + K + + L++MY+K+ +
Sbjct: 295 KSSGLVPNVFMLSSILKACAGAGAFDLGRQIHGFMIKANADSDDYIGVGLVDMYAKNHFL 354
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
D + VF RD+ WN++I G SH G EA +F ++ G N T VL +
Sbjct: 355 DDAMKVFDWMSHRDLILWNALISGCSHGGRHDEAFSIFYGLRKEGLGVNRTTLAAVLKST 414
Query: 445 AHL 447
A L
Sbjct: 415 ASL 417
Score = 126 bits (316), Expect = 3e-27
Identities = 101/367 (27%), Positives = 164/367 (44%), Gaps = 11/367 (2%)
Query: 147 GMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNV 206
G H L K G ++ +++ L + Y++C A +V + ++SL+
Sbjct: 23 GAHLHANLLKSGFLAS--LRNHLISFYSKCRRPCCARRVFDEIPDPCHVS---WSSLVTA 77
Query: 207 LVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFD 266
+G AI M E V N + V+ C + D QLG +VHA + G D
Sbjct: 78 YSNNGLPRSAIQAFHGMRAEG-VCCNEFALPVVLKC--VPDAQLGAQVHAMAMATGFGSD 134
Query: 267 EFVGSKLIDMYGKCCHVLNARKVFDGL-QNRNVVVWTSLMSAYLQNRYFEETLDLLTCMD 325
FV + L+ MYG + +AR+VFD RN V W LMSAY++N + + + M
Sbjct: 135 VFVANALVAMYGGFGFMDDARRVFDEAGSERNAVSWNGLMSAYVKNDQCGDAIQVFGEMV 194
Query: 326 QEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDID 385
P E F ++ AC G + G +HA + ++G++ V NAL++MY K G +D
Sbjct: 195 WSGIQPTEFGFSCVVNACTGSRNIDAGRQVHAMVVRMGYEKDVFTANALVDMYVKMGRVD 254
Query: 386 SSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACA 445
+ +F DV +WN++I G +G A+ + MKS+G PN +L ACA
Sbjct: 255 IASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQMKSSGLVPNVFMLSSILKACA 314
Query: 446 HLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAW 505
D G ++ M + +V +Y + L+ A + D++ W
Sbjct: 315 GAGAFDLGRQ-IHGFMIKANADSDDYIGVGLVDMYAKNHFLDDAMKVFDWMSHR-DLILW 372
Query: 506 RTLLNSC 512
L++ C
Sbjct: 373 NALISGC 379
Score = 68.9 bits (167), Expect = 5e-10
Identities = 45/164 (27%), Positives = 77/164 (46%), Gaps = 8/164 (4%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
LL A GK +HA L+ R SD N+L++ Y KCG + A F +P
Sbjct: 511 LLNACASLSAYEQGKQVHAHLIKR---QFMSDAFAGNALVYTYAKCGSIEDAELAFSSLP 567
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
R V SW+ ++ G G+ L LF +V PN T+VL +C+H+G V
Sbjct: 568 ERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGIN----PNHITMTSVLCACNHAGLVD 623
Query: 146 EGMQCHGYLFK-YGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
E + + + +G+ + S + ++ R ++ A++++N+
Sbjct: 624 EAKRYFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVNS 667
>UniRef100_Q9LW63 Emb|CAB37460.1 [Arabidopsis thaliana]
Length = 715
Score = 436 bits (1122), Expect = e-121
Identities = 237/716 (33%), Positives = 388/716 (54%), Gaps = 41/716 (5%)
Query: 23 IVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFD 82
I L+K K + K +HAQ + S S + +I +Y L A +F
Sbjct: 8 IKTLIKNPTRIKSKSQAKQLHAQFIRTQSLSHTSASI----VISIYTNLKLLHEALLLFK 63
Query: 83 KMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSG 142
+ V +W ++ + + + L F + +S CP+ VF +VL SC+
Sbjct: 64 TLKSPPVLAWKSVIRCFTDQSLFSKALASFVEMRASGR----CPDHNVFPSVLKSCTMMM 119
Query: 143 RVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLN--------TESGGDE 194
+R G HG++ + G+ + +AL NMYA+ L + + V N T + GDE
Sbjct: 120 DLRFGESVHGFIVRLGMDCDLYTGNALMNMYAKLLGMGSKISVGNVFDEMPQRTSNSGDE 179
Query: 195 N-------------------------DIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMV 229
+ D+ YN+++ +SG +E+A+ ++R MG +
Sbjct: 180 DVKAETCIMPFGIDSVRRVFEVMPRKDVVSYNTIIAGYAQSGMYEDALRMVREMGTTDLK 239
Query: 230 WDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKV 289
D+ T V+ + ++ D+ G +H ++R G+ D ++GS L+DMY K + ++ +V
Sbjct: 240 PDSFTLSSVLPIFSEYVDVIKGKEIHGYVIRKGIDSDVYIGSSLVDMYAKSARIEDSERV 299
Query: 290 FDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALL 349
F L R+ + W SL++ Y+QN + E L L M P F ++ ACA +A L
