

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0122.7
(125 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
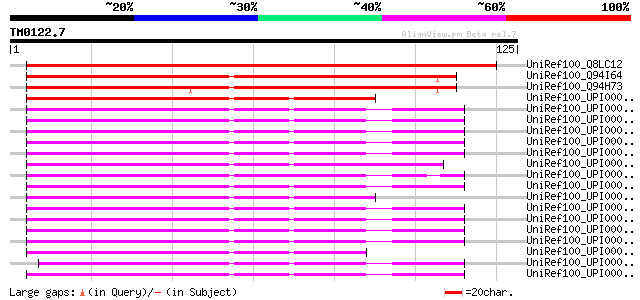
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LC12 Hypothetical protein [Arabidopsis thaliana] 158 3e-38
UniRef100_Q94I64 Hypothetical protein OSJNBa0084C09.7 [Oryza sat... 134 6e-31
UniRef100_Q94H73 Hypothetical protein OSJNBb0057P11.6 [Oryza sat... 126 1e-28
UniRef100_UPI00002D7DAD UPI00002D7DAD UniRef100 entry 78 5e-14
UniRef100_UPI00002B8772 UPI00002B8772 UniRef100 entry 77 7e-14
UniRef100_UPI00002B0BCA UPI00002B0BCA UniRef100 entry 77 7e-14
UniRef100_UPI00002A7823 UPI00002A7823 UniRef100 entry 76 2e-13
UniRef100_UPI000028004F UPI000028004F UniRef100 entry 75 3e-13
UniRef100_UPI00003435E8 UPI00003435E8 UniRef100 entry 75 4e-13
UniRef100_UPI0000301F7B UPI0000301F7B UniRef100 entry 72 2e-12
UniRef100_UPI0000342515 UPI0000342515 UniRef100 entry 72 3e-12
UniRef100_UPI000031D623 UPI000031D623 UniRef100 entry 72 4e-12
UniRef100_UPI000030AE26 UPI000030AE26 UniRef100 entry 72 4e-12
UniRef100_UPI000030DAB9 UPI000030DAB9 UniRef100 entry 71 5e-12
UniRef100_UPI00002FB182 UPI00002FB182 UniRef100 entry 71 5e-12
UniRef100_UPI00002C494F UPI00002C494F UniRef100 entry 71 6e-12
UniRef100_UPI0000309560 UPI0000309560 UniRef100 entry 70 8e-12
UniRef100_UPI00003138E4 UPI00003138E4 UniRef100 entry 69 2e-11
UniRef100_UPI00002E6442 UPI00002E6442 UniRef100 entry 68 4e-11
UniRef100_UPI000033A7B6 UPI000033A7B6 UniRef100 entry 67 9e-11
>UniRef100_Q8LC12 Hypothetical protein [Arabidopsis thaliana]
Length = 122
Score = 158 bits (399), Expect = 3e-38
Identities = 72/116 (62%), Positives = 91/116 (78%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+YRIST +EWEE + NGS++G +DKS+ + HLSKLDQV+ TL+ F+++ KE LYLLQ+
Sbjct: 6 FIYRISTEQEWEEFKKNGSSYGAEIDKSTCYYHLSKLDQVQLTLKNFFVDVKEYLYLLQV 65
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSDGRFTCCLL 120
D KKLGDGL+YE VD NSFPHFYGP ++F PL LD V KAEKL+ +G FTC L
Sbjct: 66 DPKKLGDGLIYEAVDEVNSFPHFYGPDKTFVPLPLDSVVKAEKLTFINGNFTCSFL 121
>UniRef100_Q94I64 Hypothetical protein OSJNBa0084C09.7 [Oryza sativa]
Length = 144
Score = 134 bits (336), Expect = 6e-31
Identities = 67/107 (62%), Positives = 79/107 (73%), Gaps = 2/107 (1%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVYRISTA EW +LQ G T GG+LD+S+ IHLS L QVR TL+ F++ + +LYLLQ+
Sbjct: 9 FVYRISTADEWAQLQRTGGTLGGDLDRSTGCIHLSDLSQVRKTLKNFFLG-RNDLYLLQV 67
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTK-AEKLSL 110
D KL DGLVYE D SN FPHFYGP RSF+PL LD V K AEK+ L
Sbjct: 68 DTSKLSDGLVYEAADDSNYFPHFYGPGRSFAPLQLDAVIKEAEKIVL 114
>UniRef100_Q94H73 Hypothetical protein OSJNBb0057P11.6 [Oryza sativa]
Length = 133
Score = 126 bits (316), Expect = 1e-28
Identities = 67/116 (57%), Positives = 79/116 (67%), Gaps = 11/116 (9%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQ---------VRSTLERFYMNC 55
FVYRISTA EW +LQ G T GG+LD+S+ IHLS L Q VR TL+ F++
Sbjct: 9 FVYRISTADEWAQLQRTGGTLGGDLDRSTGCIHLSDLSQGIHNVRKKQVRKTLKNFFLG- 67
