

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.3
(700 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
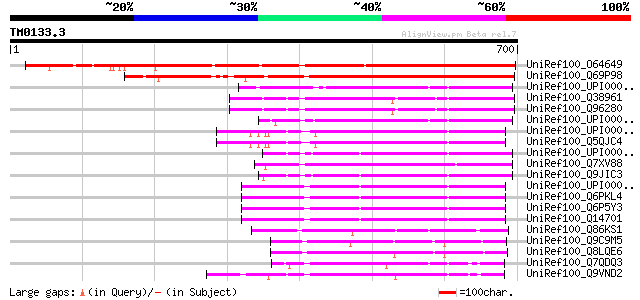
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O64649 Hypothetical protein At2g45700 [Arabidopsis tha... 655 0.0
UniRef100_Q69P98 Putative SNM1 [Oryza sativa] 627 e-178
UniRef100_UPI00003632EB UPI00003632EB UniRef100 entry 239 2e-61
UniRef100_Q38961 Hypothetical protein At3g26680/MLJ15_8 [Arabido... 228 4e-58
UniRef100_Q96280 Orf12 protein [Arabidopsis thaliana] 228 4e-58
UniRef100_UPI00004364BC UPI00004364BC UniRef100 entry 223 2e-56
UniRef100_UPI00003AE3F5 UPI00003AE3F5 UniRef100 entry 219 3e-55
UniRef100_Q5QJC4 SNM1A [Gallus gallus] 218 4e-55
UniRef100_UPI00001CEC53 UPI00001CEC53 UniRef100 entry 214 9e-54
UniRef100_Q7XV88 OSJNBb0012E08.8 protein [Oryza sativa] 212 3e-53
UniRef100_Q9JIC3 SNM1 protein [Mus musculus] 208 5e-52
UniRef100_UPI000036E972 UPI000036E972 UniRef100 entry 206 2e-51
UniRef100_Q6PKL4 DNA cross-link repair 1A [Homo sapiens] 206 2e-51
UniRef100_Q6P5Y3 DNA-crosslink repair gene SNM1 [Homo sapiens] 206 2e-51
UniRef100_Q14701 KIAA0086 protein [Homo sapiens] 206 2e-51
UniRef100_Q86KS1 Similar to Arabidopsis thaliana (Mouse-ear cres... 196 1e-48
UniRef100_Q9C9M5 DNA ligase I, putative [Arabidopsis thaliana] 181 5e-44
UniRef100_Q8LQE6 Putative DNA ligase [Oryza sativa] 176 3e-42
UniRef100_Q7QDQ3 ENSANGP00000016032 [Anopheles gambiae str. PEST] 170 1e-40
UniRef100_Q9VND2 CG10018-PA [Drosophila melanogaster] 167 7e-40
>UniRef100_O64649 Hypothetical protein At2g45700 [Arabidopsis thaliana]
Length = 723
Score = 655 bits (1689), Expect = 0.0
Identities = 372/731 (50%), Positives = 464/731 (62%), Gaps = 71/731 (9%)
Query: 22 DDDEFEIPLTQTATIHRPLKKHKLETTAFGGS----------GKENVPPNASSSTIHDDG 71
DDD+F+IP + +I +PL + GKENV P S
Sbjct: 8 DDDDFQIPPSSQLSIRKPLHPTNANNISHRPPNKKPRLCRYPGKENVTPPPSPDPDLFCS 67
Query: 72 YEIENCSLDFIPSTIDSVGTALTDSSSSSSSASVSAFVISSPVKVKMKGTAYFGNSIESK 131
+C LD IPS++D +L D + SS +K+ Y NS+E++
Sbjct: 68 SSTPHCILDCIPSSVDC---SLGDFNGPISSLGEEDKEDKDDC-IKVNREGYLCNSMEAR 123
Query: 132 LVVSRA-----NAINN-----AEAGSEL----NLSDELDG-----------SVRCPLCEV 166
L+ SR + I+ E+ SEL NL E +G S++CPLC +
Sbjct: 124 LLKSRICLGFDSGIHEDDEGFVESNSELDVLINLCSESEGRSGEFSLGKDDSIQCPLCSM 183
Query: 167 DISNLTEEQRHLHTNDCLDRGDDVAVRDDNVDA-------------------QFAPKVSR 207
DIS+L+EEQR +H+N CLD+ + D++ Q +S
Sbjct: 184 DISSLSEEQRQVHSNTCLDKSYNQPSEQDSLRKCENLSSLIKESIDDPVQLPQLVTDLSP 243
Query: 208 VVDWLRDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGI 267
V+ WLR LGL KYEDVF+REE+DWDTLQ LTEEDLLS+G+T+LGPRKKIV+AL R
Sbjct: 244 VLKWLRSLGLAKYEDVFIREEIDWDTLQSLTEEDLLSIGITSLGPRKKIVNALSGVRDPF 303
Query: 268 AASNEKPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFATNGKKVS 327
A+S E + + + + + RK KP ANK ITE+FPG AT G K+
Sbjct: 304 ASSAEVQAQSHCT---SGHVTERQRDKSTTRKASEPKKPTANKLITEFFPGQATEGTKIR 360
Query: 328 TAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGDCSHW 387
TAP+ E S S S R + + N K + +P W I GTPFRVDAFKYL DC HW
Sbjct: 361 TAPKPVAEKSPSDSSSRR--AVRRNGNNGKSKVIPHWNCIPGTPFRVDAFKYLTRDCCHW 418
Query: 388 FLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEI 447
FLTHFHLD + L T G + A+LVNM IGIP+++L +L L QKV I
Sbjct: 419 FLTHFHLDHYQGL------TKSFSHGKIYCSLVTAKLVNMKIGIPWERLQVLDLGQKVNI 472
Query: 448 AGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTT 507
+GIDVTC DANHCPGSI+ILF+P NGKAVLHTGDFR+S+E++ L + +LILDTT
Sbjct: 473 SGIDVTCFDANHCPGSIMILFEPANGKAVLHTGDFRYSEEMS--NWLIGSHISSLILDTT 530
Query: 508 YCNPQYDFPKQEAVIQFVIDAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVN 567
YCNPQYDFPKQEAVIQFV++AIQAEAFNP TLFLIGSYTIGKERLFLEVAR LR K+Y+N
Sbjct: 531 YCNPQYDFPKQEAVIQFVVEAIQAEAFNPKTLFLIGSYTIGKERLFLEVARVLREKIYIN 590
Query: 568 AAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAF 627
AKL+LL+CL F ++D+QWFT+ E ES+IHV P+WTLAS KRLKHV+++Y +R+SLIVAF
Sbjct: 591 PAKLKLLECLGFSKDDIQWFTVKEEESHIHVVPLWTLASFKRLKHVANRYTNRYSLIVAF 650
Query: 628 SPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDG 687
