

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0134.4
(311 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
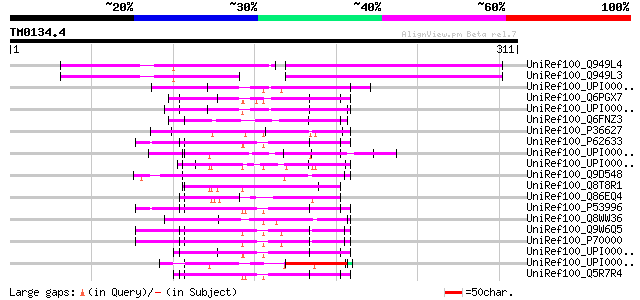
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q949L4 Putative polyprotein [Cicer arietinum] 101 2e-20
UniRef100_Q949L3 Putative polyprotein [Cicer arietinum] 98 3e-19
UniRef100_UPI000036346C UPI000036346C UniRef100 entry 82 2e-14
UniRef100_Q6PGX7 Hypothetical protein zff9 [Brachydanio rerio] 79 1e-13
UniRef100_UPI0000318F83 UPI0000318F83 UniRef100 entry 78 3e-13
UniRef100_Q6FNZ3 Similar to sp|P53849 Saccharomyces cerevisiae Y... 78 3e-13
UniRef100_P36627 Cellular nucleic acid binding protein homolog [... 78 3e-13
UniRef100_P62633 Cellular nucleic acid binding protein [Homo sap... 77 5e-13
UniRef100_UPI000042E8CA UPI000042E8CA UniRef100 entry 76 1e-12
UniRef100_UPI00003C130E UPI00003C130E UniRef100 entry 76 1e-12
UniRef100_Q9D548 Mus musculus adult male testis cDNA, RIKEN full... 76 1e-12
UniRef100_Q8T8R1 GM14667p [Drosophila melanogaster] 76 1e-12
UniRef100_Q86EQ4 Clone ZZD1536 mRNA sequence [Schistosoma japoni... 76 1e-12
UniRef100_P53996 Cellular nucleic acid binding protein [Mus musc... 76 1e-12
UniRef100_Q8WW36 Zinc finger, CCHC domain containing 13 [Homo sa... 75 2e-12
UniRef100_Q9W6Q5 Cellular nucleic acid binding protein [Bufo are... 75 2e-12
UniRef100_P70000 Cellular nucleic acid binding protein [Xenopus ... 75 2e-12
UniRef100_UPI00003AA82A UPI00003AA82A UniRef100 entry 74 4e-12
UniRef100_UPI000042FF1E UPI000042FF1E UniRef100 entry 74 4e-12
UniRef100_Q5R7R4 Hypothetical protein DKFZp470E2414 [Pongo pygma... 74 4e-12
>UniRef100_Q949L4 Putative polyprotein [Cicer arietinum]
Length = 318
Score = 101 bits (252), Expect = 2e-20
Identities = 53/134 (39%), Positives = 72/134 (53%), Gaps = 1/134 (0%)
Query: 170 CYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGKVYTM 229
C+RCG GH+AS C+A P C NC + H+ C PKVEP VN AR P ARG++Y M
Sbjct: 75 CFRCGGEGHYASACTANVPTCHNCRKSEHMERDCTAPKVEPVVNVARATRPTARGRIYCM 134
Query: 230 DGEEAEGADELVKGERKNDGNLLTILSHSSIIRSITLVAHVTHL-FYLYCLSIDFIVITP 288
+ EE + L++ + + GN LT L S S + V L + L D +V TP
Sbjct: 135 NAEEGNPSGNLIQRDCEIAGNTLTALFDSGATHSFIAMDCVNRLKLSVSALPFDLVVSTP 194
Query: 289 NNTLFDRCSCLNYR 302
+ TL +CL+ R
Sbjct: 195 DKTLTVNSACLHCR 208
Score = 55.5 bits (132), Expect = 2e-06
Identities = 38/136 (27%), Positives = 62/136 (44%), Gaps = 13/136 (9%)
Query: 32 EAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEY-RQLG-PYQHPKGRGFTPRSFKS 89
+A E+ + + + + SR+ +GGP R+ Q +Q+ PY P+ +
Sbjct: 3 QALVEKCREVEDMKNKRASRVGNFNSGGPSRANYQNKGKQVAKPYNRPQNN--------N 54
Query: 90 STTATPGARS--QSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAG 147
+ PG ++ + + CFRCG GHYA+AC P C NC + H+ +C +
Sbjct: 55 GGQSRPGNQNFGRQLGQVLRCFRCGGEGHYASACTANVPTCHNCRKSEHMERDCTAPKVE 114
Query: 148 PSDNKMKGKFPAIARG 163
P N + P ARG
Sbjct: 115 PVVNVARATRPT-ARG 129
>UniRef100_Q949L3 Putative polyprotein [Cicer arietinum]
Length = 318
Score = 97.8 bits (242), Expect = 3e-19
Identities = 51/134 (38%), Positives = 71/134 (52%), Gaps = 1/134 (0%)
Query: 170 CYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAARGKVYTM 229
C+RCG GH+AS C+ P C NC ++GH+ C PKVE VN AR P ARG++Y +
Sbjct: 75 CFRCGGEGHYASACTTNIPICHNCRKLGHMTRDCTAPKVELVVNVARATRPTARGRIYCV 134
Query: 230 DGEEAEGADELVKGERKNDGNLLTILSHSSIIRSITLVAHVTHL-FYLYCLSIDFIVITP 288
+ EE + L++ + + GN LT L S S + V L + L D +V TP
Sbjct: 135 NAEEGNPSGNLIQRDCEIAGNTLTALFDSGATHSFIAMGCVNRLKLSVSALPFDLVVSTP 194
Query: 289 NNTLFDRCSCLNYR 302
TL +CL+ R
Sbjct: 195 AKTLTVNSACLHCR 208
Score = 55.8 bits (133), Expect = 1e-06
Identities = 33/114 (28%), Positives = 56/114 (48%), Gaps = 12/114 (10%)
Query: 32 EAARERPQPLIEASKYQQSRLDRAGTGGPVRSGSQEY-RQLG-PYQHPKGRGFTPRSFKS 89
+A E+ + + + + SR+ +GGP R+ Q +Q+ PY P+ +
Sbjct: 3 QALVEKCREVEDMKNKRASRVGNFNSGGPSRANYQNKGKQVAKPYNRPQNN--------N 54
Query: 90 STTATPGARS--QSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNC 141
