

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0118c.6
(278 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
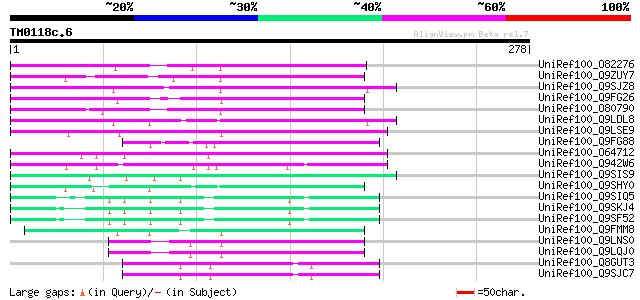
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 96 1e-18
UniRef100_Q9ZUY7 Putative reverse transcriptase [Arabidopsis tha... 93 9e-18
UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcrip... 92 2e-17
UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like... 87 4e-16
UniRef100_O80790 Putative non-LTR retroelement reverse transcrip... 85 2e-15
UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis tha... 79 1e-13
UniRef100_Q9LSE9 Reverse transcriptase-like protein [Arabidopsis... 74 3e-12
UniRef100_Q9FG88 Non-LTR retrolelement reverse transcriptase-lik... 73 9e-12
UniRef100_O64712 Putative reverse transcriptase [Arabidopsis tha... 71 4e-11
UniRef100_Q942W6 P0506E04.5 protein [Oryza sativa] 69 2e-10
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 67 5e-10
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 66 9e-10
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 66 1e-09
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 65 2e-09
UniRef100_Q9SF52 Putative non-LTR reverse transcriptase [Arabido... 65 2e-09
UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like... 64 4e-09
UniRef100_Q9LNS0 F1L3.4 [Arabidopsis thaliana] 63 7e-09
UniRef100_Q9LQJ0 F28G4.15 protein [Arabidopsis thaliana] 63 7e-09
UniRef100_Q8GUT3 Hypothetical protein T1O3.17 [Arabidopsis thali... 63 1e-08
UniRef100_Q9SJC7 Putative non-LTR retrolelement reverse transcri... 62 2e-08
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 95.5 bits (236), Expect = 1e-18
Identities = 67/207 (32%), Positives = 92/207 (44%), Gaps = 25/207 (12%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN-FGVGDHGSWIRSQARSRNSV---R 56
C C+ EE ILH LRDCP + IW RL L + F W+ + +
Sbjct: 944 CSVCNGAEETILHVLRDCPAMEPIWRRLLPLRRHHEFFSQSLLEWLFTNMDPVKGIWPTL 1003
Query: 57 FLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFIT--------FSNGVAGDHGSW 108
F G+W WKWR VF + R+IC + +FI G G+ +
Sbjct: 1004 FGMGIWWAWKWRCCDVFGE---------RKICRDRLKFIKDMAEEVRRVHVGAVGNRPNG 1054
Query: 109 LSS----RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALI 164
+ RWQ P G VK+ D + AGG IR+ QG WLGGF + A +
Sbjct: 1055 VRVERMIRWQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAPL 1114
Query: 165 AEATALLLGLELVWDLGYRQVMVEVDC 191
AE GL + WD G+R+V +++DC
Sbjct: 1115 AELWGAYYGLLIAWDKGFRRVELDLDC 1141
>UniRef100_Q9ZUY7 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 314
Score = 92.8 bits (229), Expect = 9e-18
Identities = 61/211 (28%), Positives = 89/211 (41%), Gaps = 30/211 (14%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRL------------GALAWTNFGVGDHGSWIRSQ 48
C C E+ I+H LRDCP + IW+RL L W +GD R