Query: 56 KEELYLLQIDAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTK-AEKLSL 110
+ +LYLLQ+D KL DGLVYE D SN FPHFYGP RSF+PL LD V K AEK+ L
Sbjct: 68 RNDLYLLQVDTSKLSDGLVYEAADDSNYFPHFYGPGRSFAPLQLDAVIKEAEKIVL 123
>UniRef100_UPI00002D7DAD UPI00002D7DAD UniRef100 entry
Length = 115
Score = 77.8 bits (190), Expect = 5e-14
Identities = 39/86 (45%), Positives = 53/86 (61%), Gaps = 2/86 (2%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
+VY+I T +EWEE + G G DK FIH S +Q++ TL++F+ N ++ L LL+I
Sbjct: 5 YVYKICTYEEWEEAKKKGKYEGSKKDKEDGFIHFSDKEQLKGTLKKFFFN-QKNLILLKI 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGP 90
DA KL L+YE V N FPH Y P
Sbjct: 64 DALKL-KNLIYEQVSDGNMFPHLYEP 88
>UniRef100_UPI00002B8772 UPI00002B8772 UniRef100 entry
Length = 114
Score = 77.4 bits (189), Expect = 7e-14
Identities = 46/108 (42%), Positives = 58/108 (53%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW E + N G LD +IHLS DQV+ TL +FY+N + L +L++
Sbjct: 5 FVYKICTKSEWSEAKQNNIFKGTALDLKDGYIHLSDKDQVKQTLNKFYIN-QNNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L LD V K + L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLDLDFVVKEYMIDLKD 104
>UniRef100_UPI00002B0BCA UPI00002B0BCA UniRef100 entry
Length = 114
Score = 77.4 bits (189), Expect = 7e-14
Identities = 46/108 (42%), Positives = 58/108 (53%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW E + N G LD +IHLS DQV+ TL +FY+N + L +L++
Sbjct: 5 FVYKICTKSEWSEAKQNNVFKGTALDLKDGYIHLSDKDQVKQTLNKFYIN-QNNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L LD V K + L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLDLDFVVKEYMIDLKD 104
>UniRef100_UPI00002A7823 UPI00002A7823 UniRef100 entry
Length = 114
Score = 75.9 bits (185), Expect = 2e-13
Identities = 44/108 (40%), Positives = 60/108 (54%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW E + N G +LD +IH S DQ++ TL +FY+N + L +L++
Sbjct: 5 FVYKICTKSEWSEAKQNNIFKGTSLDLKDGYIHFSDKDQIKQTLNKFYVN-QTNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L+LD V K ++L D
Sbjct: 64 DAIKL-KNLVWEQSIDGNMFPHLY------SDLNLDFVVKEYMINLKD 104
>UniRef100_UPI000028004F UPI000028004F UniRef100 entry
Length = 114
Score = 75.5 bits (184), Expect = 3e-13
Identities = 45/108 (41%), Positives = 57/108 (52%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW E + N G LD +IH S DQV+ TL +FY+N + L +L++
Sbjct: 5 FVYKICTKSEWSEAKQNNVFKGTALDLKDGYIHFSDKDQVKQTLNKFYIN-QTNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L LD V K + L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLDLDFVVKEYMIDLKD 104
>UniRef100_UPI00003435E8 UPI00003435E8 UniRef100 entry
Length = 114
Score = 74.7 bits (182), Expect = 4e-13
Identities = 43/108 (39%), Positives = 58/108 (52%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW + N G LD +IH S DQ++ TL +FY+N + L +L++
Sbjct: 5 FVYKICTKSEWSNAKQNNIFKGTTLDLKDGYIHFSDKDQIKQTLNKFYVN-QTNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L+LD V K ++L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLNLDFVVKEYMINLKD 104
>UniRef100_UPI0000301F7B UPI0000301F7B UniRef100 entry
Length = 115
