

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0114.1
(661 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
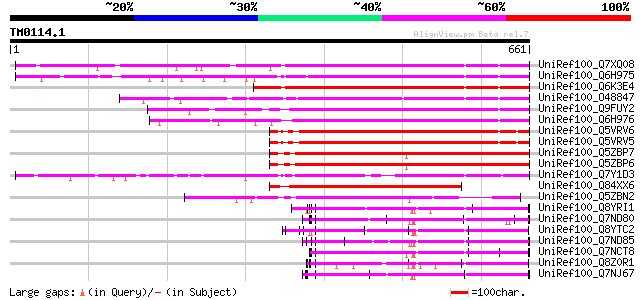
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XQ08 OSJNBb0065L13.11 protein [Oryza sativa] 369 e-100
UniRef100_Q6H975 STY-L protein [Antirrhinum majus] 365 2e-99
UniRef100_Q6K3E4 Putative LEUNIG [Oryza sativa] 365 2e-99
UniRef100_O48847 Expressed protein [Arabidopsis thaliana] 341 5e-92
UniRef100_Q9FUY2 Transcriptional corepressor LEUNIG [Arabidopsis... 327 7e-88
UniRef100_Q6H976 STYLOSA protein [Antirrhinum majus] 326 1e-87
UniRef100_Q5VRV6 Putative transcriptional corepressor LEUNIG [Or... 315 4e-84
UniRef100_Q5VRV5 Putative transcriptional corepressor LEUNIG [Or... 315 4e-84
UniRef100_Q5ZBP7 Putative LEUNIG [Oryza sativa] 299 2e-79
UniRef100_Q5ZBP6 LEUNIG-like [Oryza sativa] 299 2e-79
UniRef100_Q7Y1D3 Putative WD repeat protein [Oryza sativa] 297 6e-79
UniRef100_Q84XX6 Leunig [Brassica rapa subsp. pekinensis] 225 4e-57
UniRef100_Q5ZBN2 Putative LEUNIG [Oryza sativa] 219 2e-55
UniRef100_Q8YRI1 Hypothetical WD-repeat protein alr3466 [Anabaen... 138 5e-31
UniRef100_Q7ND80 WD-repeat protein [Gloeobacter violaceus] 136 2e-30
UniRef100_Q8YTC2 Hypothetical WD-repeat protein alr2800 [Anabaen... 135 4e-30
UniRef100_Q7ND85 WD-repeat protein [Gloeobacter violaceus] 129 2e-28
UniRef100_Q7NCT8 WD-repeat protein [Gloeobacter violaceus] 127 8e-28
UniRef100_Q8Z0R1 WD-40 repeat protein [Anabaena sp.] 126 2e-27
UniRef100_Q7NJ67 WD-repeat protein [Gloeobacter violaceus] 125 5e-27
>UniRef100_Q7XQ08 OSJNBb0065L13.11 protein [Oryza sativa]
Length = 793
Score = 369 bits (946), Expect = e-100
Identities = 246/770 (31%), Positives = 367/770 (46%), Gaps = 137/770 (17%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H + + ++ Q+ +
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH----SEIAAAYLEAQQTKAREHQQQMQMQQLQL 100
Query: 68 NEHRPQQFQANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENV-----------EPSLHD 116
+ R Q Q + + + P + L + LA + +
Sbjct: 101 IQQRHAQLQRTNATHPSLNGPISGLNSDGILGPSTASVLAAKMYEERLKHSHPMDSDGSQ 160
Query: 117 LLRENNQKLLSGTNSNHLPHPQDVGKQVQKYVFKVVTQFTSIYTN--ENHKLSGSGVTMR 174
LL + LL ++NH G+ V V T I +N + G
Sbjct: 161 LLDASRLALLKSASTNHS------GQSVPGTPGSVSTTLQQIQARNQQNIDIKSEGNMSV 214
Query: 175 VGRDVPRDPPDVV-QKTMLPLDGPHQTNNNEALNLVP------------QPLNGGPVNTP 221
R +P DP + Q + P G N+ ++ +P +P GG + P
Sbjct: 215 AQRSMPMDPSSLYGQGIIQPKPGLGGGVLNQGVSGLPLKGWPLTGIDQLRPNLGGQMQKP 274
Query: 222 --NSHQQCQVLKTQNQD--------------------------AALTQA--------PAC 245
++ Q Q++ Q Q +ALT++ PA
Sbjct: 275 FLSTQSQFQLMSPQQQQQFLAQAQAQGNLGNSTNLGDMDPRRLSALTRSVLNGKDGQPAG 334
Query: 246 TPNRQTSTVPGTKRKNKHPK--IPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQNV 303
T TS + + K + + + KT S + Q +Q H Q QQ Q ++
Sbjct: 335 TDGCITSPMQSSSPKVRPDQEYLMKTSSQQTQEQLQQQ------HNQQQQQQTQQGNRKR 388
Query: 304 EKSTKRMITESTRPVESAQDCADAKPA--------------------------------- 330
++ T ST + ++ P+
Sbjct: 389 KQPTSSGAANSTGTANTVGPSTNSPPSTPSTHTPGDGLGMTGNMRHVPKNLMMYGVEGTG 448
Query: 331 -------------------DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFS 371
D+NVESFL+ D A + F+ ++ A + +KGF+
Sbjct: 449 LPSSSNLDDLEQFGDMGSLDDNVESFLANGDGDAR---DIFAAPEKSPAEPNPVASKGFT 505
Query: 372 LQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVR 431
E+ C + +KV+ CHFSSDGK++ASAGHEKK +WNME FQ +A+ H+ +ITDVR
Sbjct: 506 FSEVNCWRTNNSKVVCCHFSSDGKILASAGHEKKAVLWNMETFQTQYTAEEHAVIITDVR 565
Query: 432 FQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDV 491
F+P S ATSSFDR+++LW+A++P SL GH + SLDFHP++ DLLCS DSN
Sbjct: 566 FRPNSNQLATSSFDRTIKLWNAADPGFSLHTFAGHCSGITSLDFHPKKTDLLCSCDSNGE 625
Query: 492 IRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKD 551
IR WNV+Q CM +GG+ QVRFQP GQ LA A N ++I DVET+ + L+GH+ +
Sbjct: 626 IRYWNVSQLSCMRAMKGGTAQVRFQPNTGQFLAAATENVVSIFDVETNGKKYTLQGHNSE 685
Query: 552 VLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
V S+CWD SG +LASVS+D ++WS +S G C E+ S GNKF SC+FHPGY LL+IGG
Sbjct: 686 VQSVCWDSSGQYLASVSQDLVKVWSISS-GDCTHEVSSNGNKFHSCVFHPGYTDLLVIGG 744
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
YQSLE W+ + +++ ++ AH+GLIA LA SP+ ++ASASHDNSVK+WK
Sbjct: 745 YQSLELWNMVK-NQSMTVQAHEGLIAALAQSPITGMVASASHDNSVKLWK 793
>UniRef100_Q6H975 STY-L protein [Antirrhinum majus]
Length = 777
Score = 365 bits (938), Expect = 2e-99
Identities = 251/756 (33%), Positives = 360/756 (47%), Gaps = 125/756 (16%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGP------------ESSCKVPMNMMPAGNSN 55
+D+P GFL++WWS+F+++F +R H E ++ M +
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNDKHSEAAAAYIETQQIKAREQQQQMQMQQLQLLQQR 104
Query: 56 NASPHRIPQIPMNEHRPQQFQANS-GFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSL 114
NA R H P NS MI QP+A ++ +++ + + E S
Sbjct: 105 NAQLQRRDP----NHPPLGGPMNSMNSEGMIGQPSASVLAMKMYEERMKHPHSMDSETS- 159
Query: 115 HDLLRENNQKLLSGTNSNHLPHPQDVGKQVQKYVFKVVTQFTSIYTNENHKLSGSGVTMR 174
L+ N LL ++ +Q Q + ++ + + +
Sbjct: 160 PGLIDANRMALLKSASN----------QQGQLMQGNTGSMSAALQQMQGRPQMANDIKGE 209
Query: 175 VG-----RDVPRDPPDVVQKTMLP----LDGPHQTNNNEALNLVPQPLNGGP-------- 217
VG + +P DP + + +L L G L L PL G
Sbjct: 210 VGLGSTQKSLPMDPSSIYGQAILQSKSGLGGAGLNQGVTGLPLKGWPLTGIDQLRPSLGL 269
Query: 218 -VNTPNSHQQCQVLKTQNQDAALTQAPAC-----TPNRQTSTVPGTKRKNKHPKIPKTGS 271
V PN Q Q L Q L QA A +PN +P K + P+
Sbjct: 270 QVQKPNLQTQNQFLLASQQQQVLAQAQAQGSLGNSPNYGYGGLPRGNSNAKDGQPPRNDG 329
Query: 272 S----------KNDSQKMEQIISTGEHQCNQDQQIQMQS--------------------- 300
S K +M+Q S + Q Q QQ Q+
Sbjct: 330 SICSPVQANSPKMKMAQMQQASSQQQDQLQQQQQQLQQNNRKRKTHSSSGPANSTGTGNT 389
Query: 301 ------------------------QNVEKSTKRM----------ITESTRPVESAQDCAD 326
Q+V +K M I ST ++ ++ D
Sbjct: 390 VGPSPGTPQSTHTPGDGMATASSLQHVNSVSKSMMMYGADAAGGIASSTNQLDDLENFGD 449
Query: 327 AKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVL 386
++NVESFLS + D + + +LK+ +KGFS E+GC+ ++ NKV
Sbjct: 450 VGSLEDNVESFLSHD---GDGNL--YGSLKQTLTEHKTETSKGFSFGEVGCI-RTRNKVT 503
Query: 387 TCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDR 446
CHFSSDGK++ASAGH+KK +WNM+ Q + + H +LITDVRF+P ST AT+SFD+
Sbjct: 504 CCHFSSDGKLLASAGHDKKAVLWNMDTLQTETTPEEHQYLITDVRFRPNSTQLATASFDK 563
Query: 447 SVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTT 506
SVRLWDA+NP L TGH VMSLDFHP++ DL C DSN+ IR W+++ C +
Sbjct: 564 SVRLWDAANPSYCLNAYTGHASHVMSLDFHPKKNDLFCFCDSNNEIRYWSISPFACTRIS 623
Query: 507 R-GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
+ GGS QVRFQP G LLA A ++I DVE D H +GH V +CWD +G+ LA
Sbjct: 624 KQGGSAQVRFQPITGHLLAAASDKVVSIYDVENDRQTHSFQGHSGVVNYLCWDLNGDLLA 683
Query: 566 SVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
SVSEDS ++WS AS G+CI EL+S GN+F SC+FHP Y +LL+IGG +SLE W+ E +K
Sbjct: 684 SVSEDSIKVWSLAS-GECIHELNSNGNQFHSCVFHPSYSALLVIGGLRSLELWNMVE-NK 741
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +++AH+ +IA LA SP+ ++ASASHD+SVK+WK
Sbjct: 742 SMTVSAHENIIAALAQSPLTGMVASASHDSSVKLWK 777
>UniRef100_Q6K3E4 Putative LEUNIG [Oryza sativa]
Length = 802
Score = 365 bits (938), Expect = 2e-99
Identities = 179/351 (50%), Positives = 243/351 (68%), Gaps = 5/351 (1%)
Query: 311 ITESTRPVESAQDCADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGF 370
+ S+ ++ + D ++NVESFL+ +D A + F+ LKR A + +KGF
Sbjct: 457 LASSSNQMDDLEPFGDVGSLEDNVESFLANDDGDAR---DIFAALKRSPAEPNPAASKGF 513
Query: 371 SLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDV 430
+ E+ CL + +KV+ CHFSSDGK++ASAGHEKK +WNM+ FQ +++ HS +ITDV
Sbjct: 514 TFNEVNCLRTNNSKVVCCHFSSDGKILASAGHEKKAVLWNMDTFQSQYTSEEHSLIITDV 573
Query: 431 RFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSND 490
RF+P S+ ATSSFDR+++LW+A++P L GH+ QV SLDFHP++ DLLCS D N
Sbjct: 574 RFRPNSSQLATSSFDRTIKLWNAADPGFCLHTFVGHNVQVTSLDFHPKKTDLLCSCDGNG 633
Query: 491 VIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDK 550
IR WN+ Q CM +GG+ QVRFQP GQ LA A + I DVET S + L+GH+
Sbjct: 634 EIRYWNLTQLSCMRAMKGGTAQVRFQPNTGQFLAAAAETMVAIFDVETHSKKYTLQGHNT 693
Query: 551 DVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIG 610
DV S+CWD SG +LASVS+D ++WS +S G+CI E+ S GNKF SC+FHP Y +LL+IG
Sbjct: 694 DVQSVCWDSSGEYLASVSQDLVKVWSISS-GECIHEVSSNGNKFHSCVFHPSYANLLVIG 752
Query: 611 GYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
GYQSLE W+ + +++ +I AH+GLIA LA SPV +ASASHDNSVK+WK
Sbjct: 753 GYQSLELWNMVK-NQSMTIQAHEGLIAALAQSPVNGTVASASHDNSVKLWK 802
>UniRef100_O48847 Expressed protein [Arabidopsis thaliana]
Length = 787
Score = 341 bits (874), Expect = 5e-92
Identities = 204/541 (37%), Positives = 303/541 (55%), Gaps = 38/541 (7%)
Query: 141 GKQVQKYVFKVVTQFTSIYTNENHKLSG--------SGVTMRVGRDVPRDPPDVVQKTML 192
G QVQK + +QF + H++ + M G PR + + +
Sbjct: 265 GPQVQKSFLQNQSQFQLSPQQQQHQMLAQVQAQGNMTNSPMYGGDMDPRRFTGLPRGNLN 324
Query: 193 PLDGPHQTNNNEALNLVPQPLNGG--------PVNTPNSHQQCQVLKTQNQ-DAALTQAP 243
P DG N+ + P+ PV +S QQ +L Q+Q + + P
Sbjct: 325 PKDGQQNANDGS----IGSPMQSSSSKHISMPPVQQSSSQQQDHLLSQQSQQNNRKRKGP 380
Query: 244 ACT-PNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGEHQCNQDQQIQMQSQN 302
+ + P T T N P P T + + I+ H N + M +
Sbjct: 381 SSSGPANSTGTGNTVGPSNSQPSTPSTHTPVDGVA-----IAGNMHHVNSMPKGPMMYGS 435
Query: 303 VEKSTKRMITESTRPVESAQD-CADAKPADENVESFLSFEDEPADNRIEPFSNLKRISAT 361
+ + + + ++ D D ++NVESFLS +D + F LKR S+
Sbjct: 436 --DGIGGLASSANQLLQDDMDQFGDVGALEDNVESFLSQDDGDGGSL---FGTLKRNSSV 490
Query: 362 CSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASAD 421
+ +K FS E+ C+ KS +KV+ C FS DGK++ASAGH+KKVFIWNME Q ++ +
Sbjct: 491 HTET-SKPFSFNEVSCIRKSASKVICCSFSYDGKLLASAGHDKKVFIWNMETLQVESTPE 549
Query: 422 THSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMD 481
H+H+ITDVRF+P ST ATSSFD+++++WDAS+P L ++GH VMS+DFHP++ +
Sbjct: 550 EHAHIITDVRFRPNSTQLATSSFDKTIKIWDASDPGYFLRTISGHAAPVMSIDFHPKKTE 609
Query: 482 LLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLATAIGNSITIIDVETDS- 540
LLCS DSN+ IR W++N S C+ +G S QVRFQP+ GQ LA A N+++I D+E ++
Sbjct: 610 LLCSCDSNNDIRFWDINAS-CVRAVKGASTQVRFQPRTGQFLAAASENTVSIFDIENNNK 668
Query: 541 HLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFH 600
++ KGH +V S+CW +G +ASVSED+ ++WS +S G CI EL + GNKF S +FH
Sbjct: 669 RVNIFKGHSSNVHSVCWSPNGELVASVSEDAVKLWSLSS-GDCIHELSNSGNKFHSVVFH 727
Query: 601 PGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
P Y LL+IGGYQ++E W+ E +K ++A H+ +I+ LA SP ++ASASHD SVK+W
Sbjct: 728 PSYPDLLVIGGYQAIELWNTME-NKCMTVAGHECVISALAQSPSTGVVASASHDKSVKIW 786
Query: 661 K 661
K
Sbjct: 787 K 787
>UniRef100_Q9FUY2 Transcriptional corepressor LEUNIG [Arabidopsis thaliana]
Length = 931
Score = 327 bits (838), Expect = 7e-88
Identities = 187/498 (37%), Positives = 265/498 (52%), Gaps = 37/498 (7%)
Query: 176 GRDVPRDPPDVVQKTMLPLDGPHQTNNNEALNLVPQPLNGGPVNTPNSHQQC-----QVL 230
G + P+ P L L P ++N +++ + GG + S QVL
Sbjct: 459 GGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGGGSITMDGSISNSFRGNEQVL 518
Query: 231 KTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQIISTGE--H 288
K Q+ + P + T N P + S + +IS H
Sbjct: 519 KNQSGRKRKQPVSSSGPANSSGTA------NTAGPSPSSAPSTPSTHTPGDVISMPNLPH 572
Query: 289 QCNQDQQIQM---QSQNVEKSTKRMITESTRPVESAQDCADAKPADENVESFLSFEDEPA 345
+ + M + S + + R VE D+NVESFLS ED
Sbjct: 573 SGGSSKSMMMFGTEGTGTLTSPSNQLADMDRFVEDGS-------LDDNVESFLSQEDGD- 624
Query: 346 DNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSDGKVMASAGHEKK 405
+R + T + +KGF+ E+ + S KV CHFSSDGK++ASAGH+KK
Sbjct: 625 ----------QRDAVTRCMDVSKGFTFTEVNSVRASTTKVTCCHFSSDGKMLASAGHDKK 674
Query: 406 VFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTG 465
+W + + + + H+ +ITD+RF P ATSSFD++VR+WDA N SL G
Sbjct: 675 AVLWYTDTMKPKTTLEEHTAMITDIRFSPSQLRLATSSFDKTVRVWDADNKGYSLRTFMG 734
Query: 466 HDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVRFQPQYGQLLAT 525
H V SLDFHP + DL+CS D+++ IR W++N C +GGS Q+RFQP+ G+ LA
Sbjct: 735 HSSMVTSLDFHPIKDDLICSCDNDNEIRYWSINNGSCTRVYKGGSTQIRFQPRVGKYLAA 794
Query: 526 AIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSARIWS--AASDGKC 583
+ N + ++DVET + H L+GH + S+CWD SG+FLASVSED ++W+ S+G+C
Sbjct: 795 SSANLVNVLDVETQAIRHSLQGHANPINSVCWDPSGDFLASVSEDMVKVWTLGTGSEGEC 854
Query: 584 IGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSP 643
+ EL GNKFQSC+FHP Y SLL+IG YQSLE W+ +E +KT ++ AH+GLI LA S
Sbjct: 855 VHELSCNGNKFQSCVFHPAYPSLLVIGCYQSLELWNMSE-NKTMTLPAHEGLITSLAVST 913
Query: 644 VGELIASASHDNSVKVWK 661
L+ASASHD VK+WK
Sbjct: 914 ATGLVASASHDKLVKLWK 931
Score = 43.1 bits (100), Expect = 0.026
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H E + M + PQ+
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH---SEVAASYIETQMIKAREQQLQQSQHPQVSQ 101
Query: 68 NEHRPQQFQ 76
+ + QQ Q
Sbjct: 102 QQQQQQQQQ 110
>UniRef100_Q6H976 STYLOSA protein [Antirrhinum majus]
Length = 915
Score = 326 bits (836), Expect = 1e-87
Identities = 198/511 (38%), Positives = 277/511 (53%), Gaps = 42/511 (8%)
Query: 179 VPRDPPDVVQK---TMLPLDGPHQTNNNEALNLVPQPLNGG-PVNTPNSHQQCQVLKTQN 234
+PR P+++ K L Q N+N+ L+G P ++ ++ QQ +++ T +
Sbjct: 419 LPRADPEMLMKLKIAQLQQQQQQQQNSNQTQQQQHHTLSGQQPQSSNHNLQQDKMMGTSS 478
Query: 235 QDAALTQAPACTPNRQTSTVPGTKRKNKHP-----KIPKTGSSKNDSQKMEQIISTGEHQ 289
+ + + N Q S T RK K P +G++ ST
Sbjct: 479 AAGEGSMSNSFRGNDQASKNQ-TGRKRKQPVSSSGPANSSGTANTAGPSPSSAPSTPSTH 537
Query: 290 CNQDQQIQMQSQNVEKSTKRMI-------TESTRPVESAQDCADAKPAD---------EN 333
D + S+K ++ T P D D PAD +N
Sbjct: 538 TPGDVMSMPALPHSGSSSKPLMMFGADNNATLTSPSNQLWDDKDLVPADMDRFVDDVEDN 597
Query: 334 VESFLSFED-EPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSS 392
VESFLS +D +P D + + +KGF+ E+ + S +KV+ CHFS
Sbjct: 598 VESFLSNDDADPRD------------AVGRCMDVSKGFTFTEVSYVRASASKVVCCHFSP 645
Query: 393 DGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
DGK++AS GH+KK +W + + + + HS LITDVRF P ATSSFD++VR+WD
Sbjct: 646 DGKLLASGGHDKKAVLWYTDTLKPKTTLEEHSSLITDVRFSPSMARLATSSFDKTVRVWD 705
Query: 453 ASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQ 512
A NP S+ TGH VMSLDFHP + DL+CS D + IR W++ C +GG+ Q
Sbjct: 706 ADNPGYSIRTFTGHSAGVMSLDFHPVKEDLICSCDGDGEIRYWSIKNGSCARVFKGGTAQ 765
Query: 513 VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA 572
VRFQP+ G+ LA A N ++I+D ET + H LKGH K + S+CWD SG LASVSEDS
Sbjct: 766 VRFQPRLGRYLAAAAENVVSILDSETLACRHSLKGHTKPIHSVCWDPSGELLASVSEDSV 825
Query: 573 RIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIA 630
R+W+ + S+G C+ EL GNKF SC+FHP Y SLL+IG YQSLE W+ +E +KT +++
Sbjct: 826 RVWTLRSGSEGDCLHELSCNGNKFHSCVFHPTYSSLLVIGCYQSLELWNMSE-NKTMTLS 884
Query: 631 AHQGLIAGLADSPVGELIASASHDNSVKVWK 661
AH+GLIA LA S L+ASASHD VK+WK
Sbjct: 885 AHEGLIASLAVSTGAGLVASASHDKIVKLWK 915
Score = 42.7 bits (99), Expect = 0.034
Identities = 22/84 (26%), Positives = 38/84 (45%), Gaps = 8/84 (9%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H S + + A + Q P
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKHSEVAASYIETQL--------MKAREQQQQQQPQ 96
Query: 68 NEHRPQQFQANSGFNNMIPQPAAR 91
+PQQ Q ++ Q A+
Sbjct: 97 QSQQPQQQQQQLHMQQLLLQRQAQ 120
>UniRef100_Q5VRV6 Putative transcriptional corepressor LEUNIG [Oryza sativa]
Length = 875
Score = 315 bits (806), Expect = 4e-84
Identities = 161/333 (48%), Positives = 220/333 (65%), Gaps = 17/333 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
+++V+SFLS +D AD R + R+ +T KGF +E+ + S NKV+ CHF
Sbjct: 558 EDHVDSFLSHDD--ADRR-----DGSRMEST------KGFIFREVSSVQASTNKVVCCHF 604
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK++A+ GH+KKV +W+ E + + + HS LITDVRF P ATSSFD++VR+
Sbjct: 605 SSDGKLLATGGHDKKVVLWHAETLKQKSVLEEHSLLITDVRFSPSIPRLATSSFDKTVRV 664
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N S+ TGH VMSLDFHP + DL+CS D ++ IR W++N + +GGS
Sbjct: 665 WDADNQGYSIRTFTGHSASVMSLDFHPNKDDLICSCDGDNEIRFWSINNGNIVRIFKGGS 724
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q+RFQP++G LA A N+++I+DVET + L +GH K V S+CWD SG ++ SVSED
Sbjct: 725 SQLRFQPRHGGYLAVASENAVSILDVETQACLRRFEGHTKHVDSVCWDPSGEYVVSVSED 784
Query: 571 SARIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWS 628
+ ++WS A SD +C+ EL G+KF SC FHP Y S+LIIG YQSLE W +E ++T +
Sbjct: 785 TVKVWSVNAGSDDRCVQELSCTGSKFHSCAFHPSYSSMLIIGCYQSLELWDMSE-NRTMT 843
Query: 629 IAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+AAH LI LA S G L+AS SHD VK+WK
Sbjct: 844 LAAHDSLITALASSSSG-LVASTSHDKFVKLWK 875
Score = 40.0 bits (92), Expect = 0.22
Identities = 22/81 (27%), Positives = 38/81 (46%), Gaps = 9/81 (11%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPES---SCKV------PMNMMPAGNSNNAS 58
+D+P GFL +WWS+F+++F +R H S S K + A + ++
Sbjct: 50 IDAPGGFLLEWWSVFWDIFIARTNEKHSDVAASYIESIKAREQQPSQLQQQEAHSQQSSQ 109
Query: 59 PHRIPQIPMNEHRPQQFQANS 79
++ Q+ + H QQ Q S
Sbjct: 110 QIQMQQLLLQRHAQQQQQQQS 130
>UniRef100_Q5VRV5 Putative transcriptional corepressor LEUNIG [Oryza sativa]
Length = 877
Score = 315 bits (806), Expect = 4e-84
Identities = 161/333 (48%), Positives = 220/333 (65%), Gaps = 17/333 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
+++V+SFLS +D AD R + R+ +T KGF +E+ + S NKV+ CHF
Sbjct: 560 EDHVDSFLSHDD--ADRR-----DGSRMEST------KGFIFREVSSVQASTNKVVCCHF 606
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
SSDGK++A+ GH+KKV +W+ E + + + HS LITDVRF P ATSSFD++VR+
Sbjct: 607 SSDGKLLATGGHDKKVVLWHAETLKQKSVLEEHSLLITDVRFSPSIPRLATSSFDKTVRV 666
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N S+ TGH VMSLDFHP + DL+CS D ++ IR W++N + +GGS
Sbjct: 667 WDADNQGYSIRTFTGHSASVMSLDFHPNKDDLICSCDGDNEIRFWSINNGNIVRIFKGGS 726
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q+RFQP++G LA A N+++I+DVET + L +GH K V S+CWD SG ++ SVSED
Sbjct: 727 SQLRFQPRHGGYLAVASENAVSILDVETQACLRRFEGHTKHVDSVCWDPSGEYVVSVSED 786
Query: 571 SARIWS--AASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWS 628
+ ++WS A SD +C+ EL G+KF SC FHP Y S+LIIG YQSLE W +E ++T +
Sbjct: 787 TVKVWSVNAGSDDRCVQELSCTGSKFHSCAFHPSYSSMLIIGCYQSLELWDMSE-NRTMT 845
Query: 629 IAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+AAH LI LA S G L+AS SHD VK+WK
Sbjct: 846 LAAHDSLITALASSSSG-LVASTSHDKFVKLWK 877
Score = 39.7 bits (91), Expect = 0.29
Identities = 20/83 (24%), Positives = 37/83 (44%), Gaps = 11/83 (13%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKV-----------PMNMMPAGNSNN 56