Sbjct: 22 CQICKGAEKTIIHILRDCPAMEGIWIRLVPAGKRREFFTQSLLEWLFANLGDR----RKT 77
Query: 49 ARSRNSVRFLAGVWGVWKWRNNMVFEDSPWLVDEACRR-------ICHEHDEFITFSNGV 101
S S F +W WKWR +F V + CR + E +
Sbjct: 78 CESTWSTLFALSIWWAWKWRCGNIFG-----VQDKCRDRVRFLKDLARETSMAHVIVRTL 132
Query: 102 AGDHGSWLSS--RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSG 159
+G HG + W P++G KLN D + AGG++RD++G W GGF +
Sbjct: 133 SGGHGERVERLIAWSKPEEGWWKLNTDGASRGNPGLASAGGVLRDEEGAWRGGFALNIGV 192
Query: 160 GNALIAEATALLLGLELVWDLGYRQVMVEVD 190
+A +AE + GL + W+ ++ +EVD
Sbjct: 193 CSAPLAELWGVYYGLYIAWERRVTRLEIEVD 223
>UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 321
Score = 91.7 bits (226), Expect = 2e-17
Identities = 70/219 (31%), Positives = 92/219 (41%), Gaps = 14/219 (6%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTN-FGVGDHGSWIRSQARSRNS--VRF 57
C C EE ILH LRDCP IW RL F WI R R S F
Sbjct: 35 CQICQGGEETILHVLRDCPAMAGIWSRLVPRDQIRQFFTASLLEWIYKNLRERGSWPTVF 94
Query: 58 LAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDHGSWLSS-----R 112
+ VW WKWR +F + D + + E + +N + +S
Sbjct: 95 VMAVWWGWKWRCGNIFGGNGKCRDRV--KFIKDLAEEVAIANAFVKGNEVRVSRVERLVS 152
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLL 172
W P+ G VKLN D + AGG++RD G W+GGF + +A +AE +
Sbjct: 153 WVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGGFAVNIGVCSAPLAELWGVYY 212
Query: 173 GLELVWDLGYRQVMVEVD----CGELLQVMADEESCRFL 207
GL + W G R+V +EVD G L +AD FL
Sbjct: 213 GLFIAWGRGARRVELEVDSKMVVGFLTTGIADSHPLSFL 251
>UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 87.4 bits (215), Expect = 4e-16
Identities = 61/212 (28%), Positives = 89/212 (41%), Gaps = 33/212 (15%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRL-GALAWTNFGVGDHGSWIRSQARSRNSVR--- 56
CP C E ++H LRDCP IW+R+ + F W+ + R+
Sbjct: 385 CPLCKGASESLIHVLRDCPAMMGIWMRVVPVMEQRRFFETSLLEWMYGNLKERSDSERRS 444
Query: 57 ----FLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDHGSWLSSR 112
F VW WKWR VF + D CR D + VA + L++
Sbjct: 445 WPTLFALTVWWGWKWRCGYVFGE-----DSRCR------DRVKFLKSAVAEVEAAHLAAN 493
Query: 113 --------------WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRS 158
W+ P +G V +N D + AGG+IRD+ G WL GF +
Sbjct: 494 GDAREDVLVERMIAWRKPAEGWVTMNTDGASHGNPGQATAGGVIRDEHGSWLVGFALNIG 553
Query: 159 GGNALIAEATALLLGLELVWDLGYRQVMVEVD 190
+A +AE + GL + W+ G+R+V +EVD
Sbjct: 554 VCSAPLAELWGVYYGLVVAWERGWRRVRLEVD 585
>UniRef100_O80790 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 970
Score = 85.1 bits (209), Expect = 2e-15
Identities = 62/205 (30%), Positives = 87/205 (42%), Gaps = 25/205 (12%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQ---ARSRNSVRF 57
C C+ +E ILH LRDCP IW RL N W+ + A+ F
Sbjct: 685 CSVCNGADESILHVLRDCPAMTPIWQRLLPQRRQNEFFSQF-EWLFTNLDPAKGDWPTLF 743
Query: 58 LAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFIT-FSNGVAGDHGSWLSS----- 111
G+W WKWR VF + R++C + +FI + V H L++
