

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0117b.14
(458 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
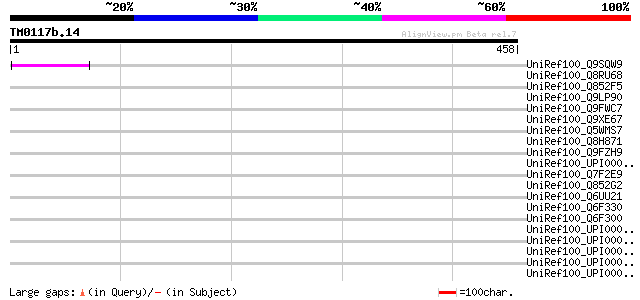
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidop... 57 1e-06
UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type... 46 0.002
UniRef100_Q852F5 Putative polyprotein [Oryza sativa] 45 0.006
UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana] 44 0.013
UniRef100_Q9FWC7 Putative plant disease resistance polyprotein [... 44 0.013
UniRef100_Q9XE67 Hypothetical protein [Sorghum bicolor] 43 0.021
UniRef100_Q5WMS7 Putative polyprotein [Oryza sativa] 42 0.028
UniRef100_Q8H871 Putative polyprotein [Oryza sativa] 42 0.037
UniRef100_Q9FZH9 F1O19.7 protein [Arabidopsis thaliana] 41 0.081
UniRef100_UPI000032FFF4 UPI000032FFF4 UniRef100 entry 39 0.24
UniRef100_Q7F2E9 Putative polyprotein [Oryza sativa] 39 0.24
UniRef100_Q852G2 Putative polyprotein [Oryza sativa] 39 0.24
UniRef100_Q6UU21 Putative polyprotein [Oryza sativa] 39 0.24
UniRef100_Q6F330 Putative polyprotein [Oryza sativa] 39 0.24
UniRef100_Q6F300 Putative polyprotein [Oryza sativa] 39 0.31
UniRef100_UPI00003B00C5 UPI00003B00C5 UniRef100 entry 39 0.40
UniRef100_UPI00003B00C4 UPI00003B00C4 UniRef100 entry 39 0.40
UniRef100_UPI00003B00C3 UPI00003B00C3 UniRef100 entry 39 0.40
UniRef100_UPI00003B00C2 UPI00003B00C2 UniRef100 entry 39 0.40
UniRef100_UPI00003B00C1 UPI00003B00C1 UniRef100 entry 39 0.40
>UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 57.0 bits (136), Expect = 1e-06
Identities = 27/71 (38%), Positives = 36/71 (50%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
+P ++G A WL +EQ + EEEK A LTG + WW C + R Q TW E
Sbjct: 233 YPAYEGGNADDWLFRLEQCFLSNRTLEEEKLEKAVSCLTGASVTWWRCSKDREQIYTWRE 292
Query: 62 FVEALLRKFEP 72
F E + +F P
Sbjct: 293 FQEKFMLRFRP 303
>UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 46.2 bits (108), Expect = 0.002
Identities = 34/109 (31%), Positives = 49/109 (44%), Gaps = 18/109 (16%)
Query: 3 PEFDGR----EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQK-- 56
P F G EA W++ +E+ EA G +++EK A L AF WW ++ +
Sbjct: 80 PTFSGTANPLEAEEWIVAMEKSFEAMGCTDKEKIIYATYMLQSSAFEWWDAHKKSYSERI 139
Query: 57 -ATWWEFVEALLRKFEPELEPYMPEPVQDSEEEEI----PGKQEVLEAE 100
TW F EA +K Y PE V+ +E+E G + V E E
Sbjct: 140 FITWELFKEAFYKK-------YFPESVKRMKEKEFLELKQGNKSVAEYE 181
>UniRef100_Q852F5 Putative polyprotein [Oryza sativa]
Length = 1155
Score = 44.7 bits (104), Expect = 0.006
Identities = 22/74 (29%), Positives = 39/74 (51%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
FP FDG + WL M E + + +++ K S A ++G A W+ ++ N+ +W +
Sbjct: 124 FPRFDGSDVRIWLDMCETYFDMYQITQNFKVSAAVLHMSGNAAQWYHSYKLVNEVNSWDQ 183
Query: 62 FVEALLRKFEPELE 75
F A+ +FE +E
Sbjct: 184 FRMAVATEFECVVE 197
>UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 43.5 bits (101), Expect = 0.013
Identities = 20/68 (29%), Positives = 34/68 (49%)
Query: 3 PEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEF 62
P FDG YGW+ VE+ + G +E E+ + +++GEA W+ R +W +
Sbjct: 133 PMFDGSGIYGWIARVERFFRSGGYNEAEQLALVSVSVSGEALSWYNWAISRGDFVSWLKL 192
Query: 63 VEALLRKF 70
L+ +F
Sbjct: 193 KSGLMLRF 200
>UniRef100_Q9FWC7 Putative plant disease resistance polyprotein [Oryza sativa]
Length = 894
Score = 43.5 bits (101), Expect = 0.013
Identities = 21/74 (28%), Positives = 38/74 (50%)
