

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0111a.1
(863 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
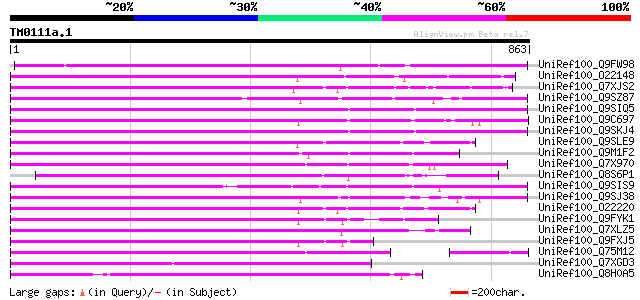
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 568 e-160
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 529 e-148
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 523 e-146
UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabi... 507 e-142
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 505 e-141
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 503 e-140
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 502 e-140
UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcrip... 482 e-134
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 481 e-134
UniRef100_Q7X970 BZIP-like protein [Oryza sativa] 476 e-132
UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa] 476 e-132
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 472 e-131
UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcrip... 471 e-131
UniRef100_O22220 Putative non-LTR retroelement reverse transcrip... 466 e-129
UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana] 459 e-127
UniRef100_Q7XLZ5 OSJNBa0086O06.14 protein [Oryza sativa] 450 e-124
UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana] 444 e-123
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 436 e-120
UniRef100_Q7XGD3 Putative retroelement [Oryza sativa] 430 e-118
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 423 e-116
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 568 bits (1463), Expect = e-160
Identities = 311/868 (35%), Positives = 477/868 (54%), Gaps = 22/868 (2%)
Query: 8 LPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLATQAQSA 67
+P + ++VV L+PKV+ ++ + RPIS CN LYK+ +K+L N+LKP + + ++ QSA
Sbjct: 496 IPEGLCDSVVVLIPKVNNASHLSKFRPISLCNVLYKIASKVLANRLKPFLPDIVSEFQSA 555
Query: 68 FISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACLTSFGFS 127
F+ GR I D+ L+A+E H ++ K F A+K++M KAYDR+EW +L CL+ GFS
Sbjct: 556 FVPGRLITDSALVAYECLHTIRKQ-HNKNPFFALKIDMMKAYDRVEWAYLSGCLSKLGFS 614
Query: 128 EKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSHLLLKAQ 187
+ W++ +M+CV+ + K+NGE++K VVP RGIRQG P+SPYLF+L E +S LL K +
Sbjct: 615 QDWINTVMRCVSSVRYAVKINGELTKPVVPSRGIRQGDPISPYLFLLCTEGLSCLLHKKE 674
Query: 188 EEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQMISIEKS 247
G +QG++ PPI+HLLFADDS+FFAKA+ L + L Y ASGQ I++ KS
Sbjct: 675 VAGELQGIKNGRHGPPISHLLFADDSIFFAKADSRNVQALKNTLRSYCSASGQKINLHKS 734
Query: 248 GIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVLDKIDGW 307
I F + + K + S L++ YLG+P E G + N ++ R+ +++GW
Sbjct: 735 SIFFGKRCPDAVKISVKSCLQVDNEVLQDSYLGMPTEIGLATTNFFKFLPERIWKRVNGW 794
Query: 308 KEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKERGIHWK 367
++ L++AG E ++KAV QA+ Y MS R P + C+ + IA WW ++ +HWK
Sbjct: 795 TDRPLSRAGMETMLKAVAQAIPNYVMSCFRIPVSICEKMKTCIADHWWGFEDGKKKMHWK 854
Query: 368 DWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNSSFLQSG 427
W ++S K GG+GF++F+T N A+L +Q WR+ T PD+L R LKG YFPNSSF ++
Sbjct: 855 SWSWLSTPKFLGGMGFREFTTFNQAMLGRQCWRLLTDPDSLCSRVLKGRYFPNSSFWEAA 914
Query: 428 RTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWV---SDQVLQAPEAGDAELT 484
+ S+ W+S+L G+E L + W +GDG +K++ D W+ Q++ + T
Sbjct: 915 QPKSPSFTWRSLLFGRELLAKGVRWGVGDGKTIKIFSDNWIPGFRPQLVTTLSPFPTDAT 974
Query: 485 VNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTVKTGYQ 544
V+ L+ D + W+ + S FP IA ++ QIP+S D WPH K G Y+V++ Y
Sbjct: 975 VSCLMNEDARCWDGDLIRSLFPVDIAKEILQIPISRHGDADFASWPHDKLGLYSVRSAYN 1034
Query: 545 VAYS------LKEQARGPQPHIL-NDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLF 597