+ PG ++ + + CFRCG GHYA+AC P C NC ++GH+ +C
Sbjct: 55 GGQSRPGNQNFGRQMGQVLRCFRCGGEGHYASACTTNIPICHNCRKLGHMTRDC 108
>UniRef100_UPI000036346C UPI000036346C UniRef100 entry
Length = 167
Score = 82.0 bits (201), Expect = 2e-14
Identities = 46/141 (32%), Positives = 64/141 (44%), Gaps = 13/141 (9%)
Query: 88 KSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAG 147
KSS G R + E C+RCG++GH A C C+NC++ GH++ +CK
Sbjct: 24 KSSGPRGRGPRGRGRVKEQFCYRCGEHGHIARDCDQPEDSCYNCHKSGHISRDCKE---- 79
Query: 148 PSDNKMK-----GKFPAIARGTE--KQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLA 200
P + GK +AR E + CY CGE GH C +C+ C +GH+A
Sbjct: 80 PKREREHLCYNCGKAGHVARDCEHANEQKCYSCGEFGHIQKLCDKV--KCYRCGEIGHVA 137
Query: 201 VVCKKPKVEPSVNTARGKHPA 221
V C K N + H A
Sbjct: 138 VQCSKASETNCYNCGKAGHVA 158
Score = 73.6 bits (179), Expect = 7e-12
Identities = 35/88 (39%), Positives = 46/88 (51%), Gaps = 7/88 (7%)
Query: 122 LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFAS 181
LG CF C + GH +C P + +G+ P RG K+ CYRCGE GH A
Sbjct: 3 LGGNNECFGCGRTGHWIKDC------PKSSGPRGRGPR-GRGRVKEQFCYRCGEHGHIAR 55
Query: 182 RCSATGPRCFNCHRVGHLAVVCKKPKVE 209
C C+NCH+ GH++ CK+PK E
Sbjct: 56 DCDQPEDSCYNCHKSGHISRDCKEPKRE 83
>UniRef100_Q6PGX7 Hypothetical protein zff9 [Brachydanio rerio]
Length = 161
Score = 79.3 bits (194), Expect = 1e-13
Identities = 46/129 (35%), Positives = 65/129 (49%), Gaps = 24/129 (18%)
Query: 98 RSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCK---------TFQAGP 148
R + ++ C+RCG+ GH A C C+NC++ GH++ +CK + G
Sbjct: 28 RGRGRGKDLFCYRCGEQGHIARDCEQTEDACYNCHRSGHISRDCKEPKKEREQCCYNCGK 87
Query: 149 SD---------NKMK----GKFPAIARGTEKQVSCYRCGEIGHFASRCS-ATGPRCFNCH 194
+ N+ K G F I + +K V CYRCGEIGH A +CS AT C+NC
Sbjct: 88 AGHVARDCDHANEQKCYSCGGFGHIQKLCDK-VKCYRCGEIGHVAVQCSKATEVNCYNCG 146
Query: 195 RVGHLAVVC 203
+ GHLA C
Sbjct: 147 KTGHLARDC 155
Score = 75.1 bits (183), Expect = 2e-12
Identities = 35/82 (42%), Positives = 45/82 (54%), Gaps = 10/82 (12%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
CF C + GH NC P+ + +G+ RG K + CYRCGE GH A C T
Sbjct: 6 CFGCGRSGHWIKNC------PNAGRGRGR----GRGRGKDLFCYRCGEQGHIARDCEQTE 55
Query: 188 PRCFNCHRVGHLAVVCKKPKVE 209
C+NCHR GH++ CK+PK E
Sbjct: 56 DACYNCHRSGHISRDCKEPKKE 77
Score = 52.0 bits (123), Expect = 2e-05
Identities = 27/84 (32%), Positives = 39/84 (46%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C GH+ C K ++ G + A+
Sbjct: 79 EQCCYNCGKAGHVARDCDHANEQKCYSCGGFGHIQKLCDKVKCYRCGEIGHV------AV 132
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CY CG+ GH A CS
Sbjct: 133 QCSKATEVNCYNCGKTGHLARDCS 156
>UniRef100_UPI0000318F83 UPI0000318F83 UniRef100 entry
Length = 167
Score = 77.8 bits (190), Expect = 3e-13
Identities = 35/88 (39%), Positives = 49/88 (54%), Gaps = 7/88 (7%)
Query: 122 LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFAS 181
LG CF C + GH NC P+ + ++G+ P RG K++ CYRCG+ GH
Sbjct: 3 LGGNSDCFGCGRPGHWVKNC------PTSSGLRGRGPR-GRGRGKELFCYRCGDQGHMVK 55
Query: 182 RCSATGPRCFNCHRVGHLAVVCKKPKVE 209
C T C+NCH+ GH++ CK+PK E
Sbjct: 56 DCDQTEDSCYNCHKSGHISRDCKEPKRE 83
Score = 73.9 bits (180), Expect = 5e-12
Identities = 35/113 (30%), Positives = 52/113 (45%), Gaps = 19/113 (16%)
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKG 155
G R + E+ C+RCG GH C C+NC++ GH++ +CK +
Sbjct: 32 GPRGRGRGKELFCYRCGDQGHMVKDCDQTEDSCYNCHKSGHISRDCKEPK---------- 81
Query: 156 KFPAIARGTEKQVSCYRCGEIGHFASRCS-ATGPRCFNCHRVGHLAVVCKKPK 207
E++ CY CG+ GH A C A +CF C +GH+ +C K K
Sbjct: 82 --------REREQQCYNCGKAGHMARECDHANEQKCFTCGTLGHIQKLCDKVK 126
Score = 51.2 bits (121), Expect = 3e-05
Identities = 26/84 (30%), Positives = 38/84 (44%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +CF C +GH+ C K ++ G + A+
Sbjct: 85 EQQCYNCGKAGHMARECDHANEQKCFTCGTLGHIQKLCDKVKCYRCGGIGHV------AL 138
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+ +CY CG+ GH A C+
Sbjct: 139 QCSKASETTCYNCGKAGHVAKDCT 162
>UniRef100_Q6FNZ3 Similar to sp|P53849 Saccharomyces cerevisiae YNL255c GIS2 [Candida