Sbjct: 300 FSRLYCRDGISWNSLVAGYVQNGRYNEALRLFRQMVTAKVKPGAVAFSSVIPACAHLATL 359
Query: 350 KHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGY 409
G LH + + GF + + + +AL++MYSK G+I ++ +F D +W ++I G+
Sbjct: 360 HLGKQLHGYVLRGGFGSNIFIASALVDMYSKCGNIKAARKIFDRMNVLDEVSWTAIIMGH 419
Query: 410 SHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPG 469
+ HG G EA+ +F++MK G PN V FV VL+AC+H+ LVD+ Y + K + +
Sbjct: 420 ALHGHGHEAVSLFEEMKRQGVKPNQVAFVAVLTACSHVGLVDEAWGYFNSMTKVYGLNQE 479
Query: 470 LEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESIL 529
LEHY + L RAG LE+A NF+ V+ W TLL+SC VH+N +L +++AE I
Sbjct: 480 LEHYAAVADLLGRAGKLEEAYNFISKMCVEPTGSVWSTLLSSCSVHKNLELAEKVAEKIF 539
Query: 530 QMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEG 589
+D ++MG Y L+ NM+A+ W + ++R M+++ ++K+P SW+E++N H FVS
Sbjct: 540 TVDSENMGAYVLMCNMYASNGRWKEMAKLRLRMRKKGLRKKPACSWIEMKNKTHGFVSGD 599
Query: 590 CKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMK 649
HP ++IN+ ++ ++ ++ GYV + S VLHDV++E K L HSE+LA+A+G++
Sbjct: 600 RSHPSMDKINEFLKAVMEQMEKEGYVADTSGVLHDVDEEHKRELLFGHSERLAVAFGIIN 659
Query: 650 MPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
IR+ KN+R+C DCH A+K ISK+T R IIVRD +RFHHF G+C+C D+W
Sbjct: 660 TEPGTTIRVTKNIRICTDCHVAIKFISKITEREIIVRDNSRFHHFNRGNCSCGDYW 715
>UniRef100_Q9LHZ4 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 734
Score = 436 bits (1121), Expect = e-120
Identities = 247/697 (35%), Positives = 393/697 (55%), Gaps = 11/697 (1%)
Query: 10 VVVNSRYLPP-LEYIVKLLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLY 68
V ++S PP L LLKL A L +G+A+HAQL R S+ + +L ++Y
Sbjct: 48 VAMSSAGAPPVLRTFTSLLKLCAARGDLATGRAVHAQLAAR---GIDSEALAATALANMY 104
Query: 69 VKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNE 128
KC R AR +FD+MP+RD +WN L+ GY +G + + +V + P+
Sbjct: 105 AKCRRPADARRVFDRMPVRDRVAWNALVAGYARNGL---ARMAMEMVVRMQEEEGERPDS 161
Query: 129 YVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
+VL +C+++ + + H + + GL V +A+ + Y +C + A V +
Sbjct: 162 ITLVSVLPACANARALAACREAHAFAIRSGLEELVNVATAILDAYCKCGDIRAARVVFDW 221
Query: 189 ESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDL 248
+N + +N++++ ++G EA+ + RM E + + + + + C ++G L
Sbjct: 222 MP--TKNSVS-WNAMIDGYAQNGDSREALALFNRMVEEGVDVTDVSVLAALQACGELGCL 278
Query: 249 QLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAY 308
G+RVH L+R GL + V + LI MY KC V A VFD L R V W +++
Sbjct: 279 DEGMRVHELLVRIGLDSNVSVMNALITMYSKCKRVDLASHVFDELDRRTQVSWNAMILGC 338
Query: 309 LQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYV 368
QN E+ + L T M E+ P+ T ++ A A I+ +H +L V
Sbjct: 339 AQNGCSEDAVRLFTRMQLENVKPDSFTLVSVIPALADISDPLQARWIHGYSIRLHLDQDV 398
Query: 369 IVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSA 428
V ALI+MY+K G ++ + +F+ R V TWN+MI GY HG G+ A+ +F++MKS
Sbjct: 399 YVLTALIDMYAKCGRVNIARILFNSARERHVITWNAMIHGYGSHGFGKAAVELFEEMKSI 458
Query: 429 GESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEK 488
G PN TF+ VLSAC+H LVD+G +Y + +++ +EPG+EHY MV L RAG L++
Sbjct: 459 GIVPNETTFLSVLSACSHAGLVDEGREYFTSMKEDYGLEPGMEHYGTMVDLLGRAGKLDE 518
Query: 489 AENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLSNMHAT 548
A F++ + + + +L +C++H+N +L + A+ I ++ PQ+ + LL+N++A
Sbjct: 519 AWAFIQKMPMDPGLSVYGAMLGACKLHKNVELAEESAQKIFELGPQEGVYHVLLANIYAN 578
Query: 549 ANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQQLLAM 608
A+MW V +R M++ ++K PG S ++++N H F S H ++ EI ++ +L+
Sbjct: 579 ASMWKDVARVRTAMEKNGLQKTPGWSIIQLKNEIHTFYSGSTNHQQAKEIYSRLAKLIEE 638
Query: 609 IKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDC 668
IK +GYVP+ ++ HDVED+ K LN HSEKLAIA+GL++ I+I KNLR+C+DC