+A S RG +L + W +W I P K+K+ LW+A +E L L
Sbjct: 1035 LARSEAFFADQSNSGRGMASRLLESQKDWKGLWKINAPGKMKITLWRAAHECLATGFQLR 1094
Query: 598 RRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVW--LSSQLHVQHLHQGSLTVKDWLTQL 655
RRHI C CN+D ++V H+ L CP+ +W + + V+ G T++ W+
Sbjct: 1095 RRHIPSTDGCVFCNRD-DTVEHVFLFCPFAAQIWEEIKGKCAVKLGRNGFSTMRQWIFDF 1153
Query: 656 VIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNGMEDNT 715
+ R S T++A W IW+ RN P I+I + + ++ NT
Sbjct: 1154 LKR------GSSHANTLLAVTFWHIWEARNNTKNNNGTVHPQRVVIKILSYVDMILKHNT 1207
Query: 716 PSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNVLSHSKT 775
+ +G RW PP +N DA F + + ++ D G +++ S+
Sbjct: 1208 KTVDGQRGGNTQAIPRWQPPPASVWMINSDAAIFSSSRTMGVGALIRDNTGKCLVACSEM 1267
Query: 776 IW-AASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWK-VEAVLRDI 833
I P AEALA+R AL LA L I + SDCL ++ I+ D V V+ DI
Sbjct: 1268 ISDVVLPELAEALAIRRALGLAKEEGLEHIVMASDCLTVIRRIQTSGRDRSGVGCVIEDI 1327
Query: 834 QD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
+ + F +C F V R NL AH +A+
Sbjct: 1328 KKLASTFVLCSFMHVNRLSNLAAHSLAR 1355
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 529 bits (1363), Expect = e-148
Identities = 295/862 (34%), Positives = 457/862 (52%), Gaps = 33/862 (3%)
Query: 1 SFFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
+FF+ G + +N+T + L+PK+ K E + RPIS CN +YKVI K++ N+LK + L
Sbjct: 473 AFFRSGSIEEGMNKTNICLIPKILKAEKMTDFRPISLCNVIYKVIGKLMANRLKKILPSL 532
Query: 61 ATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKAC 120
++ Q+AF+ GR I DNILIAHE H L + K EF+AIK +++KAYDR+EW FL+
Sbjct: 533 ISETQAAFVKGRLISDNILIAHELLHALSSNNKCSEEFIAIKTDISKAYDRVEWPFLEKA 592
Query: 121 LTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMS 180
+ GF++ W+ IM+CV ++ +NG ++P RG+RQG PLSPYLFV+ E +
Sbjct: 593 MRGLGFADHWIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLV 652
Query: 181 HLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQ 240
+L A+++ +I GL++A PPI+HLLFADDS+F+ K N E Q+I I+ +Y+LASGQ
Sbjct: 653 KMLQSAEQKNQITGLKVARGAPPISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQ 712
Query: 241 MISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRV 300
++ KS I F ++ + + L + G YLGLP + S+ TL+++K R+
Sbjct: 713 RVNYLKSSIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRL 772
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGK 360
K+ GW+ L+ GKE+L+KAV A+ YTMS + P+ CQ I + +A FWW++ +
Sbjct: 773 GKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFWWKNKKE 832
Query: 361 ERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPN 420
RG+HWK W +S+ K GGLGFK+ N+ALL KQ WR+ T+ D+L + K YF
Sbjct: 833 GRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSK 892
Query: 421 SSFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPE--- 477
S L + +R S+ W+SI + + +K+ +IG+G + VW D W+ + +A +
Sbjct: 893 SDPLNAPLGSRPSFAWKSIYEAQVLIKQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVK 952
Query: 478 --------AGDAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCW 529
A ++ V L+LPD + WN + FP + + + D+ W
Sbjct: 953 RSHLVSQYAANSIHVVKDLLLPDGRDWNWNLVSLLFPDNTQENILALRPGGKETRDRFTW 1012
Query: 530 PHTKDGEYTVKTGYQVAYSLKEQARGPQPHILN---DPIWNQIWGIQVP*KIKMFLWKAC 586
+++ G Y+VK+GY V + Q PQ +L DPI+ QIW + VP KI FLW+
Sbjct: 1013 EYSRSGHYSVKSGYWVMTEIINQRNNPQ-EVLQPSLDPIFQQIWKLDVPPKIHHFLWRCV 1071
Query: 587 NEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSL 646
N L V ++L RH+A + C C E+V HLL CP+ R W S L
Sbjct: 1072 NNCLSVASNLAYRHLAREKSCVRCPSHGETVNHLLFKCPFARLTWAISPLPAP------- 1124
Query: 647 TVKDWLTQL------VIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTT 700
+W L V+ + ++ + S ++ + LW +WK RN+ F+ R