Score = 72.4 bits (176), Expect = 2e-12
Identities = 39/103 (37%), Positives = 57/103 (54%), Gaps = 2/103 (1%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
+VY+I T +EW+ + G G DK FIH S DQ++ T ++F+ N + L LL+I
Sbjct: 5 YVYKICTKEEWKIAKKIGKFEGSKKDKEDGFIHFSDKDQLKETFKKFFFN-QNNLILLKI 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEK 107
DA KL L+YE + N FPH Y P + ++ +T EK
Sbjct: 64 DALKL-KNLIYEQISDGNMFPHLYEPFEVKNVVTEHEITLNEK 105
>UniRef100_UPI0000342515 UPI0000342515 UniRef100 entry
Length = 102
Score = 72.0 bits (175), Expect = 3e-12
Identities = 43/108 (39%), Positives = 58/108 (52%), Gaps = 10/108 (9%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVYR+ +KEWE Q + G LD+SS F+HLS Q++ T+ Y L +L+I
Sbjct: 4 FVYRVLYSKEWERFQKDRIYKGNELDRSSGFLHLSNKQQLKKTINIHYKK-NNNLVILKI 62
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
++KKL LV+E G FPH Y L LD VT ++SL D
Sbjct: 63 ESKKLNKNLVWEFSRGGEKFPHLY------DELHLDDVT---EISLPD 101
>UniRef100_UPI000031D623 UPI000031D623 UniRef100 entry
Length = 114
Score = 71.6 bits (174), Expect = 4e-12
Identities = 45/108 (41%), Positives = 57/108 (52%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T E E + N G LD +IHLS DQV+ TL +FY+N + L +L++
Sbjct: 5 FVYKICTKSELSEAKQNNVFKGTALDLKDGYIHLSDKDQVKQTLNKFYLN-QTNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L LD V K + L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLDLDFVVKEYIIDLKD 104
>UniRef100_UPI000030AE26 UPI000030AE26 UniRef100 entry
Length = 115
Score = 71.6 bits (174), Expect = 4e-12
Identities = 36/86 (41%), Positives = 49/86 (56%), Gaps = 2/86 (2%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
+VY+I T +EWEE + G G DK FIH S +Q++ T +F+ ++ L LL+I
Sbjct: 5 YVYKICTYEEWEEAKEKGKYEGSKKDKEDGFIHFSDKEQLKGTFRKFFSK-QKNLILLKI 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGP 90
DA KL L+YE N FPH Y P
Sbjct: 64 DALKL-KNLIYEQASDGNMFPHLYEP 88
>UniRef100_UPI000030DAB9 UPI000030DAB9 UniRef100 entry
Length = 114
Score = 71.2 bits (173), Expect = 5e-12
Identities = 43/108 (39%), Positives = 54/108 (49%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW E + N G LD +IH S DQ+ TL +FY + L +L++
Sbjct: 5 FVYKICTKSEWSEAKQNNVFKGTALDLKDGYIHFSDKDQINQTLNKFYEK-QSNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L LD V K + L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLDLDFVVKEYMIDLKD 104
>UniRef100_UPI00002FB182 UPI00002FB182 UniRef100 entry
Length = 114
Score = 71.2 bits (173), Expect = 5e-12
Identities = 43/108 (39%), Positives = 54/108 (49%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW E + N G LD +IH S DQ+ TL +FY + L +L++
Sbjct: 5 FVYKICTKSEWNEAKQNNIFKGTALDLKDGYIHFSDKDQINQTLNKFYEK-QSNLIILKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E N FPH Y S L LD V K + L D
Sbjct: 64 DALKL-KNLVWEQSTDGNMFPHLY------SDLDLDFVVKEYMIDLKD 104
>UniRef100_UPI00002C494F UPI00002C494F UniRef100 entry
Length = 115
Score = 70.9 bits (172), Expect = 6e-12
Identities = 42/108 (38%), Positives = 57/108 (51%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW + + N G LD FIH S ++QV+ TL +FY+N + L LL++