Sbjct: 639 IKAVGYVPDTDSI-HDVEDDVKAQLLNTHSEKLAIAFGLIRTAPGTTIQIKKNLRVCNDC 697
Query: 669 HTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
H A KLIS VT R II+RD RFHHF+DG C+C D+W
Sbjct: 698 HNATKLISLVTGREIIMRDIQRFHHFKDGKCSCGDYW 734
>UniRef100_Q5ZDP1 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 877
Score = 434 bits (1115), Expect = e-120
Identities = 237/670 (35%), Positives = 381/670 (56%), Gaps = 10/670 (1%)
Query: 36 LTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDL 95
+ +G+ +HA ++VR K DV N+L+ +Y+K GR+ +A +F+KMP DV SWN L
Sbjct: 218 IEAGRQVHA-MVVRMGYDK--DVFTANALVDMYMKMGRVDIASVIFEKMPDSDVVSWNAL 274
Query: 96 MTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLF 155
++G + +G+ + L + S PN + +++L +CS +G G Q HG++
Sbjct: 275 ISGCVLNGHDHRAIELLLQMKYSGLV----PNVFTLSSILKACSGAGAFDLGRQIHGFMI 330
Query: 156 KYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEE 215
K S ++ L +MYA+ ++ A +V + D+ L N+L++ GR +E
Sbjct: 331 KANADSDDYIGVGLVDMYAKNHFLDDARKVFDWMF---HRDLILCNALISGCSHGGRHDE 387
Query: 216 AIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLID 275
A+ + + E + + T V+ A + +VHA ++ G +FD V + LID
Sbjct: 388 ALSLFYELRKEGLGVNRTTLAAVLKSTASLEAASTTRQVHALAVKIGFIFDAHVVNGLID 447
Query: 276 MYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECT 335
Y KC + +A +VF+ + +++ TS+++A Q + E + L M ++ P+
Sbjct: 448 SYWKCSCLSDANRVFEECSSGDIIACTSMITALSQCDHGEGAIKLFMEMLRKGLEPDPFV 507
Query: 336 FKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTM 395
LL ACA ++ + G +HA L K F + NAL+ Y+K G I+ + FS
Sbjct: 508 LSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGNALVYTYAKCGSIEDAELAFSSLP 567
Query: 396 CRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLD 455
R V +W++MI G + HG G+ AL +F M G +PN++T VL AC H LVD+
Sbjct: 568 ERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGINPNHITMTSVLCACNHAGLVDEAKR 627
Query: 456 YLYNEMKNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
Y + + I+ EHY+CM+ L RAG L+ A + + + + W LL + RVH
Sbjct: 628 YFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVNSMPFQANASIWGALLGASRVH 687
Query: 516 RNYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASW 575
++ +LGK AE + ++P+ GT+ LL+N +A+A MW+ V ++RK+MK+ NIKKEP SW
Sbjct: 688 KDPELGKLAAEKLFILEPEKSGTHVLLANTYASAGMWNEVAKVRKLMKDSNIKKEPAMSW 747
Query: 576 LEIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLN 635
+E+++ H F+ HP + EI K+ +L ++ G+VPN+ LHD++ +KE L+
Sbjct: 748 IEVKDKVHTFIVGDKSHPMTKEIYAKLVELGDLMSKAGFVPNVDVDLHDLDRSEKELLLS 807
Query: 636 CHSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFR 695
HSE+LA+A+ L+ P APIR+ KNLR+C DCH A K ISK+ +R II+RD NRFHHFR
Sbjct: 808 HHSERLAVAFALLSTPPGAPIRVKKNLRICRDCHVAFKFISKIVSREIIIRDINRFHHFR 867
Query: 696 DGSCTCADHW 705
DG+C+C D+W
Sbjct: 868 DGTCSCGDYW 877
Score = 155 bits (393), Expect = 3e-36
Identities = 124/423 (29%), Positives = 192/423 (45%), Gaps = 26/423 (6%)
Query: 31 ADAKCLTSGKAIHAQLLVRDQ-ASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDV 89
A A+ L G +HA LL AS R N LI Y KC R AR +FD++P
Sbjct: 15 AAAQALLPGAHLHASLLKSGSLASFR------NHLISFYSKCRRPCCARRVFDEIPDPCH 68
Query: 90 FSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQ 149
SW+ L+T Y ++G + F + + C NE+ VL C R+ G Q
Sbjct: 69 VSWSSLVTAYSNNGLPRSAIQAFHGM----RAEGVCCNEFALPVVLK-CVPDARL--GAQ 121
Query: 150 CHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVE 209
H G S FV +AL MY ++ A +V N E + +N L++ V+
Sbjct: 122 VHAMAMATGFGSDVFVANALVAMYGGFGFMDDARRVFN--EADSERNAVSWNGLMSAYVK 179
Query: 210 SGRWEEAIGVLRRMGNECMVWDNAT-----YVGVMGLCAQIGDLQLGLRVHARLLRGGLL 264
+ + +AI V M VW + V+ C +++ G +VHA ++R G
