

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0140.2
(123 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
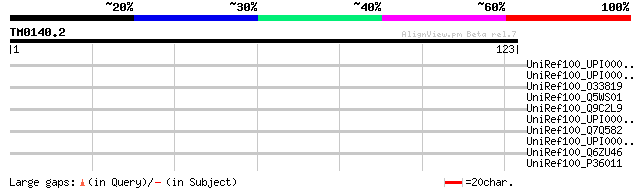
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI00002DA779 UPI00002DA779 UniRef100 entry 34 0.87
UniRef100_UPI0000457652 UPI0000457652 UniRef100 entry 33 1.1
UniRef100_O33819 4-hydroxybenzoyl-CoA reductase alpha subunit [T... 32 2.5
UniRef100_Q5WS01 Hypothetical protein [Legionella pneumophila st... 32 4.3
UniRef100_Q9C2L9 Hypothetical protein 17E5.290 [Neurospora crassa] 32 4.3
UniRef100_UPI00002D1E61 UPI00002D1E61 UniRef100 entry 31 5.6
UniRef100_Q7Q582 ENSANGP00000002585 [Anopheles gambiae str. PEST] 31 5.6
UniRef100_UPI00002BB3F4 UPI00002BB3F4 UniRef100 entry 30 9.6
UniRef100_Q6ZU46 Hypothetical protein FLJ44004 [Homo sapiens] 30 9.6
UniRef100_P36011 Cell pattern formation-associated protein stuA ... 30 9.6
>UniRef100_UPI00002DA779 UPI00002DA779 UniRef100 entry
Length = 909
Score = 33.9 bits (76), Expect = 0.87
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Query: 52 CVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEP-PKTALHNV 97
C+ + + P Q H SC P RAP P K V P+P P+ H++
Sbjct: 187 CIKHMINVDPKQGHLSCSPLPTRAPSPTPRKDTVSPQPHPQPHSHSL 233
>UniRef100_UPI0000457652 UPI0000457652 UniRef100 entry
Length = 701
Score = 33.5 bits (75), Expect = 1.1
Identities = 18/55 (32%), Positives = 24/55 (42%), Gaps = 7/55 (12%)
Query: 55 MIASTPPSQHHHSCKPT-------VNRAPGWFPEKRGVFPEPPKTALHNVTLLQH 102
++ PPS H HSC+ + V RAP G P+PP T L + H
Sbjct: 352 LLQGEPPSHHPHSCRESPVTTHTPVGRAPATTHTPAGRAPQPPPTLLQGESPSHH 406
Score = 32.7 bits (73), Expect = 1.9
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 7/50 (14%)
Query: 51 PCVIMIASTPPSQHHHSCKPT-------VNRAPGWFPEKRGVFPEPPKTA 93
P ++ PPS H HSC+ + V RAP G P+PP T+
Sbjct: 144 PPPTLLQGEPPSHHPHSCRESPVTTHTPVGRAPATTHTPAGRVPQPPPTS 193
>UniRef100_O33819 4-hydroxybenzoyl-CoA reductase alpha subunit [Thauera aromatica]
Length = 769
Score = 32.3 bits (72), Expect = 2.5
Identities = 23/77 (29%), Positives = 32/77 (40%), Gaps = 5/77 (6%)
Query: 45 NLMPRTPCVIMIASTPPSQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLR 104
N++P+ P V M A S C V A GW E++G P+ + L H
Sbjct: 391 NMLPQIPYVTMYAQRVMSYGVPECLEKVKAASGW-EERKGKLPKGRGLGI----ALSHFV 445
Query: 105 NTTQSDIHRTGSSDTTV 121
+ T + H TG TV
Sbjct: 446 SGTSTPKHWTGEPHATV 462
>UniRef100_Q5WS01 Hypothetical protein [Legionella pneumophila str. Lens]
Length = 387
Score = 31.6 bits (70), Expect = 4.3
Identities = 25/84 (29%), Positives = 35/84 (40%), Gaps = 16/84 (19%)
Query: 16 YLASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTPCVIMIASTPPS----QHHHSCKPT 71
Y+ + TNAR + LYQ L+ ++ C I+ PPS + H C
Sbjct: 263 YIQFIETNARPPGIGLNKLYQRKYSISLETIL----CCIVCGVEPPSMVENKSHFVC--- 315
Query: 72 VNRAPGWFPEKRGVFPEPPKTALH 95
G+FP K GV + K LH
Sbjct: 316 -----GYFPTKMGVIKKINKPELH 334
>UniRef100_Q9C2L9 Hypothetical protein 17E5.290 [Neurospora crassa]
Length = 479