+D+P GFL +WWS+F+++F +R H S + + A + +
Sbjct: 50 IDAPGGFLLEWWSVFWDIFIARTNEKHSDVAASYIETQSIKAREQQPSQLQQQEAHSQQS 109
Query: 57 ASPHRIPQIPMNEHRPQQFQANS 79
+ ++ Q+ + H QQ Q S
Sbjct: 110 SQQIQMQQLLLQRHAQQQQQQQS 132
>UniRef100_Q5ZBP7 Putative LEUNIG [Oryza sativa]
Length = 876
Score = 299 bits (766), Expect = 2e-79
Identities = 162/338 (47%), Positives = 213/338 (62%), Gaps = 18/338 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
DENVESFLS +D ++P +L R S + +KGF E+ S KV CHF
Sbjct: 550 DENVESFLSQDD------MDPRDSLGR-----SMDASKGFGFAEVAKARASATKVTCCHF 598
Query: 391 SSDGKVMASAGHEKKVFIWNMEN-FQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
SSDGK++A+ GH+KKV +W E + +S + HS LITDVRF P + ATSSFD++VR
Sbjct: 599 SSDGKLLATGGHDKKVLLWCTEPALKPTSSLEEHSALITDVRFSPSMSRLATSSFDKTVR 658
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM---HTT 506
+WDA N SL TGH VMSLDFHP + D++CS D + +R W++N C+
Sbjct: 659 VWDADNTSYSLRTFTGHSASVMSLDFHPNKEDMICSCDGDGEVRSWSINNGSCLTFVKVF 718
Query: 507 RGGSKQVRFQPQYGQLLATAIGNSITIIDVETD-SHLHDLKGHDKDVLSICWDRSGNFLA 565
+GG+ Q+RFQPQ G+ LA A +I I+D ET + + L+GH K++ S+CWD +G+ LA
Sbjct: 719 KGGATQMRFQPQKGKYLAAASEKAIYILDGETQLACRNPLQGHTKNIHSLCWDSTGDNLA 778
Query: 566 SVSEDSARIWSAA--SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEG 623
SVSEDS RIWS A DG+ + EL GNKFQSC+FHP Y LL+IG Y+SLE W E
Sbjct: 779 SVSEDSVRIWSFAPGHDGEFVNELSCSGNKFQSCVFHPSYPYLLVIGCYESLELWDIREK 838
Query: 624 SKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +AH GL+A LA S +AS SHD VK+WK
Sbjct: 839 NAMTVHSAHDGLVAALAASSATGKVASVSHDRFVKLWK 876
Score = 42.7 bits (99), Expect = 0.034
Identities = 35/138 (25%), Positives = 58/138 (41%), Gaps = 10/138 (7%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWS+F+++F +R H ++ + M A + PQ
Sbjct: 45 IDAPGGFLFEWWSVFWDIFIARTNEKH--SDVAASYIETQHMKAREQQQQQQQQPPQ--Q 100
Query: 68 NEHRPQQFQANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSLHDLLRENNQKLLS 127
+ +PQ Q M+ Q AA+ Q Q + + R + LL+
Sbjct: 101 RQQQPQHIQ----MQQMLLQRAAQQ-QQQQQQQQQQQQQQQQQQQQQQQQQRRDGTHLLN 155
Query: 128 GTNSNHLPHPQDVGKQVQ 145
GT S LP + +Q Q
Sbjct: 156 GTASG-LPGNNPLMRQNQ 172
>UniRef100_Q5ZBP6 LEUNIG-like [Oryza sativa]
Length = 538
Score = 299 bits (766), Expect = 2e-79
Identities = 162/338 (47%), Positives = 213/338 (62%), Gaps = 18/338 (5%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
DENVESFLS +D ++P +L R S + +KGF E+ S KV CHF
Sbjct: 212 DENVESFLSQDD------MDPRDSLGR-----SMDASKGFGFAEVAKARASATKVTCCHF 260
Query: 391 SSDGKVMASAGHEKKVFIWNMEN-FQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
SSDGK++A+ GH+KKV +W E + +S + HS LITDVRF P + ATSSFD++VR
Sbjct: 261 SSDGKLLATGGHDKKVLLWCTEPALKPTSSLEEHSALITDVRFSPSMSRLATSSFDKTVR 320
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM---HTT 506
+WDA N SL TGH VMSLDFHP + D++CS D + +R W++N C+
Sbjct: 321 VWDADNTSYSLRTFTGHSASVMSLDFHPNKEDMICSCDGDGEVRSWSINNGSCLTFVKVF 380
Query: 507 RGGSKQVRFQPQYGQLLATAIGNSITIIDVETD-SHLHDLKGHDKDVLSICWDRSGNFLA 565
+GG+ Q+RFQPQ G+ LA A +I I+D ET + + L+GH K++ S+CWD +G+ LA
Sbjct: 381 KGGATQMRFQPQKGKYLAAASEKAIYILDGETQLACRNPLQGHTKNIHSLCWDSTGDNLA 440
Query: 566 SVSEDSARIWSAA--SDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEG 623
SVSEDS RIWS A DG+ + EL GNKFQSC+FHP Y LL+IG Y+SLE W E
Sbjct: 441 SVSEDSVRIWSFAPGHDGEFVNELSCSGNKFQSCVFHPSYPYLLVIGCYESLELWDIREK 500
Query: 624 SKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ +AH GL+A LA S +AS SHD VK+WK
Sbjct: 501 NAMTVHSAHDGLVAALAASSATGKVASVSHDRFVKLWK 538
>UniRef100_Q7Y1D3 Putative WD repeat protein [Oryza sativa]
Length = 684
Score = 297 bits (761), Expect = 6e-79
Identities = 226/689 (32%), Positives = 348/689 (49%), Gaps = 84/689 (12%)
Query: 8 LDSPDGFLYDWWSIFYEVFASRYGRGHCTGPESSCKVPMNMMPAGNSNNASPHRIPQIPM 67
+D+P GFL++WWSIF+++F +R P P +P + R+ +
Sbjct: 45 IDAPGGFLFEWWSIFWDIFDARTR----DKPPQPQPQPPPPIPIDIKSREQQMRLQLLQQ 100
Query: 68 NEHRPQQFQ---ANSGFNNMIPQPAARLIPQAVFDNGQLGNLAENVEPSLHDLLRENNQK 124
++ QQ + N+ + + +A L + + D + N ++ + H LL N
Sbjct: 101 QQNAHQQRRDPPLNAAMDALNSDVSAVLASKMMQDRMRNPNPTDS--DASHQLLDANRIA 158
Query: 125 LLS-GTN---------SNHLPHPQDVGKQVQK--------------YVFKVVTQFTSIYT 160
LL TN S ++ Q + + Q+ Y ++T +I+
Sbjct: 159 LLKPATNQTGQLVQGASVNMSALQQIHSRNQQPVIPFHLLKLHWFMYRLSLLTLCPTIFK 218
Query: 161 NENHKLSGSGVTMRVGRDVPRDPPDVVQKTML-PLDGPHQTNNNEALNLVPQPLNGGPVN 219
+ + G + R +P DP + M+ P G T N+ + VP L G P+
Sbjct: 219 D----MKGDAAMSQ--RSMPTDPSTLYGSGMMQPKSGLVSTGLNQGVGSVP--LKGWPL- 269
Query: 220 TPNSHQQCQVLKTQNQDAALTQAPACTPNRQTSTVPGT-KRKNKHPKIPKTGSSKNDSQK 278
T + C +LK D + +Q P + + ++ S+ND +
Sbjct: 270 TKSLPTSC-LLKVPGIDQLRSNLGV---QKQLMASPNQFQLLSPQQQLIAQAQSQNDLAR 325
Query: 279 MEQIISTGEHQCNQDQQIQMQ----SQNVEKSTKRMITESTRPVESAQDCADA-KPADEN 333
M +G + D+ M +Q + S R++ + + QD D+ DEN
Sbjct: 326 MGSPAPSGSPKVRPDESDYMMKLKMAQMQQPSGHRLMELQQQQQQLQQDNLDSFVDFDEN 385
Query: 334 VESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHFSSD 393
V+SFLS +D D R + F++LK+ S+ ++ KG SL E G S NKV+ CHFS+D
Sbjct: 386 VDSFLSNDD--GDGR-DIFASLKKGSS--EQDSLKGLSLSEFGNNRTSNNKVVCCHFSTD 440
Query: 394 GKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDA 453
GK++ASAGHEKKVF+WNM+N + H++ ITD+RF+P ST ATSS D +VRLW+A
Sbjct: 441 GKLLASAGHEKKVFLWNMDNLNMDTKIEEHTNFITDIRFKPNSTQLATSSSDGTVRLWNA 500
Query: 454 SNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV 513
+ I T + ++ +L V + + + +GG+ +V
Sbjct: 501 ------IEIFTRNLQRFYAL-----------------VTTMEKFVSGKLVKMKQGGTGRV 537
Query: 514 RFQPQYGQLLATAIGNSITIIDVETDSHLHDL-KGHDKDVLSICWDRSGNFLASVSEDSA 572
RFQPQ GQLLA A G+ + I+DVE ++ LH L K H +V ICWD G +ASVS+D+
Sbjct: 538 RFQPQIGQLLAVATGSIVNIVDVEKEASLHSLPKVHTNEVNCICWDEKGERVASVSQDTV 597
Query: 573 RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAH 632
++WS AS G CI EL S GN++QSCIFHP Y ++LI+GGYQ++E WS ++ + ++ AH
Sbjct: 598 KVWSVAS-GACIHELRSHGNQYQSCIFHPRYPNVLIVGGYQTMELWSLSDNHRN-TVQAH 655
Query: 633 QGLIAGLADSPVGELIASASHDNSVKVWK 661
+GLIA LA S +IASASHD SVK+WK
Sbjct: 656 EGLIAALAHSQFTGMIASASHDRSVKLWK 684
>UniRef100_Q84XX6 Leunig [Brassica rapa subsp. pekinensis]
Length = 249
Score = 225 bits (573), Expect = 4e-57
Identities = 107/245 (43%), Positives = 152/245 (61%), Gaps = 11/245 (4%)
Query: 331 DENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINKVLTCHF 390
D+NVESFLS ED +R + + +KGF+ E+ + S +KV CHF
Sbjct: 16 DDNVESFLSHEDGD-----------QRDTVGRCMDVSKGFTFAEVNSVRASTSKVTCCHF 64
Query: 391 SSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRL 450
S DGK++ASAGH+KK IW+ + + + + H+ +ITDVRF P ATSSFD++VR+
Sbjct: 65 SLDGKMLASAGHDKKAVIWHTDTMKPKTTLEEHTAMITDVRFSPNLPRLATSSFDKTVRV 124
Query: 451 WDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGS 510
WDA N SL GH + S+DFHP + DL+CS D + IR W++N C +GGS
Sbjct: 125 WDADNKGYSLRNFIGHSSMITSVDFHPNKDDLICSCDDDGEIRYWSINNGTCSRVYKGGS 184
Query: 511 KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED 570
Q RFQP+ G+ LA + N ++++DVET + H L+GH + S+CWD SG+FLA+VSED
Sbjct: 185 TQTRFQPRVGKYLAASSANVVSVLDVETQACRHSLQGHTNQINSVCWDASGDFLATVSED 244
Query: 571 SARIW 575
++W
Sbjct: 245 MVKVW 249
>UniRef100_Q5ZBN2 Putative LEUNIG [Oryza sativa]
Length = 774
Score = 219 bits (558), Expect = 2e-55
Identities = 150/451 (33%), Positives = 211/451 (46%), Gaps = 76/451 (16%)
Query: 223 SHQQCQVLKTQNQDAALTQAPACTPNRQTSTVPGTKRKNKHPKIPKTGSSKNDSQKMEQI 282
SHQQ Q L + A + + S + K KI + N S +
Sbjct: 369 SHQQAQSLNQLHHQQAQSIGSMLDGSIPNSFGLANRASKKRKKIVSSSERANSSGTSNNV 428
Query: 283 ISTGE--------HQCNQDQQIQMQSQNVEKS----------TKRMITESTRPVESAQDC 324
S+ H + + N KS TK +I+ T P+
Sbjct: 429 GSSSSSAPSTPFTHTPRDEMSMPQLKYNGGKSKSLSMFGYDDTKSLISP-TNPLGDVDQL 487
Query: 325 ADAKPADENVESFLSFEDEPADNRIEPFSNLKRISATCSRNENKGFSLQEIGCLHKSINK 384
+ DENVESFLS ED ++P + + +KGF E+ S NK
Sbjct: 488 QEDGSLDENVESFLSQED------MDPQETMGHCM-----DASKGFGFIEVAKARASTNK 536
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V CHFSSDGK++A+ GH+KKV +W ++ A + HS +ITDVRF T ATSSF
Sbjct: 537 VDCCHFSSDGKLLATGGHDKKVVLWFTDDLNIKAIFEEHSMIITDVRFSSIMTRLATSSF 596
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D+++R+WDA+NP SL GH V+SLDFHP + D++CS DS+ +R W+++ C++
Sbjct: 597 DKTIRVWDANNPEYSLHTFIGHSTSVVSLDFHPNKEDIICSCDSDGEVRCWSIDNGSCVN 656
Query: 505 TTR---GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLH--DLKGHDKDVLSICWDR 559
R GG+ Q+RFQP +G+ LA I+I+D ET H++ DL+GH K++ S+CWD
Sbjct: 657 CVRVFNGGAIQLRFQPHHGKYLAVVSEKMISILDAET-LHIYRSDLQGHLKNIHSVCWDA 715
Query: 560 SGNFLASVSEDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
+G +LASVSEDS + SLE W
Sbjct: 716 TGGYLASVSEDSIK----------------------------------------SLELWD 735
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIAS 650
E + AH G+I LA S LIAS
Sbjct: 736 IREKNIVTINNAHDGMIPSLAASNASGLIAS 766
Score = 43.1 bits (100), Expect = 0.026
Identities = 14/30 (46%), Positives = 23/30 (76%)
Query: 5 DSVLDSPDGFLYDWWSIFYEVFASRYGRGH 34
D +D+P GFL++WWSIF+++F ++ R H
Sbjct: 4 DKAIDAPGGFLFEWWSIFWDIFIAQTDREH 33
>UniRef100_Q8YRI1 Hypothetical WD-repeat protein alr3466 [Anabaena sp.]