Sbjct: 180 NDQCGDAIQVFGEM-----VWSGIQPTEFGFSCVVNACTGSRNIEAGRQVHAMVVRMGYD 234
Query: 265 FDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCM 324
D F + L+DMY K V A +F+ + + +VV W +L+S + N + ++LL M
Sbjct: 235 KDVFTANALVDMYMKMGRVDIASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQM 294
Query: 325 DQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDI 384
+PN T +L AC+G G +H + K + + L++MY+K+ +
Sbjct: 295 KYSGLVPNVFTLSSILKACSGAGAFDLGRQIHGFMIKANADSDDYIGVGLVDMYAKNHFL 354
Query: 385 DSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSAC 444
D + VF RD+ N++I G SH G EAL +F +++ G N T VL +
Sbjct: 355 DDARKVFDWMFHRDLILCNALISGCSHGGRHDEALSLFYELRKEGLGVNRTTLAAVLKST 414
Query: 445 AHL 447
A L
Sbjct: 415 ASL 417
Score = 119 bits (297), Expect = 4e-25
Identities = 90/308 (29%), Positives = 142/308 (45%), Gaps = 9/308 (2%)
Query: 147 GMQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNV 206
G H L K G S ++ L + Y++C A +V + ++SL+
Sbjct: 23 GAHLHASLLKSG--SLASFRNHLISFYSKCRRPCCARRVFDEIPDPCHVS---WSSLVTA 77
Query: 207 LVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFD 266
+G AI M E V N + V+ C + D +LG +VHA + G D
Sbjct: 78 YSNNGLPRSAIQAFHGMRAEG-VCCNEFALPVVLKC--VPDARLGAQVHAMAMATGFGSD 134
Query: 267 EFVGSKLIDMYGKCCHVLNARKVFDGLQN-RNVVVWTSLMSAYLQNRYFEETLDLLTCMD 325
FV + L+ MYG + +AR+VF+ + RN V W LMSAY++N + + + M
Sbjct: 135 VFVANALVAMYGGFGFMDDARRVFNEADSERNAVSWNGLMSAYVKNDQCGDAIQVFGEMV 194
Query: 326 QEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDID 385
P E F ++ AC G ++ G +HA + ++G+ V NAL++MY K G +D
Sbjct: 195 WSGIQPTEFGFSCVVNACTGSRNIEAGRQVHAMVVRMGYDKDVFTANALVDMYMKMGRVD 254
Query: 386 SSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACA 445
+ +F DV +WN++I G +G A+ + MK +G PN T +L AC+
Sbjct: 255 IASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQMKYSGLVPNVFTLSSILKACS 314
Query: 446 HLALVDKG 453
D G
Sbjct: 315 GAGAFDLG 322
Score = 68.9 bits (167), Expect = 5e-10
Identities = 45/164 (27%), Positives = 77/164 (46%), Gaps = 8/164 (4%)
Query: 26 LLKLSADAKCLTSGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMP 85
LL A GK +HA L+ R SD N+L++ Y KCG + A F +P
Sbjct: 511 LLNACASLSAYEQGKQVHAHLIKR---QFMSDAFAGNALVYTYAKCGSIEDAELAFSSLP 567
Query: 86 LRDVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVR 145
R V SW+ ++ G G+ L LF +V PN T+VL +C+H+G V
Sbjct: 568 ERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGIN----PNHITMTSVLCACNHAGLVD 623
Query: 146 EGMQCHGYLFK-YGLVSYQFVKSALFNMYARCLHVNLALQVLNT 188
E + + + +G+ + S + ++ R ++ A++++N+
Sbjct: 624 EAKRYFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVNS 667
>UniRef100_Q9SUH6 Hypothetical protein T10C21.50 [Arabidopsis thaliana]
Length = 792
Score = 428 bits (1100), Expect = e-118
Identities = 226/669 (33%), Positives = 378/669 (55%), Gaps = 14/669 (2%)
Query: 38 SGKAIHAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMT 97
+G+ IH Q +V S+++ ++++ +Y K R+ AR +FD+MP +D WN +++
Sbjct: 137 AGRVIHGQAVVD---GCDSELLLGSNIVKMYFKFWRVEDARKVFDRMPEKDTILWNTMIS 193
Query: 98 GYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKY 157
GY + Y+E + +F++L++ S T+ + +L + + +R GMQ H K
Sbjct: 194 GYRKNEMYVESIQVFRDLINESCTRL---DTTTLLDILPAVAELQELRLGMQIHSLATKT 250
Query: 158 GLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAI 217
G S+ +V + ++Y++C + + + + DI YN++++ +G E ++
Sbjct: 251 GCYSHDYVLTGFISLYSKCGKIKMGSALFREFR---KPDIVAYNAMIHGYTSNGETELSL 307
Query: 218 GVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMY 277
+ + + ++T V ++ + G L L +H L+ L V + L +Y
Sbjct: 308 SLFKELMLSGARLRSSTLVSLVPVS---GHLMLIYAIHGYCLKSNFLSHASVSTALTTVY 364
Query: 278 GKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFK 337
