

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111b.4
(467 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
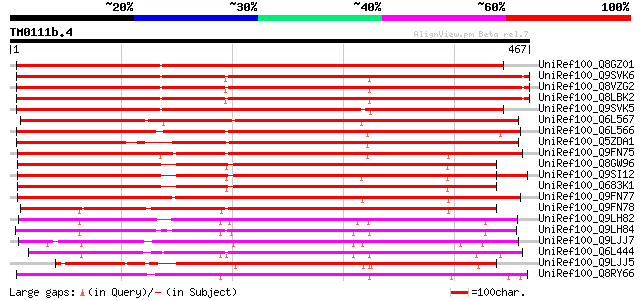
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GZ01 Putative BCS1 protein [Arabidopsis thaliana] 475 e-132
UniRef100_Q9SVK6 Putative mitochondrial protein [Arabidopsis tha... 464 e-129
UniRef100_Q8VZG2 AT3g50930/F18B3_210 [Arabidopsis thaliana] 464 e-129
UniRef100_Q8LBK2 BCS1 protein-like protein [Arabidopsis thaliana] 464 e-129
UniRef100_Q9SVK5 Putative mitochondrial protein [Arabidopsis tha... 464 e-129
UniRef100_Q6L567 Putative AAA-type ATPase [Oryza sativa] 413 e-114
UniRef100_Q6L566 Putative AAA-type ATPase [Oryza sativa] 410 e-113
UniRef100_Q5ZDA1 BCS1 protein-like [Oryza sativa] 393 e-108
UniRef100_Q9FN75 AAA-type ATPase-like protein [Arabidopsis thali... 384 e-105
UniRef100_Q8GW96 AAA-type ATPase like protein [Arabidopsis thali... 377 e-103
UniRef100_Q9SI12 Putative AAA-type ATPase [Arabidopsis thaliana] 377 e-103
UniRef100_Q683K1 AAA-type ATPase like protein [Arabidopsis thali... 375 e-102
UniRef100_Q9FN77 AAA-type ATPase-like protein [Arabidopsis thali... 355 2e-96
UniRef100_Q9FN78 AAA-type ATPase-like protein [Arabidopsis thali... 353 8e-96
UniRef100_Q9LH82 Similarity to AAA-type ATPase [Arabidopsis thal... 348 1e-94
UniRef100_Q9LH84 Similarity to AAA-type ATPase [Arabidopsis thal... 337 6e-91
UniRef100_Q9LJJ7 Mitochondrial protein-like [Arabidopsis thaliana] 331 3e-89
UniRef100_Q6L444 Putative ATPase protein [Solanum demissum] 329 9e-89
UniRef100_Q9LJJ5 Mitochondrial protein-like [Arabidopsis thaliana] 328 2e-88
UniRef100_Q8RY66 AT4g25830/F14M19_110 [Arabidopsis thaliana] 328 3e-88
>UniRef100_Q8GZ01 Putative BCS1 protein [Arabidopsis thaliana]
Length = 451
Score = 475 bits (1223), Expect = e-132
Identities = 243/438 (55%), Positives = 318/438 (72%), Gaps = 1/438 (0%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K+ L+A AS+AA A+L RS+ D++P+E+ ++I+ F FS Q T VIEEF G
Sbjct: 13 KTALTAVASVAAAAILARSVVQDYMPNEVHEYISHGFRRFFSYFSYQMTAVIEEFGGFEH 72
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
N+V+EAAEAYLSTK + S +R+K K E S +RDEEV D F+GV+++W L C
Sbjct: 73 NQVFEAAEAYLSTKISNSTRRIKVNKLEKQSNYSVTVERDEEVVDIFDGVKLSWILVCRH 132
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
VD R +++S+L SEVRSYELSF KK K + SYLP+V+E+A +IKQ+ +KI
Sbjct: 133 VDKKDFRNP-RDLNSTLKSEVRSYELSFRKKFKNMVLESYLPFVVEQAASIKQKFKTLKI 191
Query: 187 YTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYL 246
+TV+ + DHP TF+TLA+D +KK LV DLDRF+Q K FY R GKAWKRGYL
Sbjct: 192 FTVDSYSVEWTSVTLDHPSTFRTLALDPEVKKNLVEDLDRFVQRKGFYGRVGKAWKRGYL 251
Query: 247 LYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSI 306
LYGPPGTGKSSLIAA+AN+L++DIYDLDLT+LN+N +L+ LL++ +NRSILV+EDIDCSI
Sbjct: 252 LYGPPGTGKSSLIAAIANHLNFDIYDLDLTSLNNNAELRRLLMSTANRSILVVEDIDCSI 311
Query: 307 KLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGR 366
+L +R D+E +TLSGLLN +DGLWS CG ERII+FTTN+RE+LDPALLRPGR
Sbjct: 312 ELKDRSTDQENNDPLHKTVTLSGLLNFVDGLWSSCGNERIIVFTTNYREKLDPALLRPGR 371
Query: 367 MDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTKIADVEE 426
MDMH+++SYCT +AF+ LA YL I H LFEQI +R ++VTP+EVA QL + V++
Sbjct: 372 MDMHIHMSYCTPAAFKVLASNYLEIQDHILFEQIEEFIREIEVTPSEVAEQLMRSDSVDK 431
Query: 427 CLHGLVKFLQDKTQNDQN 444
L GLV+FL+ K Q D +
Sbjct: 432 VLQGLVEFLKAKKQIDNS 449
>UniRef100_Q9SVK6 Putative mitochondrial protein [Arabidopsis thaliana]
Length = 534
Score = 464 bits (1194), Expect = e-129
Identities = 239/476 (50%), Positives = 336/476 (70%), Gaps = 19/476 (3%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K++L+ AAS+AATAML RS+ D++P E+ +I+ F ++ FS+Q T++IEEF+G
Sbjct: 17 KTVLTTAASVAATAMLARSLVQDYLPDEVHHYISYGFRSIFGYFSSQMTIIIEEFEGFAH 76
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
NEV+EAAEAYL+TK + S +R+K K E + + +RDEEV D + GV+ W L C
Sbjct: 77 NEVFEAAEAYLATKISPSNKRIKVSKHEKENNYNVTVERDEEVVDTYNGVKFQWILHCRH 136
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
V+S +++S+L SEVRS+EL+FHKK K+ SYLP++++RA +KQE +KI
Sbjct: 137 VESKHFHNP-RDLNSTLRSEVRSFELNFHKKFKDVALESYLPFMVKRATLMKQEKKTLKI 195
Query: 187 YTVEYE--------AWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTG 238
+T+ E AW DHP TFKTLA+D+ +K ++ DLD+F++ ++FY R G
Sbjct: 196 FTLSPENMYGNYSDAW--TSVTLDHPSTFKTLAMDSDVKTSVMEDLDKFVKRRDFYKRVG 253
Query: 239 KAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILV 298
KAWKRGYLLYGPPGTGKSSLIAAMAN+L++DIYDL+LTA+N+N +L+ LL+A +NRSIL+
Sbjct: 254 KAWKRGYLLYGPPGTGKSSLIAAMANHLNFDIYDLELTAVNNNSELRRLLIATANRSILI 313
Query: 299 IEDIDCSIKLPNREEDEEAAKNGD------NKMTLSGLLNVIDGLWSCCGEERIIIFTTN 352
+EDIDCS++L +R DE ++ D K+TLSGLLN IDGLWS CG+ERIIIFTTN
Sbjct: 314 VEDIDCSLELKDRTSDEPPRESDDIEDPRYKKVTLSGLLNFIDGLWSSCGDERIIIFTTN 373
Query: 353 HRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPA 412
++E+LD ALLRPGRMDMH+++SYCT S F+ LA YL I +H+LF +I + +VTPA
Sbjct: 374 YKEKLDAALLRPGRMDMHIHMSYCTPSTFKALALNYLEIKEHRLFSKIEEGIEATEVTPA 433