SPTGWT GK KKKSPGRR Q GTIIRYEVPYSEH SFTELK+FV+ +SP IIPSVNNDG
Sbjct: 651 SPTGWTSGKTKKKSPGRRLQQGTIIRYEVPYSEHSSFTELKEFVQKVSPEVIIPSVNNDG 710
Query: 688 PESANAMISLM 698
P+SA AM+SL+
Sbjct: 711 PDSAAAMVSLL 721
>UniRef100_Q69P98 Putative SNM1 [Oryza sativa]
Length = 1024
Score = 627 bits (1618), Expect = e-178
Identities = 319/549 (58%), Positives = 400/549 (72%), Gaps = 40/549 (7%)
Query: 159 VRCPLCEVDISNLTEEQRHLHTNDCLDRGDDVAVRDDNVDAQFAP------KVSRVVDWL 212
V+CPLC +IS+L+EE R +HTN CLD D ++ N D Q P + RV++WL
Sbjct: 500 VQCPLCGSNISDLSEELRLVHTNSCLD--GDKPAKEPNSDNQNEPCGESNVEKRRVMEWL 557
Query: 213 RDLGLGKYEDVFVREEVDWDTLQWLTEEDLLSMGVTALGPRKKIVHALCEFRKGIAASNE 272
R+LGL KYE++F++EEVDW+TLQWLTEEDLL MG+T+LGPRKKI HALCE RK +N+
Sbjct: 558 RNLGLSKYEEIFIKEEVDWETLQWLTEEDLLGMGITSLGPRKKIAHALCELRKKNNDAND 617
Query: 273 KPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFATNGK-----KVS 327
D L +N K + + ++G NK ITEYF +++ + KV+
Sbjct: 618 LAADML------NLENTK----KAKIPMNG------NKLITEYFRCPSSDQRQKKACKVN 661
Query: 328 TAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGDCSHW 387
T + KS+ +G K K++D P WC I GTPFRVDAF+YLRGDC HW
Sbjct: 662 TPSNLNSQKKSNAKATGGRRTVKG-----KVKDTPIWCCIPGTPFRVDAFRYLRGDCCHW 716
Query: 388 FLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEI 447
FLTHFH+D + L + G + A LV+ IGIP+D+LH+LPLN+K+ I
Sbjct: 717 FLTHFHVDHYQGLTKSFCH------GKIYCSSVTANLVHYKIGIPWDRLHVLPLNEKITI 770
Query: 448 AGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTT 507
AG+++TC DANHCPG++IILF+P NGKAVLHTGDFRFS E+A N VL+ P+HTLILDTT
Sbjct: 771 AGVNLTCFDANHCPGAVIILFEPSNGKAVLHTGDFRFSSEMANNRVLQSSPIHTLILDTT 830
Query: 508 YCNPQYDFPKQEAVIQFVIDAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVN 567
YCNP+YDFP QE VIQFVI+AIQAEAFNP TLFLIGSYTIGKERL++EVAR L++K+YV
Sbjct: 831 YCNPRYDFPTQEIVIQFVIEAIQAEAFNPKTLFLIGSYTIGKERLYMEVARLLQKKIYVG 890
Query: 568 AAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAF 627
AAKL++LK L +E M WFT NE ES+IHV PMWTLAS KR+K++S+QYA RF LIVAF
Sbjct: 891 AAKLQILKHLGLPQEIMHWFTANEAESHIHVVPMWTLASFKRMKYLSTQYADRFDLIVAF 950
Query: 628 SPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDG 687
PTGW+FGKGKK++PGR+WQ G IIRYEVPYSEH SFTEL++FV+ ISP +IIPSVNNDG
Sbjct: 951 CPTGWSFGKGKKRTPGRKWQQGAIIRYEVPYSEHSSFTELREFVRFISPEHIIPSVNNDG 1010
Query: 688 PESANAMIS 696
P+SANAM++
Sbjct: 1011 PDSANAMLA 1019
>UniRef100_UPI00003632EB UPI00003632EB UniRef100 entry
Length = 385
Score = 239 bits (611), Expect = 2e-61
Identities = 145/383 (37%), Positives = 217/383 (55%), Gaps = 19/383 (4%)
Query: 317 PGFATNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDVPKWCSIEGTPFRVDA 376
P + G V++ ++Q E+K+ R +N+ +L P + I GT F +DA
Sbjct: 9 PAGTSRGLGVNSLVDDQGELKNHRR---RRWNRRNADGEVELPRCPFYKKIPGTKFAIDA 65
Query: 377 FKY-LRGDCSHWFLTHFHLDRTR-LLDLNIFDTDLRQM-GYLIKFECFARLVNMNIGIPY 433
F+Y + + +FLTHFH D L + F ++ G L+K + + + PY
Sbjct: 66 FRYGMIEGITAYFLTHFHSDHYGGLTKSSTFPVYCNKITGNLVKSK-------LKVAEPY 118
Query: 434 DKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPV 493
+H+LP+N +V + G+ V L+ANHCPG+ ++LF P+G+ VLHTGDFR + + P
Sbjct: 119 --IHVLPMNTQVTVEGVTVVLLEANHCPGAAMLLFFLPDGQIVLHTGDFRADPSMELYPE 176
Query: 494 LRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERL 552
L C V TL LDTTYC+P+Y FP Q+ VI F A + A NP TL + GSY++GKE++
Sbjct: 177 LLSCRVQTLYLDTTYCSPEYTFPTQQEVINFAASTAFELVALNPRTLVVCGSYSVGKEKV 236
Query: 553 FLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKH 612
F +A L KV ++ K + CLE E+ Q T + + +HV PM L + K+L+
Sbjct: 237 FFALADVLGSKVSLSRDKYNTMCCLE-SEQVKQCITTDWKAARVHVLPMMQL-TFKKLEQ 294
Query: 613 VSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFV 671
++++S++ +VAF PTGWTF + + G +G I Y +PYSEH SF E+K FV
Sbjct: 295 HLARFSSQYDQLVAFKPTGWTFSQQVESVGGIEPDVSGNISIYGIPYSEHSSFVEMKRFV 354
Query: 672 KHISPANIIPSVNNDGPESANAM 694
+ + P IIP+VNN ES AM
Sbjct: 355 QWLQPLKIIPTVNNGSWESRRAM 377
>UniRef100_Q38961 Hypothetical protein At3g26680/MLJ15_8 [Arabidopsis thaliana]
Length = 484
Score = 228 bits (582), Expect = 4e-58
