

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0121b.2
(253 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
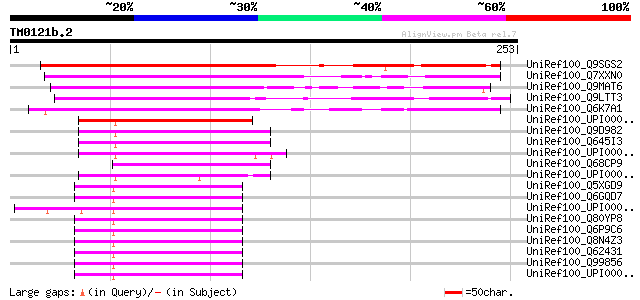
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SGS2 T23E18.4 [Arabidopsis thaliana] 207 2e-52
UniRef100_Q7XXN0 Glutathione S-transferase GST 16-like protein [... 169 6e-41
UniRef100_Q9MAT6 F13M7.13 protein [Arabidopsis thaliana] 140 3e-32
UniRef100_Q9LTT3 High mobility group protein-like [Arabidopsis t... 112 1e-23
UniRef100_Q6K7A1 Glutathione S-transferase GST16-like protein [O... 112 1e-23
UniRef100_UPI00003A9660 UPI00003A9660 UniRef100 entry 85 2e-15
UniRef100_Q9D982 Mus musculus adult male testis cDNA, RIKEN full... 85 2e-15
UniRef100_Q645I3 ARID2 [Homo sapiens] 85 2e-15
UniRef100_UPI0000360A2D UPI0000360A2D UniRef100 entry 83 8e-15
UniRef100_Q68CP9 Hypothetical protein DKFZp779P0222 [Homo sapiens] 70 7e-11
UniRef100_UPI00002CEE47 UPI00002CEE47 UniRef100 entry 66 1e-09
UniRef100_Q5XGD9 Hypothetical protein [Xenopus tropicalis] 60 4e-08
UniRef100_Q6GQD7 MGC80148 protein [Xenopus laevis] 60 4e-08
UniRef100_UPI0000365B23 UPI0000365B23 UniRef100 entry 60 5e-08
UniRef100_Q80YP8 Dead ringer homolog 1 [Mus musculus] 59 9e-08
UniRef100_Q6P9C6 AT rich interactive domain 3A (BRIGHT-like) pro... 59 9e-08
UniRef100_Q8N4Z3 ARID3A protein [Homo sapiens] 59 9e-08
UniRef100_Q62431 AT-rich interactive domain-containing protein 3... 59 9e-08
UniRef100_Q99856 AT-rich interactive domain-containing protein 3... 59 9e-08
UniRef100_UPI00003AFFE8 UPI00003AFFE8 UniRef100 entry 59 1e-07
>UniRef100_Q9SGS2 T23E18.4 [Arabidopsis thaliana]
Length = 338
Score = 207 bits (528), Expect = 2e-52
Identities = 117/233 (50%), Positives = 142/233 (60%), Gaps = 43/233 (18%)
Query: 16 KFYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGY 75
K YP PLA HEV+V D+ +F DTLR+FH M +K+MIPVIGG++LDLH LYVEVTRR GY
Sbjct: 25 KEYPEPLALHEVVVKDSSVFWDTLRRFHSIMSTKFMIPVIGGKELDLHVLYVEVTRRGGY 84
Query: 76 EKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVL 135
EKVV EKKWREVG VF FS TTTSAS+VL+KHY NLL+++EQVH F +GP+ P T
Sbjct: 85 EKVVVEKKWREVGGVFRFSATTTSASFVLRKHYLNLLFHYEQVHLFTARGPLLHPIAT-- 142
Query: 136 NSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAP---KHANNG 192
HA + S ++ALVEY P ++ N
Sbjct: 143 -------------------FHA--------------NPSTSKEMALVEYTPPSIRYHNTH 169
Query: 193 PESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
P S + G IE KFDCGYLV VKLGSE+L GV++H + P PSS
Sbjct: 170 PPSQGSSSFT---AIGTIEGKFDCGYLVKVKLGSEILNGVLYHSAQ--PGPSS 217
>UniRef100_Q7XXN0 Glutathione S-transferase GST 16-like protein [Oryza sativa]
Length = 306
Score = 169 bits (428), Expect = 6e-41
Identities = 97/228 (42%), Positives = 134/228 (58%), Gaps = 31/228 (13%)
Query: 18 YPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEK 77
YPPPL SHE + ND F+DTLR+FH MG+K+MIPVIGG+++DLH LYVEVT R G K
Sbjct: 7 YPPPLLSHEEVANDRAAFMDTLRRFHSLMGTKFMIPVIGGKEMDLHALYVEVTSRGGLAK 66
Query: 78 VVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNS 137
V+ E+KWREV + F+F TTTSASYVL+++Y +LL+++EQV+FF+ G + PA + L
Sbjct: 67 VMEERKWREVMARFSFPATTTSASYVLRRYYLSLLHHYEQVYFFRAHGALLRPAASALTK 126
Query: 138 FTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNV 197
M S D ++ +GK +AL P+ P S
Sbjct: 127 TPRRKMRGTS------------------DQSPAAAEAGK-RMAL----PERLGGEPCSFS 163
Query: 198 EAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSS 245
