

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0126.13
(168 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
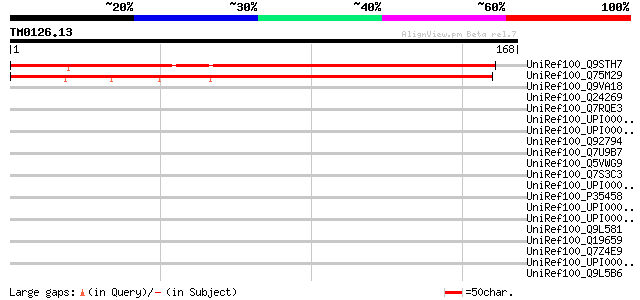
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9STH7 Hypothetical protein T4C9.180 [Arabidopsis thal... 148 5e-35
UniRef100_Q75M29 Unknow protein [Oryza sativa] 141 7e-33
UniRef100_Q9VA18 CG1715-PA [Drosophila melanogaster] 45 0.001
UniRef100_Q24269 D.melanogaster coding region cloned by GAL4 enh... 44 0.002
UniRef100_Q7RQE3 Hypothetical protein [Plasmodium yoelii yoelii] 40 0.030
UniRef100_UPI000036E2C6 UPI000036E2C6 UniRef100 entry 39 0.039
UniRef100_UPI000033F494 UPI000033F494 UniRef100 entry 39 0.039
UniRef100_Q92794 MYST histone acetyltransferase 3 [Homo sapiens] 39 0.039
UniRef100_Q7U9B7 Putative chromosome segregation protein, SMC AT... 39 0.051
UniRef100_Q5VWG9 OTTHUMP00000045008 (TAF3 RNA polymerase II, TAT... 39 0.051
UniRef100_Q7S3C3 Predicted protein [Neurospora crassa] 39 0.066
UniRef100_UPI00002D5A83 UPI00002D5A83 UniRef100 entry 38 0.087
UniRef100_P35458 Dynactin 1 [Gallus gallus] 38 0.087
UniRef100_UPI0000075B68 UPI0000075B68 UniRef100 entry 37 0.15
UniRef100_UPI000023F0C2 UPI000023F0C2 UniRef100 entry 37 0.15
UniRef100_Q9L581 PspA [Streptococcus pneumoniae] 37 0.15
UniRef100_Q19659 Hypothetical protein F55D12.5 [Caenorhabditis e... 37 0.15
UniRef100_Q7Z4E9 MSTP089 [Homo sapiens] 37 0.15
UniRef100_UPI000036B7F3 UPI000036B7F3 UniRef100 entry 37 0.19
UniRef100_Q9L5B6 PspA [Streptococcus pneumoniae] 37 0.19
>UniRef100_Q9STH7 Hypothetical protein T4C9.180 [Arabidopsis thaliana]
Length = 170
Score = 148 bits (374), Expect = 5e-35
Identities = 73/162 (45%), Positives = 114/162 (70%), Gaps = 3/162 (1%)
Query: 1 MGEFTIQISNELVNQLVD-DPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQ 59
MG+F+IQIS++L+NQL + + PK++ ++ + KV+ ++++ ++ K N A P+Q
Sbjct: 1 MGDFSIQISSKLINQLAEGNDQPKRRAKKTKPKVSPQSKQKTNQDEEKKPNPVAE-LPMQ 59
Query: 60 SPLFLPAKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKL 119
P F P PP A E++ I+SV++ESEKVLE L+ QE+N+++EVT++AKDL +KE+K+
Sbjct: 60 PPFFFPI-PPQGAASTELESIKSVVKESEKVLEKLELQEKNIVREVTERAKDLREKEFKI 118
Query: 120 PNPKPEPCIAERLATLSCYKEHIKDPLKCSAFVTNFAGCLRK 161
P PKP PC ++ A + CYKE+I PLKCS FV +F C R+
Sbjct: 119 PEPKPMPCSSDHEAWMKCYKENIGSPLKCSGFVKSFQDCARR 160
>UniRef100_Q75M29 Unknow protein [Oryza sativa]
Length = 180
Score = 141 bits (355), Expect = 7e-33
Identities = 78/168 (46%), Positives = 102/168 (60%), Gaps = 8/168 (4%)
Query: 1 MGEFTIQISNELVNQLV-DDPVPKKKIRRNRRK--VAKETEKPQSKVTGKP--ENAAAPG 55
MG++TIQIS +L++QL DD KKK R+ + K V + ++PQ P E A PG
Sbjct: 1 MGDYTIQISTKLIDQLARDDEKIKKKTRKPKPKKTVKQHQQEPQDNSRELPASEPKAPPG 60
Query: 56 WPVQSPLFLP---AKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDL 112
WP+Q P++LP A PP PA E++ IR+VL ESEKV E L KQ M E+ +K+KDL
Sbjct: 61 WPLQPPMYLPVTPAPPPPPPAFSELEAIRAVLEESEKVQEKLDKQHAGMRDELIKKSKDL 120
