

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.7
(402 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
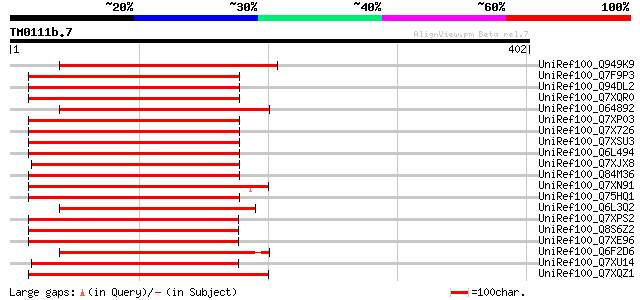
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q949K9 Putative polyprotein [Cicer arietinum] 236 1e-60
UniRef100_Q7F9P3 OSJNBa0049H08.6 protein [Oryza sativa] 194 4e-48
UniRef100_Q94DL2 Putative polyprotein [Oryza sativa] 194 5e-48
UniRef100_Q7XQR0 OSJNBa0091D06.9 protein [Oryza sativa] 193 6e-48
UniRef100_O64892 Polyprotein [Ananas comosus] 193 8e-48
UniRef100_Q7XP03 OSJNBb0013J13.17 protein [Oryza sativa] 192 1e-47
UniRef100_Q7X726 OSJNBa0089N06.1 protein [Oryza sativa] 191 2e-47
UniRef100_Q7XSU3 OSJNBa0039K24.13 protein [Oryza sativa] 191 2e-47
UniRef100_Q6L494 Putative polyprotein [Oryza sativa] 191 2e-47
UniRef100_Q7XJX8 OSJNBa0063C18.15 protein [Oryza sativa] 191 3e-47
UniRef100_Q84M36 Putative polyprotein [Oryza sativa] 191 3e-47
UniRef100_Q7XN91 OSJNBa0011F23.1 protein [Oryza sativa] 191 3e-47
UniRef100_Q75HQ1 Putative polyprotein [Oryza sativa] 191 4e-47
UniRef100_Q6L3Q2 Putative polyprotein, 3'-partial [Solanum demis... 191 4e-47
UniRef100_Q7XPS2 OSJNBa0065O17.9 protein [Oryza sativa] 190 7e-47
UniRef100_Q8S6Z2 Putative polyprotein [Oryza sativa] 189 9e-47
UniRef100_Q7XE96 Putative gag-pol protein [Oryza sativa] 189 9e-47
UniRef100_Q6F2D6 Putative polyprotein [Solanum demissum] 189 1e-46
UniRef100_Q7XU14 OSJNBa0091D06.10 protein [Oryza sativa] 187 3e-46
UniRef100_Q7XQZ1 OSJNBb0045P24.14 protein [Oryza sativa] 187 3e-46
>UniRef100_Q949K9 Putative polyprotein [Cicer arietinum]
Length = 655
Score = 236 bits (601), Expect = 1e-60
Identities = 115/170 (67%), Positives = 140/170 (81%), Gaps = 1/170 (0%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
+FQ +K++LT +PVL LP +EP+EVYC+AS QGLGCVLMQH++ +AYAS QLK+HERNY
Sbjct: 93 NFQLLKRKLTTSPVLVLPQSEEPHEVYCDASHQGLGCVLMQHKKVVAYASRQLKIHERNY 152
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
THDLELA VVFALKIWRHYLYG TFTVFS+HK LKYLFD K+LNM QR W+ ++D+DF
Sbjct: 153 PTHDLELAAVVFALKIWRHYLYGCTFTVFSDHKSLKYLFDQKELNMRQRRWIETLKDFDF 212
Query: 159 TLLYHPGKANVVADALSHQAIHVSSL-MVKKLELIETFRDLSLDAKIKLG 207
TL YHPGKANVVADALS +++ VSSL M ++ EL E FRDL L+ + G
Sbjct: 213 TLQYHPGKANVVADALSRRSVSVSSLIMARQQELWEAFRDLHLNVEFAPG 262
>UniRef100_Q7F9P3 OSJNBa0049H08.6 protein [Oryza sativa]
Length = 1523
Score = 194 bits (493), Expect = 4e-48
Identities = 96/164 (58%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E FH SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1034 KLARPMTELLKKEKKFHWSAACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1093
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK K
Sbjct: 1094 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSFK 1153
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1154 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 1197
>UniRef100_Q94DL2 Putative polyprotein [Oryza sativa]
Length = 1572
Score = 194 bits (492), Expect = 5e-48
Identities = 95/164 (57%), Positives = 119/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQE+KKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 895 KLARPMTELLKKEKKFQWSAACEDSFQELKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 954
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ ++ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 955 CVLMQEQKVVAYASRQLRPHEANYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1014
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALSH+A
Sbjct: 1015 YIFTQTELNMRQRKWLELIKDYDLGIHYHPGKANVVADALSHKA 1058
>UniRef100_Q7XQR0 OSJNBa0091D06.9 protein [Oryza sativa]
Length = 1762
Score = 193 bits (491), Expect = 6e-48
Identities = 96/164 (58%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1085 KLARPMTELLKKEKKFQWSAACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1144
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1145 CVLMQERKVVAYASRQLRPHEANYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1204