G I+ KF+ GYLV+VK+ +E LRGV++ PPP++
Sbjct: 164 VT--------GSIDGKFEHGYLVTVKIAAETLRGVLYRVAPPPPPPAA 203
>UniRef100_Q9MAT6 F13M7.13 protein [Arabidopsis thaliana]
Length = 448
Score = 140 bits (353), Expect = 3e-32
Identities = 85/222 (38%), Positives = 131/222 (58%), Gaps = 30/222 (13%)
Query: 21 PLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVA 80
P A++E +V D +LF+ +L + H +G+K+M+P+IGGR LDLH L+VEVT R G K++
Sbjct: 21 PEATYEAVVADPRLFMTSLERLHSLLGTKFMVPIIGGRDLDLHKLFVEVTSRGGINKILN 80
Query: 81 EKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTL 140
E++W+EV + F F T T+ASYVL+K+Y++LL N+EQ++FF+ G I PP S
Sbjct: 81 ERRWKEVTATFVFPPTATNASYVLRKYYFSLLNNYEQIYFFRSNGQI-PPDSMQSPSARP 139
Query: 141 CFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAK 200
CF +QG I E +++ + + +P + E+ + SNV
Sbjct: 140 CF-----IQGA---IRPSQELQAL-------TFTPQPKINTAEFL---GGSLAGSNV--- 178
Query: 201 GYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFH--PEETV 240
G I+ KF+ GYLV+V +GSE L+GV++ P+ TV
Sbjct: 179 ------VGVIDGKFESGYLVTVTIGSEQLKGVLYQLLPQNTV 214
>UniRef100_Q9LTT3 High mobility group protein-like [Arabidopsis thaliana]
Length = 319
Score = 112 bits (279), Expect = 1e-23
Identities = 73/228 (32%), Positives = 114/228 (49%), Gaps = 49/228 (21%)
Query: 23 ASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEK 82
A ++ +V ++ LF + LR F +P +GG LDLH L++EVT R G E+VV ++
Sbjct: 34 AKYDDLVRNSALFWEKLRAFLGLTSKTLKVPTVGGNTLDLHRLFIEVTSRGGIERVVKDR 93
Query: 83 KWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPPAGTVLNSFTLCF 142
KW+EV F+F TT TSAS+VL+K+Y L+ E V++ ++ P++
Sbjct: 94 KWKEVIGAFSFPTTITSASFVLRKYYLKFLFQLEHVYY--LEKPVS-------------- 137
Query: 143 MEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEYAPKHANNGPESNVEAKGY 202
SLQ S+ LA P+ + P+ E +G+
Sbjct: 138 ----SLQ---------------------STDEALKSLANESPNPEEGIDEPQVGYEVQGF 172
Query: 203 PRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPSSRVVGT 250
I+ KFD GYLV++KLGS+ L+GV++H +T P S + + T
Sbjct: 173 -------IDGKFDSGYLVTMKLGSQELKGVLYHIPQT-PSQSQQTMET 212
>UniRef100_Q6K7A1 Glutathione S-transferase GST16-like protein [Oryza sativa]
Length = 467
Score = 112 bits (279), Expect = 1e-23
Identities = 76/241 (31%), Positives = 122/241 (50%), Gaps = 42/241 (17%)
Query: 10 SGGDDGK-----FYPPPLASHEVIVNDAKLFLDTLRQFHFHMGSKYMIPVIGGRKLDLHT 64
SGG G YP +A ++ +V DA +F L H MG+K +P+IGG+ LDLH
Sbjct: 51 SGGSGGGRRTLVAYPARVAGYKDVVADAAVFRRALEGLHAQMGTKLKVPIIGGKDLDLHQ 110
Query: 65 LYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFEQVHFFKVQ 124
L+ EVT R G +KV ++ +WREV + F F T T+AS++LKK+Y +LLY+FE+++ F+ Q
Sbjct: 111 LFKEVTSRGGIDKVKSDNRWREVTASFIFPATATNASFMLKKYYMSLLYHFERLYLFEAQ 170
Query: 125 GPITPPAGTVLNSFTLCFMEMDSLQGMYPNIHAVTEFESICDVLSGSSSSGKPDLALVEY 184
G + E DS S ++ + +S K
Sbjct: 171 G---------------WYQETDS------------RSISCIEMKAEGQASRK-------- 195
Query: 185 APKHANNGPESNVEAKGYPRYGEGRIEEKFDCGYLVSVKLGSEVLRGVIFHPEETVPPPS 244
K +N S++ A + I+ KF+ GY+V+V +GS+ + V+++ E P+
Sbjct: 196 -RKRGSNSCSSDL-AASLDNDVQVIIDGKFEHGYIVTVIMGSKSTKAVLYNCTEEPAVPT 253
Query: 245 S 245
+
Sbjct: 254 A 254
>UniRef100_UPI00003A9660 UPI00003A9660 UniRef100 entry
Length = 141
Score = 84.7 bits (208), Expect = 2e-15
Identities = 40/88 (45%), Positives = 57/88 (64%), Gaps = 1/88 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IPV+GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPVVGGKELDLHALYTRVTTLGGFGKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFF 121