Identities = 142/403 (35%), Positives = 224/403 (55%), Gaps = 24/403 (5%)
Query: 304 SKPAANKKI--TEYFPGFATNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDV 361
+ P KK+ T+ F + T+P K++ ++ R +S +N R
Sbjct: 88 NSPDTTKKMKQTDLFQSWGLQKPSPFTSPASNSAKKTTSALGKRRRD--SSFSNDSPRPC 145
Query: 362 PKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFE 419
P + + GTPF VDAF+Y ++G CS +FLTHFH D L T G +
Sbjct: 146 PFYKKLPGTPFTVDAFRYGCVQG-CSAYFLTHFHADHYIGL------TKAWSHGPIYCSS 198
Query: 420 CFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHT 479
+RL+ +++ + +H L L+ + I GI VT ++ANHCPG+ +I F+ +G LHT
Sbjct: 199 LTSRLLRLSLSVNPSSIHPLELDVEYTINGIKVTLIEANHCPGAALIHFRLLDGTCYLHT 258
Query: 480 GDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVI----DAIQAEAFN 535
GDFR S ++ +P+L VH L LDTTYCNP+Y FP +E V+ +V+ D ++ +
Sbjct: 259 GDFRASKQMQTHPLLFNQRVHVLYLDTTYCNPRYKFPSKEDVLSYVVRITKDFLRKQ--- 315
Query: 536 PGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESN 595
P TL ++GSY+IGKE ++L +A++L K++ NA++ R+L+ +++ T + +
Sbjct: 316 PKTLIVVGSYSIGKECVYLAIAKALGVKIFANASRRRILQSFGWDDISKNLST-DGKATC 374
Query: 596 IHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGK--GKKKSPGRRWQNGTIIR 653
+HV PM +L ++RL Y ++ ++AF PTGWT+ + G+ + G I
Sbjct: 375 LHVLPMSSL-KVERLDEHLKIYREQYGAVLAFRPTGWTYSEKIGEHLDLIKPTSRGKITI 433
Query: 654 YEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAMIS 696
Y VPYSEH SFTEL++FV+ + P IIP+VNN + M S
Sbjct: 434 YGVPYSEHSSFTELREFVQFLRPDKIIPTVNNGNAGTREKMQS 476
>UniRef100_Q96280 Orf12 protein [Arabidopsis thaliana]
Length = 484
Score = 228 bits (582), Expect = 4e-58
Identities = 142/403 (35%), Positives = 224/403 (55%), Gaps = 24/403 (5%)
Query: 304 SKPAANKKI--TEYFPGFATNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSSTNRKLRDV 361
+ P KK+ T+ F + T+P K++ ++ R +S +N R
Sbjct: 88 NSPDTTKKMKQTDLFQSWGLQKPSPFTSPASNSAKKTTSALGKRRRD--SSFSNDSPRPC 145
Query: 362 PKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFE 419
P + + GTPF VDAF+Y ++G CS +FLTHFH D L T G +
Sbjct: 146 PFYKKLPGTPFTVDAFRYGCVQG-CSAYFLTHFHADHYIGL------TKAWSHGPIYCSS 198
Query: 420 CFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHT 479
+RL+ +++ + +H L L+ + I GI VT ++ANHCPG+ +I F+ +G LHT
Sbjct: 199 LTSRLLRLSLSVNPSSIHPLELDVEYTINGIKVTLIEANHCPGAALIHFRLLDGTCYLHT 258
Query: 480 GDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVI----DAIQAEAFN 535
GDFR S ++ +P+L VH L LDTTYCNP+Y FP +E V+ +V+ D ++ +
Sbjct: 259 GDFRASKQMQTHPLLFNQRVHVLYLDTTYCNPRYKFPSKEDVLSYVVRITKDFLRKQ--- 315
Query: 536 PGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESN 595
P TL ++GSY+IGKE ++L +A++L K++ NA++ R+L+ +++ T + +
Sbjct: 316 PKTLIVVGSYSIGKECVYLAIAKALGVKIFANASRRRILQSFGWDDISKNLST-DGKATC 374
Query: 596 IHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGK--GKKKSPGRRWQNGTIIR 653
+HV PM +L ++RL Y ++ ++AF PTGWT+ + G+ + G I
Sbjct: 375 LHVLPMSSL-KVERLDEHLKIYREQYGAVLAFRPTGWTYSEKIGEHLDLIKPTSRGKITI 433
Query: 654 YEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAMIS 696
Y VPYSEH SFTEL++FV+ + P IIP+VNN + M S
Sbjct: 434 YGVPYSEHSSFTELREFVQFLRPDKIIPTVNNGNAGTREKMQS 476
>UniRef100_UPI00004364BC UPI00004364BC UniRef100 entry
Length = 355
Score = 223 bits (567), Expect = 2e-56
Identities = 135/357 (37%), Positives = 196/357 (54%), Gaps = 16/357 (4%)
Query: 344 GRNHKGKNSSTNRKLRDVPKWC----SIEGTPFRVDAFKY-LRGDCSHWFLTHFHLDRTR 398
GR + +T+ ++ PK C I GT F VDAF+Y + + +FLTHFH D
Sbjct: 1 GRKRWNRGKATDGDPKE-PKRCPFYKKIPGTGFAVDAFQYGVVEGVTAYFLTHFHSDHYG 59
Query: 399 LLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDAN 458
L D Y K + LV + + +H+LP+N + + G+ VT LDAN
Sbjct: 60 GLK-----KDSAVPIYCNKVT--SNLVKSKLKVDEQYIHVLPMNTECIVQGVKVTLLDAN 112
Query: 459 HCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQ 518
HCPG++++LF P+G+ VLHTGDFR + P L+ + TL LDTTYC+P+Y FP Q
Sbjct: 113 HCPGAVMLLFVLPDGQTVLHTGDFRADPSMERYPELQGLRIQTLYLDTTYCSPEYTFPTQ 172
Query: 519 EAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCL 577
+ V+ F ++ A + NP TL + G+Y++GKE++FL V+ L KV ++ K + CL
Sbjct: 173 QEVVTFAVNTAFERVTLNPRTLVVCGTYSVGKEKVFLAVSEVLSSKVCLSKDKYNTMCCL 232