Query: 413 EVAGQLTKIADVEECLHGLVKFLQ-DKTQNDQNINQDNASQKEKKNQDHEYGIDNL 467
EVA QL + V++ L GL++FL+ K +N+Q+ + + E K + E G D++
Sbjct: 434 EVAEQLMRNDSVDKVLEGLIEFLKVKKIENEQDKAKTEKQELENKKKTKE-GTDSV 488
>UniRef100_Q8VZG2 AT3g50930/F18B3_210 [Arabidopsis thaliana]
Length = 576
Score = 464 bits (1194), Expect = e-129
Identities = 239/476 (50%), Positives = 336/476 (70%), Gaps = 19/476 (3%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K++L+ AAS+AATAML RS+ D++P E+ +I+ F ++ FS+Q T++IEEF+G
Sbjct: 59 KTVLTTAASVAATAMLARSLVQDYLPDEVHHYISYGFRSIFGYFSSQMTIIIEEFEGFAH 118
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
NEV+EAAEAYL+TK + S +R+K K E + + +RDEEV D + GV+ W L C
Sbjct: 119 NEVFEAAEAYLATKISPSNKRIKVSKHEKENNYNVTVERDEEVVDTYNGVKFQWILHCRH 178
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
V+S +++S+L SEVRS+EL+FHKK K+ SYLP++++RA +KQE +KI
Sbjct: 179 VESKHFHNP-RDLNSTLRSEVRSFELNFHKKFKDVALESYLPFMVKRATLMKQEKKTLKI 237
Query: 187 YTVEYE--------AWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTG 238
+T+ E AW DHP TFKTLA+D+ +K ++ DLD+F++ ++FY R G
Sbjct: 238 FTLSPENMYGNYSDAW--TSVTLDHPSTFKTLAMDSDVKTSVMEDLDKFVKRRDFYKRVG 295
Query: 239 KAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILV 298
KAWKRGYLLYGPPGTGKSSLIAAMAN+L++DIYDL+LTA+N+N +L+ LL+A +NRSIL+
Sbjct: 296 KAWKRGYLLYGPPGTGKSSLIAAMANHLNFDIYDLELTAVNNNSELRRLLIATANRSILI 355
Query: 299 IEDIDCSIKLPNREEDEEAAKNGD------NKMTLSGLLNVIDGLWSCCGEERIIIFTTN 352
+EDIDCS++L +R DE ++ D K+TLSGLLN IDGLWS CG+ERIIIFTTN
Sbjct: 356 VEDIDCSLELKDRTSDEPPRESDDIEDPRYKKVTLSGLLNFIDGLWSSCGDERIIIFTTN 415
Query: 353 HRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPA 412
++E+LD ALLRPGRMDMH+++SYCT S F+ LA YL I +H+LF +I + +VTPA
Sbjct: 416 YKEKLDAALLRPGRMDMHIHMSYCTPSTFKALALNYLEIKEHRLFSKIEEGIEATEVTPA 475
Query: 413 EVAGQLTKIADVEECLHGLVKFLQ-DKTQNDQNINQDNASQKEKKNQDHEYGIDNL 467
EVA QL + V++ L GL++FL+ K +N+Q+ + + E K + E G D++
Sbjct: 476 EVAEQLMRNDSVDKVLEGLIEFLKVKKIENEQDKAKTEKQELENKKKTKE-GTDSV 530
>UniRef100_Q8LBK2 BCS1 protein-like protein [Arabidopsis thaliana]
Length = 534
Score = 464 bits (1194), Expect = e-129
Identities = 239/476 (50%), Positives = 336/476 (70%), Gaps = 19/476 (3%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K++L+ AAS+AATAML RS+ D++P E+ +I+ F ++ FS+Q T++IEEF+G
Sbjct: 17 KTVLTTAASVAATAMLARSLVQDYLPDEVHHYISYGFRSIFGYFSSQMTIIIEEFEGFAH 76
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
NEV+EAAEAYL+TK + S +R+K K E + + +RDEEV D + GV+ W L C
Sbjct: 77 NEVFEAAEAYLATKISPSNKRIKVSKHEKENNYNVTVERDEEVVDTYNGVKFQWILHCRH 136
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
V+S +++S+L SEVRS+EL+FHKK K+ SYLP++++RA +KQE +KI
Sbjct: 137 VESKHFHNP-RDLNSTLRSEVRSFELNFHKKFKDVALESYLPFMVKRATLMKQEKKTLKI 195
Query: 187 YTVEYE--------AWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTG 238
+T+ E AW DHP TFKTLA+D+ +K ++ DLD+F++ ++FY R G
Sbjct: 196 FTLSPENMYGNYSDAW--TSVTLDHPSTFKTLAMDSDVKTSVMEDLDKFVKRRDFYKRVG 253
Query: 239 KAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILV 298
KAWKRGYLLYGPPGTGKSSLIAAMAN+L++DIYDL+LTA+N+N +L+ LL+A +NRSIL+
Sbjct: 254 KAWKRGYLLYGPPGTGKSSLIAAMANHLNFDIYDLELTAVNNNSELRRLLIATANRSILI 313
Query: 299 IEDIDCSIKLPNREEDEEAAKNGD------NKMTLSGLLNVIDGLWSCCGEERIIIFTTN 352
+EDIDCS++L +R DE ++ D K+TLSGLLN IDGLWS CG+ERIIIFTTN
Sbjct: 314 VEDIDCSLELKDRTSDEPPRESDDIEDPRYKKVTLSGLLNFIDGLWSSCGDERIIIFTTN 373
Query: 353 HRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPA 412
++E+LD ALLRPGRMDMH+++SYCT S F+ LA YL I +H+LF +I + +VTPA
Sbjct: 374 YKEKLDAALLRPGRMDMHIHMSYCTPSTFKALALNYLEIKEHRLFSKIEEGIEATEVTPA 433
Query: 413 EVAGQLTKIADVEECLHGLVKFLQ-DKTQNDQNINQDNASQKEKKNQDHEYGIDNL 467
EVA QL + V++ L GL++FL+ K +N+Q+ + + E K + E G D++
Sbjct: 434 EVAEQLMRNDSVDKVLEGLIEFLKVKKIENEQDKAKTEKQELENKKRTKE-GTDSV 488
>UniRef100_Q9SVK5 Putative mitochondrial protein [Arabidopsis thaliana]
Length = 480
Score = 464 bits (1193), Expect = e-129
Identities = 246/470 (52%), Positives = 321/470 (67%), Gaps = 36/470 (7%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K+ L+A AS+AA A+L RS+ D++P+E+ ++I+ F FS Q T VIEEF G
Sbjct: 13 KTALTAVASVAAAAILARSVVQDYMPNEVHEYISHGFRRFFSYFSYQMTAVIEEFGGFEH 72
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
N+V+EAAEAYLSTK + S +R+K K E S +RDEEV D F+GV+++W L C
Sbjct: 73 NQVFEAAEAYLSTKISNSTRRIKVNKLEKQSNYSVTVERDEEVVDIFDGVKLSWILVCRH 132
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
VD R +++S+L SEVRSYELSF KK K + SYLP+V+E+A +IKQ+ +KI
Sbjct: 133 VDKKDFRNP-RDLNSTLKSEVRSYELSFRKKFKNMVLESYLPFVVEQAASIKQKFKTLKI 191
Query: 187 YTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYL 246
+TV+ + DHP TF+TLA+D +KK LV DLDRF+Q K FY R GKAWKRGYL
Sbjct: 192 FTVDSYSVEWTSVTLDHPSTFRTLALDPEVKKNLVEDLDRFVQRKGFYGRVGKAWKRGYL 251
Query: 247 LYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSI 306
LYGPPGTGKSSLIAA+AN+L++DIYDLDLT+LN+N +L+ LL++ +NRSILV+EDIDCSI
Sbjct: 252 LYGPPGTGKSSLIAAIANHLNFDIYDLDLTSLNNNAELRRLLMSTANRSILVVEDIDCSI 311
Query: 307 KLPNREEDEEAAKNGDN--------------------------------KMTLSGLLNVI 334
+L +R D+E N D ++TLSGLLN +
