

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0112.2
(150 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
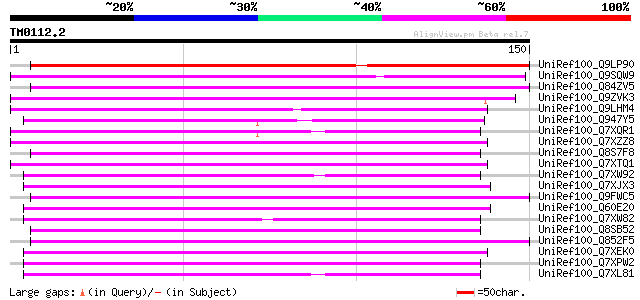
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana] 123 7e-28
UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidop... 110 8e-24
UniRef100_Q84ZV5 Polyprotein [Glycine max] 107 4e-23
UniRef100_Q9ZVK3 Putative retroelement pol polyprotein [Arabidop... 106 1e-22
UniRef100_Q9LHM4 Retroelement pol polyprotein [Arabidopsis thali... 102 1e-21
UniRef100_Q947Y5 Putative retroelement [Oryza sativa] 102 1e-21
UniRef100_Q7XQR1 OSJNBa0091D06.3 protein [Oryza sativa] 101 4e-21
UniRef100_Q7XZZ8 Putative polyprotein [Oryza sativa] 100 5e-21
UniRef100_Q8S7F8 Putative polyprotein [Oryza sativa] 100 7e-21
UniRef100_Q7XTQ1 OSJNBa0041A02.23 protein [Oryza sativa] 99 1e-20
UniRef100_Q7XW92 OSJNBb0043H09.2 protein [Oryza sativa] 99 1e-20
UniRef100_Q7XJX3 OSJNBa0004L19.16 protein [Oryza sativa] 99 2e-20
UniRef100_Q9FWC5 Putative plant disease resistance polyprotein [... 98 4e-20
UniRef100_Q60E20 Putative polyprotein [Oryza sativa] 98 4e-20
UniRef100_Q7XW82 OSJNBa0019J05.12 protein [Oryza sativa] 98 4e-20
UniRef100_Q8SB52 Putative polyprotein [Oryza sativa] 97 6e-20
UniRef100_Q852F5 Putative polyprotein [Oryza sativa] 97 9e-20
UniRef100_Q7XEK0 Hypothetical protein [Oryza sativa] 96 2e-19
UniRef100_Q7XPW2 OSJNBa0032F06.19 protein [Oryza sativa] 95 3e-19
UniRef100_Q7XL81 OSJNBb0014D23.8 protein [Oryza sativa] 95 3e-19
>UniRef100_Q9LP90 T32E20.30 [Arabidopsis thaliana]
Length = 1397
Score = 123 bits (309), Expect = 7e-28
Identities = 61/144 (42%), Positives = 91/144 (62%), Gaps = 3/144 (2%)
Query: 7 VFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 66
VF+KLR R+NSV R C +L A Y+GP+ I++RIG VAYRL+LPEG ++H VFH S LK
Sbjct: 1249 VFLKLRPYRQNSVTKRVCQKLAAKYFGPFEIMERIGKVAYRLKLPEGSKIHLVFHVSQLK 1308
Query: 67 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 126
+ +G++ + L + LT + V P +V+ +R+ E L + + WQG P E TW+
Sbjct: 1309 QVLGDHHQVIPLPEVLTADNEFVVVPEAVLETRY---NEDGLLEALVHWQGLPVHEDTWE 1365
Query: 127 DTLNIRSQFPVFNLEDKVDLSAGG 150
+++ QFP L+DK+ + GG
Sbjct: 1366 IAKDLKKQFPGLALQDKLHVEGGG 1389
>UniRef100_Q9SQW9 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1661
Score = 110 bits (274), Expect = 8e-24
Identities = 60/149 (40%), Positives = 85/149 (56%), Gaps = 2/149 (1%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE EWV++KLR R++SV R +L+ Y+GP+ ++ RIG VAY+LQLPE +HPVF
Sbjct: 1477 FEIDEWVYLKLRPYRQSSVAHRKNEKLSQRYFGPFKVLHRIGQVAYKLQLPEHSTIHPVF 1536
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H S LK AV + +L L+ + P ++ R + P+V +QW G
Sbjct: 1537 HVSQLKRAVPPSFTPQELPKILSPTLEWNTGPEKLLDIRQSNTNSG--PEVLVQWSGLST 1594
Query: 121 DEPTWKDTLNIRSQFPVFNLEDKVDLSAG 149
E TW+ L + Q+P F+LEDKV L G
Sbjct: 1595 LESTWEPLLTLVQQYPDFDLEDKVSLLRG 1623
>UniRef100_Q84ZV5 Polyprotein [Glycine max]
Length = 1552
Score = 107 bits (268), Expect = 4e-23
Identities = 53/144 (36%), Positives = 80/144 (54%)