Score = 31.6 bits (70), Expect = 4.3
Identities = 24/95 (25%), Positives = 38/95 (39%), Gaps = 28/95 (29%)
Query: 37 PMLV-CHLDNLMPRTPCVIMIASTPPS----------QHHHSCKPTVNRAPGWFPEKRGV 85
P++V CH + +++ TPPS QHHHS + + + +F K+ V
Sbjct: 65 PVIVPCHSQMTLDDFAAALLVPVTPPSSPLPNNSNNPQHHHSHRSVPSGSGSYFSSKQNV 124
Query: 86 -----------------FPEPPKTALHNVTLLQHL 103
+PP T + NV L Q+L
Sbjct: 125 PGSSSSAHPPPPSQQQQQQQPPATQIANVVLAQNL 159
>UniRef100_UPI00002D1E61 UPI00002D1E61 UniRef100 entry
Length = 274
Score = 31.2 bits (69), Expect = 5.6
Identities = 21/58 (36%), Positives = 28/58 (48%), Gaps = 2/58 (3%)
Query: 62 SQHHHSCKPTVNRAPGWFPEKRGVFPEPPKTALHNVTLLQHLRN-TTQSDI-HRTGSS 117
S+ HSC+ + +A W+PE V N + LQ + N TQS I HR G S
Sbjct: 47 SKSQHSCREELGQACEWYPEPSCVVAVQDDECKVNTSALQSMLNCPTQSFIAHRLGCS 104
>UniRef100_Q7Q582 ENSANGP00000002585 [Anopheles gambiae str. PEST]
Length = 880
Score = 31.2 bits (69), Expect = 5.6
Identities = 15/47 (31%), Positives = 20/47 (41%)
Query: 30 HILHLYQPMLVCHLDNLMPRTPCVIMIASTPPSQHHHSCKPTVNRAP 76
H++ LY P +V P P + PS HH+ TV R P
Sbjct: 211 HVMGLYLPAMVRSTPTPSPSPPAPATVGGHVPSAKHHALHQTVLREP 257
>UniRef100_UPI00002BB3F4 UPI00002BB3F4 UniRef100 entry
Length = 1209
Score = 30.4 bits (67), Expect = 9.6
Identities = 21/75 (28%), Positives = 31/75 (41%), Gaps = 9/75 (12%)
Query: 9 SAIDITTYL---ASMSTNARTLSPHILHLYQPMLVCHLDNLMPRTP--CVIMIASTPP-- 61
+ +++T Y+ MS + TL +L P+ C N P CV+ + P
Sbjct: 128 NTVNLTVYVRPTTQMSLSTPTLVARSSNLKMPVATCVSANGKPPATIRCVVFLCCPWPLN 187
Query: 62 --SQHHHSCKPTVNR 74
HHH C P NR
Sbjct: 188 SVDAHHHLCFPCTNR 202
>UniRef100_Q6ZU46 Hypothetical protein FLJ44004 [Homo sapiens]
Length = 910
Score = 30.4 bits (67), Expect = 9.6
Identities = 24/64 (37%), Positives = 29/64 (44%), Gaps = 6/64 (9%)
Query: 40 VCHLDNLMPRTPCVIMIASTPPSQHHHSC-KPTVNRAPGWFPE--KRGV---FPEPPKTA 93
V HL P T + P + H C +P R P PE K GV FPEPPKT
Sbjct: 226 VSHLCPEPPETRVSPLRQLPPEAGVSHLCPEPPKTRVPPLRPETPKNGVSPLFPEPPKTR 285
Query: 94 LHNV 97
+ N+
Sbjct: 286 ISNL 289
>UniRef100_P36011 Cell pattern formation-associated protein stuA [Emericella
nidulans]
Length = 622
Score = 30.4 bits (67), Expect = 9.6
Identities = 17/42 (40%), Positives = 22/42 (51%), Gaps = 5/42 (11%)
Query: 57 ASTPPSQHHHSCK---PTVNRAPGWFP--EKRGVFPEPPKTA 93
A PPS HHHS + P+ PG P ++ FP PP +A
Sbjct: 263 AQQPPSLHHHSLQTPVPSHMSQPGGRPSLDRAHTFPTPPASA 304
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.131 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 202,384,476
Number of Sequences: 2790947
Number of extensions: 7995676
Number of successful extensions: 16977
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 16962
Number of HSP's gapped (non-prelim): 21
length of query: 123
length of database: 848,049,833
effective HSP length: 99
effective length of query: 24
effective length of database: 571,746,080
effective search space: 13721905920
effective search space used: 13721905920
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0140.2