Query: 2 FPEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWE 61
FP FDG + WL M E + + +++ K S ++G A W+ ++ N+ +W +
Sbjct: 124 FPRFDGSDVRIWLNMCETYFDMYQITQNFKVSAVVLHMSGNAAQWYHSYKLVNEVNSWDQ 183
Query: 62 FVEALLRKFEPELE 75
F A+ +FE +E
Sbjct: 184 FRMAVATEFEGVVE 197
>UniRef100_Q9XE67 Hypothetical protein [Sorghum bicolor]
Length = 416
Score = 42.7 bits (99), Expect = 0.021
Identities = 46/189 (24%), Positives = 80/189 (41%), Gaps = 31/189 (16%)
Query: 10 AYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQK---ATWWEFVEAL 66
A WL +E+ + ++EE A LTG A WW + ++ TW EF EA
Sbjct: 132 AEDWLREIEKKLDLTTCTDEECVGVAAHQLTGAARAWWDSYSDTHENPGGITWAEFTEAF 191
Query: 67 LRKFEPELEPYMPEPVQDSEEEEI----PGKQEVLE-AERRTVVVEAKPEANSVLLAKSE 121
E ++PE V D++ EE G Q+V E A R T + P+ ++
Sbjct: 192 -------REQHVPEGVMDAKVEEFRNISQGTQKVQEYATRFTRTMRYAPDESNT------ 238
Query: 122 IKPYSPENSEMEASRKDSEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKIDAEGEIK 181
E +M +K ++ +S T+ + I+ L + ++++A+ + K
Sbjct: 239 ------EKKKMYFFKK----GLSTRLKVALSGHTCYTLREMINKALEMERDRLEADAQYK 288
Query: 182 IKRLQEGSS 190
K+ + SS
Sbjct: 289 EKKRRSESS 297
>UniRef100_Q5WMS7 Putative polyprotein [Oryza sativa]
Length = 1131
Score = 42.4 bits (98), Expect = 0.028
Identities = 24/67 (35%), Positives = 32/67 (46%), Gaps = 3/67 (4%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRR---NQKATWWEFVEA 65
EA WL VE+ + +E+EK S A L G A WW +R+ + TWWEF A
Sbjct: 73 EAGDWLHAVEKKLDLIQCTEQEKVSFASHQLHGPAAEWWDHFRQNRAAGEPITWWEFTAA 132
Query: 66 LLRKFEP 72
+ P
Sbjct: 133 FKKTHIP 139
>UniRef100_Q8H871 Putative polyprotein [Oryza sativa]
Length = 1802
Score = 42.0 bits (97), Expect = 0.037
Identities = 28/89 (31%), Positives = 39/89 (43%), Gaps = 8/89 (8%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRR---RNQKATWWEFVEA 65
+A WL V + + S+EEK A L G A +WW ++ Q TW F A
Sbjct: 419 DALDWLHAVGKKLDTVQCSDEEKVIFAAHQLQGPASLWWDHFQATQPEGQPITWARFTAA 478
Query: 66 LLRKFEP-----ELEPYMPEPVQDSEEEE 89
R P L Y PE V++ EE++
Sbjct: 479 FRRTHVPAGVFNNLARYAPEDVREDEEKQ 507
>UniRef100_Q9FZH9 F1O19.7 protein [Arabidopsis thaliana]
Length = 659
Score = 40.8 bits (94), Expect = 0.081
Identities = 30/125 (24%), Positives = 49/125 (39%), Gaps = 7/125 (5%)
Query: 3 PEFDGREAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEF 62
P FDG Y W VE+ + +K +L G A W+ + W F
Sbjct: 113 PVFDGSGVYEWFSKVERFFRVGRYQDSDKLDLVALSLEGVALKWFLREMSTLEFRDWNSF 172
Query: 63 VEALLRKFEPELEPYMPEPVQD-------SEEEEIPGKQEVLEAERRTVVVEAKPEANSV 115
+ LL +F+P PE + + E E +E++E + +V ++ PE +
Sbjct: 173 EQRLLARFDPVKIHSSPEVLMPTTLIHVVAHETESRCTEELIEEDESSVRKKSNPEIVLI 232
Query: 116 LLAKS 120
KS
Sbjct: 233 TQKKS 237
>UniRef100_UPI000032FFF4 UPI000032FFF4 UniRef100 entry
Length = 200
Score = 39.3 bits (90), Expect = 0.24
Identities = 36/102 (35%), Positives = 50/102 (48%), Gaps = 16/102 (15%)
Query: 62 FVEALLRKFEPEL--EPYMPEPVQDSEEEEIPGKQ-----EVLEAERRTVVVEAKPEANS 114
FVE R+ E EL E + E V+ E+++ K E +EA+ T VEAKPE
Sbjct: 87 FVELTCREAETELADESVIEEKVEKDLEDKVSAKTKEPETEEVEAKPETEEVEAKPETEE 146
Query: 115 VLLAKSEIKPYSPENSEMEASRKDSEFAMAAEQTAEVSRCQP 156
V E K PE E+EA + ++E A +T EV +P
Sbjct: 147 V-----EAK---PETEEVEA-KPETEEVEAKPETEEVEAKEP 179
>UniRef100_Q7F2E9 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 39.3 bits (90), Expect = 0.24