Sbjct: 744 SMGIWWAWKWRCGDVFGE---------RKLCRDRLKFIKDIAEEVRKAHVGTLNNHVKRA 794
Query: 112 ------RWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIA 165
RW+ P VKL D + A G I + QG WLGGF + +A +A
Sbjct: 795 RVERMIRWKAPSDRWVKLTTDGASRGHQGLAAASGAILNLQGEWLGGFALNIGSCDAPLA 854
Query: 166 EATALLLGLELVWDLGYRQVMVEVD 190
E GL + WD G+R+V + +D
Sbjct: 855 ELWGAYYGLLIAWDKGFRRVELNLD 879
>UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis thaliana]
Length = 947
Score = 79.3 bits (194), Expect = 1e-13
Identities = 67/225 (29%), Positives = 92/225 (40%), Gaps = 21/225 (9%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWT-NFGVGDHGSWIRSQARSRNS----- 54
C C +E ILH L+DCP IW RL + + +F G W+ +N+
Sbjct: 718 CQVCKGGDETILHVLKDCPSIAGIWRRLVQVQRSYDFFNGSLFGWLYVNLGMKNAETGYA 777
Query: 55 --VRFLAGVWGVWKWRNNMVF------EDSPWLVDEACRRICHEHDEFITFSNGVAGDHG 106
F VW WKWR VF D + + H H I NG
Sbjct: 778 WATLFAIVVWWSWKWRCGYVFGEVGKCRDRVKFFRDLAAEVSHAHA--IHSQNGGLRTRV 835
Query: 107 SWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAE 166
L + W+PP VKLN D + GG++RD G W GGF +A +AE
Sbjct: 836 ERLVA-WKPPDGEWVKLNTDGASRGNLGLATTGGVLRDGIGHWCGGFALDIGVCSAPLAE 894
Query: 167 ATALLLGLELVWDLGYRQVMVEVD----CGELLQVMADEESCRFL 207
+ GL + W+ + +V +EVD G L ++D S FL
Sbjct: 895 LWGVYYGLYMAWERRFTRVELEVDSELVVGFLTTGISDTHSLSFL 939
>UniRef100_Q9LSE9 Reverse transcriptase-like protein [Arabidopsis thaliana]
Length = 343
Score = 74.3 bits (181), Expect = 3e-12
Identities = 57/221 (25%), Positives = 99/221 (44%), Gaps = 19/221 (8%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGA----LAWTNFGVGDHGSWIRSQARSRNSVR 56
C RC +E H DC ++Q++W G L T + + S + +
Sbjct: 56 CHRCCQEDETSQHLFFDCFYAQQVWRASGIPHQELRTTGITMETKMELLLSSCLANRQPQ 115
Query: 57 F----LAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDHGSWLSSR 112
+ +W +WK RN +VF+ +R ++ E+ + V + SSR
Sbjct: 116 LFNLAIWILWRLWKSRNQLVFQQKSISWQNTLQRARNDVQEWEDTNTYVQSLNQQVHSSR 175
Query: 113 ----------WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRS-GGN 161
WQ P +K N D +F Q R AG L+RD+ G+++G A S +
Sbjct: 176 HQQPTMARTKWQRPPSTWIKYNYDGAFNHQTRNAKAGWLMRDENGVYMGSGQAIGSTTSD 235
Query: 162 ALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMADEE 202
+L +E AL++ ++ W GYR+V+ E D ++ ++M +E+
Sbjct: 236 SLESEFQALIIAMQHAWSQGYRKVIFEGDSKQVEELMNNEK 276
>UniRef100_Q9FG88 Non-LTR retrolelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 228
Score = 72.8 bits (177), Expect = 9e-12
Identities = 50/150 (33%), Positives = 80/150 (53%), Gaps = 16/150 (10%)
Query: 61 VWGVWKWRNNMVFE--DSPWLVDEACRRICHEHDEFITFSNGVAGD-------HGSW--L 109
+W +WK RN +VF+ + W D R +E +E+IT NG+ +G+
Sbjct: 10 LWRIWKSRNKLVFQRKQNDWWRD--IRDATNEAEEWIT--NGLCAQVPTPSTTNGTTRQF 65
Query: 110 SSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYA-HRSGGNALIAEAT 168
S+WQ P G +K N D SF + R A +IRDD G++ G A + +A AE