K + +ARK+FD +++ W +++S Y QN E+ + L M + + PN T
Sbjct: 365 SKLNEIESARKLFDESPEKSLPSWNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTIT 424
Query: 338 VLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCR 397
+L ACA + L G +H + F++ + V+ ALI MY+K G I + +F +
Sbjct: 425 CILSACAQLGALSLGKWVHDLVRSTDFESSIYVSTALIGMYAKCGSIAEARRLFDLMTKK 484
Query: 398 DVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYL 457
+ TWN+MI GY HG GQEAL +F +M ++G +P VTF+ VL AC+H LV +G D +
Sbjct: 485 NEVTWNTMISGYGLHGQGQEALNIFYEMLNSGITPTPVTFLCVLYACSHAGLVKEG-DEI 543
Query: 458 YNEM-KNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHR 516
+N M + EP ++HY CMV + RAG L++A F++ ++ W TLL +CR+H+
Sbjct: 544 FNSMIHRYGFEPSVKHYACMVDILGRAGHLQRALQFIEAMSIEPGSSVWETLLGACRIHK 603
Query: 517 NYDLGKRIAESILQMDPQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKEPGASWL 576
+ +L + ++E + ++DP ++G + LLSN+H+ + +R+ K+R + K PG + +
Sbjct: 604 DTNLARTVSEKLFELDPDNVGYHVLLSNIHSADRNYPQAATVRQTAKKRKLAKAPGYTLI 663
Query: 577 EIRNVNHVFVSEGCKHPESNEINKKVQQLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNC 636
EI HVF S HP+ EI +K+++L ++ GY P LHDVE+E++E +
Sbjct: 664 EIGETPHVFTSGDQSHPQVKEIYEKLEKLEGKMREAGYQPETELALHDVEEEERELMVKV 723
Query: 637 HSEKLAIAYGLMKMPSPAPIRIIKNLRMCDDCHTAVKLISKVTNRFIIVRDANRFHHFRD 696
HSE+LAIA+GL+ IRIIKNLR+C DCHT KLISK+T R I+VRDANRFHHF+D
Sbjct: 724 HSERLAIAFGLIATEPGTEIRIIKNLRVCLDCHTVTKLISKITERVIVVRDANRFHHFKD 783
Query: 697 GSCTCADHW 705
G C+C D+W
Sbjct: 784 GVCSCGDYW 792
Score = 161 bits (407), Expect = 7e-38
Identities = 117/474 (24%), Positives = 227/474 (47%), Gaps = 15/474 (3%)
Query: 43 HAQLLVRDQASKRSDVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHS 102
HAQ+++ R+D+ L L G + AR +F + DVF +N LM G+ +
Sbjct: 40 HAQIILH---GFRNDISLLTKLTQRLSDLGAIYYARDIFLSVQRPDVFLFNVLMRGFSVN 96
Query: 103 GNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSY 162
+ L +F +L S+ + PN + +S+ S R G HG G S
Sbjct: 97 ESPHSSLSVFAHLRKSTDLK---PNSSTYAFAISAASGFRDDRAGRVIHGQAVVDGCDSE 153
Query: 163 QFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRR 222
+ S + MY + V A +V + E D L+N++++ ++ + E+I V R
Sbjct: 154 LLLGSNIVKMYFKFWRVEDARKVFDRM---PEKDTILWNTMISGYRKNEMYVESIQVFRD 210
Query: 223 MGNE-CMVWDNATYVGVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCC 281
+ NE C D T + ++ A++ +L+LG+++H+ + G ++V + I +Y KC
Sbjct: 211 LINESCTRLDTTTLLDILPAVAELQELRLGMQIHSLATKTGCYSHDYVLTGFISLYSKCG 270
Query: 282 HVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLG 341
+ +F + ++V + +++ Y N E +L L + T L+
Sbjct: 271 KIKMGSALFREFRKPDIVAYNAMIHGYTSNGETELSLSLFKELMLSGARLRSSTLVSLVP 330
Query: 342 ACAGIALLKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTT 401
+ L+ +H K F ++ V+ AL +YSK +I+S+ +F ++ + + +
Sbjct: 331 VSGHLMLIY---AIHGYCLKSNFLSHASVSTALTTVYSKLNEIESARKLFDESPEKSLPS 387
Query: 402 WNSMICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEM 461
WN+MI GY+ +GL ++A+ +F++M+ + SPN VT +LSACA L + G ++++ +
Sbjct: 388 WNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTITCILSACAQLGALSLG-KWVHDLV 446
Query: 462 KNHKIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVH 515
++ E + T ++ +Y + G + +A K + V W T+++ +H
Sbjct: 447 RSTDFESSIYVSTALIGMYAKCGSIAEARRLF-DLMTKKNEVTWNTMISGYGLH 499
Score = 101 bits (251), Expect = 9e-20
Identities = 91/408 (22%), Positives = 176/408 (42%), Gaps = 36/408 (8%)
Query: 196 DIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNA-TYVGVMGLCAQIGDLQLGLRV 254
D+FL+N L+ + ++ V + + N+ TY + + D + G +
Sbjct: 82 DVFLFNVLMRGFSVNESPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFRDDRAGRVI 141