Sbjct: 1125 PGGEWAESLFRNMHHVLSVHKSQPEESDHHALIPWILWRLWKNRNDLVFKGREFTAPQVI 1184
Query: 701 IRIRASMLNGMEDNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVV 760
++ M P P T + +W P +K N D + K+ + V
Sbjct: 1185 LKATEDMDAWNNRKEPQPQVTSSTRDRC-VKWQPPSHGWVKCNTDGAWSKDLGNCGVGWV 1243
Query: 761 VFDEKGTNVLSHSKTIWA-ASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRE 819
+ + G + + + + S L E A+R A+L ++ SD LVSLI
Sbjct: 1244 LRNHTGRLLWLGLRALPSQQSVLETEVEALRWAVLSLSRFNYRRVIFESDSQYLVSLI-- 1301
Query: 820 RRTDWKVEAVLRDIQD*RNLFR 841
+ + + ++ IQD RNL R
Sbjct: 1302 -QNEMDIPSLAPRIQDIRNLLR 1322
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 523 bits (1346), Expect = e-146
Identities = 305/848 (35%), Positives = 468/848 (54%), Gaps = 30/848 (3%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ G LP N T + L+PK+ P+ + LRPIS C+ LYK+I+KIL +LK H+ +
Sbjct: 452 FFETGVLPQDWNHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIV 511
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ QSAF+ R I DNIL+AHE H L+ + E MA K +M+KAYDR+EW FL+ +
Sbjct: 512 STTQSAFVPQRLISDNILVAHEMIHSLRTNDRISKEHMAFKTDMSKAYDRVEWPFLETMM 571
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
T+ GF+ KW+ IM CVT S+ +NG+ ++P RGIRQG PLSP LFVL E + H
Sbjct: 572 TALGFNNKWISWIMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIH 631
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
+L KA++ G+I G++ + + HLLFADD+L KA ++E +L+ L+ Y SGQM
Sbjct: 632 ILNKAEQAGKITGIQFQDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQM 691
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I++ KS I F +N K I S +S GKYLGLP S+ + +IK ++
Sbjct: 692 INLNKSAITFGKNVDIQIKDWIKSRSGISLEGGTGKYLGLPECLSGSKRDLFGFIKEKLQ 751
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
++ GW K L+Q GKEVL+K++ A+ Y MS + P+N CQ + + FWW S ++
Sbjct: 752 SRLTGWYAKTLSQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNSMQQK 811
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R IHW W ++ K +GG GFKD N ALLAKQAWR+ + +L+ R + YF NS
Sbjct: 812 RKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNS 871
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSD----QVLQAPE 477
FL + R +R S+ W+SIL G+E L + +IG+G + VW DKW+ D + L
Sbjct: 872 DFLSATRGSRPSYAWRSILFGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRR 931
Query: 478 AGDAELTVNSLILPDVKQWNLPKLLSTFP-RQIAFKVFQIPLSLTQVPDKLCWPHTKDGE 536
+ +L V+ LI P + WNL L FP + + + Q PL + D CW H+ +G
Sbjct: 932 FINVDLKVSQLIDPTSRNWNLNMLRDLFPWKDVEIILKQRPLFFKE--DSFCWLHSHNGL 989
Query: 537 YTVKTGY-----QVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALP 591
Y+VKTGY QV + L ++A+ +P + + ++++IW + KI++FLWKA + A+P
Sbjct: 990 YSVKTGYEFLSKQVHHRLYQEAK-VKPSV--NSLFDKIWNLHTAPKIRIFLWKALHGAIP 1046
Query: 592 VKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDW 651
V+ L R I D C +C+ + E++ H+L CP R VW + L + S +V
Sbjct: 1047 VEDRLRTRGIRSDDGCLMCDTENETINHILFECPLARQVWAITHLSSAG-SEFSNSVYTN 1105
Query: 652 LTQLVIRLQENSEDLSRGLTIVA-YYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNG 710
+++L+ Q+N DL L V+ + LW +WK RN F+ + + TT+ +A
Sbjct: 1106 MSRLIDLTQQN--DLPHHLRFVSPWILWFLWKNRNALLFEGK--GSITTTLVDKA--YEA 1159
Query: 711 MEDNTPSPIPTQGRKPNYQ-ARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNV 769
+ + Q + + + +W P P LK N+ + K+ + VV D +G V
Sbjct: 1160 YHEWFSAQTHMQNDEKHLKITKWCPPLPGELKCNIGFAWSKQHHFSGASWVVRDSQG-KV 1218
Query: 770 LSHSKTIW--AASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWKVE 827
L HS+ + SP +A+ + AL + ++ S ++ + + +W +
Sbjct: 1219 LLHSRRSFNEVHSPYSAKIRSWEWALESMTHHHFDRVIFASSTHEIIQALHKPH-EWPL- 1276
Query: 828 AVLRDIQD 835
+L DI +
Sbjct: 1277 -LLGDISE 1283