Query: 578 EFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKG 637
E E+ Q T N + +HV PM + + K L+ +++ ++ +VAF PTGWTF +
Sbjct: 233 E-SEDIGQRITTNWQSAQVHVLPMMQI-NFKNLQTHLKKFSKKYDQLVAFKPTGWTFNQT 290
Query: 638 KKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
G I Y +PYSEH SF ELK FV+ + P IIP+VN S AM
Sbjct: 291 VGVDDILPQTQGNISIYGIPYSEHSSFLELKRFVQWLRPKKIIPTVNVGSWRSRKAM 347
>UniRef100_UPI00003AE3F5 UPI00003AE3F5 UniRef100 entry
Length = 967
Score = 219 bits (557), Expect = 3e-55
Identities = 151/441 (34%), Positives = 225/441 (50%), Gaps = 55/441 (12%)
Query: 286 NQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFA--TNGKKV--STAPE----EQQEMK 337
+ N K P + +++D K+ E G A + GK++ S AP QQ+ K
Sbjct: 520 SMNAKSSPAKELKQMDIGVFFGLKPKVKEESKGEACLSEGKQIPSSVAPSGKRPRQQKRK 579
Query: 338 SSGSV--------------------SGRNHKGKN----SSTN---RKLRDVPKWCSIEGT 370
+ GSV SG K + SST + + P + I GT
Sbjct: 580 AEGSVEDLEAVEESSNKDGGVANVTSGGQRKWRKRFRESSTTDEGARKKQCPFYKKIPGT 639
Query: 371 PFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFEC---FARLV 425
F VDAF+Y + G C+ +FLTHFH D L N ++ C LV
Sbjct: 640 GFTVDAFQYGEIEG-CTAYFLTHFHSDHYCGLTKN----------FVFPLYCNKITGNLV 688
Query: 426 NMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFS 485
+ + +++LP++ + + GI V LDANHCPG+ +ILF P+G A+LHTGDFR
Sbjct: 689 KSKLRVKEQYINVLPMDTECIVNGIKVLLLDANHCPGATMILFYLPSGTAILHTGDFRAD 748
Query: 486 DELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGS 544
+ P L +HTL LDTTYC+P+Y FP Q+ VIQF ++ A + NP TL + G+
Sbjct: 749 PSMERYPALIGQKIHTLYLDTTYCSPEYTFPSQQEVIQFAVNTAFEMVTLNPRTLVVCGT 808
Query: 545 YTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTL 604
Y+IGKE++FL +A L K ++ K + L+CLE + T+N + +H+ PM +
Sbjct: 809 YSIGKEKVFLAIAEVLGSKASMSRDKYKTLQCLESAAVN-SLITMNWDGTLLHILPMMQI 867
Query: 605 ASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCS 663
+ K L+ ++++ F ++AF PTGWT+ + Q G I Y +PYSEH S
Sbjct: 868 -NFKGLQEHLNKFSENFDQVLAFKPTGWTYSDSCLSVMDIKPQTRGNITIYGIPYSEHSS 926
Query: 664 FTELKDFVKHISPANIIPSVN 684
+ E+K FV+ + P IIP+VN
Sbjct: 927 YLEMKRFVQWLKPQKIIPTVN 947
>UniRef100_Q5QJC4 SNM1A [Gallus gallus]
Length = 972
Score = 218 bits (556), Expect = 4e-55
Identities = 151/441 (34%), Positives = 225/441 (50%), Gaps = 55/441 (12%)
Query: 286 NQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFA--TNGKKV--STAPE----EQQEMK 337
+ N K P + +++D K+ E G A + GK++ S AP QQ+ K
Sbjct: 520 SMNAKSSPAKELKQMDIGVFFGLKPKVKEESKGEACLSEGKQIPSSVAPSGKRPRQQKRK 579
Query: 338 SSGSV--------------------SGRNHKGKN----SSTN---RKLRDVPKWCSIEGT 370
+ GSV SG K + SST + + P + I GT
Sbjct: 580 AEGSVEDLEAVEESSNKDGGDANVTSGGQRKWRKRFRESSTTDEGARKKQCPFYKKIPGT 639
Query: 371 PFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFEC---FARLV 425
F VDAF+Y + G C+ +FLTHFH D L N ++ C LV
Sbjct: 640 GFTVDAFQYGEIEG-CTAYFLTHFHSDHYCGLTKN----------FVFPLYCNKITGNLV 688
Query: 426 NMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFS 485
+ + +++LP++ + + GI V LDANHCPG+ +ILF P+G A+LHTGDFR
Sbjct: 689 KSKLRVKEQYINVLPMDTECIVNGIKVLLLDANHCPGATMILFYLPSGTAILHTGDFRAD 748
Query: 486 DELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGS 544
+ P L +HTL LDTTYC+P+Y FP Q+ VIQF ++ A + NP TL + G+
Sbjct: 749 PSMERYPALIGQKIHTLYLDTTYCSPEYTFPSQQEVIQFAVNTAFEMVTLNPRTLVVCGT 808
Query: 545 YTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTL 604
Y+IGKE++FL +A L K ++ K + L+CLE + T+N + +H+ PM +
Sbjct: 809 YSIGKEKVFLAIAEVLGSKASMSRDKYKTLQCLESAAVN-SLITMNWDGTLLHILPMMQI 867
Query: 605 ASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQ-NGTIIRYEVPYSEHCS 663
+ K L+ ++++ F ++AF PTGWT+ + Q G I Y +PYSEH S
Sbjct: 868 -NFKGLQDHLNKFSENFDQVLAFKPTGWTYSDSCLSVMDIKPQTRGNITIYGIPYSEHSS 926
Query: 664 FTELKDFVKHISPANIIPSVN 684
+ E+K FV+ + P IIP+VN
Sbjct: 927 YLEMKRFVQWLKPQKIIPTVN 947
>UniRef100_UPI00001CEC53 UPI00001CEC53 UniRef100 entry
Length = 1026
Score = 214 bits (544), Expect = 9e-54
Identities = 135/349 (38%), Positives = 187/349 (52%), Gaps = 15/349 (4%)
Query: 350 KNSSTNRKLRDVPKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDRTRLLDLNIFDT 407
++S R P + I GT F VDAF+Y + G C+ +FLTHFH D L
Sbjct: 676 ESSGGGESRRTCPFYKRIPGTGFTVDAFQYGEIEG-CTAYFLTHFHSDHYAGLS-----K 729
Query: 408 DLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIIL 467
D + Y E L+ + + +H LP++ + + G+ V LDANHCPG+ +IL
Sbjct: 730 DFTRPIYCS--EITGSLLKKKLRVQEQYIHQLPMDTECIVDGVKVVLLDANHCPGATMIL 787
Query: 468 FQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVID 527
FQ PNG LHTGDFR +D +L VHTL LDTTYC+P+Y FP Q+ IQF I+
Sbjct: 788 FQLPNGAVTLHTGDFR-ADPSMERSLLASRKVHTLFLDTTYCSPEYTFPSQQEAIQFAIN 846
Query: 528 -AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQW 586
A +A NP L + G+Y IGKE++FL +A L KV ++ K + L+CL E
Sbjct: 847 TAFEAVTLNPRALIVCGTYCIGKEKVFLAIADVLGSKVGMSQEKYKTLQCLNIPEVS-SL 905
Query: 587 FTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRW 646
T + S +H+ PM + + K L++ + +F I+AF PTGWT
Sbjct: 906 ITTDMCNSLVHLLPMMQI-NFKGLQNHLKKCGGKFDQILAFRPTGWTHSNNITSIADITP 964
Query: 647 Q-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
Q G I Y +PYSEH S+ E+K FV+ + P IIP+VN +S N M
Sbjct: 965 QTKGNIAIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGTFQSRNTM 1013
>UniRef100_Q7XV88 OSJNBb0012E08.8 protein [Oryza sativa]
Length = 481
Score = 212 bits (539), Expect = 3e-53
Identities = 131/363 (36%), Positives = 204/363 (56%), Gaps = 15/363 (4%)
Query: 339 SGSVSGRNHKGKN---SSTNRKLRDVPKWCSIEGTPFRVDAFKYLRGD-CSHWFLTHFHL 394
SGS GR + ++ +RK P + I GTPF VDAF+Y + C+ +FL+HFH
Sbjct: 116 SGSWPGRKRRRGGEVEAAADRKPLACPFYKKIPGTPFTVDAFRYGAVEGCNAYFLSHFHH 175
Query: 395 DRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTC 454
D L T G + ARLV M + + + + L L+++ I G+ VT
Sbjct: 176 DHYGGL------TKKWCHGPIYCTALTARLVKMCLSVNPEYICPLELDKEYVIEGVSVTL 229
Query: 455 LDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYD 514
L+ANHCPG+ +I F+ +GK LHTGDFR S + + P+L+ ++ L LDTTYCNP+Y
Sbjct: 230 LEANHCPGAALIHFRLGDGKKYLHTGDFRASKSMQLYPLLQRGQINLLYLDTTYCNPKYK 289
Query: 515 FPKQEAVIQFVI-DAIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRL 573
FP +E VI F + A + P TL ++G+Y+IGKE ++L ++++L+ +Y +A++ R+
Sbjct: 290 FPPKEDVIDFAVRTAKRYLQKEPKTLIVVGAYSIGKENVYLAISKALQVPIYTDASRRRI 349
Query: 574 LKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWT 633
L + + + + S++HV P+ +L K++ + RF ++AF PTGWT
Sbjct: 350 LHAFGWSDLS-KMICSDSQSSSLHVLPLSSLRHENLQKYLET-LKQRFLAVLAFRPTGWT 407
Query: 634 FGK--GKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESA 691
F + G + + G I Y VPYSEH SF+EL++FV + P +IP+VN S
Sbjct: 408 FSEETGNQLDLIKPSSRGKITIYGVPYSEHSSFSELREFVMFLRPQKVIPTVNVGNAASR 467
Query: 692 NAM 694
+ M
Sbjct: 468 DKM 470
>UniRef100_Q9JIC3 SNM1 protein [Mus musculus]
Length = 1023
Score = 208 bits (529), Expect = 5e-52
Identities = 135/360 (37%), Positives = 190/360 (52%), Gaps = 20/360 (5%)
Query: 344 GRNHKG-----KNSSTNRKLRDVPKWCSIEGTPFRVDAFKY--LRGDCSHWFLTHFHLDR 396
GR +G ++S R P + I GT F VDAF+Y + G C+ +FLTHFH D
Sbjct: 662 GRTQRGNMNISESSGAGEVRRTCPFYKRIPGTGFTVDAFQYGEIEG-CTAYFLTHFHSDH 720
Query: 397 TRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLD 456
L D + Y E L+ + + + LP++ + + + V +D
Sbjct: 721 YAGLS-----KDFTRPVYCS--EITGNLLKKKLRVQEQYIRQLPMDTECVVDSVKVVFVD 773
Query: 457 ANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFP 516
ANHCPG+ +ILFQ PNG +LHTGDFR +D L VHTL LDTTYC+P+Y FP
Sbjct: 774 ANHCPGATMILFQLPNGAVILHTGDFR-ADPSMERSRLAGRKVHTLFLDTTYCSPEYTFP 832
Query: 517 KQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLK 575
Q+ VIQF I+ A +A NP L + G+Y IGKE++FL +A L KV ++ K + L+
Sbjct: 833 SQQEVIQFAINTAFEAVTLNPRALVVCGTYCIGKEKVFLAIADVLGSKVGMSQEKYKTLQ 892
Query: 576 CLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFG 635
CL E T + +S +H+ PM + + K L+ + ++ I+AF PTGWT
Sbjct: 893 CLNIPEVS-SLITTDMCDSLVHLLPMMQI-NFKGLQSHLKKCGGKYDQILAFRPTGWTHS 950
Query: 636 KGKKKSPGRRWQ-NGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDGPESANAM 694
+ Q G I Y +PYSEH S+ E+K FV+ + P IIP+VN S N M
Sbjct: 951 NNITSTADIIPQTRGNISIYGIPYSEHSSYLEMKRFVQWLKPQKIIPTVNVGSFRSRNTM 1010