Query: 255 HARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNRYF 314
H + + G + +GS ++ MY K V +ARKVFD + ++ ++W +++S Y +N +
Sbjct: 142 HGQAVVDGCDSELLLGSNIVKMYFKFWRVEDARKVFDRMPEKDTILWNTMISGYRKNEMY 201
Query: 315 EETLDLLTCMDQED-TLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVNNA 373
E++ + + E T + T +L A A + L+ G +H+ K G ++ V
Sbjct: 202 VESIQVFRDLINESCTRLDTTTLLDILPAVAELQELRLGMQIHSLATKTGCYSHDYVLTG 261
Query: 374 LINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDMKSAGESPN 433
I++YSK G I +F + D+ +N+MI GY+ +G + +L +F+++ +G
Sbjct: 262 FISLYSKCGKIKMGSALFREFRKPDIVAYNAMIHGYTSNGETELSLSLFKELMLSGARLR 321
Query: 434 NVTFVGVLSACAHLAL------------------VDKGLDYLYNEMK---------NHKI 466
+ T V ++ HL L V L +Y+++ +
Sbjct: 322 SSTLVSLVPVSGHLMLIYAIHGYCLKSNFLSHASVSTALTTVYSKLNEIESARKLFDESP 381
Query: 467 EPGLEHYTCMVVLYCRAGLLEKAENF---MKTTKVKWDIVAWRTLLNSCRVHRNYDLGKR 523
E L + M+ Y + GL E A + M+ ++ + V +L++C LGK
Sbjct: 382 EKSLPSWNAMISGYTQNGLTEDAISLFREMQKSEFSPNPVTITCILSACAQLGALSLGKW 441
Query: 524 IAESILQMD-PQDMGTYTLLSNMHATANMWDGVVEIRKMMKERNIKKE 570
+ + + D + T L M+A + E R++ K E
Sbjct: 442 VHDLVRSTDFESSIYVSTALIGMYAKCG---SIAEARRLFDLMTKKNE 486
Score = 68.9 bits (167), Expect = 5e-10
Identities = 58/266 (21%), Positives = 118/266 (43%), Gaps = 16/266 (6%)
Query: 253 RVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSLMSAYLQNR 312
+ HA+++ G D + +KL + AR +F +Q +V ++ LM + N
Sbjct: 38 QTHAQIILHGFRNDISLLTKLTQRLSDLGAIYYARDIFLSVQRPDVFLFNVLMRGFSVNE 97
Query: 313 YFEETLDLLTCMDQE-DTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGFKNYVIVN 371
+L + + + D PN T+ + A +G + G ++H + G + +++
Sbjct: 98 SPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFRDDRAGRVIHGQAVVDGCDSELLLG 157
Query: 372 NALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQDM-KSAGE 430
+ ++ MY K ++ + VF +D WN+MI GY + + E+++VF+D+ +
Sbjct: 158 SNIVKMYFKFWRVEDARKVFDRMPEKDTILWNTMISGYRKNEMYVESIQVFRDLINESCT 217
Query: 431 SPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHY------TCMVVLYCRAG 484
+ T + +L A A L + G M+ H + Y T + LY + G
Sbjct: 218 RLDTTTLLDILPAVAELQELRLG-------MQIHSLATKTGCYSHDYVLTGFISLYSKCG 270
Query: 485 LLEKAENFMKTTKVKWDIVAWRTLLN 510
++ + + K DIVA+ +++
Sbjct: 271 KIKMGSALFREFR-KPDIVAYNAMIH 295
Score = 43.9 bits (102), Expect = 0.016
Identities = 44/204 (21%), Positives = 86/204 (41%), Gaps = 3/204 (1%)
Query: 349 LKHGDLLHARLEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICG 408
+ H HA++ GF+N + + L S G I + ++F DV +N ++ G
Sbjct: 33 ISHLAQTHAQIILHGFRNDISLLTKLTQRLSDLGAIYYARDIFLSVQRPDVFLFNVLMRG 92
Query: 409 YSHHGLGQEALRVFQDM-KSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIE 467
+S + +L VF + KS PN+ T+ +SA + D+ ++ + +
Sbjct: 93 FSVNESPHSSLSVFAHLRKSTDLKPNSSTYAFAISAASGFR-DDRAGRVIHGQAVVDGCD 151
Query: 468 PGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAES 527
L + +V +Y + +E A K D + W T+++ R + Y ++
Sbjct: 152 SELLLGSNIVKMYFKFWRVEDARKVFDRMPEK-DTILWNTMISGYRKNEMYVESIQVFRD 210
Query: 528 ILQMDPQDMGTYTLLSNMHATANM 551
++ + T TLL + A A +
Sbjct: 211 LINESCTRLDTTTLLDILPAVAEL 234
>UniRef100_Q9SY02 Hypothetical protein T5J8.5 [Arabidopsis thaliana]
Length = 781
Score = 427 bits (1098), Expect = e-118
Identities = 245/702 (34%), Positives = 379/702 (53%), Gaps = 71/702 (10%)
Query: 59 VQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFK----- 113
V N +I Y++ G LAR +FD+MP RD+ SWN ++ GY+ + N + LF+
Sbjct: 96 VSYNGMISGYLRNGEFELARKLFDEMPERDLVSWNVMIKGYVRNRNLGKARELFEIMPER 155
Query: 114 -----NLVSSSTTQHAC-------------PNEYVFTTVLSSCSHSGRVREGMQCHGYLF 155
N + S Q+ C N+ + +LS+ + ++ E
Sbjct: 156 DVCSWNTMLSGYAQNGCVDDARSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSRE 215
