

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0131.12
(78 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
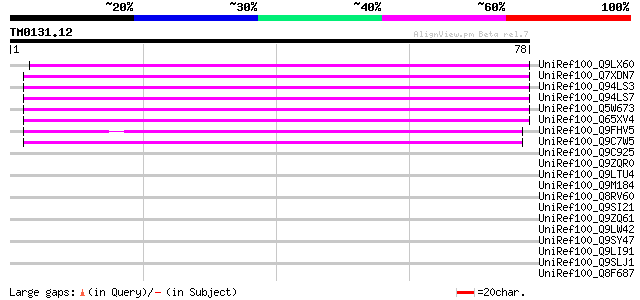
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 54 1e-06
UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa] 52 4e-06
UniRef100_Q94LS3 Putative helicase [Oryza sativa] 52 4e-06
UniRef100_Q94LS7 Putative helicase [Oryza sativa] 51 6e-06
UniRef100_Q5W673 Putative helicase [Oryza sativa] 47 9e-05
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 47 1e-04
UniRef100_Q9FHV5 Helicase [Arabidopsis thaliana] 46 2e-04
UniRef100_Q9C7W5 Hypothetical protein F15H21.18 [Arabidopsis tha... 45 5e-04
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 44 0.001
UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana] 43 0.002
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 43 0.002
UniRef100_Q9M184 Hypothetical protein T5C2_50 [Arabidopsis thali... 41 0.007
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 39 0.025
UniRef100_Q9SI21 Putative helicase [Arabidopsis thaliana] 39 0.033
UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana] 38 0.056
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 35 0.36
UniRef100_Q9SY47 Hypothetical protein T5L23.19 [Arabidopsis thal... 35 0.47
UniRef100_Q9LI91 Similarity to unknown protein [Arabidopsis thal... 34 1.1
UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana] 33 1.4
UniRef100_Q8F687 Serine/threonine protein kinases [Leptospira in... 33 2.4
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 53.5 bits (127), Expect = 1e-06
Identities = 27/75 (36%), Positives = 43/75 (57%)
Query: 4 GNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPIL 63
G+ LR FA LL+++ + P+HVW+ TW LLA DI+++ R P + T + L
Sbjct: 1155 GDYLRNFFAMLLLSDSLARPEHVWSETWHLLAEDIENKKREDFKNPDLKLTLAEIRNYTL 1214
Query: 64 LKIDNHLQSNGKSLK 78
+I+ + NG +LK
Sbjct: 1215 QEIEKIMLRNGATLK 1229
>UniRef100_Q7XDN7 Hypothetical protein [Oryza sativa]
Length = 1477
Score = 52.0 bits (123), Expect = 4e-06
Identities = 28/76 (36%), Positives = 44/76 (57%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SG +LR+LFA++L + +P +W +W+ L+ DIQH L P T S K
Sbjct: 922 SGMKLRQLFATVLCHCEVTDPKRLWESSWEKLSKDIQHTQSWALNFPTSCLTPSHRRKCA 981
Query: 63 LLKIDNHLQSNGKSLK 78
L++I+ +++ GKSLK
Sbjct: 982 LIEIEKNMRQAGKSLK 997
>UniRef100_Q94LS3 Putative helicase [Oryza sativa]
Length = 1501
Score = 52.0 bits (123), Expect = 4e-06
Identities = 28/76 (36%), Positives = 44/76 (57%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SG +LR+LFA++L + +P +W +W+ L+ DIQH L P T S K
Sbjct: 960 SGMKLRQLFATVLCHCEVTDPKRLWESSWEKLSKDIQHTQSWALNFPTSCLTPSHRRKCA 1019
Query: 63 LLKIDNHLQSNGKSLK 78
L++I+ +++ GKSLK
Sbjct: 1020 LIEIEKNMRQAGKSLK 1035
>UniRef100_Q94LS7 Putative helicase [Oryza sativa]
Length = 1573
Score = 51.2 bits (121), Expect = 6e-06
Identities = 28/76 (36%), Positives = 43/76 (55%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SG EL++LFA++L + +P +W W+ L+ DIQH L P T S K
Sbjct: 976 SGIELQQLFATILCHCEVTDPKSLWESIWEELSKDIQHTQSWLLNFPASCLTPSHKRKCA 1035
Query: 63 LLKIDNHLQSNGKSLK 78
L++I+ +++ GKSLK
Sbjct: 1036 LIEIEKNMRQAGKSLK 1051
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 47.4 bits (111), Expect = 9e-05