>UniRef100_Q9SZ87 RNA-directed DNA polymerase-like protein [Arabidopsis thaliana]
Length = 1274
Score = 507 bits (1306), Expect = e-142
Identities = 319/881 (36%), Positives = 460/881 (52%), Gaps = 54/881 (6%)
Query: 1 SFFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
SFF + L P +NET V L+PK+ P + RPI+ CN YK++ KIL +L+P ++ L
Sbjct: 414 SFFVDSCLSPRLNETHVTLIPKISAPRKVSDYRPIALCNVQYKIVAKILTRRLQPWLSEL 473
Query: 61 ATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKAC 120
+ QSAF+ GR I DN+LI HE HFL+ + K MAIK +M+KAYDR++W+FL+
Sbjct: 474 ISLHQSAFVPGRAIADNVLITHEILHFLRVSGAKKYCSMAIKTDMSKAYDRIKWNFLQEV 533
Query: 121 LTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMS 180
L GF +KW+ +MQCV S+ F +NG VVP RG+RQG PLSPYLF+L E +S
Sbjct: 534 LMRLGFHDKWIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLS 593
Query: 181 HLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQ 240
L KAQE+G + G+R+A P + HLLFADD++FF K N L +IL Y LASGQ
Sbjct: 594 GLCRKAQEKGVMVGIRVARGSPQVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQ 653
Query: 241 MISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRV 300
I++ KS I FS T K ++ LR+ GKYLGLP +G+ + + + I R+
Sbjct: 654 SINLAKSAITFSSKTPQDIKRRVKLSLRIDNEGGIGKYLGLPEHFGRRKRDIFSSIVDRI 713
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGK 360
+ W + L+ AGK++L+KAVL +M +Y M + P + C+ I + + RFWW S
Sbjct: 714 RQRSHSWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFWWDSKPD 773
Query: 361 ERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPN 420
+R + W W ++ +GGLGF++ AK +WRI +P +L R L G Y
Sbjct: 774 KRKMAWVSWDKLTLPINEGGLGFREIE-------AKLSWRILKEPHSLLSRVLLGKYCNT 826
Query: 421 SSFLQ-SGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAG 479
SSF+ S + S GW+ IL G++ L++ W IG G + VW + W+S Q P
Sbjct: 827 SSFMDCSASPSFASHGWRGILAGRDLLRKGLGWSIGQGDSINVWTEAWLSPSSPQTPIGP 886
Query: 480 DAE----LTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDG 535
E L+V+ LI DVK WN+ + P Q ++ +I ++ + D L W K G
Sbjct: 887 PTETNKDLSVHDLICHDVKSWNVEAIRKHLP-QYEDQIRKITINALPLQDSLVWLPVKSG 945
Query: 536 EYTVKTGYQVAYSLKEQARGPQPHILNDPIWNQ-IWGIQVP*KIKMFLWKACNEALPVKA 594
EYT KTGY +A P D W + IW I K+K FLWKA ALPV
Sbjct: 946 EYTTKTGYALA------KLNSFPASQLDFNWQKNIWKIHTSPKVKHFLWKAMKGALPVGE 999
Query: 595 SLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQ 654
+L RR+I + C C Q E S+ HL+L CP+ + VW + + L S +
Sbjct: 1000 ALSRRNIEAEVTCKRCGQTESSL-HLMLLCPYAKKVWELAPV----LFNPSEATHSSVAL 1054
Query: 655 LVIRLQENSEDLSRGLTIVAYY---LWGIWKERNECHFQQRPPDPLGTTIR----IRASM 707
L++ + GL Y LW +WK RN F G ++ RA M
Sbjct: 1055 LLVDAKRMVALPPTGLGSAPLYPWLLWHLWKARNRLIFDNHSCSEEGLVLKAILDARAWM 1114
Query: 708 LNGMEDNTPSPIPTQGRKPNYQARWDVPPPW-TLKLN---VDATYFKETCKGSIAVVVFD 763
+ + PSPI D P P LK+ VDA + T G + F
Sbjct: 1115 EAQLLIHHPSPIS------------DYPSPTPNLKVTSCFVDAAW---TTSGYCGMGWFL 1159
Query: 764 EKGTNVL---SHSKTIWAASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRER 820
+ V + S + + S L AE LAV AL+ A++ + ++ + SDC L+SL+
Sbjct: 1160 QDPYKVKIKENQSSSSFVGSALMAETLAVHLALVDALSTGVRQLNVFSDCKELISLLNSG 1219
Query: 821 RTDWKVEAVLRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
++ ++ +L DI++ F FF++PR N+ A +AK
Sbjct: 1220 KSIVELRGLLHDIRELSVSFTHLCFFFIPRLSNVVADSLAK 1260
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 505 bits (1301), Expect = e-141
Identities = 294/872 (33%), Positives = 456/872 (51%), Gaps = 17/872 (1%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ ++ P IN T + ++PK+ P + RPI+ CN LYKVI+K LVN+LK H+N +
Sbjct: 630 FFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIV 689
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ +Q+AFI GR I DN++IAHE H LK + +MA+K +++KAYDR+EWDFL+ +