Query: 113 HDKEYKLPNPKPEPCIAERLATLSCYKEHIKDPLKCSAFVTNFAGCLR 160
DKE+KLP P PC ER CY + +DPLKC+ V F C+R
Sbjct: 121 RDKEFKLPYQNPTPCTEERSNCRQCYVSNAQDPLKCAEAVKRFEACVR 168
>UniRef100_Q9VA18 CG1715-PA [Drosophila melanogaster]
Length = 223
Score = 44.7 bits (104), Expect = 0.001
Identities = 40/176 (22%), Positives = 68/176 (37%), Gaps = 27/176 (15%)
Query: 6 IQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLP 65
I IS+++V +L + + + +K KP +K KP ++ P P
Sbjct: 25 IDISDDVVKRLKAGISQQAREHAAAAEESKPVPKPTTKAAAKPAASSPAASPAPKVSSYP 84
Query: 66 AKPPV---------QPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKE 116
A P+ AD + + +++ E E + K EEN+ +K + +KE
Sbjct: 85 AAVPIYVQGGGHTISAADVQRQMNQELIKNDELWKERMAKLEENL-----KKTNTILEKE 139
Query: 117 YK-------------LPNPKPEPCIAERLATLSCYKEHIKDPLKCSAFVTNFAGCL 159
Y + K PC + L+CY+ H + LKC V F C+
Sbjct: 140 YANAVENVHKRFVSTASSHKVPPCQDLKSQLLACYRAHPGETLKCIEEVAQFRQCI 195
>UniRef100_Q24269 D.melanogaster coding region cloned by GAL4 enhancer trap line
[Drosophila melanogaster]
Length = 223
Score = 43.9 bits (102), Expect = 0.002
Identities = 40/176 (22%), Positives = 67/176 (37%), Gaps = 27/176 (15%)
Query: 6 IQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLP 65
I IS+++V +L + + +K KP +K KP ++ P P
Sbjct: 25 IDISDDVVKRLKAGISQQAHEHAAAAEESKPVPKPTTKAAAKPAASSPAASPAPKVSSYP 84
Query: 66 AKPPV---------QPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKE 116
A P+ AD + + +++ E E + K EEN+ +K + +KE
Sbjct: 85 AAVPIYVQGGGHTISAADVQRQMNQELIKNDELWKERMAKLEENL-----KKTNTILEKE 139
Query: 117 YK-------------LPNPKPEPCIAERLATLSCYKEHIKDPLKCSAFVTNFAGCL 159
Y + K PC + L+CY+ H + LKC V F C+
Sbjct: 140 YANAVENVHKRFVSTASSHKVPPCQDLKSQLLACYRAHPGETLKCMEEVAQFRQCI 195
>UniRef100_Q7RQE3 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 470
Score = 39.7 bits (91), Expect = 0.030
Identities = 30/124 (24%), Positives = 57/124 (45%), Gaps = 8/124 (6%)
Query: 5 TIQISNELVNQLVDDPVPKKKIRRNRRKVA---KETEKPQSKVTGKPENAAAPGWPVQSP 61
T I EL + + KK++ +++V +E E Q +V K + + V+S
Sbjct: 181 TEHIKKELEGKNKEVEDKKKEVESKQKEVESKQREVESKQKEVESKQKEVESKQKEVESK 240
Query: 62 LFLPAKPPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPN 121
+ V+ E++ + + +K +ES QK+ E+ +EV K KD+ ++E +
Sbjct: 241 -----QKEVETKQKEVESKQKEVETQQKEVESKQKEVESKQKEVESKQKDIENREKESKE 295
Query: 122 PKPE 125
K E
Sbjct: 296 TKVE 299
Score = 32.7 bits (73), Expect = 3.6
Identities = 21/102 (20%), Positives = 49/102 (47%), Gaps = 8/102 (7%)
Query: 23 KKKIRRNRRKVA---KETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQPADPEIDG 79
+K++ +R+V KE E Q +V K + + V++ + V+ E++
Sbjct: 206 QKEVESKQREVESKQKEVESKQKEVESKQKEVESKQKEVETK-----QKEVESKQKEVET 260
Query: 80 IRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPN 121
+ + +K +ES QK+ E+ +++ + K+ + + + PN
Sbjct: 261 QQKEVESKQKEVESKQKEVESKQKDIENREKESKETKVETPN 302
>UniRef100_UPI000036E2C6 UPI000036E2C6 UniRef100 entry