Identities = 23/76 (30%), Positives = 42/76 (55%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SGN+LR LF ++L + +P +W W+ L+ DI+++ R L P + T
Sbjct: 1038 SGNQLRHLFTTILCHCEVTDPKRIWESCWEDLSEDIEYKQRKNLNYPTLRLTEQQKKGHA 1097
Query: 63 LLKIDNHLQSNGKSLK 78
L++I+ ++ GK+L+
Sbjct: 1098 LIEIEKLMRQAGKTLE 1113
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 47.0 bits (110), Expect = 1e-04
Identities = 23/76 (30%), Positives = 41/76 (53%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SGN+LR LF ++L + +P +W W+ L DI+++ R L P + T
Sbjct: 929 SGNQLRHLFTTILCHCEVTDPKRIWESCWEDLGEDIEYKQRKNLNYPTLRLTEQQKKGHA 988
Query: 63 LLKIDNHLQSNGKSLK 78
L++I+ ++ GK+L+
Sbjct: 989 LIEIEKLMRQAGKTLE 1004
>UniRef100_Q9FHV5 Helicase [Arabidopsis thaliana]
Length = 1523
Score = 46.2 bits (108), Expect = 2e-04
Identities = 27/75 (36%), Positives = 43/75 (57%), Gaps = 2/75 (2%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
SG LR+LF +L + + +P++VW TW+ L+ DIQ+ R +P +I +
Sbjct: 954 SGGYLRQLFVIML--DALISPENVWAATWQHLSEDIQNNKRKYFNRPDLILSDEEKKLYA 1011
Query: 63 LLKIDNHLQSNGKSL 77
L +ID+ L+ NG SL
Sbjct: 1012 LQEIDHILRRNGTSL 1026
>UniRef100_Q9C7W5 Hypothetical protein F15H21.18 [Arabidopsis thaliana]
Length = 753
Score = 45.1 bits (105), Expect = 5e-04
Identities = 24/75 (32%), Positives = 40/75 (53%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
S +RRLF +L++ + P+ VW+ TWK+L+ DI+ + R +P + + +
Sbjct: 405 SAKYVRRLFVIMLLSESLTKPEMVWDETWKILSEDIERKKRKEWKRPDLQLSDEERQQYC 464
Query: 63 LLKIDNHLQSNGKSL 77
L +I L NG SL
Sbjct: 465 LQEIARLLTKNGVSL 479
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 43.5 bits (101), Expect = 0.001
Identities = 23/75 (30%), Positives = 39/75 (51%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPI 62
S +RRLF +L++ + P+ VW+ TW++L+ DI+ R +P + + +
Sbjct: 427 SAKYVRRLFVIMLLSESLTKPEMVWDETWRILSEDIERRKRKEWKRPDLQLSDEERQQYC 486
Query: 63 LLKIDNHLQSNGKSL 77
L +I L NG SL
Sbjct: 487 LQEIARLLTKNGVSL 501
>UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana]
Length = 1265
Score = 43.1 bits (100), Expect = 0.002
Identities = 23/61 (37%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 8 RRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKID 67
+ FA LL+++ ++ P HVW+ TW +LA DI + R P I F KP + ID
Sbjct: 717 QEFFAMLLLSDSLSRPAHVWSQTWHILAEDILKKKRDEFKNPEDIDEF---PKPTIDGID 773
Query: 68 N 68
N
Sbjct: 774 N 774
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/75 (29%), Positives = 40/75 (53%)
Query: 4 GNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPIL 63
G+ LR F+ +L+++ + P+HVW+ TW LL+ DI + R + T + L
Sbjct: 832 GDYLRNFFSMMLLSDSLARPEHVWSETWHLLSEDILIKKRDEFKNQELTLTEAQIQNYTL 891
Query: 64 LKIDNHLQSNGKSLK 78
+I+ + NG +L+
Sbjct: 892 QEIEKIMLFNGATLE 906
>UniRef100_Q9M184 Hypothetical protein T5C2_50 [Arabidopsis thaliana]
Length = 830
Score = 41.2 bits (95), Expect = 0.007
Identities = 15/38 (39%), Positives = 26/38 (67%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQH 40
S +RRLF +L++ + P+ VW+ TW++L+ DI+H
Sbjct: 94 SAKYVRRLFVIMLLSESLTKPEMVWDETWRILSKDIEH 131
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 39.3 bits (90), Expect = 0.025
Identities = 17/35 (48%), Positives = 23/35 (65%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLAND 37
SG +RRLFA LL + ++ P HVW TW L++D
Sbjct: 728 SGRYMRRLFAQLLTSESLSTPYHVWLNTWNQLSDD 762