glabrata]
Length = 155
Score = 77.8 bits (190), Expect = 3e-13
Identities = 38/96 (39%), Positives = 51/96 (52%), Gaps = 14/96 (14%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C+ CG+ GH + C Q RCFNCNQ GH++ C KG+F A + K
Sbjct: 49 CYNCGETGHVKSECSIQ--RCFNCNQTGHVSRECP--------EPRKGRFGAAS----KN 94
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVC 203
VSCY+CG H A C T +C++C R GH++ C
Sbjct: 95 VSCYKCGGPNHVARDCMQTDTKCYSCGRFGHVSRDC 130
Score = 70.9 bits (172), Expect = 4e-11
Identities = 36/100 (36%), Positives = 49/100 (49%), Gaps = 21/100 (21%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
C+ CGK GH A+ C + C+NCNQ GH+ C + T +
Sbjct: 6 CYVCGKIGHLADDCDSER-LCYNCNQPGHVQSECTMPR------------------TVEH 46
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPK 207
CY CGE GH S CS RCFNC++ GH++ C +P+
Sbjct: 47 KQCYNCGETGHVKSECSIQ--RCFNCNQTGHVSRECPEPR 84
Score = 50.1 bits (118), Expect = 8e-05
Identities = 23/86 (26%), Positives = 38/86 (43%), Gaps = 20/86 (23%)
Query: 98 RSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKF 157
R +A V+C++CG H A C+ +C++C + GH++ +C GP++
Sbjct: 87 RFGAASKNVSCYKCGGPNHVARDCMQTDTKCYSCGRFGHVSRDCPN---GPNEK------ 137
Query: 158 PAIARGTEKQVSCYRCGEIGHFASRC 183
CY C E GH + C
Sbjct: 138 -----------VCYNCNETGHISRDC 152
Score = 42.7 bits (99), Expect = 0.012
Identities = 17/41 (41%), Positives = 25/41 (60%), Gaps = 1/41 (2%)
Query: 167 QVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPK 207
Q +CY CG+IGH A C + C+NC++ GH+ C P+
Sbjct: 3 QKACYVCGKIGHLADDCDSE-RLCYNCNQPGHVQSECTMPR 42
>UniRef100_P36627 Cellular nucleic acid binding protein homolog [Schizosaccharomyces
pombe]
Length = 179
Score = 77.8 bits (190), Expect = 3e-13
Identities = 42/140 (30%), Positives = 64/140 (45%), Gaps = 17/140 (12%)
Query: 87 FKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLG-----QGPRCFNCNQIGHLAVNC 141
+ + T + E TC+ CG GH C QG C+ C ++GH+A +C
Sbjct: 39 YNCNQTGHKASECTEPQQEKTCYACGTAGHLVRDCPSSPNPRQGAECYKCGRVGHIARDC 98
Query: 142 KT------FQAGPSDNKMK----GKFPAIARGTEKQVSCYRCGEIGHFASRC--SATGPR 189
+T + G + M G + AR V CY CG+IGH + C ++ G
Sbjct: 99 RTNGQQSGGRFGGHRSNMNCYACGSYGHQARDCTMGVKCYSCGKIGHRSFECQQASDGQL 158
Query: 190 CFNCHRVGHLAVVCKKPKVE 209
C+ C++ GH+AV C P +E
Sbjct: 159 CYKCNQPGHIAVNCTSPVIE 178
Score = 73.2 bits (178), Expect = 9e-12
Identities = 36/110 (32%), Positives = 50/110 (44%), Gaps = 26/110 (23%)
Query: 100 QSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
Q+ P C+ CG+NGH A C +G C+NCNQ GH A C
Sbjct: 11 QTTRPGPRCYNCGENGHQARECT-KGSICYNCNQTGHKASECTE---------------- 53
Query: 160 IARGTEKQVSCYRCGEIGHFASRCSAT-----GPRCFNCHRVGHLAVVCK 204
+++ +CY CG GH C ++ G C+ C RVGH+A C+
Sbjct: 54 ----PQQEKTCYACGTAGHLVRDCPSSPNPRQGAECYKCGRVGHIARDCR 99
Score = 48.5 bits (114), Expect = 2e-04
Identities = 21/52 (40%), Positives = 29/52 (55%), Gaps = 1/52 (1%)
Query: 158 PAIARGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
P + + T CY CGE GH A C+ G C+NC++ GH A C +P+ E
Sbjct: 7 PTVPQTTRPGPRCYNCGENGHQARECT-KGSICYNCNQTGHKASECTEPQQE 57
>UniRef100_P62633 Cellular nucleic acid binding protein [Homo sapiens]
Length = 177
Score = 77.4 bits (189), Expect = 5e-13
Identities = 49/149 (32%), Positives = 67/149 (44%), Gaps = 25/149 (16%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHL 137
+GRG R + T+ G + S+ C+RCG++GH A C Q C+NC + GH+
Sbjct: 25 RGRGMRSRG-RGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHI 83
Query: 138 AVNCK---------TFQAGPSDNKMK-------------GKFPAIARGTEKQVSCYRCGE 175
A +CK + G + + G+F I + K V CYRCGE
Sbjct: 84 AKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCGE 142
Query: 176 IGHFASRCSATGP-RCFNCHRVGHLAVVC 203
GH A CS T C+ C GHLA C
Sbjct: 143 TGHVAINCSKTSEVNCYRCGESGHLAREC 171
Score = 65.1 bits (157), Expect = 2e-09
Identities = 36/102 (35%), Positives = 45/102 (43%), Gaps = 14/102 (13%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R G F + + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGR-----GGFTSDRGFQFVSSSLPDI------- 53