Query: 7 VFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 66
V +KL+ R++S V R +L+ Y+GP+ ++ +IG VAY+L+LP R+HPVFH S LK
Sbjct: 1358 VLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLELPSAARIHPVFHVSQLK 1417
Query: 67 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 126
G L E + P ++ SR R + + Q+ +QW+ DE TW+
Sbjct: 1418 PFNGTAQDPYLPLPLTVTEMGPVMQPVKILASRIIIRGHNQIEQILVQWENGLQDEATWE 1477
Query: 127 DTLNIRSQFPVFNLEDKVDLSAGG 150
D +I++ +P FNLEDKV G
Sbjct: 1478 DIEDIKASYPTFNLEDKVVFKGEG 1501
>UniRef100_Q9ZVK3 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 945
Score = 106 bits (264), Expect = 1e-22
Identities = 51/148 (34%), Positives = 87/148 (58%), Gaps = 2/148 (1%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
F+ +WV ++++ R+ ++ R +L+ +YGP+ + + G VAYRL LPEG R+HPVF
Sbjct: 798 FQVGDWVLLRIQPYRQKTLFRRSSQKLSHRFYGPFQVASKHGEVAYRLTLPEGTRIHPVF 857
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H SLLK VG+ ++ L L + P +V+ R+ ++ + + + +QW+G
Sbjct: 858 HVSLLKPWVGDGEPDMGQLPPLRNNGELKLQPTAVLEVRWRSQDKKRVADLLVQWEGLHI 917
Query: 121 DEPTWKDTLNIRSQFP--VFNLEDKVDL 146
++ TW++ + + FP V NLEDKV L
Sbjct: 918 EDATWEEYDQLAASFPEFVLNLEDKVRL 945
>UniRef100_Q9LHM4 Retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1499
Score = 102 bits (255), Expect = 1e-21
Identities = 53/138 (38%), Positives = 80/138 (57%), Gaps = 2/138 (1%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE ++V++KL+ R+ SVV R +L+ Y+GPY II R G VAY+L LP +VHPVF
Sbjct: 1362 FEIGDYVYVKLQPYRQQSVVMRANQKLSPKYFGPYKIIDRCGEVAYKLALPSYSQVHPVF 1421
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H S LK VGN S + L + ++V P V+ + RQ + +V ++W +P
Sbjct: 1422 HVSQLKVLVGNVSTTVHLPSVM--QDVFEKVPEKVVERKMVNRQGKAVTKVLVKWSNEPL 1479
Query: 121 DEPTWKDTLNIRSQFPVF 138
+E TW+ +++ FP F
Sbjct: 1480 EEATWEFLFDLQKTFPEF 1497
>UniRef100_Q947Y5 Putative retroelement [Oryza sativa]
Length = 1923
Score = 102 bits (255), Expect = 1e-21
Identities = 49/134 (36%), Positives = 81/134 (59%), Gaps = 5/134 (3%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+WVF+KL+ + SV +R C +L +YGP+ I+ R+G VAY+LQLP+ +HPVFH S
Sbjct: 841 DWVFLKLQPYVQKSVASRACHKLAFKFYGPFQILARVGTVAYKLQLPDDSTIHPVFHVSQ 900
Query: 65 LKEAVG-NNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEP 123
LK A G + V+ +L L ++V+P ++ R + T+ QV + W A++
Sbjct: 901 LKVAHGFKHQVQSRLPKFLK----STVYPLQILDQRLIRKGNRTVSQVLVYWSDSVAEDA 956
Query: 124 TWKDTLNIRSQFPV 137
TW+D +++ +FP+
Sbjct: 957 TWEDREDLQQRFPM 970
>UniRef100_Q7XQR1 OSJNBa0091D06.3 protein [Oryza sativa]
Length = 753
Score = 101 bits (251), Expect = 4e-21
Identities = 54/137 (39%), Positives = 77/137 (55%), Gaps = 5/137 (3%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE +WV++KL+ + SV R +L Y+GPY + +IG VAY+LQLP +HPVF
Sbjct: 307 FEVGDWVYLKLQPYVQTSVAVRANHKLAFKYFGPYQVTAKIGTVAYQLQLPSSTSIHPVF 366
Query: 61 HASLLKEAVG-NNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKP 119
H S LK A+G V+ QL +V P ++ R T R S + QV IQW