Length = 1526
Score = 138 bits (348), Expect = 5e-31
Identities = 87/283 (30%), Positives = 145/283 (50%), Gaps = 9/283 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V FS DG +AS ++ V +W++ + + H+ + V F P + A+
Sbjct: 1159 NWVNAVAFSPDGATLASGSGDQTVRLWDISSSKCLYILQGHTSWVNSVVFNPDGSTLASG 1218
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S D++VRLW+ N + L GH V S+ F+P +L S S+ +RLW+++ S+C
Sbjct: 1219 SSDQTVRLWEI-NSSKCLCTFQGHTSWVNSVVFNPDG-SMLASGSSDKTVRLWDISSSKC 1276
Query: 503 MHTTRGGSK---QVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWD 558
+HT +G + V F P G +LA+ G+ ++ + ++ + LH +GH V S+ +
Sbjct: 1277 LHTFQGHTNWVNSVAFNPD-GSMLASGSGDQTVRLWEISSSKCLHTFQGHTSWVSSVTFS 1335
Query: 559 RSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA 617
G LAS S+D + R+WS +S G+C+ N S IF P L G Q++
Sbjct: 1336 PDGTMLASGSDDQTVRLWSISS-GECLYTFLGHTNWVGSVIFSPDGAILASGSGDQTVRL 1394
Query: 618 WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
WS + G +++ H + + SP G L+AS S D +V++W
Sbjct: 1395 WSISSGKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQTVRLW 1437
Score = 129 bits (325), Expect = 2e-28
Identities = 79/275 (28%), Positives = 136/275 (48%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F+ DG +AS ++ V +W + + + + H+ + V F P +M A+ S D++VR
Sbjct: 1208 FNPDGSTLASGSSDQTVRLWEINSSKCLCTFQGHTSWVNSVVFNPDGSMLASGSSDKTVR 1267
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD S+ + L GH V S+ F+P +L S + +RLW ++ S+C+HT +G
Sbjct: 1268 LWDISSS-KCLHTFQGHTNWVNSVAFNPDG-SMLASGSGDQTVRLWEISSSKCLHTFQGH 1325
Query: 510 SK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P L + + ++ + + + L+ GH V S+ + G LAS
Sbjct: 1326 TSWVSSVTFSPDGTMLASGSDDQTVRLWSISSGECLYTFLGHTNWVGSVIFSPDGAILAS 1385
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S D + R+WS +S GKC+ L N S +F P L Q++ W+ + G
Sbjct: 1386 GSGDQTVRLWSISS-GKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQTVRLWNISSGEC 1444
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++ H + +A S G ++AS S D ++K+W
Sbjct: 1445 LYTLHGHINSVRSVAFSSDGLILASGSDDETIKLW 1479
Score = 122 bits (305), Expect = 4e-26
Identities = 77/275 (28%), Positives = 136/275 (49%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DGK++AS ++ V +W++ + Q + H+ + V F P S M A+ S D++VR
Sbjct: 914 FSQDGKMLASGSDDQTVRLWDISSGQCLKTFKGHTSRVRSVVFSPNSLMLASGSSDQTVR 973
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD S+ L I GH V S+ F+ +L + + +RLW+++ S+C + +G
Sbjct: 974 LWDISSG-ECLYIFQGHTGWVYSVAFN-LDGSMLATGSGDQTVRLWDISSSQCFYIFQGH 1031
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ VR F L + + ++ + D+ + + L+ L+GH V S+ + G LAS
Sbjct: 1032 TSCVRSVVFSSDGAMLASGSDDQTVRLWDISSGNCLYTLQGHTSCVRSVVFSPDGAMLAS 1091
Query: 567 VSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
+D R+W +S G C+ L + + +F P +L Q + W +
Sbjct: 1092 GGDDQIVRLWDISS-GNCLYTLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLWDISSKKC 1150
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++ H + +A SP G +AS S D +V++W
Sbjct: 1151 LYTLQGHTNWVNAVAFSPDGATLASGSGDQTVRLW 1185
Score = 121 bits (303), Expect = 8e-26
Identities = 82/286 (28%), Positives = 138/286 (47%), Gaps = 9/286 (3%)
Query: 380 KSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMF 439
K + VLT FS DGK+ A+ V W + + H+ + V F M
Sbjct: 862 KILGSVLTVAFSPDGKLFATGDSGGIVRFWEAATGKELLTCKGHNSWVNSVGFSQDGKML 921
Query: 440 ATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
A+ S D++VRLWD S+ + L GH +V S+ F P + +L S S+ +RLW+++
Sbjct: 922 ASGSDDQTVRLWDISSG-QCLKTFKGHTSRVRSVVFSPNSL-MLASGSSDQTVRLWDISS 979
Query: 500 SECMHTTRGGS---KQVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSI 555
EC++ +G + V F G +LAT G+ ++ + D+ + + +GH V S+
Sbjct: 980 GECLYIFQGHTGWVYSVAFNLD-GSMLATGSGDQTVRLWDISSSQCFYIFQGHTSCVRSV 1038
Query: 556 CWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQS 614
+ G LAS S+D + R+W +S G C+ L + +S +F P L G Q
Sbjct: 1039 VFSSDGAMLASGSDDQTVRLWDISS-GNCLYTLQGHTSCVRSVVFSPDGAMLASGGDDQI 1097
Query: 615 LEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ W + G+ +++ + + L SP G +A+ S D V++W
Sbjct: 1098 VRLWDISSGNCLYTLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLW 1143
Score = 118 bits (295), Expect = 6e-25
Identities = 74/276 (26%), Positives = 135/276 (48%), Gaps = 7/276 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FSSDG ++AS ++ V +W++ + + H+ + V F P M A+ D+ VR
Sbjct: 1040 FSSDGAMLASGSDDQTVRLWDISSGNCLYTLQGHTSCVRSVVFSPDGAMLASGGDDQIVR 1099
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD S+ L L G+ V L F P + L + S+ ++RLW+++ +C++T +G
Sbjct: 1100 LWDISSG-NCLYTLQGYTSWVRFLVFSPNGVTL-ANGSSDQIVRLWDISSKKCLYTLQGH 1157
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P L + + ++ + D+ + L+ L+GH V S+ ++ G+ LAS
Sbjct: 1158 TNWVNAVAFSPDGATLASGSGDQTVRLWDISSSKCLYILQGHTSWVNSVVFNPDGSTLAS 1217
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S D + R+W S KC+ + S +F+P L +++ W +
Sbjct: 1218 GSSDQTVRLWEINSS-KCLCTFQGHTSWVNSVVFNPDGSMLASGSSDKTVRLWDISSSKC 1276
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+ H + +A +P G ++AS S D +V++W+
Sbjct: 1277 LHTFQGHTNWVNSVAFNPDGSMLASGSGDQTVRLWE 1312
Score = 118 bits (295), Expect = 6e-25
Identities = 78/280 (27%), Positives = 136/280 (47%), Gaps = 7/280 (2%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + F+ DG ++A+ ++ V +W++ + Q F H+ + V F M A+ S
Sbjct: 993 VYSVAFNLDGSMLATGSGDQTVRLWDISSSQCFYIFQGHTSCVRSVVFSSDGAMLASGSD 1052
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++VRLWD S+ L L GH V S+ F P +L S + ++RLW+++ C++
Sbjct: 1053 DQTVRLWDISSG-NCLYTLQGHTSCVRSVVFSPDGA-MLASGGDDQIVRLWDISSGNCLY 1110
Query: 505 TTRGGSKQVRFQPQYGQLLATAIGNSITII---DVETDSHLHDLKGHDKDVLSICWDRSG 561
T +G + VRF + A G+S I+ D+ + L+ L+GH V ++ + G
Sbjct: 1111 TLQGYTSWVRFLVFSPNGVTLANGSSDQIVRLWDISSKKCLYTLQGHTNWVNAVAFSPDG 1170
Query: 562 NFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
LAS S D + R+W +S KC+ L + S +F+P +L Q++ W
Sbjct: 1171 ATLASGSGDQTVRLWDISS-SKCLYILQGHTSWVNSVVFNPDGSTLASGSSDQTVRLWEI 1229
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ H + + +P G ++AS S D +V++W
Sbjct: 1230 NSSKCLCTFQGHTSWVNSVVFNPDGSMLASGSSDKTVRLW 1269
Score = 96.3 bits (238), Expect = 3e-18
Identities = 68/241 (28%), Positives = 118/241 (48%), Gaps = 13/241 (5%)
Query: 359 SATCSRNENKGFSLQEIG---CLHK---SINKVLTCHFSSDGKVMASAGHEKKVFIWNME 412
S S + +K L +I CLH N V + F+ DG ++AS ++ V +W +
Sbjct: 1255 SMLASGSSDKTVRLWDISSSKCLHTFQGHTNWVNSVAFNPDGSMLASGSGDQTVRLWEIS 1314
Query: 413 NFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMS 472
+ + + H+ ++ V F P TM A+ S D++VRLW S+ L GH V S
Sbjct: 1315 SSKCLHTFQGHTSWVSSVTFSPDGTMLASGSDDQTVRLWSISSG-ECLYTFLGHTNWVGS 1373
Query: 473 LDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGN 529
+ F P +L S + +RLW+++ +C++T +G + V F P L + +
Sbjct: 1374 VIFSPDGA-ILASGSGDQTVRLWSISSGKCLYTLQGHNNWVGSIVFSPDGTLLASGSDDQ 1432
Query: 530 SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELH 588
++ + ++ + L+ L GH V S+ + G LAS S+D + ++W + G+CI L
Sbjct: 1433 TVRLWNISSGECLYTLHGHINSVRSVAFSSDGLILASGSDDETIKLWDVKT-GECIKTLK 1491
Query: 589 S 589
S
Sbjct: 1492 S 1492
>UniRef100_Q7ND80 WD-repeat protein [Gloeobacter violaceus]
Length = 1188
Score = 136 bits (342), Expect = 2e-30
Identities = 87/292 (29%), Positives = 141/292 (47%), Gaps = 7/292 (2%)
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
+ +G L ++VL+ FS DG V+AS H++ + +W + + H+ I + F
Sbjct: 728 ERLGTLTGHTDQVLSVAFSPDGGVLASGSHDQTLKLWEVTTGTCLTTLTGHTGRIRAISF 787
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P A+SS D +V+LWDA+ L TGH QV S+ F P L S + +
Sbjct: 788 SPDGEWLASSSLDCTVKLWDAATG-ECLRTFTGHSGQVWSVSFAPDGQTL-ASGSLDQTV 845
Query: 493 RLWNVNQSECMHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
R+W+ +C+ T +G + V F P L + ++ ++ I DV + + L GH
Sbjct: 846 RIWDAATGQCLRTLQGNAGWIWSVAFAPDGQTLASGSLDRTVRIWDVPSGRCVRTLTGHG 905
Query: 550 KDVLSICWDRSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLI 608
V S+ + G LAS S D ++W AA+ G+C+ L N +S F P R+L
Sbjct: 906 SWVWSVAFSPDGRTLASGSFDQTIKLWDAAT-GQCLRTLSGHNNWVRSVAFSPDGRTLAS 964
Query: 609 IGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
Q+++ W + G ++ H + +A SP G +AS S D +V+VW
Sbjct: 965 GSHDQTVKLWEVSSGQCLRTLTGHSSWVWSVAFSPDGRTVASGSFDQTVRVW 1016
Score = 128 bits (321), Expect = 6e-28
Identities = 81/276 (29%), Positives = 137/276 (49%), Gaps = 7/276 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG +A+A ++ V +W++ + + H+ + V F P + A+ S D++++
Sbjct: 703 FSPDGHTLAAASLDRTVKLWDVRTGERLGTLTGHTDQVLSVAFSPDGGVLASGSHDQTLK 762
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ + L LTGH ++ ++ F P + L SS + ++LW+ EC+ T G
Sbjct: 763 LWEVTTGT-CLTTLTGHTGRIRAISFSPDG-EWLASSSLDCTVKLWDAATGECLRTFTGH 820
Query: 510 SKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
S QV F P L + ++ ++ I D T L L+G+ + S+ + G LAS
Sbjct: 821 SGQVWSVSFAPDGQTLASGSLDQTVRIWDAATGQCLRTLQGNAGWIWSVAFAPDGQTLAS 880
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S D + RIW S G+C+ L G+ S F P R+L Q+++ W G
Sbjct: 881 GSLDRTVRIWDVPS-GRCVRTLTGHGSWVWSVAFSPDGRTLASGSFDQTIKLWDAATGQC 939
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
+++ H + +A SP G +AS SHD +VK+W+
Sbjct: 940 LRTLSGHNNWVRSVAFSPDGRTLASGSHDQTVKLWE 975
Score = 121 bits (303), Expect = 8e-26
Identities = 77/280 (27%), Positives = 133/280 (47%), Gaps = 6/280 (2%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
+L +S G+++A +V +W + + Q S H+ I+ + F P ++ A+ S
Sbjct: 571 ILFVAYSPKGELLAIGDDSGEVRLWRVRDGQQQLSFRGHTDWISALAFSPDGSVLASGSE 630
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D++++LWD + + L LTGH V S+ F P + SS SN+ +RLW+ +C
Sbjct: 631 DQTIKLWDTATG-QCLRTLTGHGGWVYSVAFSPDGTLIASSSPSNETVRLWDAAGGQCTR 689
Query: 505 TTR---GGSKQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
T + G V F P L A ++ ++ + DV T L L GH VLS+ + G
Sbjct: 690 TFKSRTGRMWSVAFSPDGHTLAAASLDRTVKLWDVRTGERLGTLTGHTDQVLSVAFSPDG 749
Query: 562 NFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
LAS S D + ++W + G C+ L + ++ F P L +++ W
Sbjct: 750 GVLASGSHDQTLKLWEVTT-GTCLTTLTGHTGRIRAISFSPDGEWLASSSLDCTVKLWDA 808
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G + H G + ++ +P G+ +AS S D +V++W