Sbjct: 66 QSQWQKPHMGWIKCNYDGSFVNRVRGSTAAWIIRDDNGVFKGAAQATGATVSSAFEAECQ 125
Query: 169 ALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
+L+L ++ +W G+R+V++E DC L+ V+
Sbjct: 126 SLILTMQQLWIRGFRKVILEGDCKLLVNVL 155
>UniRef100_O64712 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 365
Score = 70.9 bits (172), Expect = 4e-11
Identities = 60/222 (27%), Positives = 99/222 (44%), Gaps = 20/222 (9%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFG----VGDHGSWI--RSQARSRNS 54
C RC EE I H + +CP++Q +W + +G D+ + + S+ ++ NS
Sbjct: 75 CQRCCIEEETIHHIMFNCPYTQSVWRSANIIIGNQWGPPSSFEDNLNRLIQLSKTQTTNS 134
Query: 55 V-RFLAG--VWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITFSNGVAGDH------ 105
+ RFL +W +WK RN +F+ D R+ + E++ + +
Sbjct: 135 LDRFLPFWIMWRLWKSRNVFLFQQKCQSPDYEARKGIQDATEWLNANETTENTNVHVATN 194
Query: 106 ----GSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQG-IWLGGFYAHRSGG 160
SS+W PP +G VK N D + + +G IR+ G I L G +S
Sbjct: 195 PIQTSRRDSSQWNPPPEGWVKCNFDSGYTQGSPYTRSGWTIRECNGHIVLCGNAKLQSST 254
Query: 161 NALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMADEE 202
+L AEA L L+++W G R V E D L+ ++ + E
Sbjct: 255 CSLHAEALGFLHALQVIWAHGLRYVWFESDSKSLVTLINNGE 296
>UniRef100_Q942W6 P0506E04.5 protein [Oryza sativa]
Length = 325
Score = 68.6 bits (166), Expect = 2e-10
Identities = 61/232 (26%), Positives = 98/232 (41%), Gaps = 33/232 (14%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLG---ALAWTNFGVGDHGSWI-----RSQARSR 52
C C EED+ H L CPH++ +W +G A++ WI R R
Sbjct: 15 CTICGTEEEDVAHALCRCPHAKYLWEAMGNTNAISCKPDSNWKDSDWILDISGRVSKEER 74
Query: 53 NSVRFLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFITF-----SNGVAGDH-- 105
+ L +W +W RN + + V+ + R I + +N + G H
Sbjct: 75 MGLMML--LWRIWYVRNGITHGKAAIPVEVSQRFIISYMASLLEIRQHPNANLIKGKHVV 132
Query: 106 --GSWL-----------SSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQG--IWL 150
GS + SS W P++G +KLN+D S++ D G G ++RD G I+
Sbjct: 133 QYGSSMPLPRQCKPSAESSSWIRPQEGWMKLNVDGSYYPSDGKGGTGAVLRDSSGNLIFA 192
Query: 151 GGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMADEE 202
HR +AL AE G+ + ++VE DC E++Q++ +E
Sbjct: 193 ACGVLHRP-ASALEAEMVDCREGISMALQWTLLPIIVETDCLEMVQLIHSDE 243
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 67.0 bits (162), Expect = 5e-10
Identities = 56/226 (24%), Positives = 90/226 (39%), Gaps = 19/226 (8%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDH---GSWIRSQARSRNSVRF 57
C RC +E I H + C ++Q +W D+ + Q + ++
Sbjct: 1426 CQRCCNADETINHIIFTCSYAQVVWRSANFSGSNRLCFTDNLEENIRLILQGKKNQNLPI 1485
Query: 58 LAGV------WGVWKWRNNMVFEDS---PWLVDEACRRICHE------HDEFITFSNGVA 102
L G+ W +WK RN +F+ PW V + + E +D I+ + +
Sbjct: 1486 LNGLMPFWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVETMVNDTAISHNTAQS 1545
Query: 103 GDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQG-IWLGGFYAHRSGGN 161