Sbjct: 690 SDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTM 749
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
FGF KW+ IM V + +NG + P RGIRQG PLSPYLF+L + +SH
Sbjct: 750 RLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSH 809
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
L+ G ++G+R+ P ITHL FADDSLFF +AN L + + Y SGQ
Sbjct: 810 LINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQK 869
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+++KS I F ST+ ++ IL + GKYLGLP ++G+ + +I RV
Sbjct: 870 INVQKSMITFGSRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVK 929
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
+ W + L+ AGKE+++K+V AM Y MS + P+ I + + FWW +
Sbjct: 930 KRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQ 989
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
RGI W W + +K +GGLGF+D + N ALLAKQAWR+ P++L+ R +K YF +
Sbjct: 990 RGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDV 1049
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQA--PEAG 479
S L + + S+GW S+L G LK+ +IGDG +++ D V + E
Sbjct: 1050 SILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTEET 1109
Query: 480 DAELTVNSLI--LPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEY 537
E+T+N+L W+ K+ + + +I L+ ++ PDK+ W + GEY
Sbjct: 1110 YKEMTINNLFERKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEY 1169
Query: 538 TVKTGYQVAY---SLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKA 594
TV++GY + S A P PH D + +IW + + K+K FLW+A ++AL
Sbjct: 1170 TVRSGYWLLTHDPSTNIPAINP-PHGSID-LKTRIWNLPIMPKLKHFLWRALSQALATTE 1227
Query: 595 SLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQ 654
L R + DP+C C+++ ES+ H L CP+ W S + S ++ ++
Sbjct: 1228 RLTTRGMRIDPICPRCHRENESINHALFTCPFATMAWWLSDSSLIRNQLMSNDFEENISN 1287
Query: 655 LVIRLQENS-EDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRA---SMLNG 710
++ +Q+ + D + L + + +W IWK RN F + P T + +A LN
Sbjct: 1288 ILNFVQDTTMSDFHKLLPV--WLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNA 1345
Query: 711 MEDNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNVL 770
+ + +P PT+ N + W PP +K N DA + + + + ++ + GT +
Sbjct: 1346 TQSHKKTPSPTRQIAEN-KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPIS 1404
Query: 771 SHS-KTIWAASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWKVEAV 829
S K ++PL AE A+ AL ++++ DC L++LI +
Sbjct: 1405 WGSMKLAHTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANH 1464
Query: 830 LRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
L DI N F F ++ R N AH +AK
Sbjct: 1465 LEDISFWANKFASIQFGFIRRKGNKLAHVLAK 1496
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 503 bits (1294), Expect = e-140
Identities = 302/880 (34%), Positives = 453/880 (51%), Gaps = 32/880 (3%)
Query: 1 SFFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
SF QEG +N T + L+PK ++P + +LRPIS CN YKVI+KIL +LK + L
Sbjct: 257 SFLQEGVFDKRLNTTNICLIPKTERPTRMTELRPISLCNVGYKVISKILCQRLKTVLPNL 316
Query: 61 ATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKAC 120
++ QSAF+ GR I DNILIA E FH L+ K +FMAIK +M+KAYD++EW+F++A
Sbjct: 317 ISETQSAFVDGRLISDNILIAQEMFHGLRTNSSCKDKFMAIKTDMSKAYDQVEWNFIEAL 376
Query: 121 LTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMS 180
L GF EKW+ IM C+T ++ +NG+ ++P+RG+RQG PLSPYLF+L E +
Sbjct: 377 LRKMGFCEKWISWIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLI 436
Query: 181 HLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQ 240
+ KA+ + I G+++A P ++HLLFADDSLFF KAN+E+ ++ IL Y SGQ
Sbjct: 437 ANIRKAERQNLITGIKVATPSPAVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQ 496
Query: 241 MISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRV 300
I+ KS I F S K I IL + G YLGLP G S+ ++++ R+