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1205 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 1248
>UniRef100_O64892 Polyprotein [Ananas comosus]
Length = 871
Score = 193 bits (490), Expect = 8e-48
Identities = 96/164 (58%), Positives = 125/164 (75%), Gaps = 1/164 (0%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
SFQE+K+RLT AP+LTLP+ Y VY +AS GLGCVLMQ + +AYAS QLK +E+NY
Sbjct: 290 SFQELKQRLTTAPILTLPVAGAGYVVYSDASLNGLGCVLMQDDKVIAYASRQLKEYEKNY 349
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
THDLELA VVFALK+WRHYLYG V+++HK LKYLF K+LN+ QR W+ ++DYD
Sbjct: 350 PTHDLELAAVVFALKLWRHYLYGERCEVYTDHKSLKYLFTQKELNLRQRRWLELLKDYDL 409
Query: 159 TLLYHPGKANVVADALSHQAI-HVSSLMVKKLELIETFRDLSLD 201
T+LYHPGKANVVADALS +++ +++ +V + LIE + L L+
Sbjct: 410 TILYHPGKANVVADALSRKSMENLAMHVVTQPRLIEQMKRLELE 453
>UniRef100_Q7XP03 OSJNBb0013J13.17 protein [Oryza sativa]
Length = 864
Score = 192 bits (489), Expect = 1e-47
Identities = 96/164 (58%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 337 KLARPMTELLKKEKKFQWSAACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 396
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 397 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 456
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 457 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 500
>UniRef100_Q7X726 OSJNBa0089N06.1 protein [Oryza sativa]
Length = 1851
Score = 191 bits (486), Expect = 2e-47
Identities = 95/164 (57%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1174 KLARPMTELLKKEKKFQWSTACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1233
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA V+ ALKIWRHYL G+ V+++HK LK
Sbjct: 1234 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVIHALKIWRHYLIGNRCEVYTDHKSLK 1293
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1294 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 1337
>UniRef100_Q7XSU3 OSJNBa0039K24.13 protein [Oryza sativa]
Length = 1851
Score = 191 bits (486), Expect = 2e-47
Identities = 96/164 (58%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1174 KLARPMTELLKKEKKFQWSVACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1233
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1234 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1293
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1294 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 1337
>UniRef100_Q6L494 Putative polyprotein [Oryza sativa]
Length = 1688
Score = 191 bits (486), Expect = 2e-47
Identities = 95/164 (57%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1011 KLARPMTELLKKEKKFQWSTACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1070
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA V+ ALKIWRHYL G+ V+++HK LK
Sbjct: 1071 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVIHALKIWRHYLIGNRCEVYTDHKSLK 1130
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1131 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 1174
>UniRef100_Q7XJX8 OSJNBa0063C18.15 protein [Oryza sativa]
Length = 1764
Score = 191 bits (485), Expect = 3e-47
Identities = 94/161 (58%), Positives = 117/161 (72%)
Query: 18 RTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVL 77
R + + +E F SFQEMKKRLT APVLTLP ++ ++++C+AS QGLGCVL
Sbjct: 1134 RPMTELLKKEKKFQWSAACEDSFQEMKKRLTTAPVLTLPDIRKDFKIFCDASRQGLGCVL 1193
Query: 78 MQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLF 137
MQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LKY+F
Sbjct: 1194 MQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLKYIF 1253
Query: 138 D*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
+LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1254 TQTELNMRQRRWLELIKDYDLRIHYHPGKANVVADALSRKA 1294
>UniRef100_Q84M36 Putative polyprotein [Oryza sativa]
Length = 1382
Score = 191 bits (485), Expect = 3e-47