Length = 2024
Score = 39.3 bits (90), Expect = 0.039
Identities = 27/94 (28%), Positives = 47/94 (49%), Gaps = 5/94 (5%)
Query: 6 IQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLP 65
IQ S E V D P+P++ R+ ++ E E+ + G+ E+AA+ P SP
Sbjct: 1201 IQESEETVEPKEDMPLPEE--RKEEEEMQAEAEEAEE---GEEEDAASSEVPAASPADSS 1255
Query: 66 AKPPVQPADPEIDGIRSVLRESEKVLESLQKQEE 99
P + +PE++ R SE+ +S ++Q+E
Sbjct: 1256 NSPETETKEPEVEEEEEKPRVSEEQRQSEEEQQE 1289
>UniRef100_UPI000033F494 UPI000033F494 UniRef100 entry
Length = 283
Score = 39.3 bits (90), Expect = 0.039
Identities = 22/71 (30%), Positives = 40/71 (55%)
Query: 73 ADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPKPEPCIAERL 132
A+ E+ R L+++E LE L+++ + +++E A L + E LP+P+PE A RL
Sbjct: 191 AEAEVARQRQALQQAEWNLERLREERDGLIEEQRSGAIRLEEMEQALPDPRPEIPEALRL 250
Query: 133 ATLSCYKEHIK 143
A L + ++
Sbjct: 251 AGLEALQADLQ 261
>UniRef100_Q92794 MYST histone acetyltransferase 3 [Homo sapiens]
Length = 2004
Score = 39.3 bits (90), Expect = 0.039
Identities = 27/94 (28%), Positives = 47/94 (49%), Gaps = 5/94 (5%)
Query: 6 IQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLP 65
IQ S E V D P+P++ R+ ++ E E+ + G+ E+AA+ P SP
Sbjct: 1204 IQESEETVEPKEDMPLPEE--RKEEEEMQAEAEEAEE---GEEEDAASSEVPAASPADSS 1258
Query: 66 AKPPVQPADPEIDGIRSVLRESEKVLESLQKQEE 99
P + +PE++ R SE+ +S ++Q+E
Sbjct: 1259 NSPETETKEPEVEEEEEKPRVSEEQRQSEEEQQE 1292
>UniRef100_Q7U9B7 Putative chromosome segregation protein, SMC ATPase superfamily
[Synechococcus sp.]
Length = 1203
Score = 38.9 bits (89), Expect = 0.051
Identities = 22/71 (30%), Positives = 39/71 (53%)
Query: 73 ADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPKPEPCIAERL 132
A+ E+ R L+++E LE L+++ E +++E A L + E LP+P+PE RL
Sbjct: 928 AEAEVGRQRQALQQAEWNLERLKEEREGLIEEQRSGAVRLQEMEQALPDPQPEIPEELRL 987
Query: 133 ATLSCYKEHIK 143
A L + ++
Sbjct: 988 AGLEALQADLQ 998
>UniRef100_Q5VWG9 OTTHUMP00000045008 (TAF3 RNA polymerase II, TATA box binding
protein (TBP)-associated factor, 140kDa
(TAF3)(TAF140,TAFII140)) [Homo sapiens]
Length = 929
Score = 38.9 bits (89), Expect = 0.051
Identities = 33/126 (26%), Positives = 56/126 (44%), Gaps = 20/126 (15%)
Query: 6 IQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWP-------V 58
+++ + LV + + KKK R +K + EK + K G+ + AP P +
Sbjct: 602 VKLKDGLVRKEKEKHKDKKKDREKGKKDKDKREKEKVKDKGREDKMKAPAPPLVLPPKEL 661
Query: 59 QSPLFLPAK--------PPVQPADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAK 110
PLF PA P + P PE + E EKV E +K+++ ++ +K K
Sbjct: 662 ALPLFSPATASRVPAMLPSLLPVLPE-----KLFEEKEKVKEKEKKKDKKEKKKKKEKEK 716
Query: 111 DLHDKE 116
+ +KE
Sbjct: 717 EKKEKE 722
>UniRef100_Q7S3C3 Predicted protein [Neurospora crassa]
Length = 2221
Score = 38.5 bits (88), Expect = 0.066
Identities = 21/57 (36%), Positives = 31/57 (53%), Gaps = 1/57 (1%)
Query: 20 PVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQPADPE 76