Sbjct: 312 ELKDRSTDQE---NNDPLHKTVMHFDSLSVMLLCDLLLISITNVLVSHFQVTLSGLLNFV 368
Query: 335 DGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQH 394
DGLWS CG ERII+FTTN+RE+LDPALLRPGRMDMH+++SYCT +AF+ LA YL I H
Sbjct: 369 DGLWSSCGNERIIVFTTNYREKLDPALLRPGRMDMHIHMSYCTPAAFKVLASNYLEIQDH 428
Query: 395 KLFEQIGGLLRHVQVTPAEVAGQLTKIADVEECLHGLVKFLQDKTQNDQN 444
LFEQI +R ++VTPAEVA QL + V++ L GLV+FL+ K Q D +
Sbjct: 429 ILFEQIEEFIREIEVTPAEVAEQLMRSDSVDKVLQGLVEFLKAKKQIDNS 478
>UniRef100_Q6L567 Putative AAA-type ATPase [Oryza sativa]
Length = 479
Score = 413 bits (1061), Expect = e-114
Identities = 210/467 (44%), Positives = 304/467 (64%), Gaps = 22/467 (4%)
Query: 10 LSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTKNEV 69
++ AA++A TAM++R + ++ +P EL + + + S+ TVVI+E +G++ N++
Sbjct: 11 MATAAAVAGTAMVVRGVVSELVPDELREMLRSAARGIRARVSSTHTVVIDETEGLSTNQI 70
Query: 70 YEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRVDS 129
Y+AA YL+ + QR++A + +D + + D+ EE+ D +GV TW+L + D+
Sbjct: 71 YDAARTYLAARINTDMQRLRASRVDDAQGIMITMDQGEEMLDVHDGVEYTWRL--VSRDT 128
Query: 130 SSSRTRYT-----------NMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIK 178
+++ T + N EV+S+E+SFHKKHKEK SYLP+V++ AKA+
Sbjct: 129 AAAATAHAAPYGIGGGGAANRRGRSRFEVKSFEVSFHKKHKEKALRSYLPFVIDTAKAMN 188
Query: 179 QENMAVKIYTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTG 238
++ +K++ +EY+AW R HP TF TLA+D LK ++ DL+RF++ K++Y R G
Sbjct: 189 DKHRNLKMHMIEYDAWTAVDLR--HPSTFDTLAMDHSLKHSVMYDLERFVKRKDYYRRIG 246
Query: 239 KAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILV 298
+AWKRGYLLYGPPGTGKSSLIAAMANYL +DIYDL+LT + N DL+ LL+ MSNRSILV
Sbjct: 247 RAWKRGYLLYGPPGTGKSSLIAAMANYLKFDIYDLELTEVKSNSDLRRLLVGMSNRSILV 306
Query: 299 IEDIDCSIKLPNREEDE-------EAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTT 351
+EDIDC+I L R+E E + + ++K+TLSGLLN +DGLWS GEERII+FTT
Sbjct: 307 VEDIDCTIDLQQRDEGEIKRAKPTYSGEENEDKVTLSGLLNFVDGLWSTSGEERIIVFTT 366
Query: 352 NHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTP 411
N+RERLDPALLRPGRMDMH+++ YCT AF LA Y + H ++ +I L+ V TP
Sbjct: 367 NYRERLDPALLRPGRMDMHIHMGYCTREAFRVLASNYHNVENHAMYPEIEQLIEEVLTTP 426
Query: 412 AEVAGQLTKIADVEECLHGLVKFLQDKTQNDQNINQDNASQKEKKNQ 458
AEVA L + DV+ L L +FL+ K +N + +K N+
Sbjct: 427 AEVAEVLMRNDDVDVALQVLAEFLKAKRNEPGETKAENKNGNQKINK 473
>UniRef100_Q6L566 Putative AAA-type ATPase [Oryza sativa]
Length = 484
Score = 410 bits (1055), Expect = e-113
Identities = 215/464 (46%), Positives = 306/464 (65%), Gaps = 18/464 (3%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K ++ AAS+AA+ ML+RS+ N+ +P+E+ D + L S+Q T++IEE +G +
Sbjct: 12 KKAITTAASVAASVMLVRSVVNELVPYEVRDVLFSGLGYLRSQISSQHTIIIEETEGWSH 71
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGK-SEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCI 125
N VY A AYL+T+ + QR++ E + + + EE+ D EG W L
Sbjct: 72 NHVYNAVRAYLATRINNNMQRLRVSSMDESSEKMVVTMEEGEELVDMHEGTEFKWCLISR 131
Query: 126 RVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVK 185
+ + + N + S EVRSYELSFH+KHKEK SYLP+++ AKAIK + ++
Sbjct: 132 SISADPN-----NGNGSGQREVRSYELSFHRKHKEKALKSYLPFIIATAKAIKDQERILQ 186
Query: 186 IYTVEY-EAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRG 244
IY EY ++W + HP TF TLA+D LK+ ++ DLDRF++ K++Y R GKAWKRG
Sbjct: 187 IYMNEYSDSW--SPIDLHHPSTFDTLAMDQKLKQSIIDDLDRFIKRKDYYKRIGKAWKRG 244
Query: 245 YLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDC 304
YLLYGPPGTGKSSLIAAMAN+L +DIYDL+LT ++ N +L+ LL+ M++RSILV+EDIDC
Sbjct: 245 YLLYGPPGTGKSSLIAAMANHLKFDIYDLELTGVHSNSELRRLLVGMTSRSILVVEDIDC 304
Query: 305 SIKLPNREEDEEAAKN-------GDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERL 357
SI+L RE EE K+ G++K+TLSGLLN +DGLWS GEERII+FTTN++ERL
Sbjct: 305 SIELKQREAGEERTKSNSTEEDKGEDKVTLSGLLNFVDGLWSTSGEERIIVFTTNYKERL 364
Query: 358 DPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQ 417
D AL+RPGRMDMH+++ YCT AF LA Y I H + +I L++ V VTPAEVA
Sbjct: 365 DQALMRPGRMDMHIHMGYCTPEAFRILASNYHSIDYHVTYPEIEELIKEVMVTPAEVAEA 424
Query: 418 LTKIADVEECLHGLVKFLQDKTQ--NDQNINQDNASQKEKKNQD 459
L + D++ L GL++ L+ K + ++ +A+++ ++N+D
Sbjct: 425 LMRNDDIDVALLGLLELLKSKIKDASETKAESKDANKQTEENKD 468
>UniRef100_Q5ZDA1 BCS1 protein-like [Oryza sativa]
Length = 453
Score = 393 bits (1010), Expect = e-108
Identities = 209/454 (46%), Positives = 287/454 (63%), Gaps = 36/454 (7%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K ++ AASLAA+AML+R + N+ +P+E+ D + L S+Q V+IEE +G T
Sbjct: 12 KRAVTTAASLAASAMLVRGVVNELVPYEVRDLLFSGVGYLRSRMSSQHMVIIEETEGWTN 71
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
N++Y+A YL+T+ QR++ + + G R G ++L
Sbjct: 72 NQLYDAVRTYLATRINTDMQRLRVSRDNSSSSNGNGNGR---------GGNGNYRL---- 118
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
EVRS+E+SFHKKHK+K NSYLP++L AK IK ++ +KI
Sbjct: 119 -------------------EVRSFEMSFHKKHKDKALNSYLPHILATAKKIKDQDRTLKI 159
Query: 187 YTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYL 246
Y E E+W HP TF TLA+D K+ ++ DL+RF++ KE+Y + GKAWKRGYL
Sbjct: 160 YMNEGESWF--AIDLHHPSTFTTLAMDHKQKQSVMDDLERFIKRKEYYKKIGKAWKRGYL 217