Sbjct: 367 HVSQLKSAIGFTGPVQHQLPASTAPLQV----PLRILDRRLTKRGNSAVAQVLIQWSASV 422
Query: 120 ADEPTWKDTLNIRSQFP 136
++ TW+D ++RS+FP
Sbjct: 423 PEDATWEDLDDLRSRFP 439
>UniRef100_Q7XZZ8 Putative polyprotein [Oryza sativa]
Length = 1246
Score = 100 bits (250), Expect = 5e-21
Identities = 48/138 (34%), Positives = 79/138 (56%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
FE + V++K++ R+N+ R +L A +YGP+ ++++IG VAY+LQLPE +HPVF
Sbjct: 1107 FELGDMVYLKMQPYRQNAFGLRGSLKLRAKFYGPFRVLEKIGKVAYKLQLPEEANIHPVF 1166
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H S LK+ VG ++ L L + + P +V+ R RQ + Q I W+
Sbjct: 1167 HVSQLKKHVGRQAIPLPNLPLVGADGTVKTEPLAVLARRVIPRQNEPVVQWLIHWENLTP 1226
Query: 121 DEPTWKDTLNIRSQFPVF 138
++ TW++ I++ FP F
Sbjct: 1227 EDATWENATFIQATFPHF 1244
>UniRef100_Q8S7F8 Putative polyprotein [Oryza sativa]
Length = 408
Score = 100 bits (249), Expect = 7e-21
Identities = 49/130 (37%), Positives = 77/130 (58%)
Query: 7 VFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 66
VF+KL+ ++SVV R P+L ++GP+ I+++IG AYRLQLP+ +HPVFH S LK
Sbjct: 257 VFLKLQPYTQSSVVNRPYPKLAFKFFGPFEILEKIGQSAYRLQLPDSSLIHPVFHVSQLK 316
Query: 67 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 126
V +++ L H + A V P ++ R + +T QV I+W PA + TW+
Sbjct: 317 AHVPDHTPVFTSLPHPVVLDAADVWPEEILDRRLVKKGNATHVQVLIKWSSLPAHQATWE 376
Query: 127 DTLNIRSQFP 136
D ++++FP
Sbjct: 377 DYEVLKTRFP 386
>UniRef100_Q7XTQ1 OSJNBa0041A02.23 protein [Oryza sativa]
Length = 1373
Score = 99.4 bits (246), Expect = 1e-20
Identities = 50/138 (36%), Positives = 79/138 (57%)
Query: 1 FEGFEWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVF 60
F+ + V++K++ R+N+ R +L A +YGP+ I++RIG VAY LQLPE +HPVF
Sbjct: 1234 FQEGDMVYLKMQPYRQNAFGLRGSLKLRAKFYGPFRILKRIGKVAYHLQLPEEAGIHPVF 1293
Query: 61 HASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPA 120
H S LK+ VGN V L L + + P +V+ R R+ + Q + W+
Sbjct: 1294 HVSQLKKHVGNLVVPLPNLPLVDEDGNIKTEPVAVLARRVVPRRNEPVTQWLVHWENLSP 1353
Query: 121 DEPTWKDTLNIRSQFPVF 138
++ TW+D+ I++ FP F
Sbjct: 1354 EDATWEDSAFIQATFPHF 1371
>UniRef100_Q7XW92 OSJNBb0043H09.2 protein [Oryza sativa]
Length = 1409
Score = 99.4 bits (246), Expect = 1e-20
Identities = 49/132 (37%), Positives = 73/132 (55%), Gaps = 3/132 (2%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+WV++KL+ + SV R +L Y+GP+ ++ R+GAVAYRLQLP +HPVFH S
Sbjct: 1211 DWVYLKLQPYIQTSVTKRANHKLEFRYFGPFQVLDRVGAVAYRLQLPSYSAIHPVFHVSQ 1270
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
L+ A G + + L GE P + R T + T+ Q+ W PA+E T
Sbjct: 1271 LRAAAGFSKPVMSQLPSSLGELQI---PLQFLDKRLTKKGNKTVVQLLTHWSDSPAEEAT 1327
Query: 125 WKDTLNIRSQFP 136
W+D +R++FP
Sbjct: 1328 WEDMEELRARFP 1339
>UniRef100_Q7XJX3 OSJNBa0004L19.16 protein [Oryza sativa]
Length = 1586
Score = 99.0 bits (245), Expect = 2e-20
Identities = 47/135 (34%), Positives = 76/135 (55%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+ V++KL+ R+ + R +L + +YGP+ I++++G VAY+LQLPEG +HPVFH S
Sbjct: 1451 DMVYLKLQPYRQTAFGIRGSLKLRSKFYGPFKIMEKVGRVAYKLQLPEGSNIHPVFHVSQ 1510