+ ++A++ LK++Y L +E+VH F
Sbjct: 79 PRSCSNAAFALKQYYLRYLEKYEKVHHF 106
>UniRef100_Q9D982 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:1700124K17 product:hypothetical AT-rich
interaction domain (ARID) containing protein, full
insert sequence [Mus musculus]
Length = 145
Score = 84.7 bits (208), Expect = 2e-15
Identities = 41/97 (42%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IP +GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP 130
+ ++A++ LK++Y L +E+VH F PP
Sbjct: 79 PRSCSNAAFALKQYYLRYLEKYEKVHHFGEDDDEVPP 115
>UniRef100_Q645I3 ARID2 [Homo sapiens]
Length = 209
Score = 84.7 bits (208), Expect = 2e-15
Identities = 41/97 (42%), Positives = 58/97 (59%), Gaps = 1/97 (1%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IP +GG++LDLH LY VT G+ KV + +W E+ FNF
Sbjct: 19 FLDELRQFHHSRGSPFKKIPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP 130
+ ++A++ LK++Y L +E+VH F PP
Sbjct: 79 PRSCSNAAFALKQYYLRYLEKYEKVHHFGEDDDEVPP 115
>UniRef100_UPI0000360A2D UPI0000360A2D UniRef100 entry
Length = 140
Score = 82.8 bits (203), Expect = 8e-15
Identities = 44/113 (38%), Positives = 65/113 (56%), Gaps = 9/113 (7%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
FLD LRQFH GS + IP +GG++LDL+ LY+ V G+ KV + +W E+G FNF
Sbjct: 19 FLDELRQFHQSRGSPFRKIPFVGGKELDLNALYIRVVSLGGFAKVSEKNQWMELGEEFNF 78
Query: 94 STTTTSASYVLKKHYWNLLYNFEQVHFF----KVQGPITP----PAGTVLNSF 138
++A++ LK++Y L +E+VH F + P P P G + NS+
Sbjct: 79 PRNCSNAAFALKQYYLRYLEKYEKVHHFGEDDEESQPGNPKASLPVGAIPNSY 131
>UniRef100_Q68CP9 Hypothetical protein DKFZp779P0222 [Homo sapiens]
Length = 1756
Score = 69.7 bits (169), Expect = 7e-11
Identities = 31/79 (39%), Positives = 47/79 (59%)
Query: 52 IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNL 111
IP +GG++LDLH LY VT G+ KV + +W E+ FNF + ++A++ LK++Y
Sbjct: 10 IPAVGGKELDLHGLYTRVTTLGGFAKVSEKNQWGEIVEEFNFPRSCSNAAFALKQYYLRY 69
Query: 112 LYNFEQVHFFKVQGPITPP 130
L +E+VH F PP
Sbjct: 70 LEKYEKVHHFGEDDDEVPP 88
>UniRef100_UPI00002CEE47 UPI00002CEE47 UniRef100 entry
Length = 301
Score = 65.9 bits (159), Expect = 1e-09
Identities = 37/98 (37%), Positives = 52/98 (52%), Gaps = 4/98 (4%)
Query: 35 FLDTLRQFHFHMGSKYM-IPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNF 93
F+ L +F GS +P + GR+LDLH LY++V +R GYEK +KW ++ N
Sbjct: 130 FMKALTEFCDLKGSPLTKLPAVSGRELDLHLLYIQVRKRGGYEKACDTRKWSDIAEAMNM 189
Query: 94 -STTTTSASYVLKKHYWNLLYNFEQVHFFKVQGPITPP 130
T T + + LKKHY LL +E+ K P PP
Sbjct: 190 HDTDTKTLAAALKKHYQQLLLPYERCE--KGLDPPPPP 225
>UniRef100_Q5XGD9 Hypothetical protein [Xenopus tropicalis]
Length = 541
Score = 60.5 bits (145), Expect = 4e-08
Identities = 33/85 (38%), Positives = 48/85 (55%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL+ LYV VT + G +V+ +K WRE+
Sbjct: 216 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLYMLYVLVTEKGGLVEVINKKLWREITKGL 275
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 276 NLPTSITSAAFTLRTQYMKYLYPYE 300
>UniRef100_Q6GQD7 MGC80148 protein [Xenopus laevis]
Length = 539
Score = 60.5 bits (145), Expect = 4e-08
Identities = 33/85 (38%), Positives = 48/85 (55%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL+ LYV VT + G +V+ +K WRE+
Sbjct: 213 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLYMLYVLVTEKGGLVEVINKKLWREITKGL 272
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 273 NLPTSITSAAFTLRTQYMKYLYPYE 297