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+PK E
Sbjct: 54 --CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKRE 93
Score = 59.3 bits (142), Expect = 1e-07
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 95 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 148
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 149 NCSKTSEVNCYRCGESGHLARECT 172
>UniRef100_UPI000042E8CA UPI000042E8CA UniRef100 entry
Length = 175
Score = 76.3 bits (186), Expect = 1e-12
Identities = 35/98 (35%), Positives = 52/98 (52%), Gaps = 6/98 (6%)
Query: 108 CFRCGKNGHYANAC--LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTE 165
C+ CG GH C QG +C+NC Q GH++ NC + + ++N K P+ G
Sbjct: 53 CYSCGDVGHIQTECPNQAQGAKCYNCGQFGHISKNCDSAPSS-TNNAPSFKRPS---GRA 108
Query: 166 KQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVC 203
+CY+CG HFA C A +C+ C +VGH++ C
Sbjct: 109 SGTTCYKCGGPNHFARDCQANTVKCYACGKVGHISKDC 146
Score = 71.2 bits (173), Expect = 3e-11
Identities = 37/119 (31%), Positives = 58/119 (48%), Gaps = 21/119 (17%)
Query: 107 TCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEK 166
TC++CG+ GH A+ C + C+NC++ GH + +C P + T K
Sbjct: 8 TCYKCGEVGHVADDCQQEERLCYNCHKPGHESNDC----------------PDPKQNTAK 51
Query: 167 QVSCYRCGEIGHFASRC--SATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAAR 223
Q CY CG++GH + C A G +C+NC + GH++ C + N K P+ R
Sbjct: 52 Q--CYSCGDVGHIQTECPNQAQGAKCYNCGQFGHISKNCDSAPSSTN-NAPSFKRPSGR 107
Score = 58.9 bits (141), Expect = 2e-07
Identities = 30/101 (29%), Positives = 47/101 (45%), Gaps = 16/101 (15%)
Query: 86 SFKSSTTATPGARSQSAHPE-VTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTF 144
S SST P + S TC++CG H+A C +C+ C ++GH++ +C +
Sbjct: 90 SAPSSTNNAPSFKRPSGRASGTTCYKCGGPNHFARDCQANTVKCYACGKVGHISKDCHS- 148
Query: 145 QAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSA 185
AG S+ K +CY CG+ GH + C+A
Sbjct: 149 SAGGSNFSAK--------------TCYNCGKSGHISKECTA 175
Score = 52.0 bits (123), Expect = 2e-05
Identities = 26/70 (37%), Positives = 37/70 (52%), Gaps = 5/70 (7%)
Query: 169 SCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVEPSVNTARGKHPAAR-GKVY 227
+CY+CGE+GH A C C+NCH+ GH + C PK NTA+ + G +
Sbjct: 8 TCYKCGEVGHVADDCQQEERLCYNCHKPGHESNDCPDPK----QNTAKQCYSCGDVGHIQ 63
Query: 228 TMDGEEAEGA 237
T +A+GA
Sbjct: 64 TECPNQAQGA 73
>UniRef100_UPI00003C130E UPI00003C130E UniRef100 entry
Length = 189
Score = 76.3 bits (186), Expect = 1e-12
Identities = 41/117 (35%), Positives = 57/117 (48%), Gaps = 22/117 (18%)
Query: 107 TCFRCGKNGHYANACL--------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFP 158
TC++C + GH + C G G C+ C Q GH+A C T AG S +G F
Sbjct: 61 TCYKCSETGHISRECPTNPAPAAGGPGGECYKCGQHGHIARACPT--AGGSS---RGGFG 115
Query: 159 AIARGTEKQVSCYRCGEIGHFASRCS------ATGPRCFNCHRVGHLAVVCKKPKVE 209
G SCY CG +GH + C+ A G RC+NC+ GH++ C KP+ +
Sbjct: 116 GARSGGR---SCYNCGGVGHLSRECTSPAGAAAGGQRCYNCNESGHISRECPKPQTK 169
Score = 55.5 bits (132), Expect = 2e-06
Identities = 38/117 (32%), Positives = 47/117 (39%), Gaps = 27/117 (23%)
Query: 115 GHYANACLGQG-PRCFNCNQIGHLAVNC----------KTFQAGPSDNKMKGKFPAIARG 163
GH A AC G P C+NC Q GH++ C K + G + PA A G
Sbjct: 26 GHNAAACPTAGNPSCYNCGQQGHISSQCGMEAQPKTCYKCSETGHISRECPTN-PAPAAG 84
Query: 164 TEKQVSCYRCGEIGHFASRCSAT--------------GPRCFNCHRVGHLAVVCKKP 206
CY+CG+ GH A C G C+NC VGHL+ C P
Sbjct: 85 GPGG-ECYKCGQHGHIARACPTAGGSSRGGFGGARSGGRSCYNCGGVGHLSRECTSP 140
Score = 52.8 bits (125), Expect = 1e-05
Identities = 32/100 (32%), Positives = 46/100 (46%), Gaps = 21/100 (21%)
Query: 104 PEVTCFRCGKNGHYANAC--------------LGQGPRCFNCNQIGHLAVNCKTFQAGPS 149
P C++CG++GH A AC G C+NC +GHL+ C T AG +
Sbjct: 86 PGGECYKCGQHGHIARACPTAGGSSRGGFGGARSGGRSCYNCGGVGHLSREC-TSPAGAA 144
Query: 150 DNKMK----GKFPAIARGTEKQV--SCYRCGEIGHFASRC 183
+ + I+R K SCYRCG+ GH ++ C
Sbjct: 145 AGGQRCYNCNESGHISRECPKPQTKSCYRCGDEGHLSAAC 184
Score = 36.6 bits (83), Expect = 0.88