>UniRef100_UPI000036E972 UPI000036E972 UniRef100 entry
Length = 1028
Score = 206 bits (523), Expect = 2e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 659 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 718
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 719 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 771
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 772 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 830
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 831 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 890
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 891 ADVLGSKVGMSREKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 948
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 949 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1008
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1009 PQKIIPTVN 1017
>UniRef100_Q6PKL4 DNA cross-link repair 1A [Homo sapiens]
Length = 1040
Score = 206 bits (523), Expect = 2e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 659 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 718
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 719 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 771
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 772 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 830
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 831 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 890
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 891 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 948
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 949 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1008
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1009 PQKIIPTVN 1017
>UniRef100_Q6P5Y3 DNA-crosslink repair gene SNM1 [Homo sapiens]
Length = 1040
Score = 206 bits (523), Expect = 2e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 659 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 718
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 719 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 771
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 772 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 830
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 831 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 890
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 891 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 948
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 949 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1008
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1009 PQKIIPTVN 1017
>UniRef100_Q14701 KIAA0086 protein [Homo sapiens]
Length = 1044
Score = 206 bits (523), Expect = 2e-51
Identities = 132/369 (35%), Positives = 197/369 (52%), Gaps = 15/369 (4%)
Query: 321 TNGKKVSTAPEEQQEMKSSGSVSGRNHKGKNSST--NRKLRDVPKWCSIEGTPFRVDAFK 378
T + V+ + + + G + N K SS + + P + I GT F VDAF+
Sbjct: 663 TESEAVNLSKVKVFTKSAHGGLQRGNKKIPESSNVGGSRKKTCPFYKKIPGTGFTVDAFQ 722
Query: 379 Y-LRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLH 437
Y + C+ +FLTHFH D L + + E L+ + + +H
Sbjct: 723 YGVVEGCTAYFLTHFHSDHYAGLSKHFTFP-------VYCSEITGNLLKNKLHVQEQYIH 775
Query: 438 ILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVLRMC 497
LPL+ + + G+ V LDANHCPG+++ILF PNG +LHTGDFR +D +L
Sbjct: 776 PLPLDTECIVNGVKVVLLDANHCPGAVMILFYLPNGTVILHTGDFR-ADPSMERSLLADQ 834
Query: 498 PVHTLILDTTYCNPQYDFPKQEAVIQFVID-AIQAEAFNPGTLFLIGSYTIGKERLFLEV 556
VH L LDTTYC+P+Y FP Q+ VI+F I+ A +A NP L + G+Y+IGKE++FL +
Sbjct: 835 KVHMLYLDTTYCSPEYTFPSQQEVIRFAINTAFEAVTLNPHALVVCGTYSIGKEKVFLAI 894
Query: 557 ARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQ 616
A L KV ++ K + L+CL E + T + S +H+ PM + + K L+ +
Sbjct: 895 ADVLGSKVGMSQEKYKTLQCLNIPEIN-SLITTDMCSSLVHLLPMMQI-NFKGLQSHLKK 952
Query: 617 YASRFSLIVAFSPTGWTF-GKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHIS 675
+++ I+AF PTGWT K + + G I Y +PYSEH S+ E+K FV+ +