Sbjct: 497 QINFSKSSIQFGHKVEDSIKADIKLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRL 556
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGK 360
+I+GW K L++ GKEV+IK+V + Y MS R P+ + +++A+FWW S G
Sbjct: 557 QSRINGWSAKFLSKGGKEVMIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNGD 616
Query: 361 ERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPN 420
RG+HW W + +K GGLGF++ N ALLAKQ WR+ T PD+L+ + KG YF
Sbjct: 617 SRGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRK 676
Query: 421 SSFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAG- 479
S+ L S ++ S+GW+S++ + + + + +G G+ + VW D W+ Q + + G
Sbjct: 677 SNPLDSIKSYSPSYGWRSMISARSLVYKGLIKRVGSGASISVWNDPWIPAQFPRPAKYGG 736
Query: 480 ---DAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGE 536
D L V SLI WN+ L F + + +P+ + D L W TK G
Sbjct: 737 SIVDPSLKVKSLIDSRSNFWNIDLLKELFDPEDVPLISALPIGNPNMEDTLGWHFTKAGN 796
Query: 537 YTVKTGYQVA-YSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKAS 595
YTVK+GY A L E P + + IW +Q P K++ FLW+ + +PV +
Sbjct: 797 YTVKSGYHTARLDLNEGTTLIGPDLTT--LKAYIWKVQCPPKLRHFLWQILSGCVPVSEN 854
Query: 596 LFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQL 655
L +R I D C C EES+ H L C R +W SQ+ S ++ L L
Sbjct: 855 LRKRGILCDKGCVSCGASEESINHTLFQCHPARQIWALSQIPTAPGIFPSNSIFTNLDHL 914
Query: 656 VIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNGMEDNT 715
R+ + + +W IWK RNE F+ DP+ + + E
Sbjct: 915 FWRIPSGVDSAP-----YPWIIWYIWKARNEKVFENVDKDPMEILLLAVKEAQSWQEAQV 969
Query: 716 PSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDE-KGT-----NV 769
G + +R V ++ D T+ C + D+ GT +
Sbjct: 970 ELHSERHG-SLSIDSRIRV-----RDVSQDTTFSGFRCFIDGSWKASDQFSGTGWFCLSS 1023
Query: 770 LSHSKTIWAA------SPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTD 823
L S T+ AA SPL E A+ A+ + + + +DC +LV ++ T+
Sbjct: 1024 LGESPTMGAANVRRSLSPLHTEMEALLWAMKCMIGADNQNVAFFTDCSDLVKMV-SSPTE 1082
Query: 824 WKVEAV-LRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAKQ 862
W +V L ++Q R F + R+ N+ A +A++
Sbjct: 1083 WPAFSVYLEELQSDREEFTNFSLSLISRSANVKADKLARK 1122
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 502 bits (1293), Expect = e-140
Identities = 293/872 (33%), Positives = 455/872 (51%), Gaps = 17/872 (1%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ ++ P IN T + ++PK+ P + RPI+ CN LYKVI+K LVN+LK H+N +
Sbjct: 856 FFETSFMKPSINHTNICMIPKITNPTTLSDYRPIALCNVLYKVISKCLVNRLKSHLNSIV 915
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ +Q+AFI GR I DN++IAHE H LK + +MA+K +++KAYDR+EWDFL+ +
Sbjct: 916 SDSQAAFIPGRIINDNVMIAHEVMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTM 975
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
FGF KW+ IM V + +NG + P RGIRQG PLSPYLF+L + +SH
Sbjct: 976 RLFGFCNKWIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSH 1035
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
L+ G ++G+R+ P ITHL FADDSLFF +AN L + + Y SGQ
Sbjct: 1036 LINGRASSGDLRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQK 1095
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+++KS I F ST+ ++ IL + GKYLGLP ++G+ + +I RV
Sbjct: 1096 INVQKSMITFGSRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVK 1155
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
+ W + L+ AGKE+++K+V AM Y MS + P+ I + + FWW +
Sbjct: 1156 KRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFWWEKASNQ 1215
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
RGI W W + +K +GGLGF+D + N ALLAKQAWR+ P++L+ R +K YF +
Sbjct: 1216 RGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARYFKDV 1275
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQA--PEAG 479