Identities = 27/94 (28%), Positives = 35/94 (36%), Gaps = 16/94 (17%)
Query: 54 RAGTGGPVRSGSQEYRQLGPYQHPKGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGK 113
R G GG RSG + G G G R S A G + C+ C +
Sbjct: 111 RGGFGG-ARSGGRSCYNCG------GVGHLSRECTSPAGAAAGGQR--------CYNCNE 155
Query: 114 NGHYANAC-LGQGPRCFNCNQIGHLAVNCKTFQA 146
+GH + C Q C+ C GHL+ C A
Sbjct: 156 SGHISRECPKPQTKSCYRCGDEGHLSAACPQIAA 189
>UniRef100_Q9D548 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4930513O09 product:similar to CELLULAR
NUCLEIC ACID BINDING PROTEIN [Mus musculus]
Length = 170
Score = 75.9 bits (185), Expect = 1e-12
Identities = 47/134 (35%), Positives = 62/134 (46%), Gaps = 13/134 (9%)
Query: 77 PKG--RGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQI 134
PKG RG T R T G + +A+ C+RCG+ GHYA C C+NC +
Sbjct: 20 PKGGTRGRTARG------RTRGPQCSTANQSDVCYRCGETGHYAKDCDLLQDTCYNCGRR 73
Query: 135 GHLAVNCKTFQAGPSDNKMKGKFPA-IARGTEKQ--VSCYRCGEIGHFASRCSATGPRCF 191
GH+A +C + P +AR +Q CY CGE GH C T +C+
Sbjct: 74 GHIAKDCTQAKREREQCCYICSQPGHLARDCNRQEEQKCYTCGEFGHIQKDC--TQIKCY 131
Query: 192 NCHRVGHLAVVCKK 205
C GH+AV C K
Sbjct: 132 RCGENGHMAVNCSK 145
Score = 61.2 bits (147), Expect = 3e-08
Identities = 34/103 (33%), Positives = 47/103 (45%), Gaps = 21/103 (20%)
Query: 107 TCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEK 166
+CF+CG +GH+A C G R G A + + + P + +
Sbjct: 5 SCFKCGHSGHWARECPKGGTR-------GRTA-------------RGRTRGPQCSTANQS 44
Query: 167 QVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
V CYRCGE GH+A C C+NC R GH+A C + K E
Sbjct: 45 DV-CYRCGETGHYAKDCDLLQDTCYNCGRRGHIAKDCTQAKRE 86
>UniRef100_Q8T8R1 GM14667p [Drosophila melanogaster]
Length = 165
Score = 75.9 bits (185), Expect = 1e-12
Identities = 31/102 (30%), Positives = 51/102 (49%), Gaps = 6/102 (5%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNC------KTFQAGPSDNKMKGKFPAIA 161
C++C + GH+A AC + RC+ CN IGH++ +C ++ + + ++ A+
Sbjct: 57 CYKCNQFGHFARACPEEAERCYRCNGIGHISKDCTQADNPTCYRCNKTGHWVRNCPEAVN 116
Query: 162 RGTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVC 203
VSCY+C GH + C T C+ C + GHL C
Sbjct: 117 ERGPTNVSCYKCNRTGHISKNCPETSKTCYGCGKSGHLRREC 158
Score = 56.2 bits (134), Expect = 1e-06
Identities = 34/128 (26%), Positives = 46/128 (35%), Gaps = 53/128 (41%)
Query: 107 TCFRCGKNGHYANACL---GQGP---------------------------RCFNCNQIGH 136
TC++C + GH+A C G GP +C+ CNQ GH
Sbjct: 6 TCYKCNRPGHFARDCSLGGGGGPGGVGGGGGGGGGGMRGNDGGGMRRNREKCYKCNQFGH 65
Query: 137 LAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCS-ATGPRCFNCHR 195
A C E+ CYRC IGH + C+ A P C+ C++
Sbjct: 66 FARAC----------------------PEEAERCYRCNGIGHISKDCTQADNPTCYRCNK 103
Query: 196 VGHLAVVC 203
GH C
Sbjct: 104 TGHWVRNC 111
Score = 50.8 bits (120), Expect = 5e-05
Identities = 27/90 (30%), Positives = 36/90 (40%), Gaps = 29/90 (32%)
Query: 107 TCFRCGKNGHYANAC----LGQGP---RCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPA 159
TC+RC K GH+ C +GP C+ CN+ GH++ NC
Sbjct: 97 TCYRCNKTGHWVRNCPEAVNERGPTNVSCYKCNRTGHISKNC------------------ 138
Query: 160 IARGTEKQVSCYRCGEIGHFASRCSATGPR 189
E +CY CG+ GH C G R
Sbjct: 139 ----PETSKTCYGCGKSGHLRRECDEKGGR 164
Score = 39.7 bits (91), Expect = 0.10
Identities = 14/47 (29%), Positives = 23/47 (48%)
Query: 95 PGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNC 141
P A ++ V+C++C + GH + C C+ C + GHL C
Sbjct: 112 PEAVNERGPTNVSCYKCNRTGHISKNCPETSKTCYGCGKSGHLRREC 158
>UniRef100_Q86EQ4 Clone ZZD1536 mRNA sequence [Schistosoma japonicum]
Length = 192
Score = 75.9 bits (185), Expect = 1e-12
Identities = 39/115 (33%), Positives = 52/115 (44%), Gaps = 35/115 (30%)
Query: 108 CFRCGKNGHYANACL----------------GQGPRCFNCNQIGHLAVNCKTFQAGPSDN 151
CF CG GH+A C G G RC+NC Q GH+ NC PS+N
Sbjct: 90 CFNCGGVGHFARECTNDGQRGDSGYNNGGGGGGGGRCYNCGQSGHVVRNC------PSNN 143
Query: 152 KMKGKFPAIARGTEKQVSCYRCGEIGHFASRCS---ATGPRCFNCHRVGHLAVVC 203
R ++ CYRC + GH+A C+ +GP+C+ C GH+A C
Sbjct: 144 ----------RNDMSEILCYRCNKYGHYAKECTESGGSGPQCYKCRGYGHIASRC 188