D S +W P +G +K N D + + G ++RD G + G + +
Sbjct: 1546 NDRPLSRSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNGRVLHSGCAKLQQSYS 1605
Query: 162 ALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVMADEESCRFL 207
AL AEA L L++VW GY V E D EL ++ E L
Sbjct: 1606 ALQAEALGFLHALQMVWIRGYCYVWFEGDNLELTNLINKTEDHHLL 1651
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 66.2 bits (160), Expect = 9e-10
Identities = 57/213 (26%), Positives = 77/213 (35%), Gaps = 35/213 (16%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRL------------GALAWTNFGVGDHGS----- 43
C C E +LH LRDCP IW+R+ W +GD
Sbjct: 438 CQVCKGGVESMLHVLRDCPAQLGIWVRVVPQRRQQGFFSKSLFEWLYDNLGDRSGCEDIP 497
Query: 44 WIRSQARSRNSVRFLAGVWGVWKWRNNMVF------EDSPWLVDEACRRICHEHDEFITF 97
W S F +W WKWR +F D V E + H +
Sbjct: 498 W---------STIFAVIIWWGWKWRCGNIFGENTKCRDRVKFVKEWAVEVYRAHSGNVLV 548
Query: 98 SNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHR 157
G+ + W P G VK+N D + AGG++RD G W GGF +
Sbjct: 549 --GITQPRVERMIG-WVSPCVGWVKVNTDGASRGNPGLASAGGVLRDCTGAWCGGFSLNI 605
Query: 158 SGGNALIAEATALLLGLELVWDLGYRQVMVEVD 190
+A AE + GL W+ +V +EVD
Sbjct: 606 GRCSAPQAELWGVYYGLYFAWEKKVPRVELEVD 638
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 65.9 bits (159), Expect = 1e-09
Identities = 65/232 (28%), Positives = 90/232 (38%), Gaps = 53/232 (22%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQARSR-------N 53
CPRC E I H L CP + W W + S IR+Q S N
Sbjct: 1240 CPRCHRENESINHALFTCPFATMAW-------WLS-----DSSLIRNQLMSNDFEENISN 1287
Query: 54 SVRFLAG--------------VWGVWKWRNNMVFE---DSPWLVDEACRRICHE------ 90
+ F+ +W +WK RNN+VF +SP + + H+
Sbjct: 1288 ILNFVQDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQ 1347
Query: 91 -HDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGI- 148
H + + + +A + W+ P VK N D F Q G +IR+ G
Sbjct: 1348 SHKKTPSPTRQIAEN-----KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTP 1402
Query: 149 --WLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
W AH S N L AE ALL L+ W GY QV +E DC L+ ++
Sbjct: 1403 ISWGSMKLAHTS--NPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLI 1452
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 65.1 bits (157), Expect = 2e-09
Identities = 66/232 (28%), Positives = 90/232 (38%), Gaps = 53/232 (22%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQARSR-------N 53
CPRC E I H L CP + W RL S IR+Q S N
Sbjct: 1466 CPRCHRENESINHALFTCPFATMAW-RLS-----------DSSLIRNQLMSNDFEENISN 1513
Query: 54 SVRFLAG--------------VWGVWKWRNNMVFE---DSPWLVDEACRRICHE------ 90
+ F+ +W +WK RNN+VF +SP + + H+
Sbjct: 1514 ILNFVQDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQ 1573
Query: 91 -HDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGI- 148
H + + + +A + W+ P VK N D F Q G +IR+ G
Sbjct: 1574 SHKKTPSPTRQIAEN-----KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTP 1628