>UniRef100_Q9SI21 Putative helicase [Arabidopsis thaliana]
Length = 1219
Score = 38.9 bits (89), Expect = 0.033
Identities = 27/81 (33%), Positives = 44/81 (53%), Gaps = 8/81 (9%)
Query: 3 SGNELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQP------GMIPTFS 56
SG LR+LF +L + + +P++VW TW+ L+ DIQ+E + +P +I +
Sbjct: 626 SGWYLRQLFVIML--DALISPENVWAATWQHLSEDIQNEKKKYFNRPVTCLFTDLILSDE 683
Query: 57 LSSKPILLKIDNHLQSNGKSL 77
L +ID+ L+ NG SL
Sbjct: 684 EKKVYALQEIDHILRRNGTSL 704
>UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana]
Length = 1230
Score = 38.1 bits (87), Expect = 0.056
Identities = 16/45 (35%), Positives = 27/45 (59%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGM 51
+R+ F +LI+ +++P VW TWK+L D Q + R L +P +
Sbjct: 734 VRKSFVIMLISESLSSPVVVWEHTWKILFEDFQRKVRDKLERPDL 778
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 35.4 bits (80), Expect = 0.36
Identities = 14/35 (40%), Positives = 23/35 (65%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHE 41
+R+LF +LI +++P VW TWK+L+ D Q +
Sbjct: 1142 VRKLFVIMLIFESLSSPAVVWEHTWKILSEDFQRK 1176
>UniRef100_Q9SY47 Hypothetical protein T5L23.19 [Arabidopsis thaliana]
Length = 570
Score = 35.0 bits (79), Expect = 0.47
Identities = 14/38 (36%), Positives = 24/38 (62%)
Query: 9 RLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTL 46
+LF +LI+ +++P VW TWK+L+ D + + R L
Sbjct: 139 KLFVIMLISERLSSPAVVWEHTWKILSEDFKRKLRNQL 176
>UniRef100_Q9LI91 Similarity to unknown protein [Arabidopsis thaliana]
Length = 619
Score = 33.9 bits (76), Expect = 1.1
Identities = 20/71 (28%), Positives = 37/71 (51%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKI 66
LR LF +LI ++ P +W+ W+ +A+D+ + + L P + K L++I
Sbjct: 95 LRSLFVLILIYCEVSEPLKLWSHCWESMADDVLRKQQRVLNFPQLELKAKELEKYTLIEI 154
Query: 67 DNHLQSNGKSL 77
+ L+ + KSL
Sbjct: 155 ETLLRQHEKSL 165
>UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana]
Length = 1250
Score = 33.5 bits (75), Expect = 1.4
Identities = 20/71 (28%), Positives = 37/71 (51%)
Query: 7 LRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILLKI 66
LR LF +LI ++ P +W+ W+ +A+D+ + + L P + K L++I
Sbjct: 726 LRSLFVLILIYCEVSEPLKLWSHCWESMADDVFRKQQKVLNFPQLELKAEELEKYTLIEI 785
Query: 67 DNHLQSNGKSL 77
+ L+ + KSL
Sbjct: 786 ETLLRQHEKSL 796
>UniRef100_Q8F687 Serine/threonine protein kinases [Leptospira interrogans]
Length = 1780
Score = 32.7 bits (73), Expect = 2.4
Identities = 23/72 (31%), Positives = 30/72 (40%), Gaps = 5/72 (6%)
Query: 5 NELRRLFASLLITNIMNNPDHVWNITWKLLANDIQHEYRGTLTQPGMIPTFSLSSKPILL 64
N R SLL M +P H+W + LL + I Y +P + P L +LL
Sbjct: 899 NLKNRSITSLLDAPTMTDPSHIWAV--NLLISAIPMAYN---KEPALFPVICLKMANLLL 953
Query: 65 KIDNHLQSNGKS 76
K N S G S
Sbjct: 954 KFGNLSDSYGYS 965
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 136,333,168
Number of Sequences: 2790947
Number of extensions: 4482111
Number of successful extensions: 9033
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 9012
Number of HSP's gapped (non-prelim): 31
length of query: 78
length of database: 848,049,833
effective HSP length: 54
effective length of query: 24
effective length of database: 697,338,695
effective search space: 16736128680
effective search space used: 16736128680
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0131.12