Query: 156 KYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEE 215
+ LVS+ + L + + + A Q ++ + D+ +N+++ +SG+ +E
Sbjct: 216 NWALVSW----NCLLGGFVKKKKIVEARQFFDSMN---VRDVVSWNTIITGYAQSGKIDE 268
Query: 216 AIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLRVHAR----------------LL 259
A R++ +E V D T+ ++ G +Q + AR +L
Sbjct: 269 A----RQLFDESPVQDVFTWTAMVS-----GYIQNRMVEEARELFDKMPERNEVSWNAML 319
Query: 260 RGGL----------LFDEF------VGSKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTS 303
G + LFD + +I Y +C + A+ +FD + R+ V W +
Sbjct: 320 AGYVQGERMEMAKELFDVMPCRNVSTWNTMITGYAQCGKISEAKNLFDKMPKRDPVSWAA 379
Query: 304 LMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLG 363
+++ Y Q+ + E L L M++E N +F L CA + L+ G LH RL K G
Sbjct: 380 MIAGYSQSGHSFEALRLFVQMEREGGRLNRSSFSSALSTCADVVALELGKQLHGRLVKGG 439
Query: 364 FKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQ 423
++ V NAL+ MY K G I+ + ++F + +D+ +WN+MI GYS HG G+ ALR F+
Sbjct: 440 YETGCFVGNALLLMYCKCGSIEEANDLFKEMAGKDIVSWNTMIAGYSRHGFGEVALRFFE 499
Query: 424 DMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRA 483
MK G P++ T V VLSAC+H LVDKG Y Y +++ + P +HY CMV L RA
Sbjct: 500 SMKREGLKPDDATMVAVLSACSHTGLVDKGRQYFYTMTQDYGVMPNSQHYACMVDLLGRA 559
Query: 484 GLLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQMDPQDMGTYTLLS 543
GLLE A N MK + D W TLL + RVH N +L + A+ I M+P++ G Y LLS
Sbjct: 560 GLLEDAHNLMKNMPFEPDAAIWGTLLGASRVHGNTELAETAADKIFAMEPENSGMYVLLS 619
Query: 544 NMHATANMWDGVVEIRKMMKERNIKKEPGASWLEIRNVNHVFVSEGCKHPESNEINKKVQ 603
N++A++ W V ++R M+++ +KK PG SW+EI+N H F HPE +EI ++
Sbjct: 620 NLYASSGRWGDVGKLRVRMRDKGVKKVPGYSWIEIQNKTHTFSVGDEFHPEKDEIFAFLE 679
Query: 604 QLLAMIKPLGYVPNISAVLHDVEDEQKEGYLNCHSEKLAIAYGLMKMPSPAPIRIIKNLR 663
+L +K GYV S VLHDVE+E+KE + HSE+LA+AYG+M++ S PIR+IKNLR
Sbjct: 680 ELDLRMKKAGYVSKTSVVLHDVEEEEKERMVRYHSERLAVAYGIMRVSSGRPIRVIKNLR 739
Query: 664 MCDDCHTAVKLISKVTNRFIIVRDANRFHHFRDGSCTCADHW 705
+C+DCH A+K ++++T R II+RD NRFHHF+DGSC+C D+W
Sbjct: 740 VCEDCHNAIKYMARITGRLIILRDNNRFHHFKDGSCSCGDYW 781
Score = 120 bits (300), Expect = 2e-25
Identities = 115/481 (23%), Positives = 208/481 (42%), Gaps = 50/481 (10%)
Query: 88 DVFSWNDLMTGYMHSGNYLEVLVLFKNLVSSSTTQHACPNEYVFTTVLSSCSHSGRVREG 147
D+ WN ++ YM +G E L +FK + S+ + ++S +G
Sbjct: 63 DIKEWNVAISSYMRTGRCNEALRVFKRMPRWSSVS--------YNGMISGYLRNGEFELA 114
Query: 148 MQCHGYLFKYGLVSYQFVKSALFNMYARCLHVNLALQVLNTESGGDENDIFLYNSLLNVL 207
+ + + LVS+ + + Y R ++ A ++ E D+ +N++L+
Sbjct: 115 RKLFDEMPERDLVSW----NVMIKGYVRNRNLGKARELFEIM---PERDVCSWNTMLSGY 167
Query: 208 VESGRWEEAIGVLRRMGNE---------------------CMVW-DNATYVGVMGLCAQI 245
++G ++A V RM + CM++ + V C
Sbjct: 168 AQNGCVDDARSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSRENWALVSWNCLLG 227
Query: 246 GDLQLGLRVHARLLRGGLLFDEFVG-SKLIDMYGKCCHVLNARKVFDGLQNRNVVVWTSL 304
G ++ V AR + + V + +I Y + + AR++FD ++V WT++
Sbjct: 228 GFVKKKKIVEARQFFDSMNVRDVVSWNTIITGYAQSGKIDEARQLFDESPVQDVFTWTAM 287
Query: 305 MSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLHARLEKLGF 364
+S Y+QNR EE +L M + NE ++ +L AG + ++ + +
Sbjct: 288 VSGYIQNRMVEEARELFDKMPER----NEVSWNAML---AGYVQGERMEMAKELFDVMPC 340
Query: 365 KNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNSMICGYSHHGLGQEALRVFQD 424
+N V N +I Y++ G I + N+F RD +W +MI GYS G EALR+F
Sbjct: 341 RN-VSTWNTMITGYAQCGKISEAKNLFDKMPKRDPVSWAAMIAGYSQSGHSFEALRLFVQ 399
Query: 425 MKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNHKIEPGLEHYTCMVVLYCRAG 484
M+ G N +F LS CA + ++ G L+ + E G ++++YC+ G
Sbjct: 400 MEREGGRLNRSSFSSALSTCADVVALELG-KQLHGRLVKGGYETGCFVGNALLLMYCKCG 458
Query: 485 LLEKAENFMKTTKVKWDIVAWRTLLNSCRVHRNYDLGKRIAESILQ--MDPQDMGTYTLL 542
