

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0128.2
(265 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
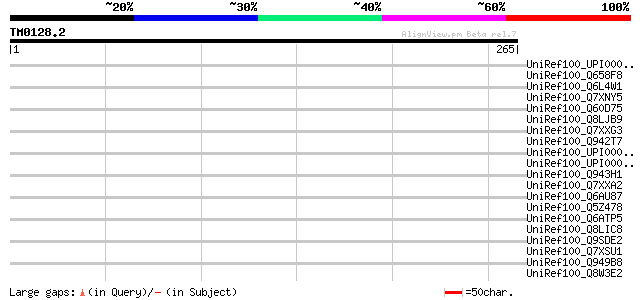
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI0000256C12 UPI0000256C12 UniRef100 entry 45 0.002
UniRef100_Q658F8 Hypothetical protein P0015E04.9 [Oryza sativa] 44 0.006
UniRef100_Q6L4W1 Hypothetical protein OSJNBb0108E17.19 [Oryza sa... 40 0.062
UniRef100_Q7XNY5 OSJNBb0015N08.6 protein [Oryza sativa] 40 0.080
UniRef100_Q60D75 Putative polyprotein [Oryza sativa] 40 0.080
UniRef100_Q8LJB9 P0505D12.16 protein [Oryza sativa] 40 0.080
UniRef100_Q7XXG3 OSJNBb0089K24.7 protein [Oryza sativa] 39 0.14
UniRef100_Q942T7 P0560B06.7 protein [Oryza sativa] 39 0.14
UniRef100_UPI00002D4CC4 UPI00002D4CC4 UniRef100 entry 39 0.18
UniRef100_UPI00002CB637 UPI00002CB637 UniRef100 entry 39 0.18
UniRef100_Q943H1 P0046E05.7 protein [Oryza sativa] 39 0.18
UniRef100_Q7XXA2 OSJNBa0019G23.14 protein [Oryza sativa] 39 0.18
UniRef100_Q6AU87 Hypothetical protein P0668F02.14 [Oryza sativa] 39 0.18
UniRef100_Q5Z478 Hypothetical protein B1130E07.22 [Oryza sativa] 39 0.18
UniRef100_Q6ATP5 Hypothetical protein OSJNBa0028F23.24 [Oryza sa... 38 0.23
UniRef100_Q8LIC8 Hypothetical protein OJ1343_D04.120 [Oryza sativa] 38 0.23
UniRef100_Q9SDE2 Hypothetical protein [Oryza sativa] 38 0.23
UniRef100_Q7XSU1 OSJNBa0039K24.15 protein [Oryza sativa] 38 0.23
UniRef100_Q949B8 Hypothetical protein C1015ERIPDK [Oryza sativa] 38 0.31
UniRef100_Q8W3E2 Putative membrane-associated protein [Oryza sat... 38 0.31
>UniRef100_UPI0000256C12 UPI0000256C12 UniRef100 entry
Length = 332
Score = 44.7 bits (104), Expect = 0.002
Identities = 50/160 (31%), Positives = 75/160 (46%), Gaps = 38/160 (23%)
Query: 56 SYYSLSLNSDSDDGDKTTVSQEV-ITVDSDSESHNSPSSSDPT---------QKRTVSIT 105
SY+++SL+S DDGD +T + V IT+ S S + SSSD + RTV++T
Sbjct: 118 SYFTVSLSSAPDDGDNSTTNDNVTITLSSSDTSEGTISSSDSSLTFTADNWNSVRTVTVT 177
Query: 106 ---SHLSTGNPS----TSEDWRTSDKCIDDKCLFRSVWNSVQSLKNNRLTARGFTRVNAE 158
+LS G+ S + D +TSD F +V S++N T +G V+
Sbjct: 178 GVADNLSDGDQSYTIVLTGDNQTSD------LRFANVDPPDVSVRNLDYTTKGGFYVSQI 231
Query: 159 TGHNDD--------LSLNIAPCD-------DNEVVTLKPS 183
+G D+ +SL+ AP D DN +TL S
Sbjct: 232 SGDTDENRSTSYFTVSLSSAPDDGDNSTTNDNVTITLSSS 271
Score = 38.1 bits (87), Expect = 0.23
Identities = 27/72 (37%), Positives = 40/72 (55%), Gaps = 13/72 (18%)
Query: 56 SYYSLSLNSDSDDGDKTTVSQEV-ITVDSDSESHNSPSSSDPT---------QKRTVSIT 105
SY+++SL+S DDGD +T + V IT+ S S + SSSD + RTV++T
Sbjct: 242 SYFTVSLSSAPDDGDNSTTNDNVTITLSSSDTSEGTISSSDSSLTFTADNWNSVRTVTVT 301
Query: 106 ---SHLSTGNPS 114
+LS G+ S
Sbjct: 302 GVADNLSDGDQS 313
Score = 35.4 bits (80), Expect = 1.5
Identities = 46/153 (30%), Positives = 68/153 (44%), Gaps = 38/153 (24%)
Query: 63 NSDSDDGDKTTVSQEV-ITVDSDSESHNSPSSSDPT---------QKRTVSIT---SHLS 109
+S DDGD +T + V IT+ S S + SSSD + RTV++T +LS
Sbjct: 1 SSAPDDGDNSTTNDNVTITLSSSDTSEGTISSSDSSLTFTADNWNSVRTVTVTGVADNLS 60
Query: 110 TGNPS----TSEDWRTSDKCIDDKCLFRSVWNSVQSLKNNRLTARGFTRVNAETGHNDD- 164
G+ S + D +TSD F +V S++N T +G V+ +G D+
Sbjct: 61 DGDQSYTIVLTGDNQTSD------LRFANVDPPDVSVRNLDYTTKGGFYVSQISGDTDEN 114
Query: 165 -------LSLNIAPCD-------DNEVVTLKPS 183
+SL+ AP D DN +TL S
Sbjct: 115 RSTSYFTVSLSSAPDDGDNSTTNDNVTITLSSS 147
>UniRef100_Q658F8 Hypothetical protein P0015E04.9 [Oryza sativa]
Length = 569
Score = 43.5 bits (101), Expect = 0.006
Identities = 21/51 (41%), Positives = 23/51 (44%)
Query: 201 ELFMKLPFSTFVCDVLTFLNVAPCQLLPNAWGFTRCFELLCEPLNFGPPYP 251
E M+LP F VL +AP QL PNAW F LC PP P
Sbjct: 95 ERLMRLPLQPFFAAVLAHFGLAPSQLTPNAWRLLAGFAALCRRAGVAPPPP 145
>UniRef100_Q6L4W1 Hypothetical protein OSJNBb0108E17.19 [Oryza sativa]
Length = 928