Identities = 95/164 (57%), Positives = 117/164 (70%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 731 KLARPMTELLKKERKFQWSAACEDSFQEMKKRLTTAPVLTLPDIQKDFEIFCDASRQGLG 790
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA V+ ALKIWRHYL G V+++HK LK
Sbjct: 791 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVIHALKIWRHYLIGKRCEVYTDHKSLK 850
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 851 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 894
>UniRef100_Q7XN91 OSJNBa0011F23.1 protein [Oryza sativa]
Length = 1787
Score = 191 bits (485), Expect = 3e-47
Identities = 103/196 (52%), Positives = 130/196 (65%), Gaps = 10/196 (5%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + + F SFQEMKKRLT APVLTLP ++ +E++C+AS QG+G
Sbjct: 1174 KLARPMSELLKKGKKFQWSAACEHSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGVG 1233
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1234 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1293
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA-IHVSSLM-------- 185
Y+F +LNM QR W+ IEDYD + YHPGKANVVADALS +A +V+ +
Sbjct: 1294 YIFTQTELNMRQRRWLELIEDYDLGIHYHPGKANVVADALSRKAYCNVAQIRPDQDRLCR 1353
Query: 186 -VKKLELIETFRDLSL 200
++KL L RDL L
Sbjct: 1354 ELEKLRLTMELRDLIL 1369
>UniRef100_Q75HQ1 Putative polyprotein [Oryza sativa]
Length = 1649
Score = 191 bits (484), Expect = 4e-47
Identities = 94/164 (57%), Positives = 118/164 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQE+KKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 971 KLARPMTELLKKEKKFQWSAACEDSFQELKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1030
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ ++ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1031 CVLMQEQKVVAYASRQLRPHEANYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1090
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA 178
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +A
Sbjct: 1091 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRKA 1134
>UniRef100_Q6L3Q2 Putative polyprotein, 3'-partial [Solanum demissum]
Length = 1622
Score = 191 bits (484), Expect = 4e-47
Identities = 90/152 (59%), Positives = 115/152 (75%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
SF +K+ LT AP+LTLP+E E + VYC+AS GLGCVLMQ +AYAS QLK+HERNY
Sbjct: 1084 SFLRLKELLTTAPILTLPVEGEGFTVYCDASGVGLGCVLMQQDRVIAYASRQLKIHERNY 1143
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
THDLELA VVFALKIWRHYLYG ++++H+ L+Y+ +DLN QR W+ ++DYD
Sbjct: 1144 PTHDLELAAVVFALKIWRHYLYGVRCEIYTDHRSLQYIMSQRDLNSRQRRWIELLKDYDL 1203
Query: 159 TLLYHPGKANVVADALSHQAIHVSSLMVKKLE 190
++LYHPGKANVVADALS +A+ + SL +E
Sbjct: 1204 SILYHPGKANVVADALSRKAVSMGSLAFLSVE 1235
>UniRef100_Q7XPS2 OSJNBa0065O17.9 protein [Oryza sativa]
Length = 1851
Score = 190 bits (482), Expect = 7e-47
Identities = 95/163 (58%), Positives = 117/163 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1174 KLARPMTELLKKEKKFQWSVACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1233
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1234 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1293
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQ 177
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +
Sbjct: 1294 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRK 1336
>UniRef100_Q8S6Z2 Putative polyprotein [Oryza sativa]
Length = 1658
Score = 189 bits (481), Expect = 9e-47
Identities = 94/163 (57%), Positives = 116/163 (70%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1134 KLARPMTELLKKEKKFEWSAACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1193
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1194 CVLMQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1253
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQ 177
Y+F +LNM QR W+ I+DYD + YHPGKANVVAD LS +
Sbjct: 1254 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADTLSRK 1296