Sbjct: 953 CGGKYNQILAFRPTGWTHSNKFTRIADVIPQTKGNISIYGIPYSEHSSYLEMKRFVQWLK 1012
Query: 676 PANIIPSVN 684
P IIP+VN
Sbjct: 1013 PQKIIPTVN 1021
>UniRef100_Q86KS1 Similar to Arabidopsis thaliana (Mouse-ear cress). At2g45700
protein [Dictyostelium discoideum]
Length = 920
Score = 196 bits (499), Expect = 1e-48
Identities = 128/365 (35%), Positives = 187/365 (51%), Gaps = 25/365 (6%)
Query: 335 EMKSSGSVSGRNHKGKNSSTNRKLRDVP-KWCSIEGTPFRVDAFKYLRGDCSHWFLTHFH 393
++K +G + + T K + +P + I+GT F VD F+Y D +H+FLTHFH
Sbjct: 233 DLKKTGKEEAKKRGYQVRKTPTKEKKIPPSFKVIDGTNFLVDGFQYKSEDFTHYFLTHFH 292
Query: 394 LDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVT 453
D + T G + E +LV+ +G+ + N+ +EI G+ V
Sbjct: 293 SDHY------VGITKTWSFGNIYCTEETGKLVSHKLGVDQRYIVKCEWNKLIEIQGVKVA 346
Query: 454 CLDANHCPGSIIILFQPP---------NGKAVLHTGDFRFSDELAINPVLRMCPVHTLIL 504
LD+NHCPGS +ILF P +++LHTGDFR++ + P+L+ + L L
Sbjct: 347 FLDSNHCPGSALILFIIPLRNKDGEIIGEESILHTGDFRYNQSMNNYPLLKGRTISKLYL 406
Query: 505 DTTYCNPQYDFPKQEAVIQFVIDAIQAEAFNPG-TLFLIGSYTIGKERLFLEVARSLRRK 563
D TYC+PQY FP Q +I+ V ++ E N G TLFL G+Y IGKER+ LE+A+ +
Sbjct: 407 DNTYCDPQYVFPPQPEIIKQVASIVRKE--NDGETLFLFGTYVIGKERILLEIAKQEGKP 464
Query: 564 VYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASRFSL 623
V+V+ K +L CL D+ FT NE + M L+ L + S +++
Sbjct: 465 VHVSNEKYAILCCLS-TCLDINKFTTNELITPFRAVTMSMLSYHNMLSLLDSS-NNKYKR 522
Query: 624 IVAFSPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSV 683
++ F PTGWT K R G Y V YSEH SF EL+D + H P IIP+V
Sbjct: 523 VIGFRPTGWTQAKKSITYLNR----GPTTFYSVAYSEHSSFNELRDCIDHFRPTQIIPTV 578
Query: 684 NNDGP 688
+ D P
Sbjct: 579 DCDSP 583
>UniRef100_Q9C9M5 DNA ligase I, putative [Arabidopsis thaliana]
Length = 1417
Score = 181 bits (460), Expect = 5e-44
Identities = 116/338 (34%), Positives = 175/338 (51%), Gaps = 23/338 (6%)
Query: 361 VPKWCSIEGTPFRVDAFKYLRGDCS-HWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFE 419
+P I T F VD F+ S +FL+HFH D L + G +
Sbjct: 54 IPNSKRIPNTNFIVDLFRLPHQSSSVAFFLSHFHSDHYSGLSSSW------SKGIIYCSH 107
Query: 420 CFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILF----QPPNGKA 475
ARLV + +P + LP+NQ V+I G +V ++ANHCPG++ LF + +
Sbjct: 108 KTARLVAEILQVPSQFVFALPMNQMVKIDGSEVVLIEANHCPGAVQFLFKVKLESSGFEK 167
Query: 476 VLHTGDFRFSDELAINPVLR-MCPVHTLILDTTYCNPQYDFPKQEAVIQFVIDAIQAEAF 534
+HTGDFRF DE+ +P L + LDTTYCNP++ FP QE + +V+ I +
Sbjct: 168 YVHTGDFRFCDEMRFDPFLNGFVGCDGVFLDTTYCNPKFVFPSQEESVGYVVSVID-KIS 226
Query: 535 NPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHES 594
LFL+ +Y +GKE++ +E+AR +RK+ V+A K+ +L L EE M FT +E+ES
Sbjct: 227 EEKVLFLVATYVVGKEKILVEIARRCKRKIVVDARKMSMLSVLGCGEEGM--FTEDENES 284
Query: 595 NIHV------APMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQN 648
++HV W +K + +V F PTGWT+ + K R +
Sbjct: 285 DVHVVGWNVLGETWPYFRPNFVKMNEIMVEKGYDKVVGFVPTGWTYEVKRNKFAVRFKDS 344
Query: 649 GTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNND 686
I + VPYSEH ++ EL++F+K + P +IP+V D
Sbjct: 345 MEI--HLVPYSEHSNYDELREFIKFLKPKRVIPTVGVD 380
>UniRef100_Q8LQE6 Putative DNA ligase [Oryza sativa]
Length = 1419
Score = 176 bits (445), Expect = 3e-42
Identities = 114/341 (33%), Positives = 177/341 (51%), Gaps = 23/341 (6%)
Query: 361 VPKWCSIEGTPFRVDAFKYLRGDCSHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFEC 420
VP I G+ F VDAF++ + +FL+HFH D L + + G +
Sbjct: 51 VPPTALIPGSRFLVDAFRHAGDFTASYFLSHFHSDHYTGLGPSW------RRGLVFCSPL 104
Query: 421 FARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQP--PNGKAVLH 478
ARL+ + +P + +L +V + G V +DANHCPG++ LF+ PN + +H
Sbjct: 105 TARLLVSVLSVPPQLVVVLDAGVRVTVDGWCVVAVDANHCPGAVQFLFRSSGPNAERYVH 164
Query: 479 TGDFRFSDELAINP-VLRMCPVHTLILDTTYCNPQYDFPKQEAVIQFVIDAI-----QAE 532
TGDFRFS + P +L + LDTTYCNP++ FP Q+ +++V+++I ++
Sbjct: 165 TGDFRFSQSMITEPNLLEFIGADAVFLDTTYCNPKFTFPPQKESLEYVVNSIKRVKEESR 224
Query: 533 AFNPGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEH 592
A L LI +Y +GKER+ LEVAR K++V++ K+ +L L ED FT +
Sbjct: 225 ASGERVLCLIATYVVGKERILLEVARRCGCKIHVDSRKMEILTLLGIGGEDGV-FTEDAA 283