Score = 40.0 bits (92), Expect = 0.062
Identities = 34/101 (33%), Positives = 46/101 (44%), Gaps = 10/101 (9%)
Query: 171 PCDDNEVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQL 226
P E+VTL +G PA Y +F F PFS+F DVL F ++ L
Sbjct: 34 PATGREIVTL----GEGRPAPCYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHL 89
Query: 227 LPNAWGFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
PNA F LCE + P LF +F+ + S++ PS
Sbjct: 90 TPNAVMTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q7XNY5 OSJNBb0015N08.6 protein [Oryza sativa]
Length = 949
Score = 39.7 bits (91), Expect = 0.080
Identities = 34/96 (35%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F N+ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYNLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q60D75 Putative polyprotein [Oryza sativa]
Length = 1093
Score = 39.7 bits (91), Expect = 0.080
Identities = 34/96 (35%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F N+ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYNLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q8LJB9 P0505D12.16 protein [Oryza sativa]
Length = 953
Score = 39.7 bits (91), Expect = 0.080
Identities = 33/96 (34%), Positives = 46/96 (47%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y ++F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRFVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q7XXG3 OSJNBb0089K24.7 protein [Oryza sativa]
Length = 1052
Score = 38.9 bits (89), Expect = 0.14
Identities = 34/96 (35%), Positives = 44/96 (45%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL G PA Y +F F PFS+F DVL F N+ L PNA
Sbjct: 39 EIVTL----GKGRPAPCYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYNLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q942T7 P0560B06.7 protein [Oryza sativa]
Length = 917
Score = 38.9 bits (89), Expect = 0.14
Identities = 32/94 (34%), Positives = 42/94 (44%), Gaps = 9/94 (9%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELF----MKLPFSTFVCDVLTFLNVAPCQLLPNAW 231
EVV L +G PA Y +F F + LPFS+F DVL F ++ L PNA
Sbjct: 39 EVVVL----GEGRPAPHYPGRSVFFLPFAMAGLVLPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKISIAKP 264
F LCE + P LF +F+ + P
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQPVSP 128
>UniRef100_UPI00002D4CC4 UPI00002D4CC4 UniRef100 entry
Length = 329
Score = 38.5 bits (88), Expect = 0.18
Identities = 45/163 (27%), Positives = 71/163 (42%), Gaps = 39/163 (23%)
Query: 58 YSLSLNSDSDDGDKTTVSQEV-ITVDSDSESHNSP------------------SSSDPTQ 98
+++ L+S DDGD +T + V IT+ S ES ++S+
Sbjct: 25 FTIRLSSAPDDGDSSTTTDNVTITLKSSDESEGKVIDISGTDVTNTDNKTLVFTASNWNS 84
Query: 99 KRTVSIT---SHLSTGNPSTSEDWRTSDKCIDDKCLFRSVWNSVQSLKNNRLTARGFTRV 155
+RTV + +LS G+ S + + +D + FR V +LKN LT +G V
Sbjct: 85 ERTVKVEGQYDNLSDGDQSYAIILSADNNSLDRR--FRFVDPPDVALKNLDLTDKGTFYV 142
Query: 156 NAETGHNDD--------LSLNIAPCD-------DNEVVTLKPS 183
+A +G D+ + L+ AP D DN +TLK S
Sbjct: 143 SAASGDTDENGVSTTFTIRLSSAPDDGDSSTTTDNVTITLKSS 185
>UniRef100_UPI00002CB637 UPI00002CB637 UniRef100 entry
Length = 200
Score = 38.5 bits (88), Expect = 0.18
Identities = 45/163 (27%), Positives = 71/163 (42%), Gaps = 39/163 (23%)
Query: 58 YSLSLNSDSDDGDKTTVSQEV-ITVDSDSESHNSP------------------SSSDPTQ 98
+++ L+S DDGD +T + V IT+ S ES ++S+
Sbjct: 29 FTIRLSSAPDDGDSSTTTDNVTITLKSSDESEGKVIDISGTDVTNTDNKTLVFTASNWNS 88
Query: 99 KRTVSIT---SHLSTGNPSTSEDWRTSDKCIDDKCLFRSVWNSVQSLKNNRLTARGFTRV 155
+RTV + +LS G+ S + + +D + FR V +LKN LT +G V
Sbjct: 89 ERTVKVEGQYDNLSDGDQSYAIILSADNNSLDRR--FRFVDPPDVALKNLDLTDKGTFYV 146
Query: 156 NAETGHNDD--------LSLNIAPCD-------DNEVVTLKPS 183
+A +G D+ + L+ AP D DN +TLK S
Sbjct: 147 SAASGDTDENGVSTTFTIRLSSAPDDGDSSTTTDNVTITLKSS 189
>UniRef100_Q943H1 P0046E05.7 protein [Oryza sativa]