Query: 149 --WLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
W AH S N L AE ALL L+ W GY QV +E DC L+ ++
Sbjct: 1629 ISWGSMKLAHTS--NPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLI 1678
>UniRef100_Q9SF52 Putative non-LTR reverse transcriptase [Arabidopsis thaliana]
Length = 484
Score = 65.1 bits (157), Expect = 2e-09
Identities = 66/232 (28%), Positives = 90/232 (38%), Gaps = 53/232 (22%)
Query: 1 CPRCSAPEEDILHCLRDCPHSQEIWLRLGALAWTNFGVGDHGSWIRSQARSR-------N 53
CPRC E I H L CP + W RL S IR+Q S N
Sbjct: 200 CPRCHRENESINHALFTCPFATMAW-RLS-----------DSSLIRNQLMSNDFEENISN 247
Query: 54 SVRFLAG--------------VWGVWKWRNNMVFE---DSPWLVDEACRRICHE------ 90
+ F+ +W +WK RNN+VF +SP + + H+
Sbjct: 248 ILNFVQDTTMSDFHKLLPVWLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNATQ 307
Query: 91 -HDEFITFSNGVAGDHGSWLSSRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGI- 148
H + + + +A + W+ P VK N D F Q G +IR+ G
Sbjct: 308 SHKKTPSPTRQIAEN-----KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTP 362
Query: 149 --WLGGFYAHRSGGNALIAEATALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
W AH S N L AE ALL L+ W GY QV +E DC L+ ++
Sbjct: 363 ISWGSMKLAHTS--NPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLI 412
>UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 308
Score = 63.9 bits (154), Expect = 4e-09
Identities = 54/198 (27%), Positives = 76/198 (38%), Gaps = 21/198 (10%)
Query: 9 EDILHCLRDCPHSQEIWLR-LGALAWTNFGVGDHGSWIRSQARSRNSVR-------FLAG 60
E +LH RDCP IW+R + F W+ R+S F
Sbjct: 25 ESVLHVFRDCPAQLGIWVRFVPRRRQQGFFSKSLFEWLYDNLCDRSSCEDIPWSTIFAVI 84
Query: 61 VWGVWKWRNNMVF------EDSPWLVDEACRRICHEHDEFITFSNGVAGDHGSWLSSR-- 112
+W WKWR + +F D V E + H N + G +
Sbjct: 85 IWWGWKWRCSNIFGENTKCRDRVKFVKEWVVEVYRAH-----LGNALVGSTQPRVERLIG 139
Query: 113 WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGNALIAEATALLL 172
W P G VK+N D + AGG++RD +G W GGF + +A AE +
Sbjct: 140 WVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGFSLNIGRCSAQHAELWGVYY 199
Query: 173 GLELVWDLGYRQVMVEVD 190
GL W+ +V +EVD
Sbjct: 200 GLYFAWEKKVPRVELEVD 217
>UniRef100_Q9LNS0 F1L3.4 [Arabidopsis thaliana]
Length = 253
Score = 63.2 bits (152), Expect = 7e-09
Identities = 42/149 (28%), Positives = 64/149 (42%), Gaps = 21/149 (14%)
Query: 54 SVRFLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFI---TFSNGVAGDHGSWLS 110
+V F +W WKWR +F ++ + C + FI A H ++
Sbjct: 23 AVTFSQAIWWGWKWRYGNIFGEN---------KKCRDRVRFIKDRALDVWKAHVHKMGVT 73
Query: 111 SR---------WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGN 161
+R W PP+ G KLN D + R AGG++RD G W GF + +
Sbjct: 74 TRTAREERLIAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICS 133
Query: 162 ALIAEATALLLGLELVWDLGYRQVMVEVD 190
A +AE GL + W+ G Q+ +E+D
Sbjct: 134 APLAELWGAYYGLNIAWERGVTQLEMEID 162