Score = 53.1 bits (126), Expect = 9e-06
Identities = 38/141 (26%), Positives = 51/141 (35%), Gaps = 51/141 (36%)
Query: 108 CFRCGKNGHYANACLGQ--------------------------GPR--CFNCNQIGHLAV 139
CF+CG+ GH+A C Q G R CFNC + H A
Sbjct: 5 CFKCGREGHFARDCQAQSRGGRGGGGGYRGRGGGGGRDRDNNDGRRDGCFNCGGLDHYAR 64
Query: 140 NCKTFQAGPSD-NKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCS-------------- 184
+C P+D G G + C+ CG +GHFA C+
Sbjct: 65 DC------PNDRGHYGGGGGGGYGGYGSRDKCFNCGGVGHFARECTNDGQRGDSGYNNGG 118
Query: 185 --ATGPRCFNCHRVGHLAVVC 203
G RC+NC + GH+ C
Sbjct: 119 GGGGGGRCYNCGQSGHVVRNC 139
Score = 50.4 bits (119), Expect = 6e-05
Identities = 19/40 (47%), Positives = 24/40 (59%), Gaps = 3/40 (7%)
Query: 105 EVTCFRCGKNGHYANACL---GQGPRCFNCNQIGHLAVNC 141
E+ C+RC K GHYA C G GP+C+ C GH+A C
Sbjct: 149 EILCYRCNKYGHYAKECTESGGSGPQCYKCRGYGHIASRC 188
>UniRef100_P53996 Cellular nucleic acid binding protein [Mus musculus]
Length = 178
Score = 75.9 bits (185), Expect = 1e-12
Identities = 49/150 (32%), Positives = 68/150 (44%), Gaps = 26/150 (17%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGH 136
+GRG R + T+ G + S+ C+RCG++GH A C L + C+NC + GH
Sbjct: 25 RGRGMRSRG-RGGFTSDRGFQFVSSSLPDICYRCGESGHLAKDCDLQEDEACYNCGRGGH 83
Query: 137 LAVNCKT---------FQAGPSDNKMK-------------GKFPAIARGTEKQVSCYRCG 174
+A +CK + G + + G+F I + K V CYRCG
Sbjct: 84 IAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCG 142
Query: 175 EIGHFASRCSATGP-RCFNCHRVGHLAVVC 203
E GH A CS T C+ C GHLA C
Sbjct: 143 ETGHVAINCSKTSEVNCYRCGESGHLAREC 172
Score = 60.8 bits (146), Expect = 4e-08
Identities = 36/103 (34%), Positives = 45/103 (42%), Gaps = 15/103 (14%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R G F + + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGR-----GGFTSDRGFQFVSSSLPDI------- 53
Query: 168 VSCYRCGEIGHFASRCS-ATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+PK E
Sbjct: 54 --CYRCGESGHLAKDCDLQEDEACYNCGRGGHIAKDCKEPKRE 94
Score = 59.3 bits (142), Expect = 1e-07
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 96 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 149
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 150 NCSKTSEVNCYRCGESGHLARECT 173
>UniRef100_Q8WW36 Zinc finger, CCHC domain containing 13 [Homo sapiens]
Length = 166
Score = 75.5 bits (184), Expect = 2e-12
Identities = 37/119 (31%), Positives = 62/119 (52%), Gaps = 13/119 (10%)
Query: 96 GARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMK- 154
G++ S TC+ CG++G A C+ G C+NC + GH+A +CK P + +
Sbjct: 35 GSQCGSTTLSYTCYCCGESGRNAKNCVLLGNICYNCGRSGHIAKDCK----DPKRERRQH 90
Query: 155 ----GKFPAIAR--GTEKQVSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPK 207
G+ +AR +K+ CY CG++GH C+ +C+ C +GH+A+ C K +
Sbjct: 91 CYTCGRLGHLARDCDRQKEQKCYSCGKLGHIQKDCAQV--KCYRCGEIGHVAINCSKAR 147
Score = 55.1 bits (131), Expect = 2e-06
Identities = 31/81 (38%), Positives = 35/81 (42%), Gaps = 1/81 (1%)
Query: 129 FNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATGP 188
F C GH A C AG G+ T +CY CGE G A C G
Sbjct: 7 FACGHSGHWARGCPRGGAGGRRGGGHGRGSQCG-STTLSYTCYCCGESGRNAKNCVLLGN 65
Query: 189 RCFNCHRVGHLAVVCKKPKVE 209
C+NC R GH+A CK PK E
Sbjct: 66 ICYNCGRSGHIAKDCKDPKRE 86
Score = 45.8 bits (107), Expect = 0.001
Identities = 33/127 (25%), Positives = 47/127 (36%), Gaps = 29/127 (22%)
Query: 109 FRCGKNGHYANACL------------GQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGK 156
F CG +GH+A C G+G +C G ++ + G S K
Sbjct: 7 FACGHSGHWARGCPRGGAGGRRGGGHGRGSQC------GSTTLSYTCYCCGESGRNAKN- 59
Query: 157 FPAIARGTEKQVSCYRCGEIGHFASRCS----ATGPRCFNCHRVGHLAVVCKKPKVEPSV 212
+ G CY CG GH A C C+ C R+GHLA C + K +
Sbjct: 60 --CVLLGN----ICYNCGRSGHIAKDCKDPKRERRQHCYTCGRLGHLARDCDRQKEQKCY 113
Query: 213 NTARGKH 219
+ + H
Sbjct: 114 SCGKLGH 120
Score = 44.7 bits (104), Expect = 0.003
Identities = 19/51 (37%), Positives = 26/51 (50%), Gaps = 2/51 (3%)
Query: 97 ARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAG 147