S L + + S+GW S+L G LK+ +IGDG +++ D V + E
Sbjct: 1276 SILDAKVRKQQSYGWASLLDGIALLKKGTRHLIGDGQNIRIGLDNIVDSHPPRPLNTEET 1335
Query: 480 DAELTVNSLI--LPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEY 537
E+T+N+L W+ K+ + + +I L+ ++ PDK+ W + GEY
Sbjct: 1336 YKEMTINNLFERKGSYYFWDDSKISQFVDQSDHGFIHRIYLAKSKKPDKIIWNYNTTGEY 1395
Query: 538 TVKTGYQVAY---SLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKA 594
TV++GY + S A P PH D + +IW + + K+K FLW+A ++AL
Sbjct: 1396 TVRSGYWLLTHDPSTNIPAINP-PHGSID-LKTRIWNLPIMPKLKHFLWRALSQALATTE 1453
Query: 595 SLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQ 654
L R + DP C C+++ ES+ H L CP+ W S + S ++ ++
Sbjct: 1454 RLTTRGMRIDPSCPRCHRENESINHALFTCPFATMAWRLSDSSLIRNQLMSNDFEENISN 1513
Query: 655 LVIRLQENS-EDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRA---SMLNG 710
++ +Q+ + D + L + + +W IWK RN F + P T + +A LN
Sbjct: 1514 ILNFVQDTTMSDFHKLLPV--WLIWRIWKARNNVVFNKFRESPSKTVLSAKAETHDWLNA 1571
Query: 711 MEDNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNVL 770
+ + +P PT+ N + W PP +K N DA + + + + ++ + GT +
Sbjct: 1572 TQSHKKTPSPTRQIAEN-KIEWRNPPATYVKCNFDAGFDVQKLEATGGWIIRNHYGTPIS 1630
Query: 771 SHS-KTIWAASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWKVEAV 829
S K ++PL AE A+ AL ++++ DC L++LI +
Sbjct: 1631 WGSMKLAHTSNPLEAETKALLAALQQTWIRGYTQVFMEGDCQTLINLINGISFHSSLANH 1690
Query: 830 LRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
L DI N F F ++ + N AH +AK
Sbjct: 1691 LEDISFWANKFASIQFGFIRKKGNKLAHVLAK 1722
>UniRef100_Q9SLE9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1319
Score = 482 bits (1240), Expect = e-134
Identities = 270/783 (34%), Positives = 415/783 (52%), Gaps = 22/783 (2%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ G LP N T + L+PK P+ + +RPIS C+ LYK+I+KIL KLK H+ +
Sbjct: 426 FFETGVLPQEWNHTHLYLIPKFTNPQRMSDIRPISLCSVLYKIISKILSFKLKKHLPSIV 485
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ +QSAF + R I DNILIAHE H L+ K EFM K +M+KAYDR+EW FL+ L
Sbjct: 486 SPSQSAFFAERLISDNILIAHEIVHSLRTNDKISKEFMVFKTDMSKAYDRVEWSFLQEIL 545
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
+ GF++KW+ IM CVT ++ +NG+ + P+RGIRQG P+SP+LFVL E + H
Sbjct: 546 VALGFNDKWISWIMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISPFLFVLCTEALIH 605
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
+L +A+ ++ G++ P + HLLF DD+ +A + + Q++ L+ Y SGQ+
Sbjct: 606 ILQQAENSKKVSGIQFNGSGPSVNHLLFVDDTQLVCRATKSDCEQMMLCLSQYGHISGQL 665
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I++EKS I F TK I + + GKYLGLP S+ + +IK ++
Sbjct: 666 INVEKSSITFGVKVDEDTKRWIKNRSGIHLEGGTGKYLGLPENLSGSKQDLFGYIKEKLQ 725
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
+ GW +K L+Q GKE+L+K++ A+ Y M+ R P+ C + + + FWW S
Sbjct: 726 SHLSGWYDKTLSQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFWWNSMEFS 785
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
IHW ++ K GG GFKD N ALLAKQAWR+++ ++ + K YF N+
Sbjct: 786 NKIHWIGGKKLTLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNT 845
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAP----E 477
FL + + R S+ W+SIL G+E L +IG+G + VW DKW+ D + P
Sbjct: 846 DFLNARQGTRPSYTWRSILYGRELLNGGLKRLIGNGEQTNVWIDKWLFDGHSRRPMNLHS 905
Query: 478 AGDAELTVNSLILPDVKQWNLPKLLSTF-PRQIAFKVFQIPLSLTQVPDKLCWPHTKDGE 536
+ + V+ LI P + WNL KL F + + + Q PL ++ D CW T +G
Sbjct: 906 LMNIHMKVSHLIDPLTRNWNLKKLTELFHEKDVQLIMHQRPLISSE--DSYCWAGTNNGL 963
Query: 537 YTVKTGYQVA--YSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKA 594
YTVK+GY+ + + K + + +P+++++W ++ KIK+F+WKA AL V+
Sbjct: 964 YTVKSGYERSSRETFKNLFKEADVYPSVNPLFDKVWSLETVPKIKVFMWKALKGALAVED 1023
Query: 595 SLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLTQ 654