>UniRef100_Q9LQJ0 F28G4.15 protein [Arabidopsis thaliana]
Length = 272
Score = 63.2 bits (152), Expect = 7e-09
Identities = 42/149 (28%), Positives = 64/149 (42%), Gaps = 21/149 (14%)
Query: 54 SVRFLAGVWGVWKWRNNMVFEDSPWLVDEACRRICHEHDEFI---TFSNGVAGDHGSWLS 110
+V F +W WKWR +F ++ + C + FI A H ++
Sbjct: 42 AVTFSQAIWWGWKWRYGNIFGEN---------KKCRDRVRFIKDRALDVWKAHVHKMGVT 92
Query: 111 SR---------WQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGGN 161
+R W PP+ G KLN D + R AGG++RD G W GF + +
Sbjct: 93 TRTAREERLIAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICS 152
Query: 162 ALIAEATALLLGLELVWDLGYRQVMVEVD 190
A +AE GL + W+ G Q+ +E+D
Sbjct: 153 APLAELWGAYYGLNIAWERGVTQLEMEID 181
>UniRef100_Q8GUT3 Hypothetical protein T1O3.17 [Arabidopsis thaliana]
Length = 221
Score = 62.8 bits (151), Expect = 1e-08
Identities = 43/151 (28%), Positives = 71/151 (46%), Gaps = 15/151 (9%)
Query: 61 VWGVWKWRNNMVFEDSPWLVDEACRRICHE------HDEFITFSNGVAGDHG----SWLS 110
+W +W+ RN +VF+ +R + E+I N G
Sbjct: 1 MWRLWRSRNQLVFQHQNLSWQSTLKRAKDDVQEWENAQEYIQSLNHYTVREGCNNRESTH 60
Query: 111 SRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGG---NALIAEA 167
+W+ P G +K N D SF + + +G LIRDD+G + G A GG NAL +E
Sbjct: 61 QKWEQPPMGWIKCNYDGSFTYRTQQTNSGWLIRDDKGFYKGA--AQAVGGTMNNALESEL 118
Query: 168 TALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
AL++ ++ W GYR+V+ E D ++ +++
Sbjct: 119 QALVMAMQHTWSQGYRKVIFEGDSKQVEELL 149
>UniRef100_Q9SJC7 Putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 221
Score = 62.0 bits (149), Expect = 2e-08
Identities = 43/151 (28%), Positives = 71/151 (46%), Gaps = 15/151 (9%)
Query: 61 VWGVWKWRNNMVFEDSPWLVDEACRRICHE------HDEFITFSNGVAGDHG----SWLS 110
+W +W+ RN +VF+ +R + E+I N G
Sbjct: 1 MWRLWRSRNQLVFQHQNLSWQSTLKRAKDDVQEWENAQEYIQSLNHYTVREGCNNRESTH 60
Query: 111 SRWQPPKQGVVKLNMDESFWEQDRCMGAGGLIRDDQGIWLGGFYAHRSGG---NALIAEA 167
+W+ P G +K N D SF + + +G LIRDD+G + G A GG NAL +E
Sbjct: 61 QKWEQPPMGWIKCNYDGSFNYRTQQTNSGWLIRDDKGFYKGA--AQAVGGTMNNALESEL 118
Query: 168 TALLLGLELVWDLGYRQVMVEVDCGELLQVM 198
AL++ ++ W GYR+V+ E D ++ +++
Sbjct: 119 QALVMAMQHTWSQGYRKVIFEGDSKQVEELL 149
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.333 0.143 0.509
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 481,472,172
Number of Sequences: 2790947
Number of extensions: 18963982
Number of successful extensions: 39386
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 48
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 39198
Number of HSP's gapped (non-prelim): 138
length of query: 278
length of database: 848,049,833
effective HSP length: 126
effective length of query: 152
effective length of database: 496,390,511
effective search space: 75451357672
effective search space used: 75451357672
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0118c.6