Query: 247 LYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSI 306
LYGPPGTGKSSLIAAMANYL +D+YDL+LT +N N L+ LL+ M+NRSILVIEDIDC++
Sbjct: 218 LYGPPGTGKSSLIAAMANYLKFDVYDLELTEVNWNSTLRRLLIGMTNRSILVIEDIDCTL 277
Query: 307 KLPNREEDEEAAKN--GDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRP 364
+L REE +E++K+ ++K+TLSGLLN +DGLWS GEERII+FTTN++ERLDPALLRP
Sbjct: 278 ELQQREEGQESSKSNPSEDKVTLSGLLNFVDGLWSTSGEERIIVFTTNYKERLDPALLRP 337
Query: 365 GRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTKIADV 424
GRMDMHV++ YC +F LA Y I H + +I L++ V VTPAEVA L + D
Sbjct: 338 GRMDMHVHMGYCCPESFRILASNYHSIDNHATYPEIEELIKEVMVTPAEVAEVLMRNDDT 397
Query: 425 EECLHGLVKFLQDKTQNDQNINQDNASQKEKKNQ 458
+ L GL++FL+ K + +N Q K +
Sbjct: 398 DVALEGLIQFLKRKKDVGKEGKAENVEQVVKAEE 431
>UniRef100_Q9FN75 AAA-type ATPase-like protein [Arabidopsis thaliana]
Length = 505
Score = 384 bits (985), Expect = e-105
Identities = 212/472 (44%), Positives = 288/472 (60%), Gaps = 22/472 (4%)
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLS-RCFSTQFTVVIEEFQGMTK 66
S+ +A AS+A M+IRS+ ++ IP L DFI +L R S+ T+ I++
Sbjct: 12 SVFTAYASMAGYMMMIRSMAHELIPAPLQDFIYRTLRSLFFRSSSSTLTLTIDDDNMGMN 71
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
NE+Y AA+ YLSTK + A R++ K DK ++ E V D +E V++ W+
Sbjct: 72 NEIYRAAQTYLSTKISPDAVRLRISKGHKDKHVNLYLSDGEIVNDVYEDVQLVWRFVTDG 131
Query: 127 VDSSSSRT----------RYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKA 176
D R M SE +ELSF KKHK+ + NSY+PY+ +AK
Sbjct: 132 GDKKGGGGGVGGRGGGGGRRGGMDDDGKSEY--FELSFDKKHKDLILNSYVPYIESKAKE 189
Query: 177 IKQENMAVKIYTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNR 236
I+ E + ++++ W +HP TF+T+A++ LK++++ DLDRF++ KEFY R
Sbjct: 190 IRDERRILMLHSLNSLRW--ESVILEHPSTFETMAMEDDLKRDVIEDLDRFIRRKEFYKR 247
Query: 237 TGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSI 296
GKAWKRGYLLYGPPGTGKSSL+AAMANYL +D+YDL L ++ + DL+ LLLA NRSI
Sbjct: 248 VGKAWKRGYLLYGPPGTGKSSLVAAMANYLKFDVYDLQLASVMRDSDLRRLLLATRNRSI 307
Query: 297 LVIEDIDCSIKLPNREEDEEAAKN---GDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNH 353
LVIEDIDC++ LPNR E KN +TLSGLLN IDGLWS CG+ERIIIFTTNH
Sbjct: 308 LVIEDIDCAVDLPNRIEQPVEGKNRGESQGPLTLSGLLNFIDGLWSSCGDERIIIFTTNH 367
Query: 354 RERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQ----HKLFEQIGGLLRHVQV 409
++RLDPALLRPGRMDMH+ + +C+F F+ LA YLG+S H+LF +I L+ +
Sbjct: 368 KDRLDPALLRPGRMDMHIYMGHCSFQGFKTLASNYLGLSDAAMPHRLFPEIERLIDGEVM 427
Query: 410 TPAEVAGQLTKIADVEECLHGLVKFLQDKTQNDQNINQDNASQKEKKNQDHE 461
TPA+VA +L K D + L GLV L+ + N QKE + + E
Sbjct: 428 TPAQVAEELMKSEDADVALEGLVNVLEKMRLKSKESNPVMMKQKESRLEMEE 479
>UniRef100_Q8GW96 AAA-type ATPase like protein [Arabidopsis thaliana]
Length = 495
Score = 377 bits (967), Expect = e-103
Identities = 201/445 (45%), Positives = 280/445 (62%), Gaps = 30/445 (6%)
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTKN 67
SL SA ASL ML RS+ +DF+P +L + + S TV+I+E G+ +N
Sbjct: 13 SLFSAYASLTGFLMLFRSMLHDFVPEKLRSYFSSLLDRFFTPKSKYLTVIIDENFGLNRN 72
Query: 68 EVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRV 127
+V++AAE YL +K +R++ GK K + +R EE+ D FE V W
Sbjct: 73 QVFDAAEMYLRSKIGPETERLRVGKIPKQKHFTISIERGEEILDTFEESEVKWSYVQSEN 132
Query: 128 DSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIY 187
+ RY YEL+F KK ++K+ NSYL +V+ ++ IK+ VK+Y
Sbjct: 133 EKGDKVKRY-------------YELTFEKKLRDKVLNSYLTHVVAESEEIKRNLRVVKLY 179
Query: 188 TVEYEA-----------W-CRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYN 235
+ + A W C N +HP TF TLA+D KK+++ DL+RFL+ KEFY
Sbjct: 180 SRDVYASDDDDGMAGGNWGCIN---LEHPSTFDTLAMDPNAKKKIIDDLERFLKRKEFYK 236
Query: 236 RTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRS 295
R GKAWKRGYLLYGPPGTGKSSLIAAMANYL +D++DL+L+++ DN +LK +LL+ +NRS
Sbjct: 237 RVGKAWKRGYLLYGPPGTGKSSLIAAMANYLKFDVFDLELSSIYDNGELKRVLLSTTNRS 296
Query: 296 ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRE 355
ILVIEDIDC+ ++ +RE + + + K+TLSG+LN IDGLWS G+ERII+FTTNH+E
Sbjct: 297 ILVIEDIDCNAEVRDREAENQEDEQIKGKVTLSGILNFIDGLWSSFGDERIIVFTTNHKE 356
Query: 356 RLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGIS--QHKLFEQIGGLLRHVQVTPAE 413
RLDPALLRPGRMD+H+N+SYCT F L YLG+ H L E+I L+ +VTPAE
Sbjct: 357 RLDPALLRPGRMDVHINMSYCTGLGFRTLVSNYLGLDGLNHPLCEEIEALVDSTEVTPAE 416
Query: 414 VAGQLTKIADVEECLHGLVKFLQDK 438
+A +L + D + L G++ F++ +
Sbjct: 417 LAEELMQDDDTDVVLRGVISFVEKR 441
>UniRef100_Q9SI12 Putative AAA-type ATPase [Arabidopsis thaliana]
Length = 996
Score = 377 bits (967), Expect = e-103
Identities = 201/445 (45%), Positives = 280/445 (62%), Gaps = 30/445 (6%)
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTKN 67
SL SA ASL ML RS+ +DF+P +L + + S TV+I+E G+ +N
Sbjct: 13 SLFSAYASLTGFLMLFRSMLHDFVPEKLRSYFSSLLDRFFTPKSKYLTVIIDENFGLNRN 72
Query: 68 EVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRV 127
+V++AAE YL +K +R++ GK K + +R EE+ D FE V W
Sbjct: 73 QVFDAAEMYLRSKIGPETERLRVGKIPKQKHFTISIERGEEILDTFEESEVKWSYVQSEN 132
Query: 128 DSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIY 187