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
LK+ +G+ +V + L + + P +V+ R R + Q + W E T
Sbjct: 1511 LKKHIGSRAVPMANLPSVGPDGQIKTEPVAVLKRRMIPRGGVAVTQWLVLWHNLSPSEAT 1570
Query: 125 WKDTLNIRSQFPVFN 139
W+D I+S FP FN
Sbjct: 1571 WEDASMIQSMFPSFN 1585
>UniRef100_Q9FWC5 Putative plant disease resistance polyprotein [Oryza sativa]
Length = 921
Score = 97.8 bits (242), Expect = 4e-20
Identities = 51/144 (35%), Positives = 79/144 (54%)
Query: 7 VFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 66
VF+KL+ ++SVV R P+L ++GP+ +++RIG AYRL LP VHPVFH S LK
Sbjct: 765 VFLKLQPYAQSSVVNRPYPKLAYKFFGPFTVLERIGVAAYRLDLPASSTVHPVFHVSQLK 824
Query: 67 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 126
V +++ L + A V P +++ R R + QV ++W PAD TW+
Sbjct: 825 AQVPDHTPVYTSLPSPVVLDAADVVPEAILERRLVKRGNAAHVQVLLKWSLLPADHATWE 884
Query: 127 DTLNIRSQFPVFNLEDKVDLSAGG 150
D ++++FP + + AGG
Sbjct: 885 DYHVVKTRFPDAPAWGQAETPAGG 908
>UniRef100_Q60E20 Putative polyprotein [Oryza sativa]
Length = 1475
Score = 97.8 bits (242), Expect = 4e-20
Identities = 45/135 (33%), Positives = 79/135 (58%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+ V++KL+ R+++ R +L + +YGP+ ++Q+IG +AY+LQLP+ ++HPVFH S
Sbjct: 1340 DMVYLKLKPYRQSAFGIRGSLKLRSKFYGPFKVLQKIGQLAYKLQLPDDAQIHPVFHVSQ 1399
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
LK+ +G +++ + L + + P +V+ R R+ + Q I WQ E T
Sbjct: 1400 LKKHLGKHAIPMSNLPSVGPDGQIKTEPLAVLQRRMVPRKGVAVTQWLILWQNLSPAEAT 1459
Query: 125 WKDTLNIRSQFPVFN 139
W+D I++ FP FN
Sbjct: 1460 WEDASVIQAMFPSFN 1474
>UniRef100_Q7XW82 OSJNBa0019J05.12 protein [Oryza sativa]
Length = 1817
Score = 97.8 bits (242), Expect = 4e-20
Identities = 48/132 (36%), Positives = 72/132 (54%), Gaps = 3/132 (2%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+WV++KL+ + SV R +L Y+GP+ I +IG VAY+LQLP +HPVFH S
Sbjct: 1475 DWVYLKLQPYVQTSVAVRANHKLAFKYFGPFQITAQIGTVAYKLQLPPTSSIHPVFHVSQ 1534
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
LK A G + HL A P+ ++ R + ST+ QV +QW P ++ T
Sbjct: 1535 LKSATGFTG---HVQSHLPSSISALQVPWRILDRRLAKKGNSTISQVLVQWSESPPEDAT 1591
Query: 125 WKDTLNIRSQFP 136
W+D + ++FP
Sbjct: 1592 WEDYEELHARFP 1603
>UniRef100_Q8SB52 Putative polyprotein [Oryza sativa]
Length = 999
Score = 97.4 bits (241), Expect = 6e-20
Identities = 45/130 (34%), Positives = 77/130 (58%)
Query: 7 VFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 66
V++KL+ + S+ R C +++ Y+GPY +++R+G VAYRL LP G ++HPV H SLLK
Sbjct: 860 VYLKLQPFAQLSIAQRSCQKISFRYFGPYTVLERVGQVAYRLDLPAGSQIHPVVHVSLLK 919
Query: 67 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 126
AV +++ L ++ + P V+ R ++ +P IQW G PA+ TW+
Sbjct: 920 AAVKPDTIVSPELPLQITDDSVLLQPEVVLRRSLIKRGKAAVPYGLIQWTGVPAELATWE 979
Query: 127 DTLNIRSQFP 136
+ +++ +FP
Sbjct: 980 NLRHLKLRFP 989
>UniRef100_Q852F5 Putative polyprotein [Oryza sativa]
Length = 1155
Score = 96.7 bits (239), Expect = 9e-20
Identities = 51/144 (35%), Positives = 78/144 (53%)
Query: 7 VFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 66