AR E C+ CGK GH C +C+ C +IGH+A+NC + G
Sbjct: 101 ARDCDRQKEQKCYSCGKLGHIQKDCAQV--KCYRCGEIGHVAINCSKARPG 149
>UniRef100_Q9W6Q5 Cellular nucleic acid binding protein [Bufo arenarum]
Length = 178
Score = 75.1 bits (183), Expect = 2e-12
Identities = 44/135 (32%), Positives = 66/135 (48%), Gaps = 13/135 (9%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHL 137
+GRG + +++ G + S+ C+RCG++GH A C Q C+NC + GH+
Sbjct: 25 RGRGGGRGRGRGGFSSSRGFQFISSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHI 84
Query: 138 AVNCKTFQAGPSDNKMK-----GKFPAIARGTE--KQVSCYRCGEIGHFASRCSATGPRC 190
A +CK P + + GK +AR E + CY CGE GH C T +C
Sbjct: 85 AKDCKE----PRKEREQCCYNCGKPGHLARDCEHADEQKCYSCGEFGHIQKDC--TKVKC 138
Query: 191 FNCHRVGHLAVVCKK 205
+ C GH+A+ C K
Sbjct: 139 YRCGDTGHVAINCSK 153
Score = 63.9 bits (154), Expect = 5e-09
Identities = 35/102 (34%), Positives = 45/102 (43%), Gaps = 13/102 (12%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG+ GH+A C G R + G F + + P I
Sbjct: 6 CFKCGRTGHWARECPTGGGR----GRGGGRGRGRGGFSSSRGFQFISSSLPDI------- 54
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+P+ E
Sbjct: 55 --CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPRKE 94
Score = 59.3 bits (142), Expect = 1e-07
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANACL-GQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 96 EQCCYNCGKPGHLARDCEHADEQKCYSCGEFGHIQKDCTKVKCYRCGDTGHV------AI 149
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 150 NCSKTSEVNCYRCGESGHLARECT 173
>UniRef100_P70000 Cellular nucleic acid binding protein [Xenopus laevis]
Length = 178
Score = 75.1 bits (183), Expect = 2e-12
Identities = 44/135 (32%), Positives = 66/135 (48%), Gaps = 13/135 (9%)
Query: 78 KGRGFTPRSFKSSTTATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHL 137
+GRG + +++ G + S+ C+RCG++GH A C Q C+NC + GH+
Sbjct: 25 RGRGGGRGRGRGGFSSSRGFQFISSSLPDICYRCGESGHLAKDCDLQEDACYNCGRGGHI 84
Query: 138 AVNCKTFQAGPSDNKMK-----GKFPAIARGTE--KQVSCYRCGEIGHFASRCSATGPRC 190
A +CK P + + GK +AR E + CY CGE GH C T +C
Sbjct: 85 AKDCKE----PRKEREQCCYNCGKPGHLARDCEHADEQKCYSCGEFGHIQKDC--TKVKC 138
Query: 191 FNCHRVGHLAVVCKK 205
+ C GH+A+ C K
Sbjct: 139 YRCGDTGHVAINCSK 153
Score = 64.3 bits (155), Expect = 4e-09
Identities = 35/102 (34%), Positives = 46/102 (44%), Gaps = 13/102 (12%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R + G F + + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGR----GRGGGRGRGRGGFSSSRGFQFISSSLPDI------- 54
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+P+ E
Sbjct: 55 --CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPRKE 94
Score = 59.3 bits (142), Expect = 1e-07
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANACL-GQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 96 EQCCYNCGKPGHLARDCEHADEQKCYSCGEFGHIQKDCTKVKCYRCGDTGHV------AI 149
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 150 NCSKTSEVNCYRCGESGHLARECT 173
>UniRef100_UPI00003AA82A UPI00003AA82A UniRef100 entry
Length = 171
Score = 74.3 bits (181), Expect = 4e-12
Identities = 44/126 (34%), Positives = 60/126 (46%), Gaps = 25/126 (19%)
Query: 101 SAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCK---------TFQAGPSDN 151
S+ P++ C+RCG++GH A C Q C+NC + GH+A +CK + G +
Sbjct: 42 SSLPDI-CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGH 100
Query: 152 KMK-------------GKFPAIARGTEKQVSCYRCGEIGHFASRCSATGP-RCFNCHRVG 197
+ G+F I + K V CYRCGE GH A CS T C+ C G
Sbjct: 101 LARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCGETGHVAINCSKTSEVNCYRCGESG 159
Query: 198 HLAVVC 203
HLA C
Sbjct: 160 HLAREC 165
Score = 65.1 bits (157), Expect = 2e-09
Identities = 31/82 (37%), Positives = 39/82 (46%)
Query: 128 CFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQVSCYRCGEIGHFASRCSATG 187
CF C + GH A C T + +G+ + CYRCGE GH A C
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 188 PRCFNCHRVGHLAVVCKKPKVE 209
C+NC R GH+A CK+PK E
Sbjct: 66 DACYNCGRGGHIAKDCKEPKRE 87