+ RY YEL+F KK ++K+ NSYL +V+ ++ IK+ VK+Y
Sbjct: 133 EKGDKVKRY-------------YELTFEKKLRDKVLNSYLTHVVAESEEIKRNLRVVKLY 179
Query: 188 TVEYEA-----------W-CRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYN 235
+ + A W C N +HP TF TLA+D KK+++ DL+RFL+ KEFY
Sbjct: 180 SRDVYASDDDDGMAGGNWGCIN---LEHPSTFDTLAMDPNAKKKIIDDLERFLKRKEFYK 236
Query: 236 RTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRS 295
R GKAWKRGYLLYGPPGTGKSSLIAAMANYL +D++DL+L+++ DN +LK +LL+ +NRS
Sbjct: 237 RVGKAWKRGYLLYGPPGTGKSSLIAAMANYLKFDVFDLELSSIYDNGELKRVLLSTTNRS 296
Query: 296 ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRE 355
ILVIEDIDC+ ++ +RE + + + K+TLSG+LN IDGLWS G+ERII+FTTNH+E
Sbjct: 297 ILVIEDIDCNAEVRDREAENQEDEQIKGKVTLSGILNFIDGLWSSFGDERIIVFTTNHKE 356
Query: 356 RLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGIS--QHKLFEQIGGLLRHVQVTPAE 413
RLDPALLRPGRMD+H+N+SYCT F L YLG+ H L E+I L+ +VTPAE
Sbjct: 357 RLDPALLRPGRMDVHINMSYCTGLGFRTLVSNYLGLDGLNHPLCEEIEALVDSTEVTPAE 416
Query: 414 VAGQLTKIADVEECLHGLVKFLQDK 438
+A +L + D + L G++ F++ +
Sbjct: 417 LAEELMQDDDTDVVLRGVISFVEKR 441
Score = 374 bits (960), Expect = e-102
Identities = 207/476 (43%), Positives = 295/476 (61%), Gaps = 33/476 (6%)
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTKN 67
SL +A ASL ML RS+ ND +P L +I S T+VI+E G +N
Sbjct: 516 SLFTAYASLTGFLMLFRSLFNDEVPERLRSYITDLLNRFFTPKSKNLTMVIDEIIGFKRN 575
Query: 68 EVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRV 127
+V++AAE YL K R++ GK K + ++ EE+ D FE + W T +
Sbjct: 576 QVFDAAEVYLRNKIGPETARLRVGKLPKQKHFTIYIEKGEEILDTFENSELRW--TYVES 633
Query: 128 DSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIY 187
++ +S+ E R YEL+F KK ++K+ NSYL +V+ ++ K++ AVK+Y
Sbjct: 634 ENEASQ-----------KEKRYYELTFEKKLRDKVMNSYLSHVVAESEETKRDLRAVKLY 682
Query: 188 TVEYEA-----------W-CRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYN 235
+ + A W C N +HP TF+TLA+D G KK+++ D++RFL+ +EFY
Sbjct: 683 SRDVRASKDDDGMAGAGWGCIN---LEHPSTFETLAMDPGAKKKIIDDMERFLKRREFYK 739
Query: 236 RTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRS 295
R GKAWKRGYLLYGPPGTGKSSLIAAMANYL +D++DL+L+++ +N LK++LL+ +NRS
Sbjct: 740 RVGKAWKRGYLLYGPPGTGKSSLIAAMANYLKFDVFDLELSSIYENAQLKSILLSTTNRS 799
Query: 296 ILVIEDIDC-SIKLPNREEDE--EAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTN 352
ILVIEDIDC S ++ +RE DE E + ++TLSGLLN +DGLWS G+ERII+FTTN
Sbjct: 800 ILVIEDIDCSSAEVVDREADEYQEYEEGYYGRVTLSGLLNFVDGLWSSFGDERIIVFTTN 859
Query: 353 HRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGIS--QHKLFEQIGGLLRHVQVT 410
H+ERLDPALLRPGRMDMH+N+SYCT F L YLG+ H L E+I L+ +VT
Sbjct: 860 HKERLDPALLRPGRMDMHINMSYCTGLGFRTLVSNYLGLGGLNHPLCEEIEALIDSTEVT 919
Query: 411 PAEVAGQLTKIADVEECLHGLVKFLQDKTQNDQNINQDNASQKEKKNQDHEYGIDN 466
PAE+A +L + D + L G+V F++++ + S K + D ++ + +
Sbjct: 920 PAELAEELMQEDDTDVVLRGVVSFVENRKVEISKTKELEGSTCRKLDGDDKHNVSS 975
>UniRef100_Q683K1 AAA-type ATPase like protein [Arabidopsis thaliana]
Length = 495
Score = 375 bits (964), Expect = e-102
Identities = 200/445 (44%), Positives = 280/445 (61%), Gaps = 30/445 (6%)
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTKN 67
SL SA ASL ML RS+ +DF+P +L + + S TV+I+E G+ +N
Sbjct: 13 SLFSAYASLTGFLMLFRSMLHDFVPEKLRSYFSSLLDRFFTPKSKYLTVIIDENFGLNRN 72
Query: 68 EVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRV 127
+V++AAE YL +K +R++ GK K + +R EE+ D FE V W
Sbjct: 73 QVFDAAEMYLRSKIGPETERLRVGKIPKQKHFTISIERGEEILDTFEESEVKWSYVQSEN 132
Query: 128 DSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIY 187
+ RY YEL+F KK ++K+ NSYL +V+ ++ IK+ VK+Y
Sbjct: 133 EKGDKVKRY-------------YELTFEKKLRDKVLNSYLTHVVAESEEIKRNLRVVKLY 179
Query: 188 TVEYEA-----------W-CRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYN 235
+ + A W C N +HP TF TLA+D K++++ DL+RFL+ KEFY
Sbjct: 180 SRDVYASDDDDGMAGGNWGCIN---LEHPSTFDTLAMDPNAKRKIIDDLERFLKRKEFYK 236
Query: 236 RTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRS 295
R GKAWKRGYLLYGPPGTGKSSLIAAMANYL +D++DL+L+++ DN +LK +LL+ +NRS
Sbjct: 237 RVGKAWKRGYLLYGPPGTGKSSLIAAMANYLKFDVFDLELSSIYDNGELKRVLLSTTNRS 296
Query: 296 ILVIEDIDCSIKLPNREEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRE 355
ILVIEDIDC+ ++ +RE + + + K+TLSG+LN IDGLWS G+ERII+FTTNH+E
Sbjct: 297 ILVIEDIDCNAEVRDREAENQEDEQIKGKVTLSGILNFIDGLWSSFGDERIIVFTTNHKE 356
Query: 356 RLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGIS--QHKLFEQIGGLLRHVQVTPAE 413
RLDPALLRPGRMD+H+N+SYCT F L YLG+ H L E+I L+ +VTPAE
Sbjct: 357 RLDPALLRPGRMDVHINMSYCTGLGFRTLVSNYLGLDGLNHPLCEEIEALVDSTEVTPAE 416
Query: 414 VAGQLTKIADVEECLHGLVKFLQDK 438
+A +L + D + L G++ F++ +
Sbjct: 417 LAEELMQDDDTDVVLRGVISFVEKR 441
>UniRef100_Q9FN77 AAA-type ATPase-like protein [Arabidopsis thaliana]
Length = 533
Score = 355 bits (911), Expect = 2e-96
Identities = 191/464 (41%), Positives = 281/464 (60%), Gaps = 14/464 (3%)
Query: 8 SLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGM-TK 66
S+ S AS+ M+I+ + N IP + +F+ + + S+ T+ I++ M
Sbjct: 12 SMFSTYASMMGYVMIIKPMINTIIPRPVQNFVFSYLKSFAGSRSSTLTLTIDQMSSMYIP 71