Sbjct: 809 ATGECLRTFTGHSGQVWSVSFAPDGQTLASGSLDQTVRIW 848
Score = 120 bits (300), Expect = 2e-25
Identities = 81/281 (28%), Positives = 133/281 (46%), Gaps = 7/281 (2%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSS 443
+V + F+ DG+ +AS ++ V IW+ Q + ++ I V F P A+ S
Sbjct: 823 QVWSVSFAPDGQTLASGSLDQTVRIWDAATGQCLRTLQGNAGWIWSVAFAPDGQTLASGS 882
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
DR+VR+WD + R + LTGH V S+ F P L S + I+LW+ +C+
Sbjct: 883 LDRTVRIWDVPSG-RCVRTLTGHGSWVWSVAFSPDGRTL-ASGSFDQTIKLWDAATGQCL 940
Query: 504 HTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRS 560
T G + VR F P L + + ++ + +V + L L GH V S+ +
Sbjct: 941 RTLSGHNNWVRSVAFSPDGRTLASGSHDQTVKLWEVSSGQCLRTLTGHSSWVWSVAFSPD 1000
Query: 561 GNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
G +AS S D + R+W+AA+ G+C+ L ++ S F P R L G ++ W
Sbjct: 1001 GRTVASGSFDQTVRVWNAAT-GECLHTLKVDSSQVWSVAFSPDGRILAGGSGNYAVWLWD 1059
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G ++ H + +A SP + S+SHD +V++W
Sbjct: 1060 TATGECLRTLTGHTSQVWSVAFSPDSRTVVSSSHDQTVRLW 1100
Score = 118 bits (295), Expect = 6e-25
Identities = 73/275 (26%), Positives = 130/275 (46%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F+ DG+ +AS ++ V IW++ + + + H + V F P A+ SFD++++
Sbjct: 871 FAPDGQTLASGSLDRTVRIWDVPSGRCVRTLTGHGSWVWSVAFSPDGRTLASGSFDQTIK 930
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWDA+ + L L+GH+ V S+ F P L S + ++LW V+ +C+ T G
Sbjct: 931 LWDAATG-QCLRTLSGHNNWVRSVAFSPDGRTL-ASGSHDQTVKLWEVSSGQCLRTLTGH 988
Query: 510 SK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
S V F P + + + ++ + + T LH LK V S+ + G LA
Sbjct: 989 SSWVWSVAFSPDGRTVASGSFDQTVRVWNAATGECLHTLKVDSSQVWSVAFSPDGRILAG 1048
Query: 567 VSEDSAR-IWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S + A +W A+ G+C+ L ++ S F P R+++ Q++ W G
Sbjct: 1049 GSGNYAVWLWDTAT-GECLRTLTGHTSQVWSVAFSPDSRTVVSSSHDQTVRLWDAATGEC 1107
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ H + +A SP G + S S D ++++W
Sbjct: 1108 LRTLTGHTSQVWSVAFSPDGRTVISGSQDETIRLW 1142
Score = 108 bits (269), Expect = 7e-22
Identities = 75/265 (28%), Positives = 127/265 (47%), Gaps = 16/265 (6%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+ +AS ++ + +W+ Q + H++ + V F P A+ S D++V+
Sbjct: 913 FSPDGRTLASGSFDQTIKLWDAATGQCLRTLSGHNNWVRSVAFSPDGRTLASGSHDQTVK 972
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ S+ + L LTGH V S+ F P + S + +R+WN EC+HT +
Sbjct: 973 LWEVSSG-QCLRTLTGHSSWVWSVAFSPDGRTV-ASGSFDQTVRVWNAATGECLHTLKVD 1030
Query: 510 SKQV---RFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
S QV F P G++LA GN ++ + D T L L GH V S+ + +
Sbjct: 1031 SSQVWSVAFSPD-GRILAGGSGNYAVWLWDTATGECLRTLTGHTSQVWSVAFSPDSRTVV 1089
Query: 566 SVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGS 624
S S D + R+W AA+ G+C+ L ++ S F P R+++ +++ W G
Sbjct: 1090 SSSHDQTVRLWDAAT-GECLRTLTGHTSQVWSVAFSPDGRTVISGSQDETIRLWDSHTGK 1148
Query: 625 KTWSIAA---HQGL----IAGLADS 642
+ A ++G+ + GL D+
Sbjct: 1149 PLELLRADRLYEGMDITGVTGLTDA 1173
Score = 92.8 bits (229), Expect = 3e-17
Identities = 58/199 (29%), Positives = 95/199 (47%), Gaps = 6/199 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V + FS DG+ +AS H++ V +W + + Q + HS + V F P A+
Sbjct: 948 NWVRSVAFSPDGRTLASGSHDQTVKLWEVSSGQCLRTLTGHSSWVWSVAFSPDGRTVASG 1007
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
SFD++VR+W+A+ L L QV S+ F P +L N + LW+ EC
Sbjct: 1008 SFDQTVRVWNAATG-ECLHTLKVDSSQVWSVAFSPDGR-ILAGGSGNYAVWLWDTATGEC 1065
Query: 503 MHTTRGGSKQ---VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
+ T G + Q V F P +++++ ++ + D T L L GH V S+ +
Sbjct: 1066 LRTLTGHTSQVWSVAFSPDSRTVVSSSHDQTVRLWDAATGECLRTLTGHTSQVWSVAFSP 1125
Query: 560 SGNFLASVSED-SARIWSA 577
G + S S+D + R+W +
Sbjct: 1126 DGRTVISGSQDETIRLWDS 1144
Score = 78.2 bits (191), Expect = 7e-13
Identities = 50/186 (26%), Positives = 85/186 (44%), Gaps = 6/186 (3%)
Query: 481 DLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVE 537
+LL D + +RLW V + + RG + + F P L + + +I + D
Sbjct: 581 ELLAIGDDSGEVRLWRVRDGQQQLSFRGHTDWISALAFSPDGSVLASGSEDQTIKLWDTA 640
Query: 538 TDSHLHDLKGHDKDVLSICWDRSGNFLASVS--EDSARIWSAASDGKCIGELHSVGNKFQ 595
T L L GH V S+ + G +AS S ++ R+W AA G+C S +
Sbjct: 641 TGQCLRTLTGHGGWVYSVAFSPDGTLIASSSPSNETVRLWDAAG-GQCTRTFKSRTGRMW 699
Query: 596 SCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDN 655
S F P +L ++++ W G + ++ H + +A SP G ++AS SHD
Sbjct: 700 SVAFSPDGHTLAAASLDRTVKLWDVRTGERLGTLTGHTDQVLSVAFSPDGGVLASGSHDQ 759
Query: 656 SVKVWK 661
++K+W+
Sbjct: 760 TLKLWE 765
>UniRef100_Q8YTC2 Hypothetical WD-repeat protein alr2800 [Anabaena sp.]
Length = 1258
Score = 135 bits (340), Expect = 4e-30
Identities = 83/291 (28%), Positives = 142/291 (48%), Gaps = 9/291 (3%)
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
I LH N+V + FS DG+ +A ++ V +WN Q + ++ V F P
Sbjct: 887 IKTLHGHTNEVCSVAFSPDGQTLACVSLDQSVRLWNCRTGQCLKAWYGNTDWALPVAFSP 946
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
+ A+ S D++V+LWD + + L GH + + + F P L S+ ++ +RL
Sbjct: 947 DRQILASGSNDKTVKLWDWQTG-KYISSLEGHTDFIYGIAFSPDSQTL-ASASTDSSVRL 1004
Query: 495 WNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDK 550
WN++ +C + V F PQ G+++AT + ++ + ++ T L L H
Sbjct: 1005 WNISTGQCFQILLEHTDWVYAVVFHPQ-GKIIATGSADCTVKLWNISTGQCLKTLSEHSD 1063
Query: 551 DVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLII 609
+L + W G LAS S D S R+W + G+C+G L N+ S IF P +
Sbjct: 1064 KILGMAWSPDGQLLASASADQSVRLWDCCT-GRCVGILRGHSNRVYSAIFSPNGEIIATC 1122
Query: 610 GGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
Q+++ W +G ++ H + +A SP G+++ASASHD +V++W
Sbjct: 1123 STDQTVKIWDWQQGKCLKTLTGHTNWVFDIAFSPDGKILASASHDQTVRIW 1173
Score = 127 bits (320), Expect = 8e-28
Identities = 82/276 (29%), Positives = 141/276 (50%), Gaps = 9/276 (3%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS D +++AS ++K V +W+ + +Y +S + H+ I + F P S A++S D SVR
Sbjct: 944 FSPDRQILASGSNDKTVKLWDWQTGKYISSLEGHTDFIYGIAFSPDSQTLASASTDSSVR 1003
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ S + IL H + V ++ FHP+ ++ + ++ ++LWN++ +C+ T
Sbjct: 1004 LWNISTG-QCFQILLEHTDWVYAVVFHPQGK-IIATGSADCTVKLWNISTGQCLKTLSEH 1061
Query: 510 SKQV---RFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
S ++ + P GQLLA+A + S+ + D T + L+GH V S + +G +A
Sbjct: 1062 SDKILGMAWSPD-GQLLASASADQSVRLWDCCTGRCVGILRGHSNRVYSAIFSPNGEIIA 1120
Query: 566 SVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGS 624
+ S D + +IW GKC+ L N F P + L Q++ W G
Sbjct: 1121 TCSTDQTVKIWDW-QQGKCLKTLTGHTNWVFDIAFSPDGKILASASHDQTVRIWDVNTGK 1179
Query: 625 KTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
H L++ +A SP GE++AS S D +V++W
Sbjct: 1180 CHHICIGHTHLVSSVAFSPDGEVVASGSQDQTVRIW 1215
Score = 120 bits (302), Expect = 1e-25
Identities = 78/297 (26%), Positives = 145/297 (48%), Gaps = 9/297 (3%)
Query: 370 FSLQEIGC--LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLI 427
F+ ++ C +++ +L+ FS +G+++A+ + V +W +++ + HS+ +
Sbjct: 628 FANSDLSCCVFTETLGNILSAAFSPEGQLLATCDTDCHVRVWEVKSGKLLLICRGHSNWV 687
Query: 428 TDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSD 487
V F P + A+ D +V+LW + + + LTGH+ +V S+ FHP + L S+
Sbjct: 688 RFVVFSPDGEILASCGADENVKLWSVRDGV-CIKTLTGHEHEVFSVAFHPDG-ETLASAS 745
Query: 488 SNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHD 544
+ I+LW++ C+ T G + VR F P L ++A ++I + DV L
Sbjct: 746 GDKTIKLWDIQDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQGKCLRT 805
Query: 545 LKGHDKDVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGY 603
LK H V S+ + G LAS S D + +IW+ + G+C+ N S + P
Sbjct: 806 LKSHTGWVRSVAFSADGQTLASGSGDRTIKIWNYHT-GECLKTYIGHTNSVYSIAYSPDS 864
Query: 604 RSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ L+ G ++++ W ++ H + +A SP G+ +A S D SV++W
Sbjct: 865 KILVSGSGDRTIKLWDCQTHICIKTLHGHTNEVCSVAFSPDGQTLACVSLDQSVRLW 921
Score = 119 bits (298), Expect = 3e-25
Identities = 88/321 (27%), Positives = 153/321 (47%), Gaps = 11/321 (3%)
Query: 348 RIEPFSNLKRISATCSRNEN-KGFSLQEIGCLHKSI---NKVLTCHFSSDGKVMASAGHE 403
R FS I A+C +EN K +S+++ C+ ++V + F DG+ +ASA +
Sbjct: 688 RFVVFSPDGEILASCGADENVKLWSVRDGVCIKTLTGHEHEVFSVAFHPDGETLASASGD 747
Query: 404 KKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGIL 463
K + +W++++ + H+ + V F P A+S+ D +++LWD S + L L
Sbjct: 748 KTIKLWDIQDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQG-KCLRTL 806
Query: 464 TGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSKQV---RFQPQYG 520
H V S+ F L S + I++WN + EC+ T G + V + P
Sbjct: 807 KSHTGWVRSVAFSADG-QTLASGSGDRTIKIWNYHTGECLKTYIGHTNSVYSIAYSPDSK 865
Query: 521 QLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSED-SARIWSAAS 579
L++ + +I + D +T + L GH +V S+ + G LA VS D S R+W+ +
Sbjct: 866 ILVSGSGDRTIKLWDCQTHICIKTLHGHTNEVCSVAFSPDGQTLACVSLDQSVRLWNCRT 925
Query: 580 DGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGL 639
G+C+ + + F P + L ++++ W G S+ H I G+
Sbjct: 926 -GQCLKAWYGNTDWALPVAFSPDRQILASGSNDKTVKLWDWQTGKYISSLEGHTDFIYGI 984
Query: 640 ADSPVGELIASASHDNSVKVW 660
A SP + +ASAS D+SV++W
Sbjct: 985 AFSPDSQTLASASTDSSVRLW 1005
Score = 96.3 bits (238), Expect = 3e-18
Identities = 61/200 (30%), Positives = 105/200 (52%), Gaps = 9/200 (4%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F GK++A+ + V +WN+ Q + HS I + + P + A++S D+SVR
Sbjct: 1028 FHPQGKIIATGSADCTVKLWNISTGQCLKTLSEHSDKILGMAWSPDGQLLASASADQSVR 1087
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD R +GIL GH +V S F P +++ + ++ +++W+ Q +C+ T G
Sbjct: 1088 LWDCCTG-RCVGILRGHSNRVYSAIFSPNG-EIIATCSTDQTVKIWDWQQGKCLKTLTGH 1145
Query: 510 SK---QVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLA 565
+ + F P G++LA+A + ++ I DV T H GH V S+ + G +A
Sbjct: 1146 TNWVFDIAFSPD-GKILASASHDQTVRIWDVNTGKCHHICIGHTHLVSSVAFSPDGEVVA 1204
Query: 566 SVSED-SARIWSAASDGKCI 584
S S+D + RIW+ + G+C+
Sbjct: 1205 SGSQDQTVRIWNVKT-GECL 1223
Score = 77.0 bits (188), Expect = 2e-12
Identities = 54/213 (25%), Positives = 95/213 (44%), Gaps = 6/213 (2%)
Query: 452 DASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSK 511
D +N S + T ++S F P LL + D++ +R+W V + + RG S
Sbjct: 627 DFANSDLSCCVFTETLGNILSAAFSPEGQ-LLATCDTDCHVRVWEVKSGKLLLICRGHSN 685
Query: 512 QVRF---QPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVS 568