>UniRef100_UPI0000365B23 UPI0000365B23 UniRef100 entry
Length = 502
Score = 60.1 bits (144), Expect = 5e-08
Identities = 41/118 (34%), Positives = 60/118 (50%), Gaps = 4/118 (3%)
Query: 3 SAARTIRSGGDDGKF-YPPPLASHEVIVNDAKL--FLDTLRQFHFHMGSKY-MIPVIGGR 58
+A + + S GD + Y L + ND K FLD L F G+ IP++ +
Sbjct: 149 AAQQVLTSPGDFADWGYDDQLKQLYELDNDPKRKEFLDELFVFMQKRGTPVNRIPIMAKQ 208
Query: 59 KLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVFNFSTTTTSASYVLKKHYWNLLYNFE 116
LDL+ LY VT + G +V+ +K WRE+ N T+ TSA++ L+ Y LY FE
Sbjct: 209 VLDLYKLYALVTEKGGLVEVINKKIWREITKGLNLPTSITSAAFTLRTQYMKYLYPFE 266
>UniRef100_Q80YP8 Dead ringer homolog 1 [Mus musculus]
Length = 599
Score = 59.3 bits (142), Expect = 9e-08
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 247 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 306
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 307 NLPTSITSAAFTLRTQYMKYLYPYE 331
>UniRef100_Q6P9C6 AT rich interactive domain 3A (BRIGHT-like) protein [Homo sapiens]
Length = 593
Score = 59.3 bits (142), Expect = 9e-08
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMEYLYPYE 326
>UniRef100_Q8N4Z3 ARID3A protein [Homo sapiens]
Length = 443
Score = 59.3 bits (142), Expect = 9e-08
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 92 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 151
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 152 NLPTSITSAAFTLRTQYMKYLYPYE 176
>UniRef100_Q62431 AT-rich interactive domain-containing protein 3A [Mus musculus]
Length = 601
Score = 59.3 bits (142), Expect = 9e-08
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 247 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 306
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 307 NLPTSITSAAFTLRTQYMKYLYPYE 331
>UniRef100_Q99856 AT-rich interactive domain-containing protein 3A [Homo sapiens]
Length = 593
Score = 59.3 bits (142), Expect = 9e-08
Identities = 33/85 (38%), Positives = 47/85 (54%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL LYV VT + G +V+ +K WRE+
Sbjct: 242 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGL 301
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 302 NLPTSITSAAFTLRTQYMKYLYPYE 326
>UniRef100_UPI00003AFFE8 UPI00003AFFE8 UniRef100 entry
Length = 284
Score = 58.9 bits (141), Expect = 1e-07
Identities = 33/85 (38%), Positives = 48/85 (55%), Gaps = 1/85 (1%)
Query: 33 KLFLDTLRQFHFHMGSKY-MIPVIGGRKLDLHTLYVEVTRRSGYEKVVAEKKWREVGSVF 91
K FLD L F G+ IP++ + LDL+TLY VT + G +V+ +K WRE+
Sbjct: 12 KEFLDDLFSFMQKRGTPVNRIPIMAKQVLDLYTLYRLVTDKGGLVEVINKKIWREITKGL 71
Query: 92 NFSTTTTSASYVLKKHYWNLLYNFE 116
N T+ TSA++ L+ Y LY +E
Sbjct: 72 NLPTSITSAAFTLRTQYMKYLYPYE 96
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.137 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 463,387,418
Number of Sequences: 2790947
Number of extensions: 19408811
Number of successful extensions: 37400
Number of sequences better than 10.0: 305
Number of HSP's better than 10.0 without gapping: 267
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 37050
Number of HSP's gapped (non-prelim): 316
length of query: 253
length of database: 848,049,833
effective HSP length: 124
effective length of query: 129
effective length of database: 501,972,405
effective search space: 64754440245
effective search space used: 64754440245
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0121b.2