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
+E+Y AA+AYLSTK + ++ R+ + +K + E V+D + G+++ W+
Sbjct: 72 DELYAAAQAYLSTKISPNSVRLIMARDPAEKKVKLYLSDGEVVSDVYNGIKLKWRFLARN 131
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
+++ + ++ E S ELSF KKH++ + NSY+PYV +AK + + +K+
Sbjct: 132 KNNTMVEEYGQSYQGNIQRE--SLELSFDKKHRDLVVNSYIPYVESKAKEVNNKRRILKM 189
Query: 187 YTVEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRGYL 246
+ + A F HP TF T+A++ LK+ ++ DLDRF+ K+FY R GKAWKRGYL
Sbjct: 190 HCYSHMAQTWQSVNFKHPSTFDTMAMNDDLKRSMIEDLDRFVGRKDFYKRVGKAWKRGYL 249
Query: 247 LYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDCSI 306
LYGPPGTGKSSL+AAMANYL +DIYDL L ++ + L++LLLA +N SIL+IEDIDCS+
Sbjct: 250 LYGPPGTGKSSLVAAMANYLKFDIYDLQLASVQGDAHLRSLLLATNNSSILLIEDIDCSV 309
Query: 307 KLPNREEDEE------AAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPA 360
LP R + A +TLSGLLN IDGLWS CG ERIIIFTTN++E+LDPA
Sbjct: 310 DLPTRLQPPTETSQPLGAVQVSKPLTLSGLLNCIDGLWSSCGNERIIIFTTNNKEKLDPA 369
Query: 361 LLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQ-----HKLFEQIGGLLRHVQVTPAEVA 415
LLRPGRMDMH+ + +C+F F+ LA YLG+S H L I L+ +TPA+VA
Sbjct: 370 LLRPGRMDMHIYMGHCSFQGFKTLASNYLGLSDENDDTHPLCPDIKHLIDGHVLTPAQVA 429
Query: 416 GQLTKIADVEECLHGLVKFLQDKTQNDQNINQDNASQKEKKNQD 459
+L K D + L GLVK L+ K + + ++ +K K+ ++
Sbjct: 430 EELMKDEDADAALEGLVKVLKRKRLEPKKCDDESKMKKLKEGEE 473
>UniRef100_Q9FN78 AAA-type ATPase-like protein [Arabidopsis thaliana]
Length = 470
Score = 353 bits (905), Expect = 8e-96
Identities = 191/438 (43%), Positives = 273/438 (61%), Gaps = 15/438 (3%)
Query: 10 LSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEF--QGMTKN 67
+SA ASL M+I+ IP L +++ + + T++I++ GM N
Sbjct: 14 VSAYASLTGYVMMIKPFLEMTIPPPLQNYMISYLNSFLHSTPSTLTLIIDDHIKNGMY-N 72
Query: 68 EVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRV 127
E+Y AA+ Y+STK +A+R++ + +K ++ E V+D ++G+ V W+ V
Sbjct: 73 ELYGAAQVYISTKVNHNAERLRISRDRSEKNVNIHFSVGEVVSDIYQGIEVKWRFC---V 129
Query: 128 DSSSSR-TRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
DS+ S Y L + ELSF KKH E + NSY+PYV +AK I E +K+
Sbjct: 130 DSNKSNMVHYFGEHFKLNPDRECVELSFEKKHTELVLNSYIPYVESKAKVINNERKILKM 189
Query: 187 YTV--EYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKRG 244
Y+ Y W +HP TF T+A++ LK+ ++ DLDRF++ K+FY R GK WKRG
Sbjct: 190 YSYCCMYLKW--QSVNLEHPSTFDTMAMNEELKRSVMGDLDRFIRRKDFYKRVGKPWKRG 247
Query: 245 YLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDIDC 304
YLLYGPPGTGK+SL+AA+ANYL +DIYDL L ++ ++ DL+ LLL +N SIL++EDIDC
Sbjct: 248 YLLYGPPGTGKTSLVAAIANYLKFDIYDLQLASVREDADLRRLLLGTTNSSILLVEDIDC 307
Query: 305 SIKLPNR-EEDEEAAKNGDNKMTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLR 363
++ L R + + G + +TLSGLL IDGLWS CG+ERI+IFTT H+ERLDPALLR
Sbjct: 308 AVDLHTRLQPKTQDDTKGSSMLTLSGLLTCIDGLWSSCGDERIVIFTTTHKERLDPALLR 367
Query: 364 PGRMDMHVNLSYCTFSAFEQLAFTYLGISQ---HKLFEQIGGLLRHVQVTPAEVAGQLTK 420
PGRMDMH+++ +C F F+ LA YLG+S H L+ +I L++ +TPA+VA +L K
Sbjct: 368 PGRMDMHIHMGHCCFDVFKTLASNYLGLSHDDPHHLYPEIERLIKGEVLTPAQVAEELMK 427
Query: 421 IADVEECLHGLVKFLQDK 438
D + L GLVK L+ K
Sbjct: 428 NEDPDVALEGLVKVLKRK 445
>UniRef100_Q9LH82 Similarity to AAA-type ATPase [Arabidopsis thaliana]
Length = 510
Score = 348 bits (894), Expect = 1e-94
Identities = 189/483 (39%), Positives = 284/483 (58%), Gaps = 45/483 (9%)
Query: 9 LLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEF---QGMT 65
L + A+ M S+ F+P+++ D++ FY + S + E+ +G+
Sbjct: 7 LFGFTGTTMASLMFFWSVYRQFVPYQIRDYLEKCFYKMFGLVSNSVHIKFTEYTEDKGLK 66
Query: 66 KNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCI 125
K++ Y+ YLS+K+T AQR+KA +S++ K+L D E V D F+GV+V W L+
Sbjct: 67 KSQAYDLIRNYLSSKSTARAQRLKANESKNSKSLVLSLDNHEAVEDVFQGVKVVWSLSVW 126
Query: 126 RVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVK 185
+ + + SE R LSFH +++E + +YL +VL K I +N K
Sbjct: 127 KSNDQADS-----------SEKRYLTLSFHNRYREMITTTYLDHVLREGKEIGLKNRERK 175
Query: 186 IYT----VEYEAWCR---NGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTG 238
+YT +Y AW + FDHP TF+TLA+D K+ + DL +F +GK++Y + G
Sbjct: 176 LYTNNSSQDYSAWREGRWSNVPFDHPATFETLAMDLEKKEGMKKDLIKFTKGKDYYRKVG 235
Query: 239 KAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILV 298
K WKRGYLL+GPPGTGKS++I+AMAN+L YD+YDL+LT + DN +LK L+L +SI+V
Sbjct: 236 KPWKRGYLLFGPPGTGKSTMISAMANFLEYDVYDLELTTVKDNSELKKLMLDTKGKSIVV 295
Query: 299 IEDIDCSIKLPNR----------EEDEEAAKNG-----------DNKMTLSGLLNVIDGL 337
IEDIDCS+ L + EE+EE K ++K+TLSGLLN IDGL
Sbjct: 296 IEDIDCSLDLTGQRKKKKEEDEDEEEEEKKKEAEKLLKRERGERESKVTLSGLLNAIDGL 355
Query: 338 WSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLF 397
WS C E+II+FTTN+ ++LDPAL+R GRMD H+ +SYC F AF+ LA YL I H LF
Sbjct: 356 WSACSGEKIIVFTTNYLDKLDPALIRRGRMDNHIEMSYCRFEAFKVLAKNYLEIESHDLF 415
Query: 398 EQIGGLLRHVQVTPAEVAGQLTKIADVEE---CLHGLVKFLQDKTQNDQNINQDNASQKE 454
+I L+ ++PA+VA L +D ++ CL LVK L+++ + + + ++ +K
Sbjct: 416 GEIKRLVEETDMSPADVAENLMPKSDEDDADICLTRLVKSLEEEKEKAKKLAEEEKMKKA 475
Query: 455 KKN 457
++
Sbjct: 476 ARD 478