Sbjct: 5 FVYKICTKSEWNDAKENKIFKGTPLDLKDGFIHFSDINQVKQTLNKFYLN-QPNLVLLKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E + FPH Y S +D V K L L +
Sbjct: 64 DALKL-QKLVWEQSSSGDMFPHLY------SDFDIDCVVKEYPLDLKE 104
>UniRef100_UPI0000309560 UPI0000309560 UniRef100 entry
Length = 123
Score = 70.5 bits (171), Expect = 8e-12
Identities = 39/108 (36%), Positives = 64/108 (59%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+++I +EW++ + G+ G + DK +IH S+ DQV TL+++Y N KE L LL++
Sbjct: 5 FIFKIIDKEEWQKAKQAGTYTGSDKDKKDGYIHFSEEDQVPETLKKYYQN-KENLILLKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
+A KL + L++E + +PH Y SPL + V +LSL+D
Sbjct: 64 NAFKL-EHLLWEQASNGDMYPHLY------SPLDIKNVEDEFELSLND 104
>UniRef100_UPI00003138E4 UPI00003138E4 UniRef100 entry
Length = 112
Score = 68.9 bits (167), Expect = 2e-11
Identities = 32/84 (38%), Positives = 50/84 (59%), Gaps = 2/84 (2%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
F+Y+I T EW + + G G D FIH S DQ++ TL ++++N +++L LL++
Sbjct: 5 FIYKICTKAEWLKAKKEGKFLGSKKDIEDGFIHFSNKDQLKGTLSKYFLN-QKDLVLLKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFY 88
+A KL + L+YE N FPH Y
Sbjct: 64 EALKL-NNLIYEQASDGNMFPHLY 86
>UniRef100_UPI00002E6442 UPI00002E6442 UniRef100 entry
Length = 107
Score = 68.2 bits (165), Expect = 4e-11
Identities = 42/105 (40%), Positives = 54/105 (51%), Gaps = 8/105 (7%)
Query: 8 RISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQIDAK 67
+I T EW E + N G LD +IH S DQV+ TL +FY+N + L +L++DA
Sbjct: 1 KICTKSEWSEAKQNNVFKGTALDLKDGYIHFSDKDQVKQTLNKFYLN-QTNLIILKVDAL 59
Query: 68 KLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
KL LV+E N FPH Y S L LD V K + L D
Sbjct: 60 KL-KNLVWEQSTDGNMFPHLY------SDLDLDFVIKEYIIDLKD 97
>UniRef100_UPI000033A7B6 UPI000033A7B6 UniRef100 entry
Length = 114
Score = 67.0 bits (162), Expect = 9e-11
Identities = 42/108 (38%), Positives = 55/108 (50%), Gaps = 8/108 (7%)
Query: 5 FVYRISTAKEWEELQANGSTFGGNLDKSSAFIHLSKLDQVRSTLERFYMNCKEELYLLQI 64
FVY+I T EW + + N G LD FIH S +QV+ TL +FY N + L LL++
Sbjct: 5 FVYKICTKSEWRDAKENKIFKGTLLDLKDGFIHFSDKNQVKQTLNKFYGN-QTNLVLLKV 63
Query: 65 DAKKLGDGLVYEMVDGSNSFPHFYGPSRSFSPLSLDVVTKAEKLSLSD 112
DA KL LV+E + FPH Y S +D V K L L +
Sbjct: 64 DALKL-QKLVWEQSSSGDMFPHLY------SDFDIDCVIKEYSLDLKE 104
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.136 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 204,962,046
Number of Sequences: 2790947
Number of extensions: 7716783
Number of successful extensions: 16149
Number of sequences better than 10.0: 126
Number of HSP's better than 10.0 without gapping: 63
Number of HSP's successfully gapped in prelim test: 63
Number of HSP's that attempted gapping in prelim test: 16010
Number of HSP's gapped (non-prelim): 126
length of query: 125
length of database: 848,049,833
effective HSP length: 101
effective length of query: 24
effective length of database: 566,164,186
effective search space: 13587940464
effective search space used: 13587940464
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0122.7