P + RR RR K + P ++ GK + P P QSP F P++PP+QP+ P+
Sbjct: 1945 PSKLRSRRRGRRTRPKVSLSPNPRLFGKG-TSPDPLQPSQSPHFQPSEPPLQPSSPQ 2000
>UniRef100_UPI00002D5A83 UPI00002D5A83 UniRef100 entry
Length = 599
Score = 38.1 bits (87), Expect = 0.087
Identities = 22/71 (30%), Positives = 39/71 (53%)
Query: 73 ADPEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPKPEPCIAERL 132
A+ E+ R L+++E LE L+++ E +++E A L + E LP+P+PE RL
Sbjct: 389 AEAEVGRQRQELQQAEWNLERLKEEREGLIEEQRSGAIRLQEMEQALPDPRPEIPEELRL 448
Query: 133 ATLSCYKEHIK 143
A L + ++
Sbjct: 449 AGLEALQADLQ 459
>UniRef100_P35458 Dynactin 1 [Gallus gallus]
Length = 1224
Score = 38.1 bits (87), Expect = 0.087
Identities = 40/149 (26%), Positives = 64/149 (42%), Gaps = 26/149 (17%)
Query: 22 PKKKIRRNRRKVAKETEKPQSKVTGKPENAAA---------PGWPVQSPLFLPAKP---- 68
PKK R + T P S G +A+A P P Q+PL P P
Sbjct: 136 PKKTTARRPKPTRTPTSAPSSGTAGPSGSASASGGEMSSSEPSTPAQTPLVAPVIPSPSL 195
Query: 69 --PVQPADP----EIDGIRSVLRESEKVLESL---QKQEENMLQEVTQKAKDLHD-KEYK 118
PV P P E + +RS +R+ E+ LE+L + +++ L+E+ + L +E+K
Sbjct: 196 TSPVAPMVPSPTKEEENLRSQVRDLEEKLETLKIKRNEDKAKLKELEKYKIQLEQVQEWK 255
Query: 119 LPNPKPEPCIAERLATLSCYKEHIKDPLK 147
+ + + RL K+ KD L+
Sbjct: 256 SKMQEQQADLQRRLKEA---KKEAKDALE 281
>UniRef100_UPI0000075B68 UPI0000075B68 UniRef100 entry
Length = 567
Score = 37.4 bits (85), Expect = 0.15
Identities = 25/90 (27%), Positives = 37/90 (40%), Gaps = 17/90 (18%)
Query: 3 EFTIQISNELVNQLVDDP-----VPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWP 57
+FT++ + E +DD VP ++ + A + KP T PE AA P W
Sbjct: 127 DFTVETNAEFNENSLDDSDEYPIVPNRQEKIEETIGAVDMRKPVETKTMSPEEAALPPWR 186
Query: 58 VQSPLF------------LPAKPPVQPADP 75
+PL +P KPP+ P P
Sbjct: 187 RMNPLNADAEWKPPPPPPMPGKPPIVPQSP 216
>UniRef100_UPI000023F0C2 UPI000023F0C2 UniRef100 entry
Length = 198
Score = 37.4 bits (85), Expect = 0.15
Identities = 32/109 (29%), Positives = 46/109 (41%), Gaps = 7/109 (6%)
Query: 23 KKKIRRNRRKVAKETEKPQSKVTGKP-ENAAAPGWPVQSPLFLPAKPPVQPADPEIDGIR 81
K++ + +K + T + T +P + AAP P +P PA P +PA PE +
Sbjct: 26 KQEAKPAEQKPTETTPADATPATTEPTKTEAAPPAPATAPA--PAAPAAEPAKPETE--- 80
Query: 82 SVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLPNPKPEPCIAE 130
V E+ K EE T+ AK +LP P PEP E
Sbjct: 81 -VKPEAPAPTTEPAKTEEAAPAPATEPAKPEAAPATELPAPVPEPAKTE 128
>UniRef100_Q9L581 PspA [Streptococcus pneumoniae]
Length = 255
Score = 37.4 bits (85), Expect = 0.15
Identities = 34/125 (27%), Positives = 48/125 (38%), Gaps = 14/125 (11%)
Query: 2 GEFTIQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSP 61
GE++ LV +K + +K E EKP + PEN A P +P
Sbjct: 128 GEYSALYLEAAEKDLVAKKAELEKTEADLKKAVNEPEKPAEE----PENPAPAPKPAPAP 183
Query: 62 LFLPAKPPVQPAD-PEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLP 120
P KP PA PE ++V E ++ E +TQ+ +K P
Sbjct: 184 Q--PEKPAPAPAPKPEKSA-------DQQVEEDYARRSEEEYNRLTQQQPPKAEKPAPAP 234
Query: 121 NPKPE 125
PKPE
Sbjct: 235 VPKPE 239