VRF P L + ++ + V + L GH+ +V S+ + G LAS S
Sbjct: 686 WVRFVVFSPDGEILASCGADENVKLWSVRDGVCIKTLTGHEHEVFSVAFHPDGETLASAS 745
Query: 569 EDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTW 627
D ++W DG C+ L + + F P +L +++ W ++G
Sbjct: 746 GDKTIKLWDI-QDGTCLQTLTGHTDWVRCVAFSPDGNTLASSAADHTIKLWDVSQGKCLR 804
Query: 628 SIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ +H G + +A S G+ +AS S D ++K+W
Sbjct: 805 TLKSHTGWVRSVAFSADGQTLASGSGDRTIKIW 837
Score = 75.5 bits (184), Expect = 5e-12
Identities = 38/133 (28%), Positives = 71/133 (52%), Gaps = 2/133 (1%)
Query: 375 IGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQP 434
+G L N+V + FS +G+++A+ ++ V IW+ + + + H++ + D+ F P
Sbjct: 1097 VGILRGHSNRVYSAIFSPNGEIIATCSTDQTVKIWDWQQGKCLKTLTGHTNWVFDIAFSP 1156
Query: 435 GSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRL 494
+ A++S D++VR+WD N + I GH V S+ F P +++ S + +R+
Sbjct: 1157 DGKILASASHDQTVRIWDV-NTGKCHHICIGHTHLVSSVAFSPDG-EVVASGSQDQTVRI 1214
Query: 495 WNVNQSECMHTTR 507
WNV EC+ R
Sbjct: 1215 WNVKTGECLQILR 1227
Score = 62.0 bits (149), Expect = 5e-08
Identities = 37/105 (35%), Positives = 57/105 (54%), Gaps = 4/105 (3%)
Query: 352 FSNLKRISATCSRNEN-KGFSLQEIGCLHK---SINKVLTCHFSSDGKVMASAGHEKKVF 407
FS I ATCS ++ K + Q+ CL N V FS DGK++ASA H++ V
Sbjct: 1112 FSPNGEIIATCSTDQTVKIWDWQQGKCLKTLTGHTNWVFDIAFSPDGKILASASHDQTVR 1171
Query: 408 IWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVRLWD 452
IW++ + H+HL++ V F P + A+ S D++VR+W+
Sbjct: 1172 IWDVNTGKCHHICIGHTHLVSSVAFSPDGEVVASGSQDQTVRIWN 1216
>UniRef100_Q7ND85 WD-repeat protein [Gloeobacter violaceus]
Length = 1184
Score = 129 bits (325), Expect = 2e-28
Identities = 84/275 (30%), Positives = 128/275 (46%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG +ASA H++ V +WN + A+ H ++ V F P AT S DR+VR
Sbjct: 741 FSPDGHRLASASHDRTVKLWNPATGRCLATLAGHGDWVSAVAFAPDGRSLATGSLDRTVR 800
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ + L L H +QV S+ FHP+ L S ++LW+ +C+ T +G
Sbjct: 801 LWETITG-QCLKTLQEHTDQVFSIAFHPQG-HTLASGSPTQTVKLWDTESGQCLRTLQGK 858
Query: 510 SKQV---RFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ V F P L++ + + + DV T L+GH + V ++ G LAS
Sbjct: 859 TVTVLAVAFSPHGQTLVSGSDDRLVRLWDVRTGECTRVLRGHLRGVTTVAVAPDGRTLAS 918
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
D S +IW A S G+C+ L +S F P R L + + W P G
Sbjct: 919 AGADLSVKIWDALS-GQCLRTLREHTGSIRSVAFAPDGRLLASGSQDGTAKLWDPGTGRC 977
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ H I +A +P G L+AS S D + ++W
Sbjct: 978 VATLRGHTSWIRSVAFAPDGGLLASGSQDGTARIW 1012
Score = 114 bits (286), Expect = 7e-24
Identities = 81/292 (27%), Positives = 134/292 (45%), Gaps = 7/292 (2%)
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
Q + L I V + F+ DG ++ASAG + V +W+ + A+ H+ ++ V F
Sbjct: 640 QCLATLRGHIGWVRSAAFAPDGSLLASAGQDSTVKLWDAATGRCLATLQGHTGVVHSVAF 699
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P ++ A++ D +V+LWDA+ R L L GH E + S+ F P L S+ + +
Sbjct: 700 APDGSLLASAGQDSTVKLWDAATG-RCLATLQGHTEPIRSVVFSPDG-HRLASASHDRTV 757
Query: 493 RLWNVNQSECMHTTRGGS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
+LWN C+ T G V F P L ++ ++ + + T L L+ H
Sbjct: 758 KLWNPATGRCLATLAGHGDWVSAVAFAPDGRSLATGSLDRTVRLWETITGQCLKTLQEHT 817
Query: 550 KDVLSICWDRSGNFLASVS-EDSARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLI 608
V SI + G+ LAS S + ++W S G+C+ L + F P ++L+
Sbjct: 818 DQVFSIAFHPQGHTLASGSPTQTVKLWDTES-GQCLRTLQGKTVTVLAVAFSPHGQTLVS 876
Query: 609 IGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ + W G T + H + +A +P G +ASA D SVK+W
Sbjct: 877 GSDDRLVRLWDVRTGECTRVLRGHLRGVTTVAVAPDGRTLASAGADLSVKIW 928
Score = 114 bits (285), Expect = 9e-24
Identities = 83/281 (29%), Positives = 134/281 (47%), Gaps = 9/281 (3%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
V + FS DG+ +A ++ +W + Q S H+ + V F P FA++S
Sbjct: 568 VFSVAFSPDGEQIAVGDDNSEIRLWRAADGQQQLSCQGHTDWVCAVAFAPNGQTFASASQ 627
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D +V+LWDA + L L GH V S F P LL S+ + ++LW+ C+
Sbjct: 628 DGTVKLWDARIG-QCLATLRGHIGWVRSAAFAPDG-SLLASAGQDSTVKLWDAATGRCLA 685
Query: 505 TTRGGS---KQVRFQPQYGQLLATAIGNS-ITIIDVETDSHLHDLKGHDKDVLSICWDRS 560
T +G + V F P G LLA+A +S + + D T L L+GH + + S+ +
Sbjct: 686 TLQGHTGVVHSVAFAPD-GSLLASAGQDSTVKLWDAATGRCLATLQGHTEPIRSVVFSPD 744
Query: 561 GNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
G+ LAS S D + ++W+ A+ G+C+ L G+ + F P RSL +++ W
Sbjct: 745 GHRLASASHDRTVKLWNPAT-GRCLATLAGHGDWVSAVAFAPDGRSLATGSLDRTVRLWE 803
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G ++ H + +A P G +AS S +VK+W
Sbjct: 804 TITGQCLKTLQEHTDQVFSIAFHPQGHTLASGSPTQTVKLW 844
Score = 100 bits (249), Expect = 1e-19
Identities = 74/280 (26%), Positives = 122/280 (43%), Gaps = 7/280 (2%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
VL FS G+ + S ++ V +W++ + H +T V P A++
Sbjct: 862 VLAVAFSPHGQTLVSGSDDRLVRLWDVRTGECTRVLRGHLRGVTTVAVAPDGRTLASAGA 921
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D SV++WDA + + L L H + S+ F P LL S + +LW+ C+
Sbjct: 922 DLSVKIWDALSG-QCLRTLREHTGSIRSVAFAPDGR-LLASGSQDGTAKLWDPGTGRCVA 979
Query: 505 TTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSG 561
T RG + +R F P G L + + + I D T L L GH + S+ + G
Sbjct: 980 TLRGHTSWIRSVAFAPDGGLLASGSQDGTARIWDTRTGECLQILAGHTYLICSVAFSLDG 1039
Query: 562 NFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSP 620
LAS S+D R+W + G C+ L S F P + L +++ W
Sbjct: 1040 QLLASGSQDQTIRLWEVQT-GACLRTLTEKTGMVFSLAFSPDGQILASGSNDMTVKLWQV 1098
Query: 621 TEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G ++ H L+ +A +P G +ASAS D +++++
Sbjct: 1099 GTGRCVKTLGPHTSLVVSIAYAPDGSTLASASLDETIRLF 1138
Score = 100 bits (249), Expect = 1e-19
Identities = 79/291 (27%), Positives = 136/291 (46%), Gaps = 13/291 (4%)
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST 437
L + ++V + F G +AS + V +W+ E+ Q + + + V F P
Sbjct: 813 LQEHTDQVFSIAFHPQGHTLASGSPTQTVKLWDTESGQCLRTLQGKTVTVLAVAFSPHGQ 872
Query: 438 MFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV 497
+ S DR VRLWD +L GH V ++ P L S+ ++ +++W+
Sbjct: 873 TLVSGSDDRLVRLWDVRTG-ECTRVLRGHLRGVTTVAVAPDGRTL-ASAGADLSVKIWDA 930
Query: 498 NQSECMHTTR---GGSKQVRFQPQYGQLLATAIGNSITII-DVETDSHLHDLKGHDKDVL 553
+C+ T R G + V F P G+LLA+ + + D T + L+GH +
Sbjct: 931 LSGQCLRTLREHTGSIRSVAFAPD-GRLLASGSQDGTAKLWDPGTGRCVATLRGHTSWIR 989
Query: 554 SICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGY 612
S+ + G LAS S+D +ARIW + G+C+ L G+ + C L+ G
Sbjct: 990 SVAFAPDGGLLASGSQDGTARIWDTRT-GECLQIL--AGHTYLICSVAFSLDGQLLASGS 1046
Query: 613 Q--SLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
Q ++ W G+ ++ G++ LA SP G+++AS S+D +VK+W+
Sbjct: 1047 QDQTIRLWEVQTGACLRTLTEKTGMVFSLAFSPDGQILASGSNDMTVKLWQ 1097
Score = 99.4 bits (246), Expect = 3e-19
Identities = 68/240 (28%), Positives = 117/240 (48%), Gaps = 9/240 (3%)
Query: 427 ITDVRFQPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSS 486
+ V F P A + +RLW A++ + L GH + V ++ F P S+
Sbjct: 568 VFSVAFSPDGEQIAVGDDNSEIRLWRAADGQQQLSC-QGHTDWVCAVAFAPNGQTF-ASA 625
Query: 487 DSNDVIRLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNS-ITIIDVETDSHL 542
+ ++LW+ +C+ T RG VR F P G LLA+A +S + + D T L
Sbjct: 626 SQDGTVKLWDARIGQCLATLRGHIGWVRSAAFAPD-GSLLASAGQDSTVKLWDAATGRCL 684
Query: 543 HDLKGHDKDVLSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHP 601
L+GH V S+ + G+ LAS +DS ++W AA+ G+C+ L +S +F P
Sbjct: 685 ATLQGHTGVVHSVAFAPDGSLLASAGQDSTVKLWDAAT-GRCLATLQGHTEPIRSVVFSP 743
Query: 602 GYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
L ++++ W+P G ++A H ++ +A +P G +A+ S D +V++W+
Sbjct: 744 DGHRLASASHDRTVKLWNPATGRCLATLAGHGDWVSAVAFAPDGRSLATGSLDRTVRLWE 803
Score = 73.6 bits (179), Expect = 2e-11
Identities = 59/212 (27%), Positives = 98/212 (45%), Gaps = 9/212 (4%)
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST 437
L + + + F+ DG+++AS + +W+ + A+ H+ I V F P
Sbjct: 939 LREHTGSIRSVAFAPDGRLLASGSQDGTAKLWDPGTGRCVATLRGHTSWIRSVAFAPDGG 998
Query: 438 MFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNV 497
+ A+ S D + R+WD L IL GH + S+ F LL S + IRLW V
Sbjct: 999 LLASGSQDGTARIWDTRTG-ECLQILAGHTYLICSVAFSLDGQ-LLASGSQDQTIRLWEV 1056
Query: 498 NQSECMHTTRGGSKQV---RFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVL 553
C+ T + V F P GQ+LA+ + ++ + V T + L H V+
Sbjct: 1057 QTGACLRTLTEKTGMVFSLAFSPD-GQILASGSNDMTVKLWQVGTGRCVKTLGPHTSLVV 1115
Query: 554 SICWDRSGNFLASVS-EDSARIWSAASDGKCI 584
SI + G+ LAS S +++ R++ A+ G C+
Sbjct: 1116 SIAYAPDGSTLASASLDETIRLFDPAT-GACL 1146
Score = 45.4 bits (106), Expect = 0.005
Identities = 22/86 (25%), Positives = 44/86 (50%), Gaps = 3/86 (3%)
Query: 370 FSLQEIGCLHKSINK---VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHL 426
+ +Q CL K V + FS DG+++AS ++ V +W + + + H+ L
Sbjct: 1054 WEVQTGACLRTLTEKTGMVFSLAFSPDGQILASGSNDMTVKLWQVGTGRCVKTLGPHTSL 1113
Query: 427 ITDVRFQPGSTMFATSSFDRSVRLWD 452
+ + + P + A++S D ++RL+D
Sbjct: 1114 VVSIAYAPDGSTLASASLDETIRLFD 1139
>UniRef100_Q7NCT8 WD-repeat protein [Gloeobacter violaceus]
Length = 1081
Score = 127 bits (320), Expect = 8e-28
Identities = 85/281 (30%), Positives = 134/281 (47%), Gaps = 7/281 (2%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSS 443
+V + FS DG+ +ASAG + V +W++ + H+ + V F PG + A+
Sbjct: 634 QVRSVAFSPDGRTLASAGVDGTVRLWDVPLGACLMVLEGHTSRVRTVAFSPGGHLLASGG 693
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
D++VRLW+ + R L +L GH QV SL FHP L S + +RLW V+ +
Sbjct: 694 HDQTVRLWEVRSG-RCLRVLPGHTGQVWSLAFHPNGRTL-ASGSMDQTVRLWEVDSGRSL 751
Query: 504 HTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRS 560
T +G S V F P L + ++ + + D T L L GH V S+ +
Sbjct: 752 KTFQGNSGWIWSVAFHPGGHLLASGSMDRLVRLWDTRTGQCLKTLAGHGCWVWSLAFHPG 811
Query: 561 GNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
G LAS S D + ++W + G+CI L N ++ F P + G Q++ W+
Sbjct: 812 GEILASGSFDQTVKLWEVDT-GRCIQSLAGHTNWIRAVAFSPDGAQIASAGVDQTIRLWA 870
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G+ T + H G + +A P G +AS S D ++K+W
Sbjct: 871 WPAGNCTAVLTGHTGWVRCVAFGPDGRQLASGSLDRTIKIW 911
Score = 119 bits (297), Expect = 4e-25
Identities = 79/283 (27%), Positives = 137/283 (47%), Gaps = 9/283 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