Identities = 35/133 (26%), Positives = 57/133 (42%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL +E+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFRHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E Q + V+E E N LLA+ Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREFRSLQ-----QGTRTVIEYLHEFN--LLAR-----Y 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>UniRef100_Q852G2 Putative polyprotein [Oryza sativa]
Length = 1079
Score = 39.3 bits (90), Expect = 0.24
Identities = 35/133 (26%), Positives = 57/133 (42%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL +E+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFRHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E Q + V+E E N LLA+ Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREFRSLQ-----QGTRTVIEYLHEFN--LLAR-----Y 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>UniRef100_Q6UU21 Putative polyprotein [Oryza sativa]
Length = 1289
Score = 39.3 bits (90), Expect = 0.24
Identities = 23/67 (34%), Positives = 31/67 (45%), Gaps = 3/67 (4%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKA---TWWEFVEA 65
EA WL +E+ + +++EK S A L G AF WW +R A TW EF A
Sbjct: 79 EAGDWLHAIEKKLDLLQCTDQEKVSFASHQLHGSAFEWWDHFRLNRTTAEPITWLEFTAA 138
Query: 66 LLRKFEP 72
+ P
Sbjct: 139 FRKTHIP 145
>UniRef100_Q6F330 Putative polyprotein [Oryza sativa]
Length = 1458
Score = 39.3 bits (90), Expect = 0.24
Identities = 35/133 (26%), Positives = 57/133 (42%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL +E+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFRHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E Q + V+E E N LLA+ Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREFRSLQ-----QGTRTVIEYLHEFN--LLAR-----Y 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>UniRef100_Q6F300 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 38.9 bits (89), Expect = 0.31
Identities = 33/133 (24%), Positives = 55/133 (40%), Gaps = 22/133 (16%)
Query: 9 EAYGWLIMVEQHCEAKGVSEEEKFSGAEKALTGEAFMWW---FCWRRRNQKATWWEFVEA 65
EA+ WL VE+ +E+EK + A L G A +WW R + TW EF +
Sbjct: 112 EAHDWLHAVEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYMVTRPAGAEVTWTEFCHS 171
Query: 66 LLRKFEPELEPYMPEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPY 125
+ +PE + ++ E L+ RTV+ L + + Y
Sbjct: 172 FNK-------AQVPEGIVAQKKREF----RSLQQGTRTVI--------EYLHEFNRLARY 212
Query: 126 SPENSEMEASRKD 138
+PE+ +A R++
Sbjct: 213 APEDVRTDAERQE 225
>UniRef100_UPI00003B00C5 UPI00003B00C5 UniRef100 entry
Length = 5527
Score = 38.5 bits (88), Expect = 0.40
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 19 QHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEFVEALLRKFEPELEPYM 78
Q EA E+++ E+A + + F W + + W E +AL+ + +
Sbjct: 3909 QQYEAVSQLNSERYARLERA---QVLVNQF-WETYEELSPWIEETQALISQ--------L 3956
Query: 79 PEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPENSEMEASRKD 138
P P D E++ +QE + R ++ E KP + +L ++K +PE EM +
Sbjct: 3957 PPPAID--HEQLKQQQEDMRQLRESIA-EHKPHIDKLLKIGPQLKDLNPEEGEMVQKKYS 4013
Query: 139 SEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKID 175
+ AM A+ EV CQ +D T +KI+
Sbjct: 4014 TAEAMYAKIKEEV--CQRALALDEAVSQSTQFHDKIE 4048
>UniRef100_UPI00003B00C4 UPI00003B00C4 UniRef100 entry
Length = 5020
Score = 38.5 bits (88), Expect = 0.40
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 19 QHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEFVEALLRKFEPELEPYM 78
Q EA E+++ E+A + + F W + + W E +AL+ + +
Sbjct: 3377 QQYEAVSQLNSERYARLERA---QVLVNQF-WETYEELSPWIEETQALISQ--------L 3424
Query: 79 PEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPENSEMEASRKD 138
P P D E++ +QE + R ++ E KP + +L ++K +PE EM +
Sbjct: 3425 PPPAID--HEQLKQQQEDMRQLRESIA-EHKPHIDKLLKIGPQLKDLNPEEGEMVQKKYS 3481
Query: 139 SEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKID 175
+ AM A+ EV CQ +D T +KI+
Sbjct: 3482 TAEAMYAKIKEEV--CQRALALDEAVSQSTQFHDKIE 3516
>UniRef100_UPI00003B00C3 UPI00003B00C3 UniRef100 entry
Length = 6981
Score = 38.5 bits (88), Expect = 0.40
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 19 QHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEFVEALLRKFEPELEPYM 78
Q EA E+++ E+A + + F W + + W E +AL+ + +
Sbjct: 5338 QQYEAVSQLNSERYARLERA---QVLVNQF-WETYEELSPWIEETQALISQ--------L 5385
Query: 79 PEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPENSEMEASRKD 138
P P D E++ +QE + R ++ E KP + +L ++K +PE EM +
Sbjct: 5386 PPPAID--HEQLKQQQEDMRQLRESIA-EHKPHIDKLLKIGPQLKDLNPEEGEMVQKKYS 5442
Query: 139 SEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKID 175
+ AM A+ EV CQ +D T +KI+
Sbjct: 5443 TAEAMYAKIKEEV--CQRALALDEAVSQSTQFHDKIE 5477
>UniRef100_UPI00003B00C2 UPI00003B00C2 UniRef100 entry
Length = 5305
Score = 38.5 bits (88), Expect = 0.40
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 19 QHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEFVEALLRKFEPELEPYM 78
Q EA E+++ E+A + + F W + + W E +AL+ + +
Sbjct: 3683 QQYEAVSQLNSERYARLERA---QVLVNQF-WETYEELSPWIEETQALISQ--------L 3730
Query: 79 PEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPENSEMEASRKD 138
P P D E++ +QE + R ++ E KP + +L ++K +PE EM +
Sbjct: 3731 PPPAID--HEQLKQQQEDMRQLRESIA-EHKPHIDKLLKIGPQLKDLNPEEGEMVQKKYS 3787
Query: 139 SEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKID 175
+ AM A+ EV CQ +D T +KI+
Sbjct: 3788 TAEAMYAKIKEEV--CQRALALDEAVSQSTQFHDKIE 3822
>UniRef100_UPI00003B00C1 UPI00003B00C1 UniRef100 entry
Length = 5410
Score = 38.5 bits (88), Expect = 0.40
Identities = 39/157 (24%), Positives = 68/157 (42%), Gaps = 17/157 (10%)
Query: 19 QHCEAKGVSEEEKFSGAEKALTGEAFMWWFCWRRRNQKATWWEFVEALLRKFEPELEPYM 78
Q EA E+++ E+A + + F W + + W E +AL+ + +
Sbjct: 3792 QQYEAVSQLNSERYARLERA---QVLVNQF-WETYEELSPWIEETQALISQ--------L 3839
Query: 79 PEPVQDSEEEEIPGKQEVLEAERRTVVVEAKPEANSVLLAKSEIKPYSPENSEMEASRKD 138
P P D E++ +QE + R ++ E KP + +L ++K +PE EM +
Sbjct: 3840 PPPAID--HEQLKQQQEDMRQLRESIA-EHKPHIDKLLKIGPQLKDLNPEEGEMVQKKYS 3896
Query: 139 SEFAMAAEQTAEVSRCQPVTVVDYIDCTLTAKGEKID 175
+ AM A+ EV CQ +D T +KI+
Sbjct: 3897 TAEAMYAKIKEEV--CQRALALDEAVSQSTQFHDKIE 3931
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 772,529,141
Number of Sequences: 2790947
Number of extensions: 31585401
Number of successful extensions: 82648
Number of sequences better than 10.0: 250
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 240
Number of HSP's that attempted gapping in prelim test: 82195
Number of HSP's gapped (non-prelim): 560
length of query: 458
length of database: 848,049,833
effective HSP length: 131
effective length of query: 327
effective length of database: 482,435,776
effective search space: 157756498752
effective search space used: 157756498752
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0117b.14