>UniRef100_Q7XE96 Putative gag-pol protein [Oryza sativa]
Length = 1829
Score = 189 bits (481), Expect = 9e-47
Identities = 94/163 (57%), Positives = 117/163 (71%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E F SFQEMKKRLT APVLTLP ++ +E++C+AS QGLG
Sbjct: 1174 KLARPMTELLKKEKKFRWSAACEDSFQEMKKRLTTAPVLTLPDIRKDFEIFCDASRQGLG 1233
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVL+Q R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LK
Sbjct: 1234 CVLIQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLK 1293
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQ 177
Y+F +LNM QR W+ I+DYD + YHPGKANVVADALS +
Sbjct: 1294 YIFTQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRK 1336
>UniRef100_Q6F2D6 Putative polyprotein [Solanum demissum]
Length = 1769
Score = 189 bits (480), Expect = 1e-46
Identities = 93/163 (57%), Positives = 119/163 (72%), Gaps = 4/163 (2%)
Query: 39 SFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVLMQHREAMAYASSQLKLHERNY 98
SF +K+ LT AP+LTLP+E E + VYC+AS GLGCVLMQ +AYAS QLK+HE NY
Sbjct: 1103 SFLRLKELLTTAPILTLPVEGEGFTVYCDASGVGLGCVLMQQGRVIAYASRQLKIHEHNY 1162
Query: 99 LTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLFD*KDLNMGQRCWMNFIEDYDF 158
THDLELA VVFALKIWRHYLYG ++++H+ L+Y+ +DLN QR W+ ++DYD
Sbjct: 1163 PTHDLELAAVVFALKIWRHYLYGVRCEIYTDHRSLQYIMSQRDLNSRQRRWIELLKDYDL 1222
Query: 159 TLLYHPGKANVVADALSHQAIHVSSLMVKKLELIETFRDLSLD 201
++LYHPGKANVVADALS +A+ + SL +E R L+LD
Sbjct: 1223 SILYHPGKANVVADALSRKAVSMGSLAFLSVE----ERPLALD 1261
>UniRef100_Q7XU14 OSJNBa0091D06.10 protein [Oryza sativa]
Length = 1831
Score = 187 bits (476), Expect = 3e-46
Identities = 93/160 (58%), Positives = 115/160 (71%)
Query: 18 RTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLGCVL 77
R + + +E F SFQEMKKRLT A VLTLP ++ +E++C+AS QGLGCVL
Sbjct: 1179 RPMTELLKKEKKFQWSAACEDSFQEMKKRLTTASVLTLPDIRKDFEIFCDASRQGLGCVL 1238
Query: 78 MQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLKYLF 137
MQ R+ +AYAS QL+ HE NY THDLELA VV ALKIWRHYL G+ V+++HK LKY+F
Sbjct: 1239 MQERKVVAYASRQLRPHEVNYPTHDLELAAVVHALKIWRHYLIGNRCEVYTDHKSLKYIF 1298
Query: 138 D*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQ 177
+LNM QR W+ I+DYD + YHPGKANVVADALS +
Sbjct: 1299 TQTELNMRQRRWLELIKDYDLGIHYHPGKANVVADALSRK 1338
>UniRef100_Q7XQZ1 OSJNBb0045P24.14 protein [Oryza sativa]
Length = 1195
Score = 187 bits (476), Expect = 3e-46
Identities = 96/187 (51%), Positives = 126/187 (67%), Gaps = 1/187 (0%)
Query: 15 KDCRTVDKTHSEESTFHMDGGMRTSFQEMKKRLTAAPVLTLPLEKEPYEVYCNAS*QGLG 74
K R + + +E + G SFQE+KKRL APVL LP ++ ++VYC+AS GLG
Sbjct: 733 KIARPMTRLLQKEVKYKWTEGCEQSFQELKKRLVTAPVLVLPDSRKGFQVYCDASRHGLG 792
Query: 75 CVLMQHREAMAYASSQLKLHERNYLTHDLELAVVVFALKIWRHYLYGSTFTVFSNHKGLK 134
CVLMQ + +AYAS QL+ HE NY THDLELA VV ALKIWRHYL+G+ ++++HK LK
Sbjct: 793 CVLMQEGKVVAYASRQLRPHENNYPTHDLELAAVVHALKIWRHYLFGNRTEIYTDHKSLK 852
Query: 135 YLFD*KDLNMGQRCWMNFIEDYDFTLLYHPGKANVVADALSHQA-IHVSSLMVKKLELIE 193
Y+F DLNM QR W+ I+DYD + YHPGKANVVADALS ++ ++S EL +
Sbjct: 853 YIFTQPDLNMRQRRWLELIKDYDMEIHYHPGKANVVADALSRKSYCNMSEGRRLPWELCQ 912
Query: 194 TFRDLSL 200
F ++L
Sbjct: 913 EFEKMNL 919
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.353 0.155 0.544
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 554,162,517
Number of Sequences: 2790947
Number of extensions: 19892437
Number of successful extensions: 91118
Number of sequences better than 10.0: 1405
Number of HSP's better than 10.0 without gapping: 1183
Number of HSP's successfully gapped in prelim test: 222
Number of HSP's that attempted gapping in prelim test: 88735
Number of HSP's gapped (non-prelim): 1763
length of query: 402
length of database: 848,049,833
effective HSP length: 130
effective length of query: 272
effective length of database: 485,226,723
effective search space: 131981668656
effective search space used: 131981668656
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0111b.7