+E+A + K K DIV+W T++ H ++ R ES+ + + P D +L
Sbjct: 459 SIEEANDLFKEMAGK-DIVSWNTMIAGYSRHGFGEVALRFFESMKREGLKPDDATMVAVL 517
Query: 543 S 543
S
Sbjct: 518 S 518
Score = 112 bits (279), Expect = 5e-23
Identities = 87/388 (22%), Positives = 178/388 (45%), Gaps = 26/388 (6%)
Query: 57 DVVQLNSLIHLYVKCGRLRLARSMFDKMPLRDVFSWNDLMTGYMHSGNYLEVLVLFKNLV 116
DVV N++I Y + G++ AR +FD+ P++DVF+W +++GY+ + E LF +
Sbjct: 249 DVVSWNTIITGYAQSGKIDEARQLFDESPVQDVFTWTAMVSGYIQNRMVEEARELFDKMP 308
Query: 117 SSSTTQHACPNEYVFTTVLSSCSHSGRVREGMQCHGYLFKYGLVSYQFVKSALFNMYARC 176
NE + +L+ E M+ LF + + YA+C
Sbjct: 309 ER--------NEVSWNAMLAGYVQG----ERMEMAKELFDVMPCRNVSTWNTMITGYAQC 356
Query: 177 LHVNLALQVLNTESGGDENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYV 236
++ A + + + D + +++ +SG EA+ + +M E + +++
Sbjct: 357 GKISEAKNLFDKM---PKRDPVSWAAMIAGYSQSGHSFEALRLFVQMEREGGRLNRSSFS 413
Query: 237 GVMGLCAQIGDLQLGLRVHARLLRGGLLFDEFVGSKLIDMYGKCCHVLNARKVFDGLQNR 296
+ CA + L+LG ++H RL++GG FVG+ L+ MY KC + A +F + +
Sbjct: 414 SALSTCADVVALELGKQLHGRLVKGGYETGCFVGNALLLMYCKCGSIEEANDLFKEMAGK 473
Query: 297 NVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGIALLKHGDLLH 356
++V W ++++ Y ++ + E L M +E P++ T +L AC+ L+ G
Sbjct: 474 DIVSWNTMIAGYSRHGFGEVALRFFESMKREGLKPDDATMVAVLSACSHTGLVDKGRQYF 533
Query: 357 ARLEKLGFKNYVIVNNA-----LINMYSKSGDIDSSYNVFSDTMCR-DVTTWNSMICGYS 410
+ ++Y ++ N+ ++++ ++G ++ ++N+ + D W +++
Sbjct: 534 YTMT----QDYGVMPNSQHYACMVDLLGRAGLLEDAHNLMKNMPFEPDAAIWGTLLGASR 589
Query: 411 HHGLGQEALRVFQDMKSAGESPNNVTFV 438
HG E D A E N+ +V
Sbjct: 590 VHG-NTELAETAADKIFAMEPENSGMYV 616
Score = 82.0 bits (201), Expect = 5e-14
Identities = 77/345 (22%), Positives = 150/345 (43%), Gaps = 53/345 (15%)
Query: 194 ENDIFLYNSLLNVLVESGRWEEAIGVLRRMGNECMVWDNATYVGVMGLCAQIGDLQLGLR 253
++DI +N ++ + +GR EA+ V +RM W + +Y G++ + G+ +L +
Sbjct: 61 DSDIKEWNVAISSYMRTGRCNEALRVFKRMPR----WSSVSYNGMISGYLRNGEFELARK 116
Query: 254 V-----HARLLRGGLLFDEFVGSK----------------------LIDMYGKCCHVLNA 286
+ L+ ++ +V ++ ++ Y + V +A
Sbjct: 117 LFDEMPERDLVSWNVMIKGYVRNRNLGKARELFEIMPERDVCSWNTMLSGYAQNGCVDDA 176
Query: 287 RKVFDGLQNRNVVVWTSLMSAYLQNRYFEETLDLLTCMDQEDTLPNECTFKVLLGACAGI 346
R VFD + +N V W +L+SAY+QN EE L + + C LLG
Sbjct: 177 RSVFDRMPEKNDVSWNALLSAYVQNSKMEEACMLFKSRENWALVSWNC----LLG----- 227
Query: 347 ALLKHGDLLHAR--LEKLGFKNYVIVNNALINMYSKSGDIDSSYNVFSDTMCRDVTTWNS 404
+K ++ AR + + ++ V+ N +I Y++SG ID + +F ++ +DV TW +
Sbjct: 228 GFVKKKKIVEARQFFDSMNVRD-VVSWNTIITGYAQSGKIDEARQLFDESPVQDVFTWTA 286
Query: 405 MICGYSHHGLGQEALRVFQDMKSAGESPNNVTFVGVLSACAHLALVDKGLDYLYNEMKNH 464
M+ GY + + +EA +F M E N G + + + + L++ M
Sbjct: 287 MVSGYIQNRMVEEARELFDKMPERNEVSWNAMLAGYVQG-ERMEMAKE----LFDVMPCR 341
Query: 465 KIEPGLEHYTCMVVLYCRAGLLEKAENFMKTTKVKWDIVAWRTLL 509
+ + M+ Y + G + +A+N K D V+W ++
Sbjct: 342 NVST----WNTMITGYAQCGKISEAKNLFDKMP-KRDPVSWAAMI 381
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.137 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,162,661,485
Number of Sequences: 2790947
Number of extensions: 47990798
Number of successful extensions: 118305
Number of sequences better than 10.0: 970
Number of HSP's better than 10.0 without gapping: 694
Number of HSP's successfully gapped in prelim test: 279
Number of HSP's that attempted gapping in prelim test: 102397
Number of HSP's gapped (non-prelim): 4733
length of query: 705
length of database: 848,049,833
effective HSP length: 135
effective length of query: 570
effective length of database: 471,271,988
effective search space: 268625033160
effective search space used: 268625033160
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0141.1