>UniRef100_Q9LH84 Similarity to AAA-type ATPase [Arabidopsis thaliana]
Length = 530
Score = 337 bits (863), Expect = 6e-91
Identities = 188/482 (39%), Positives = 286/482 (59%), Gaps = 38/482 (7%)
Query: 6 TKSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEF--QG 63
T ++ + + M +I ++P ++ F+ + S + E+ +G
Sbjct: 4 TGAIWGITGTTVTSFMFFWAIYKQYVPAHFRAYVERYFHKMIGWISYYVDIKFTEYTDEG 63
Query: 64 MTKNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLT 123
+ +++ Y++ YL++K+T A+R+KA ++++ K+L F D EE+ D+FEGV+V W
Sbjct: 64 LKRSQAYDSIRNYLASKSTALAKRLKANETKNSKSLVFSMDDHEEIEDEFEGVKVKWYSN 123
Query: 124 CIRVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMA 183
+ S+ Y SS E R + LSFH++H+ + +YL +VL KAI N
Sbjct: 124 VKVIQPQSN---YGQRSSE---ERRHFTLSFHRRHRGMIIETYLDHVLREGKAIGLMNRE 177
Query: 184 VKIYT----VEYEAWCRNG----TRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYN 235
K+YT E+ W R+G F HP TF+TLA+D K+ + DL +F +GK++Y
Sbjct: 178 RKLYTNNSSQEWYPW-RSGKWSNVPFHHPATFETLAMDPEKKEGIKKDLIKFSKGKDYYK 236
Query: 236 RTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRS 295
+ GK WKRGYLL+GPPGTGKS++IAA+AN+L YD+YDL+LT + DN +LK LLL +++S
Sbjct: 237 KVGKPWKRGYLLFGPPGTGKSTMIAAIANFLDYDVYDLELTTVKDNSELKKLLLDTTSKS 296
Query: 296 ILVIEDIDCSIKL---------PNREEDEEAAKNGD---------NKMTLSGLLNVIDGL 337
I+VIEDIDCS+ L + EED E K G+ +K+TLSGLLN IDGL
Sbjct: 297 IIVIEDIDCSLDLTGQRKKKKEEDEEEDGEEKKEGEKKPKVDDKQSKVTLSGLLNSIDGL 356
Query: 338 WSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLF 397
WS C E+II+FTTN ++LDPAL+R GRMD H+ +SYC F AF+ LA YL I H L+
Sbjct: 357 WSACSGEKIIVFTTNFVDKLDPALIRRGRMDNHIEMSYCKFEAFKVLAKNYLEIETHDLY 416
Query: 398 EQIGGLLRHVQVTPAEVAGQLTKIADVEE---CLHGLVKFLQDKTQNDQNINQDNASQKE 454
+I L ++PA+VA L +D E+ C+ LVK L+++ + + + ++ +K
Sbjct: 417 GEIERKLEETDMSPADVAETLMPKSDEEDADICIKRLVKTLEEEKEKARKLAEEEEKKKA 476
Query: 455 KK 456
+K
Sbjct: 477 EK 478
>UniRef100_Q9LJJ7 Mitochondrial protein-like [Arabidopsis thaliana]
Length = 500
Score = 331 bits (848), Expect = 3e-89
Identities = 187/485 (38%), Positives = 283/485 (57%), Gaps = 45/485 (9%)
Query: 9 LLSAAASLAATAMLIRSITNDFIP---HELIDFINLKFYNLSRCFSTQFTVVIEEFQG-- 63
L + S AT M + +I F P +L F+ Y L F + E+ G
Sbjct: 7 LWTNTGSALATLMFVYTIFKQFFPLFGPQLEPFL----YRLFGRFYPYIQITFHEYSGEH 62
Query: 64 MTKNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLT 123
++E Y ++YLS ++ A+++KA ++ K++ D EE+TDDFEG+RV W+
Sbjct: 63 FKRSEAYLGIQSYLSKDSSARAKKLKANTTKGSKSIVLSMDDKEEITDDFEGIRVWWQ-- 120
Query: 124 CIRVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMA 183
TR + +E R Y L FH++ +E + YL +V+ K I+Q+N
Sbjct: 121 ----SKKEGATRQSFSFYPEANEKRYYMLRFHRRDREVIIERYLEHVMREGKTIEQKNRE 176
Query: 184 VKIYTVEYEAWCRNGTR-----FDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTG 238
K+Y+ N ++ F+HP TF TLA++ K+E+ SDL +F + K++Y + G
Sbjct: 177 RKLYSNTPGQSHGNNSKWSHVTFEHPATFDTLAMEENKKEEIKSDLIKFSKSKDYYKKIG 236
Query: 239 KAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILV 298
KAWKRGYLL+GPPGTGKS++IAAMAN+L YD+YDL+LT + DN L+ LL+ S +SI+V
Sbjct: 237 KAWKRGYLLFGPPGTGKSTMIAAMANFLEYDVYDLELTTVKDNTHLRRLLIETSAKSIIV 296
Query: 299 IEDIDCSIKL----PNREEDEE----------------AAKNGDNKMTLSGLLNVIDGLW 338
IEDIDCS+ L +EE+EE +N ++K+TLSGLLN IDGLW
Sbjct: 297 IEDIDCSLNLTGQRKKKEEEEEDGDDKNTIEKKMMMKNEGENKESKVTLSGLLNFIDGLW 356
Query: 339 SCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKLFE 398
S CG ERII+FTTN ++LDPAL+R GRMD H+ +SYC F AF+ LA YL + + ++FE
Sbjct: 357 SACGGERIIVFTTNFVDKLDPALIRKGRMDKHIEMSYCCFEAFKVLAKNYLDVEESEMFE 416
Query: 399 QIGGLL--RHVQVTPAEVAGQLTKIADV---EECLHGLVKFLQDKTQNDQNINQDNASQK 453
+I LL +++TPA+V L ++ E CL L++ L+++ + + ++ +K
Sbjct: 417 EIKRLLEVEEIKMTPADVGENLLPKSEKEGGETCLKRLIEALKEEKEEAKKKVEEEEEEK 476
Query: 454 EKKNQ 458
++K +
Sbjct: 477 QRKKE 481
>UniRef100_Q6L444 Putative ATPase protein [Solanum demissum]
Length = 527
Score = 329 bits (844), Expect = 9e-89
Identities = 193/490 (39%), Positives = 283/490 (57%), Gaps = 55/490 (11%)
Query: 18 ATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQG---MTKNEVYEAAE 74
A M I ++ ++ PHEL I L F ++ E + +++ Y A E
Sbjct: 14 AAIMFIWTMYQNYFPHELRGHIRRYTNKLVSYFYPYMHIIFYELETEGWFERSKAYVAIE 73
Query: 75 AYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIRVDSSSSRT 134
YLS ++ A+R+KA +D ++L D EE+TD+++G +V W + S +
Sbjct: 74 RYLSKNSSTQAKRLKANAVKDGQSLVLTMDDHEEITDEYKGEKVWW------ISSQKPAS 127
Query: 135 RYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKIYT------ 188
R T +S E R ++L FHKK+++ + NSYL YVL+ KAI + K+YT
Sbjct: 128 RQT-ISFYREDEKRYFKLKFHKKNRDLITNSYLKYVLDEGKAISVKERQRKLYTNNKGDG 186
Query: 189 --VEYEA---WCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKAWKR 243
Y W +G F+HP TF TLA+D K+E++ DL+ F + K++Y + GKAWKR
Sbjct: 187 GGYRYRGGRMW--SGVVFEHPSTFDTLAMDPNKKQEIIDDLETFSKSKDYYAKIGKAWKR 244
Query: 244 GYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIEDID 303
GYLLYGPPGTGKSS+IAAMAN+L YDIYDL+LT++ DN +L+ LL+ + +SI+VIEDID
Sbjct: 245 GYLLYGPPGTGKSSMIAAMANFLKYDIYDLELTSVKDNTELRKLLIDTTGKSIIVIEDID 304