++V T FS G ++AS GH++ V +W + + + H+ + + F P A+
Sbjct: 675 SRVRTVAFSPGGHLLASGGHDQTVRLWEVRSGRCLRVLPGHTGQVWSLAFHPNGRTLASG 734
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S D++VRLW+ + RSL G+ + S+ FHP LL S + ++RLW+ +C
Sbjct: 735 SMDQTVRLWEVDSG-RSLKTFQGNSGWIWSVAFHPGG-HLLASGSMDRLVRLWDTRTGQC 792
Query: 503 MHTTRGGSKQV---RFQPQYGQLLAT-AIGNSITIIDVETDSHLHDLKGHDKDVLSICWD 558
+ T G V F P G++LA+ + ++ + +V+T + L GH + ++ +
Sbjct: 793 LKTLAGHGCWVWSLAFHPG-GEILASGSFDQTVKLWEVDTGRCIQSLAGHTNWIRAVAFS 851
Query: 559 RSGNFLASVSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEA 617
G +AS D R+W+ + G C L + F P R L ++++
Sbjct: 852 PDGAQIASAGVDQTIRLWAWPA-GNCTAVLTGHTGWVRCVAFGPDGRQLASGSLDRTIKI 910
Query: 618 WSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
W G ++ H+G I +A SP G L+ASA+ D+ VK+W
Sbjct: 911 WDAATGECVATLGGHRGQICAVAFSPDGSLLASAAEDHLVKLW 953
Score = 112 bits (280), Expect = 3e-23
Identities = 74/275 (26%), Positives = 125/275 (44%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F G ++AS ++ V +W+ Q + H + + F PG + A+ SFD++V+
Sbjct: 766 FHPGGHLLASGSMDRLVRLWDTRTGQCLKTLAGHGCWVWSLAFHPGGEILASGSFDQTVK 825
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ R + L GH + ++ F P + S+ + IRLW C G
Sbjct: 826 LWEVDTG-RCIQSLAGHTNWIRAVAFSPDGAQI-ASAGVDQTIRLWAWPAGNCTAVLTGH 883
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ VR F P QL + ++ +I I D T + L GH + ++ + G+ LAS
Sbjct: 884 TGWVRCVAFGPDGRQLASGSLDRTIKIWDAATGECVATLGGHRGQICAVAFSPDGSLLAS 943
Query: 567 VSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
+ED ++W+ A+ G+C+ L S F P L G Q + W G+
Sbjct: 944 AAEDHLVKLWNLAT-GECVATLAGHCGPVWSVAFAPDGLHLASCGHDQVVRFWDAGSGAL 1002
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
T ++ H + +A P GE +AS S D ++++W
Sbjct: 1003 TATLRGHSDQVWSVAYDPRGETLASGSQDKTIRLW 1037
Score = 111 bits (278), Expect = 6e-23
Identities = 78/281 (27%), Positives = 130/281 (45%), Gaps = 7/281 (2%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSS 443
+V + F +G+ +AS ++ V +W +++ + + +S I V F PG + A+ S
Sbjct: 718 QVWSLAFHPNGRTLASGSMDQTVRLWEVDSGRSLKTFQGNSGWIWSVAFHPGGHLLASGS 777
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
DR VRLWD + L L GH V SL FHP ++L S + ++LW V+ C+
Sbjct: 778 MDRLVRLWDTRTG-QCLKTLAGHGCWVWSLAFHPGG-EILASGSFDQTVKLWEVDTGRCI 835
Query: 504 HTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRS 560
+ G + +R F P Q+ + + +I + + L GH V + +
Sbjct: 836 QSLAGHTNWIRAVAFSPDGAQIASAGVDQTIRLWAWPAGNCTAVLTGHTGWVRCVAFGPD 895
Query: 561 GNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
G LAS S D + +IW AA+ G+C+ L + + F P L ++ W+
Sbjct: 896 GRQLASGSLDRTIKIWDAAT-GECVATLGGHRGQICAVAFSPDGSLLASAAEDHLVKLWN 954
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
G ++A H G + +A +P G +AS HD V+ W
Sbjct: 955 LATGECVATLAGHCGPVWSVAFAPDGLHLASCGHDQVVRFW 995
Score = 110 bits (276), Expect = 1e-22
Identities = 79/275 (28%), Positives = 123/275 (44%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F +G ++AS + V +W + Q A+ H+ + V F P A+ S D +VR
Sbjct: 514 FHPEGNLLASGSEDLSVKLWAAGSGQCLATLTGHTGWVYAVAFAPDGRTLASGSVDGTVR 573
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD + L IL Q S+ F P L + + I+LW V+ C + G
Sbjct: 574 LWDVGTGL-CLKILCEPGGQFWSVAFAPDGQTLATAGHGH-AIKLWQVSSGACALSLEGH 631
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ QVR F P L + + ++ + DV + L L+GH V ++ + G+ LAS
Sbjct: 632 TAQVRSVAFSPDGRTLASAGVDGTVRLWDVPLGACLMVLEGHTSRVRTVAFSPGGHLLAS 691
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
D + R+W S G+C+ L + S FHP R+L Q++ W G
Sbjct: 692 GGHDQTVRLWEVRS-GRCLRVLPGHTGQVWSLAFHPNGRTLASGSMDQTVRLWEVDSGRS 750
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ + G I +A P G L+AS S D V++W
Sbjct: 751 LKTFQGNSGWIWSVAFHPGGHLLASGSMDRLVRLW 785
Score = 102 bits (255), Expect = 3e-20
Identities = 81/282 (28%), Positives = 127/282 (44%), Gaps = 9/282 (3%)
Query: 385 VLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSF 444
+L F+ +G V+A ++ + + Q A H+ + + F P + A+ S
Sbjct: 467 ILLVAFNPEGTVLAIGDDSGEIRLLRAADGQQQARCTGHTDALCAMAFHPEGNLLASGSE 526
Query: 445 DRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMH 504
D SV+LW A + + L LTGH V ++ F P L S + +RLW+V C+
Sbjct: 527 DLSVKLWAAGSG-QCLATLTGHTGWVYAVAFAPDGR-TLASGSVDGTVRLWDVGTGLCLK 584
Query: 505 ---TTRGGSKQVRFQPQYGQLLATA-IGNSITIIDVETDSHLHDLKGHDKDVLSICWDRS 560
G V F P GQ LATA G++I + V + + L+GH V S+ +
Sbjct: 585 ILCEPGGQFWSVAFAPD-GQTLATAGHGHAIKLWQVSSGACALSLEGHTAQVRSVAFSPD 643
Query: 561 GNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWS 619
G LAS D + R+W G C+ L ++ ++ F PG L G Q++ W
Sbjct: 644 GRTLASAGVDGTVRLWDVPL-GACLMVLEGHTSRVRTVAFSPGGHLLASGGHDQTVRLWE 702
Query: 620 PTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
G + H G + LA P G +AS S D +V++W+
Sbjct: 703 VRSGRCLRVLPGHTGQVWSLAFHPNGRTLASGSMDQTVRLWE 744
Score = 88.2 bits (217), Expect = 7e-16
Identities = 60/206 (29%), Positives = 99/206 (47%), Gaps = 7/206 (3%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N + FS DG +ASAG ++ + +W A H+ + V F P A+
Sbjct: 843 NWIRAVAFSPDGAQIASAGVDQTIRLWAWPAGNCTAVLTGHTGWVRCVAFGPDGRQLASG 902
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSEC 502
S DR++++WDA+ + L GH Q+ ++ F P LL S+ + +++LWN+ EC
Sbjct: 903 SLDRTIKIWDAATG-ECVATLGGHRGQICAVAFSPDG-SLLASAAEDHLVKLWNLATGEC 960
Query: 503 MHTTRGGSK---QVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDR 559
+ T G V F P L + + D + + L+GH V S+ +D
Sbjct: 961 VATLAGHCGPVWSVAFAPDGLHLASCGHDQVVRFWDAGSGALTATLRGHSDQVWSVAYDP 1020
Query: 560 SGNFLASVSED-SARIWSAASDGKCI 584
G LAS S+D + R+W+ A+ G+C+
Sbjct: 1021 RGETLASGSQDKTIRLWNPAT-GECL 1045
Score = 81.3 bits (199), Expect = 9e-14
Identities = 62/238 (26%), Positives = 102/238 (42%), Gaps = 7/238 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F G+++AS ++ V +W ++ + S H++ I V F P A++ D+++R
Sbjct: 808 FHPGGEILASGSFDQTVKLWEVDTGRCIQSLAGHTNWIRAVAFSPDGAQIASAGVDQTIR 867
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW A +LTGH V + F P L S + I++W+ EC+ T G
Sbjct: 868 LW-AWPAGNCTAVLTGHTGWVRCVAFGPDGRQL-ASGSLDRTIKIWDAATGECVATLGGH 925
Query: 510 SKQ---VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
Q V F P L + A + + + ++ T + L GH V S+ + G LAS
Sbjct: 926 RGQICAVAFSPDGSLLASAAEDHLVKLWNLATGECVATLAGHCGPVWSVAFAPDGLHLAS 985
Query: 567 VSEDSA-RIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEG 623
D R W A S G L ++ S + P +L +++ W+P G
Sbjct: 986 CGHDQVVRFWDAGS-GALTATLRGHSDQVWSVAYDPRGETLASGSQDKTIRLWNPATG 1042
>UniRef100_Q8Z0R1 WD-40 repeat protein [Anabaena sp.]
Length = 1227
Score = 126 bits (316), Expect = 2e-27
Identities = 86/291 (29%), Positives = 140/291 (47%), Gaps = 12/291 (4%)
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGS- 436
L K+ NKV + FS DG+++ASA ++ + +W++ + H + V F P +
Sbjct: 682 LSKNTNKVYSVAFSPDGRILASASQDQTIKLWDIATGNCQQTLIGHDDWVWSVTFSPVTD 741
Query: 437 ---TMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIR 493
+ A+SS D+ ++LWD + + L L GH +V S+ F P L SS + +R
Sbjct: 742 DRPLLLASSSADQHIKLWDVATG-KCLKTLKGHTREVHSVSFSPDG-QTLASSGEDSTVR 799
Query: 494 LWNVNQSECMHTTRGGSKQ---VRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDK 550
LW+V +C G SK+ VRF P L + SI + D++ ++ L GH
Sbjct: 800 LWDVKTGQCWQIFEGHSKKVYSVRFSPDGQTLASCGEDRSIKLWDIQRGECVNTLWGHSS 859
Query: 551 DVLSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLII 609
V +I + G L S S+D +AR+W + G + L S F P + L
Sbjct: 860 QVWAIAFSPDGRTLISCSDDQTARLWDVIT-GNSLNILRGYTRDVYSVAFSPDSQILASG 918
Query: 610 GGYQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ W+ G + + HQG I +A P G+++AS S DN++K+W
Sbjct: 919 RDDYTIGLWNLKTG-ECHPLRGHQGRIRSVAFHPDGKILASGSADNTIKLW 968
Score = 119 bits (298), Expect = 3e-25
Identities = 84/284 (29%), Positives = 134/284 (46%), Gaps = 11/284 (3%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSS 443
KV + FS DG+ +AS G ++ + +W+++ + + HS + + F P + S
Sbjct: 818 KVYSVRFSPDGQTLASCGEDRSIKLWDIQRGECVNTLWGHSSQVWAIAFSPDGRTLISCS 877
Query: 444 FDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECM 503
D++ RLWD SL IL G+ V S+ F P +L S + I LWN+ EC
Sbjct: 878 DDQTARLWDVITG-NSLNILRGYTRDVYSVAFSP-DSQILASGRDDYTIGLWNLKTGEC- 934
Query: 504 HTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSH---LHDLKGHDKDVLSICW 557
H RG ++R F P L + + N+I + D+ +H + L GH V ++ +
Sbjct: 935 HPLRGHQGRIRSVAFHPDGKILASGSADNTIKLWDISDTNHSKYIRTLTGHTNWVWTVVF 994
Query: 558 DRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLE 616
+ LAS SED + R+W G C+ +L + + F P R L ++
Sbjct: 995 SPDKHTLASSSEDRTIRLWD-KDTGDCLQKLKGHSHWVWTVAFSPDGRILASGSADSEIK 1053
Query: 617 AWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
W G ++ QG+I +A S G L+ASAS D +VK+W
Sbjct: 1054 IWDVASGKCLQTLTDPQGMIWSVAFSLDGTLLASASEDQTVKLW 1097
Score = 107 bits (268), Expect = 9e-22
Identities = 77/289 (26%), Positives = 129/289 (43%), Gaps = 11/289 (3%)
Query: 380 KSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMF 439
++++ V++ FS DGK A+ ++ +W + + H+ + F P S M
Sbjct: 600 ETMSSVVSVKFSPDGKYFATGLMNGEIRLWQTSDNKQLRIYKGHTAWVWAFAFSPDSRML 659
Query: 440 ATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
A+ S D +++LWD L L+ + +V S+ F P +L S+ + I+LW++
Sbjct: 660 ASGSADSTIKLWDVHTG-ECLKTLSKNTNKVYSVAFSPDGR-ILASASQDQTIKLWDIAT 717
Query: 500 SECMHTTRGGSK---QVRFQPQYGQ----LLATAIGNSITIIDVETDSHLHDLKGHDKDV 552
C T G V F P L +++ I + DV T L LKGH ++V
Sbjct: 718 GNCQQTLIGHDDWVWSVTFSPVTDDRPLLLASSSADQHIKLWDVATGKCLKTLKGHTREV 777
Query: 553 LSICWDRSGNFLASVSEDS-ARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
S+ + G LAS EDS R+W + G+C K S F P ++L G
Sbjct: 778 HSVSFSPDGQTLASSGEDSTVRLWDVKT-GQCWQIFEGHSKKVYSVRFSPDGQTLASCGE 836
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+S++ W G ++ H + +A SP G + S S D + ++W
Sbjct: 837 DRSIKLWDIQRGECVNTLWGHSSQVWAIAFSPDGRTLISCSDDQTARLW 885
Score = 96.7 bits (239), Expect = 2e-18
Identities = 75/285 (26%), Positives = 127/285 (44%), Gaps = 53/285 (18%)
Query: 384 KVLTCHFSSDGKVMASAGHEKKVFIWNMENF---QYFASADTHSHLITDVRFQPGSTMFA 440
++ + F DGK++AS + + +W++ + +Y + H++ + V F P A
Sbjct: 943 RIRSVAFHPDGKILASGSADNTIKLWDISDTNHSKYIRTLTGHTNWVWTVVFSPDKHTLA 1002
Query: 441 TSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQS 500