>UniRef100_Q19659 Hypothetical protein F55D12.5 [Caenorhabditis elegans]
Length = 552
Score = 37.4 bits (85), Expect = 0.15
Identities = 25/90 (27%), Positives = 37/90 (40%), Gaps = 17/90 (18%)
Query: 3 EFTIQISNELVNQLVDDP-----VPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWP 57
+FT++ + E +DD VP ++ + A + KP T PE AA P W
Sbjct: 127 DFTVETNAEFNENSLDDSDEYPIVPNRQEKIEETIGAVDMRKPVETKTMSPEEAALPPWR 186
Query: 58 VQSPLF------------LPAKPPVQPADP 75
+PL +P KPP+ P P
Sbjct: 187 RMNPLNADAEWKPPPPPPMPGKPPIVPQSP 216
>UniRef100_Q7Z4E9 MSTP089 [Homo sapiens]
Length = 568
Score = 37.4 bits (85), Expect = 0.15
Identities = 28/108 (25%), Positives = 50/108 (45%), Gaps = 14/108 (12%)
Query: 18 DDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQP--ADP 75
D P+ K ++ R+ AK+ K +++ P+ A P +P P Q DP
Sbjct: 460 DKPLSKTALKNQRKHEAKKAAKQEARSAKSPDLAPTP-----APQSTPRNTVSQSISGDP 514
Query: 76 EIDGIRSVLRESEKVLESLQKQ-------EENMLQEVTQKAKDLHDKE 116
EID L++ K +E L++Q E+N L+++ ++ L + E
Sbjct: 515 EIDKKIKNLKKKLKAIEQLKEQAATGKQLEKNQLEKIQKETALLQELE 562
>UniRef100_UPI000036B7F3 UPI000036B7F3 UniRef100 entry
Length = 402
Score = 37.0 bits (84), Expect = 0.19
Identities = 28/108 (25%), Positives = 50/108 (45%), Gaps = 14/108 (12%)
Query: 18 DDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSPLFLPAKPPVQP--ADP 75
D P+ K ++ R+ AK+ K +++ P+ A P +P P Q DP
Sbjct: 294 DKPLSKTALKNQRKHEAKKAAKQEARSDKSPDLAPTP-----APQSTPRNTVSQSISGDP 348
Query: 76 EIDGIRSVLRESEKVLESLQKQ-------EENMLQEVTQKAKDLHDKE 116
EID L++ K +E L++Q E+N L+++ ++ L + E
Sbjct: 349 EIDKKIKNLKKKLKAIEQLKEQAATGKQLEKNQLEKIQKETALLQELE 396
>UniRef100_Q9L5B6 PspA [Streptococcus pneumoniae]
Length = 255
Score = 37.0 bits (84), Expect = 0.19
Identities = 35/125 (28%), Positives = 51/125 (40%), Gaps = 14/125 (11%)
Query: 2 GEFTIQISNELVNQLVDDPVPKKKIRRNRRKVAKETEKPQSKVTGKPENAAAPGWPVQSP 61
GE++ LV +K + +K E EKP + PEN A P +P
Sbjct: 128 GEYSALYLEAAEKDLVAKKAELEKTEADLKKAVNEPEKPAEE----PENPAPAPKPAPAP 183
Query: 62 LFLPAKPPVQPAD-PEIDGIRSVLRESEKVLESLQKQEENMLQEVTQKAKDLHDKEYKLP 120
P KP PA PE +S +++E E ++ E +TQ+ +K P
Sbjct: 184 Q--PEKPAPAPAPKPE----KSADQQAE---EDYARRSEEEYNRLTQQQPPKAEKPAPAP 234
Query: 121 NPKPE 125
PKPE
Sbjct: 235 VPKPE 239
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.132 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 288,392,899
Number of Sequences: 2790947
Number of extensions: 12073064
Number of successful extensions: 56759
Number of sequences better than 10.0: 392
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 347
Number of HSP's that attempted gapping in prelim test: 56251
Number of HSP's gapped (non-prelim): 748
length of query: 168
length of database: 848,049,833
effective HSP length: 118
effective length of query: 50
effective length of database: 518,718,087
effective search space: 25935904350
effective search space used: 25935904350
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0126.13