Length = 1054
Score = 38.5 bits (88), Expect = 0.18
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPERSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q7XXA2 OSJNBa0019G23.14 protein [Oryza sativa]
Length = 1041
Score = 38.5 bits (88), Expect = 0.18
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPHYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q6AU87 Hypothetical protein P0668F02.14 [Oryza sativa]
Length = 867
Score = 38.5 bits (88), Expect = 0.18
Identities = 34/96 (35%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGHPAPCYPGRSVFFLPFAMAGLVPPFSSFFMDVLKFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ I S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTIQSVSPPS 130
>UniRef100_Q5Z478 Hypothetical protein B1130E07.22 [Oryza sativa]
Length = 911
Score = 38.5 bits (88), Expect = 0.18
Identities = 33/96 (34%), Positives = 46/96 (47%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F + PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRSVFFLPFAMVGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q6ATP5 Hypothetical protein OSJNBa0028F23.24 [Oryza sativa]
Length = 1096
Score = 38.1 bits (87), Expect = 0.23
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPCYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q8LIC8 Hypothetical protein OJ1343_D04.120 [Oryza sativa]
Length = 1035
Score = 38.1 bits (87), Expect = 0.23
Identities = 34/101 (33%), Positives = 45/101 (43%), Gaps = 10/101 (9%)
Query: 171 PCDDNEVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQL 226
P E+V L +G PA Y IF F PFS+F DVL F ++ L
Sbjct: 34 PATGREIVML----GEGRPAPDYPGRSIFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHL 89
Query: 227 LPNAWGFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
PNA F LCE + P LF +F+ + S++ PS
Sbjct: 90 TPNAVMTLAIFAHLCEMFIGVRPSLQLFRWFFTVQSVSPPS 130
>UniRef100_Q9SDE2 Hypothetical protein [Oryza sativa]
Length = 965
Score = 38.1 bits (87), Expect = 0.23
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPCYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q7XSU1 OSJNBa0039K24.15 protein [Oryza sativa]
Length = 1036
Score = 38.1 bits (87), Expect = 0.23
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLQLFRWFFTVQSVSPPS 130
>UniRef100_Q949B8 Hypothetical protein C1015ERIPDK [Oryza sativa]
Length = 1012
Score = 37.7 bits (86), Expect = 0.31
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRSVFFLPFAMAGLVPPFSSFFMDVLEFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
>UniRef100_Q8W3E2 Putative membrane-associated protein [Oryza sativa]
Length = 946
Score = 37.7 bits (86), Expect = 0.31
Identities = 33/96 (34%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 176 EVVTLKPSGADGEPAWFYLHGYIFNELFMKL----PFSTFVCDVLTFLNVAPCQLLPNAW 231
E+VTL +G PA Y +F F PFS+F DVL F ++ L PNA
Sbjct: 39 EIVTL----GEGRPAPDYPGRSVFFLPFAMAGLVPPFSSFFMDVLKFYDLQMAHLTPNAV 94
Query: 232 GFTRCFELLCEP-LNFGPPYPLFLFFYKI-SIAKPS 265
F LCE + P LF +F+ + S++ PS
Sbjct: 95 MTLAIFAHLCEMFIGVRPSLRLFRWFFTVQSVSPPS 130
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 473,148,833
Number of Sequences: 2790947
Number of extensions: 19733387
Number of successful extensions: 57614
Number of sequences better than 10.0: 234
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 216
Number of HSP's that attempted gapping in prelim test: 57035
Number of HSP's gapped (non-prelim): 644
length of query: 265
length of database: 848,049,833
effective HSP length: 125
effective length of query: 140
effective length of database: 499,181,458
effective search space: 69885404120
effective search space used: 69885404120
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0128.2