+SS DR++RLWD L L GH V ++ F P +L S ++ I++W+V
Sbjct: 1003 SSSEDRTIRLWDKDTG-DCLQKLKGHSHWVWTVAFSPDGR-ILASGSADSEIKIWDVASG 1060
Query: 501 ECMHTT---RGGSKQVRFQPQYGQLLATAIGN-SITIIDVETDSHLHDLKGHDKDVLSIC 556
+C+ T +G V F G LLA+A + ++ + +++T +H LKGH+K V S+
Sbjct: 1061 KCLQTLTDPQGMIWSVAFSLD-GTLLASASEDQTVKLWNLKTGECVHTLKGHEKQVYSVA 1119
Query: 557 WDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSL 615
+ +G AS SED + ++W S G C+ L
Sbjct: 1120 FSPNGQIAASGSEDTTVKLWD-ISTGSCVDTLKH-------------------------- 1152
Query: 616 EAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
H I +A SP G L+AS S D +++W
Sbjct: 1153 ---------------GHTAAIRSVAFSPDGRLLASGSEDEKIQLW 1182
Score = 91.7 bits (226), Expect = 6e-17
Identities = 61/201 (30%), Positives = 101/201 (49%), Gaps = 13/201 (6%)
Query: 383 NKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATS 442
N V T FS D +AS+ ++ + +W+ + HSH + V F P + A+
Sbjct: 987 NWVWTVVFSPDKHTLASSSEDRTIRLWDKDTGDCLQKLKGHSHWVWTVAFSPDGRILASG 1046
Query: 443 SFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMD--LLCSSDSNDVIRLWNVNQS 500
S D +++WD ++ + L LT + S+ F +D LL S+ + ++LWN+
Sbjct: 1047 SADSEIKIWDVASG-KCLQTLTDPQGMIWSVAF---SLDGTLLASASEDQTVKLWNLKTG 1102
Query: 501 ECMHTTRGGSKQ---VRFQPQYGQLLAT-AIGNSITIIDVETDSHLHDLK-GHDKDVLSI 555
EC+HT +G KQ V F P GQ+ A+ + ++ + D+ T S + LK GH + S+
Sbjct: 1103 ECVHTLKGHEKQVYSVAFSPN-GQIAASGSEDTTVKLWDISTGSCVDTLKHGHTAAIRSV 1161
Query: 556 CWDRSGNFLASVSED-SARIW 575
+ G LAS SED ++W
Sbjct: 1162 AFSPDGRLLASGSEDEKIQLW 1182
Score = 60.1 bits (144), Expect = 2e-07
Identities = 34/118 (28%), Positives = 58/118 (48%), Gaps = 1/118 (0%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG ++ASA ++ V +WN++ + + H + V F P + A+ S D +V+
Sbjct: 1078 FSLDGTLLASASEDQTVKLWNLKTGECVHTLKGHEKQVYSVAFSPNGQIAASGSEDTTVK 1137
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTR 507
LWD S + GH + S+ F P LL S ++ I+LW++ + T +
Sbjct: 1138 LWDISTGSCVDTLKHGHTAAIRSVAFSPDGR-LLASGSEDEKIQLWDMQNCSRLKTLK 1194
>UniRef100_Q7NJ67 WD-repeat protein [Gloeobacter violaceus]
Length = 1197
Score = 125 bits (313), Expect = 5e-27
Identities = 85/288 (29%), Positives = 137/288 (47%), Gaps = 7/288 (2%)
Query: 377 CLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGS 436
CL V + FS+DG+ +AS ++ V +W+ ++ F HS+ I+ V F P
Sbjct: 772 CLQGHTGWVRSVDFSADGRTLASGSDDQTVRLWDADSGLCFRVMHGHSNWISSVVFSPDG 831
Query: 437 TMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWN 496
+ + S D SVR+W+ S+ L +L GH + S+ F L S + V RLW+
Sbjct: 832 RLLTSGSVDHSVRIWEISSG-HCLRVLQGHGSGIWSVAFRGDGKTLASGSIDHSV-RLWD 889
Query: 497 VNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVL 553
+ + M + + + VR F P L ++ +I + D ++ L L+GH V
Sbjct: 890 FSTRQPMRSLQAHTSWVRTVAFSPDGTLLASSGQDRTIKLWDPDSGRCLKTLRGHTGWVN 949
Query: 554 SICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGY 612
S+ + +G LAS S D S RIW+ + G+C+G L + +S FHP R L
Sbjct: 950 SLAFSPNGALLASSSVDHSLRIWNVET-GQCLGMLQGHTSWVRSVAFHPDGRVLASASQD 1008
Query: 613 QSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
++ W G W++ H + +A P G +AS S D +VK+W
Sbjct: 1009 KTARLWDIETGRCLWTLQGHTSWVRSVAFHPDGHTLASGSDDGTVKLW 1056
Score = 119 bits (299), Expect = 2e-25
Identities = 85/289 (29%), Positives = 137/289 (46%), Gaps = 11/289 (3%)
Query: 378 LHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGST 437
+H N + + FS DG+++ S + V IW + + H I V F+
Sbjct: 815 MHGHSNWISSVVFSPDGRLLTSGSVDHSVRIWEISSGHCLRVLQGHGSGIWSVAFRGDGK 874
Query: 438 MFATSSFDRSVRLWDASN--PIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLW 495
A+ S D SVRLWD S P+RSL H V ++ F P LL SS + I+LW
Sbjct: 875 TLASGSIDHSVRLWDFSTRQPMRSL---QAHTSWVRTVAFSPDGT-LLASSGQDRTIKLW 930
Query: 496 NVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDV 552
+ + C+ T RG + V F P L ++++ +S+ I +VET L L+GH V
Sbjct: 931 DPDSGRCLKTLRGHTGWVNSLAFSPNGALLASSSVDHSLRIWNVETGQCLGMLQGHTSWV 990
Query: 553 LSICWDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGG 611
S+ + G LAS S+D +AR+W + G+C+ L + +S FHP +L
Sbjct: 991 RSVAFHPDGRVLASASQDKTARLWDIET-GRCLWTLQGHTSWVRSVAFHPDGHTLASGSD 1049
Query: 612 YQSLEAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+++ W G S++ H + + + G+ +AS D +V++W
Sbjct: 1050 DGTVKLWDVQTGRLADSLSGHGSGVWSVVFAADGKRLASGGDDKTVRLW 1098
Score = 117 bits (294), Expect = 8e-25
Identities = 80/275 (29%), Positives = 129/275 (46%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F DGK +AS + V +W+ Q S H+ + V F P T+ A+S DR+++
Sbjct: 869 FRGDGKTLASGSIDHSVRLWDFSTRQPMRSLQAHTSWVRTVAFSPDGTLLASSGQDRTIK 928
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LWD + R L L GH V SL F P LL SS + +R+WNV +C+ +G
Sbjct: 929 LWDPDSG-RCLKTLRGHTGWVNSLAFSPNGA-LLASSSVDHSLRIWNVETGQCLGMLQGH 986
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ VR F P L + + + + D+ET L L+GH V S+ + G+ LAS
Sbjct: 987 TSWVRSVAFHPDGRVLASASQDKTARLWDIETGRCLWTLQGHTSWVRSVAFHPDGHTLAS 1046
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S+D + ++W + G+ L G+ S +F + L G +++ W T
Sbjct: 1047 GSDDGTVKLWDVQT-GRLADSLSGHGSGVWSVVFAADGKRLASGGDDKTVRLWDTTSMQC 1105
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
T + H + +A ++AS+S D ++ +W
Sbjct: 1106 THVLNRHASGVLCVAIEADSRILASSSADETITLW 1140
Score = 113 bits (282), Expect = 2e-23
Identities = 81/286 (28%), Positives = 133/286 (46%), Gaps = 7/286 (2%)
Query: 380 KSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMF 439
++++ V + FS DG+++A++ + +W + Q A H+ + + F P +
Sbjct: 565 EALSTVSSVAFSPDGQLLATSEINGTIRLWQAADAQQLAYCRGHTSWVWSIAFSPDGRVL 624
Query: 440 ATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQ 499
A+ S DR+VRLWD + L + GH+ V S+ FHP +L S + +RLW V+
Sbjct: 625 ASGSADRTVRLWDYRTG-QCLKVFQGHEGWVRSVAFHPGG-GILASGSEDAAVRLWEVDS 682
Query: 500 SECMHTTRGGS---KQVRFQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSIC 556
C+ T RG S VRF P L +++ I + E+ L ++GH V SI
Sbjct: 683 GRCLLTLRGHSGWIHAVRFSPNGQWLASSSQDGKIQLWHPESGEPLQAMQGHTGWVRSIA 742
Query: 557 WDRSGNFLASVSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSL 615
+ G L S S+D + R+W G + L +S F R+L Q++
Sbjct: 743 FAPDGQTLISGSDDQTLRLWD-VQRGLLLKCLQGHTGWVRSVDFSADGRTLASGSDDQTV 801
Query: 616 EAWSPTEGSKTWSIAAHQGLIAGLADSPVGELIASASHDNSVKVWK 661
W G + H I+ + SP G L+ S S D+SV++W+
Sbjct: 802 RLWDADSGLCFRVMHGHSNWISSVVFSPDGRLLTSGSVDHSVRIWE 847
Score = 112 bits (279), Expect = 5e-23
Identities = 83/275 (30%), Positives = 128/275 (46%), Gaps = 7/275 (2%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
FS DG+V+AS ++ V +W+ Q H + V F PG + A+ S D +VR
Sbjct: 617 FSPDGRVLASGSADRTVRLWDYRTGQCLKVFQGHEGWVRSVAFHPGGGILASGSEDAAVR 676
Query: 450 LWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGG 509
LW+ + R L L GH + ++ F P L SS + I+LW+ E + +G
Sbjct: 677 LWEVDSG-RCLLTLRGHSGWIHAVRFSPNGQ-WLASSSQDGKIQLWHPESGEPLQAMQGH 734
Query: 510 SKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLAS 566
+ VR F P L++ + ++ + DV+ L L+GH V S+ + G LAS
Sbjct: 735 TGWVRSIAFAPDGQTLISGSDDQTLRLWDVQRGLLLKCLQGHTGWVRSVDFSADGRTLAS 794
Query: 567 VSED-SARIWSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSK 625
S+D + R+W A S G C +H N S +F P R L S+ W + G
Sbjct: 795 GSDDQTVRLWDADS-GLCFRVMHGHSNWISSVVFSPDGRLLTSGSVDHSVRIWEISSGHC 853
Query: 626 TWSIAAHQGLIAGLADSPVGELIASASHDNSVKVW 660
+ H I +A G+ +AS S D+SV++W
Sbjct: 854 LRVLQGHGSGIWSVAFRGDGKTLASGSIDHSVRLW 888
Score = 102 bits (253), Expect = 5e-20
Identities = 61/211 (28%), Positives = 105/211 (48%), Gaps = 6/211 (2%)
Query: 373 QEIGCLHKSINKVLTCHFSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRF 432
Q + L + V T FS DG ++AS+G ++ + +W+ ++ + + H+ + + F
Sbjct: 894 QPMRSLQAHTSWVRTVAFSPDGTLLASSGQDRTIKLWDPDSGRCLKTLRGHTGWVNSLAF 953
Query: 433 QPGSTMFATSSFDRSVRLWDASNPIRSLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVI 492
P + A+SS D S+R+W+ + LG+L GH V S+ FHP +L S+ +
Sbjct: 954 SPNGALLASSSVDHSLRIWNVETG-QCLGMLQGHTSWVRSVAFHPDGR-VLASASQDKTA 1011
Query: 493 RLWNVNQSECMHTTRGGSKQVR---FQPQYGQLLATAIGNSITIIDVETDSHLHDLKGHD 549
RLW++ C+ T +G + VR F P L + + ++ + DV+T L GH
Sbjct: 1012 RLWDIETGRCLWTLQGHTSWVRSVAFHPDGHTLASGSDDGTVKLWDVQTGRLADSLSGHG 1071
Query: 550 KDVLSICWDRSGNFLASVSED-SARIWSAAS 579
V S+ + G LAS +D + R+W S
Sbjct: 1072 SGVWSVVFAADGKRLASGGDDKTVRLWDTTS 1102
Score = 78.2 bits (191), Expect = 7e-13
Identities = 56/206 (27%), Positives = 92/206 (44%), Gaps = 6/206 (2%)
Query: 459 SLGILTGHDEQVMSLDFHPRRMDLLCSSDSNDVIRLWNVNQSECMHTTRGGSK---QVRF 515
S I T V S+ F P LL +S+ N IRLW ++ + RG + + F
Sbjct: 559 SQSIFTEALSTVSSVAFSPDGQ-LLATSEINGTIRLWQAADAQQLAYCRGHTSWVWSIAF 617
Query: 516 QPQYGQLLATAIGNSITIIDVETDSHLHDLKGHDKDVLSICWDRSGNFLASVSEDSA-RI 574
P L + + ++ + D T L +GH+ V S+ + G LAS SED+A R+
Sbjct: 618 SPDGRVLASGSADRTVRLWDYRTGQCLKVFQGHEGWVRSVAFHPGGGILASGSEDAAVRL 677
Query: 575 WSAASDGKCIGELHSVGNKFQSCIFHPGYRSLLIIGGYQSLEAWSPTEGSKTWSIAAHQG 634
W S G+C+ L + F P + L ++ W P G ++ H G
Sbjct: 678 WEVDS-GRCLLTLRGHSGWIHAVRFSPNGQWLASSSQDGKIQLWHPESGEPLQAMQGHTG 736
Query: 635 LIAGLADSPVGELIASASHDNSVKVW 660
+ +A +P G+ + S S D ++++W
Sbjct: 737 WVRSIAFAPDGQTLISGSDDQTLRLW 762
Score = 44.7 bits (104), Expect = 0.009
Identities = 21/63 (33%), Positives = 36/63 (56%)
Query: 390 FSSDGKVMASAGHEKKVFIWNMENFQYFASADTHSHLITDVRFQPGSTMFATSSFDRSVR 449
F++DGK +AS G +K V +W+ + Q + H+ + V + S + A+SS D ++
Sbjct: 1079 FAADGKRLASGGDDKTVRLWDTTSMQCTHVLNRHASGVLCVAIEADSRILASSSADETIT 1138
Query: 450 LWD 452
LWD
Sbjct: 1139 LWD 1141
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.130 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,178,610,023
Number of Sequences: 2790947
Number of extensions: 52031724
Number of successful extensions: 173345
Number of sequences better than 10.0: 4469
Number of HSP's better than 10.0 without gapping: 1706
Number of HSP's successfully gapped in prelim test: 2799
Number of HSP's that attempted gapping in prelim test: 144042
Number of HSP's gapped (non-prelim): 18495
length of query: 661
length of database: 848,049,833
effective HSP length: 134
effective length of query: 527
effective length of database: 474,062,935
effective search space: 249831166745
effective search space used: 249831166745
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0114.1