VF+KL+ ++SVV R P+L ++GP+ +++RIG AYRL LP VHPVFH S LK
Sbjct: 999 VFLKLQPYAQSSVVNRPYPKLAYKFFGPFTVLERIGVAAYRLDLPASSTVHPVFHVSQLK 1058
Query: 67 EAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWK 126
V +++ L + A V P +++ R R + QV ++W PAD TW+
Sbjct: 1059 AQVPDHTPVYTSLPSPVVLDAADVVPEAILERRLVKRGNAAHVQVLLKWSLLPADHATWE 1118
Query: 127 DTLNIRSQFPVFNLEDKVDLSAGG 150
D ++++FP + AGG
Sbjct: 1119 DYHVVKTRFPDAPAWGQAKTPAGG 1142
>UniRef100_Q7XEK0 Hypothetical protein [Oryza sativa]
Length = 1611
Score = 95.5 bits (236), Expect = 2e-19
Identities = 46/134 (34%), Positives = 77/134 (57%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+ V++KL+ R+ ++ R +L + YYGP+ +I+++GAVAY+LQLP+G +HPVFH S
Sbjct: 1476 DMVYLKLQPYRQVAMGIRGSLKLRSKYYGPFKVIEKMGAVAYKLQLPDGAGIHPVFHVSQ 1535
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
LK+ +G ++ + L + + P +V+ R R + Q I W+ E T
Sbjct: 1536 LKKHLGARAIPMPNLPAIGPDGQIKTEPAAVLQRRMIPRHNEPVTQWLILWENLTPAEAT 1595
Query: 125 WKDTLNIRSQFPVF 138
W+D I++ FP F
Sbjct: 1596 WEDASYIQAAFPNF 1609
>UniRef100_Q7XPW2 OSJNBa0032F06.19 protein [Oryza sativa]
Length = 1575
Score = 95.1 bits (235), Expect = 3e-19
Identities = 48/132 (36%), Positives = 73/132 (54%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+WVF+KL+ + SV+TR +L+ YYGP+ ++QRIG VAYRLQLP +HPV H S
Sbjct: 1391 DWVFMKLQPYIQQSVMTRSNRKLSFKYYGPFQVLQRIGLVAYRLQLPAHSLIHPVVHVSQ 1450
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
LK+A+ + L L + P ++ R R + QV I+W G + T
Sbjct: 1451 LKKALAPSEAVQPSLPVLNPDAGHLAIPIKILDHRSIRRGSKMIKQVLIRWSGVSSSADT 1510
Query: 125 WKDTLNIRSQFP 136
W+ ++++FP
Sbjct: 1511 WESLDEMQAKFP 1522
>UniRef100_Q7XL81 OSJNBb0014D23.8 protein [Oryza sativa]
Length = 1545
Score = 95.1 bits (235), Expect = 3e-19
Identities = 50/132 (37%), Positives = 76/132 (56%), Gaps = 4/132 (3%)
Query: 5 EWVFIKLRALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASL 64
+WV++KL+ ++SV R +L Y+GP+ I+ ++G VAY+LQLP VHPVFH S
Sbjct: 848 DWVYLKLQPYIQSSVARRANHKLCFKYFGPFQILHKVGKVAYQLQLPANSAVHPVFHVSQ 907
Query: 65 LKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEPT 124
LK AVG L L+ +V P SV+ SR + ++L QV ++W E T
Sbjct: 908 LKHAVGFKHQVQPRLPTLSTFQV----PVSVLDSRQIRKGNTSLTQVLVRWSDDGYGEET 963
Query: 125 WKDTLNIRSQFP 136
W+D + ++ +FP
Sbjct: 964 WEDLVELQQRFP 975
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 264,432,732
Number of Sequences: 2790947
Number of extensions: 10326290
Number of successful extensions: 23073
Number of sequences better than 10.0: 587
Number of HSP's better than 10.0 without gapping: 425
Number of HSP's successfully gapped in prelim test: 162
Number of HSP's that attempted gapping in prelim test: 22169
Number of HSP's gapped (non-prelim): 598
length of query: 150
length of database: 848,049,833
effective HSP length: 126
effective length of query: 24
effective length of database: 496,390,511
effective search space: 11913372264
effective search space used: 11913372264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0112.2