Query: 593 ESNIHVA------PMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRW 646
+++HV W +K ++ V F PTGW + + KK+ R
Sbjct: 284 ATDVHVTGWNILGETWPYFRPNFVKMKEIMVERGYNKAVGFVPTGWMY-ETKKEGFAVRT 342
Query: 647 QNGTIIRYEVPYSEHCSFTELKDFVKHISPANIIPSVNNDG 687
++ I VPYSEH S+ EL+D+VK + P +IP+V DG
Sbjct: 343 KDSLEIHL-VPYSEHSSYNELRDYVKFLHPKRVIPTVGLDG 382
>UniRef100_Q7QDQ3 ENSANGP00000016032 [Anopheles gambiae str. PEST]
Length = 349
Score = 170 bits (431), Expect = 1e-40
Identities = 116/328 (35%), Positives = 165/328 (49%), Gaps = 24/328 (7%)
Query: 362 PKWCSIEGTPFRVDAFKYLRGDC---SHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKF 418
P + I GT F VDAF+Y GD +H+FLTHFH D L LI
Sbjct: 2 PSYKIIAGTNFAVDAFRY--GDIEGVTHYFLTHFHADHYIGLKKTFSKP-------LIMS 52
Query: 419 ECFARLVNMNIGIPYDKLHILPLNQKVEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLH 478
ARLV I + + I+ L+Q + I +++ LDANHCPG I+ LF+ PNG VLH
Sbjct: 53 SITARLVKAFINVAEEFYQIVELHQSIVIDDVEIIALDANHCPGGIMFLFRLPNGSNVLH 112
Query: 479 TGDFRFSDELAINPVLRMCPVHTLILDTTYCNPQYDFPKQ---EAVIQFVIDAIQAEAFN 535
TGDFR S E+ P + + LDTTY + +Y F Q A + + A +
Sbjct: 113 TGDFRASPEMEEYPEFWNFQIDIIYLDTTYLSSKYAFKSQWESVADARETVSAYLKKHIG 172
Query: 536 PGTLFLIGSYTIGKERLFLEVARSLRRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESN 595
L + GSY IGKE+++LE+A S KV+ + + LK + + + + + +++N
Sbjct: 173 VKVLIVCGSYLIGKEKVWLELAISTGMKVWTEPNRWKALKAIA-DSQQLSVLVADPNKAN 231
Query: 596 IHVAPMWTLASLKRLKHVSSQYASRFSLIVAFSPTGWTFGKGKKKSPGRRWQNGTIIRYE 655
IHV M L S L +Q+ R+ ++A P+GW K P R G I
Sbjct: 232 IHVLAMNKL-SYDELNEYMNQFPDRYESVIAIRPSGWE----KNSKPQYR---GRINIVG 283
Query: 656 VPYSEHCSFTELKDFVKHISPANIIPSV 683
+ YSEH SF ELK FV+ + P +I +V
Sbjct: 284 IEYSEHSSFDELKRFVQFLRPHEVISTV 311
>UniRef100_Q9VND2 CG10018-PA [Drosophila melanogaster]
Length = 763
Score = 167 bits (424), Expect = 7e-40
Identities = 118/423 (27%), Positives = 202/423 (46%), Gaps = 35/423 (8%)
Query: 272 EKPEDALAEPRRNRNQNVKMQPHRPERKVDGTSKPAANKKITEYFPGFATNGKKVSTAPE 331
++ A+A P + + PE + ++ + E PG + +P
Sbjct: 177 KRKPSAIASPCEKSSDVQATKFRTPETSMPEAKLSNSSLDLVEISPG-------PNKSPT 229
Query: 332 EQQEMKSSGSVSGRNHKGKNSSTN-----RKLRDVPKWCSIEGTPFRVDAFKY--LRGDC 384
S +++ + GKNS RK + P + +EGT F VD F++ + G
Sbjct: 230 NNFYRSSPAAITKESPLGKNSKKGTVRKQRKPKPCPPYKVVEGTSFCVDGFQFGEIEG-V 288
Query: 385 SHWFLTHFHLDRTRLLDLNIFDTDLRQMGYLIKFECFARLVNMNIGIPYDKLHILPLNQK 444
+H+FLTHFH D L L ARLV I + +H + ++Q
Sbjct: 289 THYFLTHFHADHYIGLTKKFCHP-------LYVSPISARLVRTFIKLDETHIHEIDVDQT 341
Query: 445 VEIAGIDVTCLDANHCPGSIIILFQPPNGKAVLHTGDFRFSDELAINPVL-RMCPVHTLI 503
+++ G+ VT L+ANHCPG+++ F+ +G+ +LHTGDFR S ++ P+ + L
Sbjct: 342 LDVDGVQVTALEANHCPGALMFFFKLSSGECILHTGDFRASADMESLPIFWNHSNIDLLY 401
Query: 504 LDTTYCNPQYDFPKQEAVIQFVIDAIQA---EAFNPGTLFLIGSYTIGKERLFLEVARSL 560
LDTTY N YDF Q + +D ++A + L + GSY IGKE+++L +A+
Sbjct: 402 LDTTYMNKNYDFCHQSESVDRAVDLVRAFLEKNAAKRILIVCGSYVIGKEKIWLALAKEF 461
Query: 561 RRKVYVNAAKLRLLKCLEFEEEDMQWFTLNEHESNIHVAPMWTLASLKRLKHVSSQYASR 620
+V+ + + ++CL + + D T + +N+HV M + S L +++ +
Sbjct: 462 TMRVWTESNRSTAVRCLNWPDLD-SVLTEDRSGANLHVIAMGKI-SYPSLVDYFTEFEDQ 519
Query: 621 FSLIVAFSPTGWTFGKGKKKSPGRRWQNGTIIRYEVPYSEHCSFTELKDFVKHISPANII 680
+ +++ P+GW K K S G+R I + YSEH S+ EL+ FV+ + P +I
Sbjct: 520 YDMLLGIRPSGWE--KNSKPSYGKR-----ISTIGIEYSEHSSYKELERFVRFLKPKRVI 572
Query: 681 PSV 683
+V
Sbjct: 573 STV 575
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,193,709,773
Number of Sequences: 2790947
Number of extensions: 52606931
Number of successful extensions: 145381
Number of sequences better than 10.0: 805
Number of HSP's better than 10.0 without gapping: 378
Number of HSP's successfully gapped in prelim test: 429
Number of HSP's that attempted gapping in prelim test: 143948
Number of HSP's gapped (non-prelim): 1226
length of query: 700
length of database: 848,049,833
effective HSP length: 135
effective length of query: 565
effective length of database: 471,271,988
effective search space: 266268673220
effective search space used: 266268673220
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0133.3