Query: 304 CSIKL-------PNREEDEEAAKNGD-----------------NKMTLSGLLNVIDGLWS 339
CS+ L ++E+E+ KN + +++TLSGLLN IDGLWS
Sbjct: 305 CSLDLTGQRETNKKKKEEEDKGKNEEDAIKEKMKKGGEVKEKQSEVTLSGLLNFIDGLWS 364
Query: 340 CCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLG-ISQHKLFE 398
G ER+I+FTTN+ E+LDPAL+R GRMD H+ LSYC F +F+ LA YL + H F
Sbjct: 365 AIGGERLIVFTTNYVEKLDPALIRRGRMDKHIVLSYCCFESFKVLAHNYLDVVESHVHFP 424
Query: 399 QIGGLLRHVQVTPAEVAGQL---TKIADVEECLHGLVKFLQDKTQ----NDQNINQDNAS 451
+I LL +TPA++A L + + + CL L+K L+ + + + A+
Sbjct: 425 EIRRLLEETNMTPADIAENLMPKSSKENADTCLERLIKALETAKEEAKLKAEEEERAKAA 484
Query: 452 QKEKKNQDHE 461
+KEK+ +D E
Sbjct: 485 EKEKEEKDRE 494
>UniRef100_Q9LJJ5 Mitochondrial protein-like [Arabidopsis thaliana]
Length = 474
Score = 328 bits (841), Expect = 2e-88
Identities = 180/419 (42%), Positives = 262/419 (61%), Gaps = 44/419 (10%)
Query: 42 KFYNLSRCFSTQFTVVIEEFQGMTKNEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSF 101
KF N FS + E++ N ++ E YL KAT A+ ++A + + K L
Sbjct: 52 KFINF---FSPYVQINFSEYEDYRVNHAFDPIETYLGAKATDKAKHLRASQVRESKGLVL 108
Query: 102 GADRDEEVTDDFEGVRVTWKLTCIRVDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEK 161
D + +V D++EG+RV W++ DS+ +T +L+FH++ ++
Sbjct: 109 KRD-ETKVRDEYEGIRVWWEM---ETDSAGYKT---------------LKLTFHRRSRDI 149
Query: 162 MFNSYLPYVLERAKAIKQENMAVKIYTVEYEA-WCRNGTRF------DHPMTFKTLAIDA 214
+ NSY+ YV+E K+I +N +K++T + W + T F +HP TF+TLA+D
Sbjct: 150 VTNSYIKYVVEEGKSIDAKNKKMKLFTNNPSSHWGSSKTSFWRYIDFEHPATFETLAMDP 209
Query: 215 GLKKELVSDLDRFLQGKEFYNRTGKAWKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLD 274
K+++++DL F GK++Y + GKAWKRGYLLYGPPGTGKS++IAAMAN L+Y IYDL+
Sbjct: 210 KKKEQILNDLAAFNNGKDYYKKIGKAWKRGYLLYGPPGTGKSTMIAAMANLLNYSIYDLE 269
Query: 275 LTALNDNKDLKNLLLAMSNRSILVIEDIDCSIKLPNREEDEEA-----AKNGD-----NK 324
LTA+ +N +L+ +L A SN+SI+VIEDIDCS+ L + + +E+ K+GD NK
Sbjct: 270 LTAIQNNSELRKILTATSNKSIIVIEDIDCSLDLTGKRKKKESNLMIWRKDGDQDNEENK 329
Query: 325 --MTLSGLLNVIDGLWSCCGEERIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFE 382
+TLSGLLN IDG+WS CG+ERII+FTTNH +LDPAL+R GRMDMH+ LSYCTF AF+
Sbjct: 330 SFVTLSGLLNFIDGIWSACGQERIIVFTTNHLAKLDPALIRRGRMDMHIELSYCTFEAFK 389
Query: 383 QLAFTYLGISQHKLFEQIGGLLRHVQVTPAEVAGQLTK---IADVEECLHGLVKFLQDK 438
LA YL + H LF +I L++ + PA+VA L K D + L+ L++ L+ K
Sbjct: 390 TLAKNYLDLDSHPLFSKIESLMKETNIAPADVAENLMKKNRETDADGSLNDLIESLERK 448
>UniRef100_Q8RY66 AT4g25830/F14M19_110 [Arabidopsis thaliana]
Length = 506
Score = 328 bits (840), Expect = 3e-88
Identities = 187/492 (38%), Positives = 275/492 (55%), Gaps = 38/492 (7%)
Query: 7 KSLLSAAASLAATAMLIRSITNDFIPHELIDFINLKFYNLSRCFSTQFTVVIEEFQGMTK 66
K ++ ASL +S+ N P EL I+ F + FST I E G+
Sbjct: 2 KEYWTSLASLLGVLAFCQSLMNSVFPPELRFAISKLFNKFFKLFSTFCYFDITEIDGVNT 61
Query: 67 NEVYEAAEAYLSTKATLSAQRVKAGKSEDDKTLSFGADRDEEVTDDFEGVRVTWKLTCIR 126
NE+Y A + YLS+ +++ R+ ++ + +++FG ++ + D F V V W+
Sbjct: 62 NELYNAVQLYLSSSVSIAGNRLSLTRAVNSSSVTFGLSNNDSIVDTFNSVTVVWEHIV-- 119
Query: 127 VDSSSSRTRYTNMSSSLMSEVRSYELSFHKKHKEKMFNSYLPYVLERAKAIKQENMAVKI 186
+ R T + E R + L KK K + +SYL Y++E+A I++ N +
Sbjct: 120 ----TQRQTQTFAWRPMPEEKRGFTLRIKKKDKNLILDSYLDYIMEKANEIRRLNQDRLL 175
Query: 187 YT------VEYEAWCRNGTRFDHPMTFKTLAIDAGLKKELVSDLDRFLQGKEFYNRTGKA 240
YT ++ F HP TF TLA+D K++++ DL F + + FY RTG+A
Sbjct: 176 YTNSRGGSLDSRGLPWESVPFKHPSTFDTLAMDPVKKQQIMEDLKDFAECQSFYERTGRA 235
Query: 241 WKRGYLLYGPPGTGKSSLIAAMANYLSYDIYDLDLTALNDNKDLKNLLLAMSNRSILVIE 300
WKRGYLLYGPPGTGKSS+IAAMANYL YDIYDL+LT + N +L+ LL+ S++SI+VIE
Sbjct: 236 WKRGYLLYGPPGTGKSSMIAAMANYLRYDIYDLELTEVKSNSELRKLLMKTSSKSIIVIE 295
Query: 301 DIDCSIKLPNREEDEEAAKNGD----------------NKMTLSGLLNVIDGLWSCCGEE 344
DIDCSI L NR + + + N +TLSGLLN DGLWSCCG E
Sbjct: 296 DIDCSINLTNRNKKQSTGSYNEPEMLTGSGLGDDLGDGNTITLSGLLNFTDGLWSCCGSE 355
Query: 345 RIIIFTTNHRERLDPALLRPGRMDMHVNLSYCTFSAFEQLAFTYLGISQHKL----FEQI 400
RI +FTTNH E+LDPALLR GRMDMH+++SYCTFS+ + L YLG + L +++
Sbjct: 356 RIFVFTTNHIEKLDPALLRSGRMDMHIHMSYCTFSSVKILLRNYLGFEEGDLNDVVLKEL 415
Query: 401 GGLLRHVQVTPAEVAGQLTK-IADVEECLHGLVKFLQDKTQNDQNINQ---DNASQKEKK 456
++ ++TPA+V+ L K D E + L+ L+ + + ++ + N S +E++
Sbjct: 416 AEVVDRAEITPADVSEALIKNRRDKERAVRELLVDLRSRVERNEKNGKSRVQNVSLEEQE 475
Query: 457 NQ--DHEYGIDN 466
N+ D Y +N
Sbjct: 476 NRAFDSLYAEEN 487
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.133 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 751,123,414
Number of Sequences: 2790947
Number of extensions: 31210644
Number of successful extensions: 123160
Number of sequences better than 10.0: 3207
Number of HSP's better than 10.0 without gapping: 1889
Number of HSP's successfully gapped in prelim test: 1321
Number of HSP's that attempted gapping in prelim test: 118501
Number of HSP's gapped (non-prelim): 3971
length of query: 467
length of database: 848,049,833
effective HSP length: 131
effective length of query: 336
effective length of database: 482,435,776
effective search space: 162098420736
effective search space used: 162098420736
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0111b.4