Score = 59.3 bits (142), Expect = 1e-07
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 89 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 142
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 143 NCSKTSEVNCYRCGESGHLARECT 166
>UniRef100_UPI000042FF1E UPI000042FF1E UniRef100 entry
Length = 184
Score = 74.3 bits (181), Expect = 4e-12
Identities = 42/120 (35%), Positives = 50/120 (41%), Gaps = 31/120 (25%)
Query: 93 ATPGARSQSAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNK 152
A PG+R CF+CG GH A C G C+NC + GH + NC P
Sbjct: 6 AVPGSRQG-------CFKCGNLGHIAENCQAPGRLCYNCREPGHESTNC------PQPRS 52
Query: 153 MKGKFPAIARGTEKQVSCYRCGEIGHFASRCSA------TGPRCFNCHRVGHLAVVCKKP 206
GK CY CG +GH S C + G +CF C R GHLA C P
Sbjct: 53 TDGK------------QCYACGGVGHVKSDCPSMRGAFGPGQKCFKCGRPGHLARECTVP 100
Score = 59.3 bits (142), Expect = 1e-07
Identities = 37/125 (29%), Positives = 48/125 (37%), Gaps = 23/125 (18%)
Query: 108 CFRCGKNGHYANAC------LGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIA 161
C+ CG GH + C G G +CF C + GHLA C T + +G F
Sbjct: 58 CYACGGVGHVKSDCPSMRGAFGPGQKCFKCGRPGHLAREC-TVPGFVGAFRGRGGFGGAF 116
Query: 162 RGTEKQ--------VSCYRCGEIGHFASRCSA--------TGPRCFNCHRVGHLAVVCKK 205
G + V CYRC H A C A +C+ C GH+A C +
Sbjct: 117 GGRPRPPINPDGTPVKCYRCNGENHLARDCLAPRDEAAILASKKCYKCQETGHIARDCTQ 176
Query: 206 PKVEP 210
V P
Sbjct: 177 ENVSP 181
Score = 48.9 bits (115), Expect = 2e-04
Identities = 16/38 (42%), Positives = 23/38 (60%)
Query: 170 CYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPK 207
C++CG +GH A C A G C+NC GH + C +P+
Sbjct: 14 CFKCGNLGHIAENCQAPGRLCYNCREPGHESTNCPQPR 51
>UniRef100_Q5R7R4 Hypothetical protein DKFZp470E2414 [Pongo pygmaeus]
Length = 170
Score = 74.3 bits (181), Expect = 4e-12
Identities = 44/126 (34%), Positives = 60/126 (46%), Gaps = 25/126 (19%)
Query: 101 SAHPEVTCFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKT---------FQAGPSDN 151
S+ P++ C+RCG++GH A C Q C+NC + GH+A +CK + G +
Sbjct: 41 SSLPDI-CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGH 99
Query: 152 KMK-------------GKFPAIARGTEKQVSCYRCGEIGHFASRCSATGP-RCFNCHRVG 197
+ G+F I + K V CYRCGE GH A CS T C+ C G
Sbjct: 100 LARDCDHADEQKCYSCGEFGHIQKDCTK-VKCYRCGETGHVAINCSKTSEVNCYRCGESG 158
Query: 198 HLAVVC 203
HLA C
Sbjct: 159 HLAREC 164
Score = 64.3 bits (155), Expect = 4e-09
Identities = 37/102 (36%), Positives = 46/102 (44%), Gaps = 21/102 (20%)
Query: 108 CFRCGKNGHYANACLGQGPRCFNCNQIGHLAVNCKTFQAGPSDNKMKGKFPAIARGTEKQ 167
CF+CG++GH+A C G R G + FQ + P I
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRG------RGFQF------VSSSLPDI------- 46
Query: 168 VSCYRCGEIGHFASRCSATGPRCFNCHRVGHLAVVCKKPKVE 209
CYRCGE GH A C C+NC R GH+A CK+PK E
Sbjct: 47 --CYRCGESGHLAKDCDLQEDACYNCGRGGHIAKDCKEPKRE 86
Score = 59.3 bits (142), Expect = 1e-07
Identities = 29/84 (34%), Positives = 43/84 (50%), Gaps = 10/84 (11%)
Query: 105 EVTCFRCGKNGHYANAC-LGQGPRCFNCNQIGHLAVNC---KTFQAGPSDNKMKGKFPAI 160
E C+ CGK GH A C +C++C + GH+ +C K ++ G + + AI
Sbjct: 88 EQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCTKVKCYRCGETGHV------AI 141
Query: 161 ARGTEKQVSCYRCGEIGHFASRCS 184
+V+CYRCGE GH A C+
Sbjct: 142 NCSKTSEVNCYRCGESGHLARECT 165
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.134 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 526,580,584
Number of Sequences: 2790947
Number of extensions: 22180734
Number of successful extensions: 83766
Number of sequences better than 10.0: 3457
Number of HSP's better than 10.0 without gapping: 362
Number of HSP's successfully gapped in prelim test: 3103
Number of HSP's that attempted gapping in prelim test: 67017
Number of HSP's gapped (non-prelim): 14719
length of query: 311
length of database: 848,049,833
effective HSP length: 127
effective length of query: 184
effective length of database: 493,599,564
effective search space: 90822319776
effective search space used: 90822319776
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0134.4