L R I C C ++ E++ HLL CP+ R VW S + G+ +
Sbjct: 1024 RLRSRGIRTADGCLFCKEEIETINHLLFQCPFARQVWALSLIQAPATGFGTSIFSN--IN 1081
Query: 655 LVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNGMEDN 714
VI+ +N T+ + LW IWK RN+ FQ GT + + E+
Sbjct: 1082 HVIQNSQNFGIPRHMRTVSPWLLWEIWKNRNKTLFQ-------GTGLTSSEIVAKAYEEC 1134
Query: 715 T---PSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNVLS 771
+ + G + +W+ PP LK N+ + ++ ++ V+ D G VL
Sbjct: 1135 NLWINAQEKSSGGVSPSEHKWNPPPAGELKCNIGVAWSRQKQLAGVSWVLRDSMG-QVLL 1193
Query: 772 HSK 774
HS+
Sbjct: 1194 HSR 1196
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 481 bits (1237), Expect = e-134
Identities = 267/759 (35%), Positives = 408/759 (53%), Gaps = 18/759 (2%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ ++ +N T + ++PK+ P+ + RPI+ CN LYKVI+K +VN+LK H+N +
Sbjct: 95 FFESSHMKTSVNHTNICMIPKIQNPQTLSDYRPIALCNVLYKVISKCMVNRLKAHLNSIV 154
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ +Q+AFI GR I DN++IAHE H LK + +MA+K +++KAYDR+EWDFL+ +
Sbjct: 155 SDSQAAFIPGRIINDNVMIAHEIMHSLKVRKRVSKTYMAVKTDVSKAYDRVEWDFLETTM 214
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
FGF +KW+ IM V + +NG + P RGIRQG PLSPYLF+L + +SH
Sbjct: 215 RLFGFCDKWIGWIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSH 274
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
L+ G I+G+R+ P ITHL FADDSLFF +AN L + + Y SGQ
Sbjct: 275 LIKVKASSGDIRGVRIGNGAPAITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQK 334
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+++KS I F ST+ ++ ++L + GKYLGLP ++G+ + N+I RV
Sbjct: 335 INVQKSLITFGSRVYGSTQTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVK 394
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
++ W K L+ AGKE+L+K+V AM Y MS + PQ I + + FWW +
Sbjct: 395 ERTASWSAKFLSPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFWWEKASNK 454
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
RGI W W + +K +GGLGF+D + N ALLAKQAWRI P++L+ R +K YF ++
Sbjct: 455 RGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDN 514
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAGDA 481
S + + ++ S+GW S+L G L++ ++IGDG +++ D V D P D
Sbjct: 515 SIIDAKTRSQQSYGWSSLLSGIALLRKGTRYVIGDGKTIRLGIDN-VVDSHPPRPLLTDE 573
Query: 482 ELTVNSLILPDVKQ-------WNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKD 534
+ N L L ++ Q W+ KL + + + +I LS D+L W +
Sbjct: 574 Q--HNGLSLDNLFQHRGHSRCWDNAKLQTFVDQSDHDYIKRIYLSTRSKTDRLIWSYNST 631
Query: 535 GEYTVKTGYQVAYSLKEQA--RGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPV 592
G+YTV++GY ++ +PH D + +IW + + K+K FLW+ ++ALP
Sbjct: 632 GDYTVRSGYWLSTHDPSNTIPTMAKPHGSVD-LKTKIWNLPIMPKLKHFLWRILSKALPT 690
Query: 593 KASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWL 652
L R + DP C C ++ ES+ H L CP+ W S + S ++D +
Sbjct: 691 TDRLTTRGMRIDPGCPRCRRENESINHALFTCPFATMAWRLSDTPLYRSSILSNNIEDNI 750
Query: 653 TQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRA---SMLN 709
+ +++ LQ + S+ L I + LW IWK RN F P T +R +A LN
Sbjct: 751 SNILLLLQNTTITDSQKL-IPFWLLWRIWKARNNVVFNNLRESPSITVVRAKAETNEWLN 809
Query: 710 GMEDNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATY 748
+ P +P + W P +K N DA++
Sbjct: 810 ATQTQGPRRLPKRTTAAG-NTTWVKPQMPYIKCNFDASF 847
>UniRef100_Q7X970 BZIP-like protein [Oryza sativa]
Length = 2367
Score = 476 bits (1225), Expect = e-132
Identities = 272/852 (31%), Positives = 445/852 (51%), Gaps = 33/852 (3%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FFQ G +P +N+T + L+PK ++P ++ RPIS CN +YKV++K LVN+L+P ++ L
Sbjct: 1208 FFQTGLMPEGVNDTAIVLIPKKEQPVDLRDFRPISLCNVVYKVVSKCLVNRLRPILDDLV 1267
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ QSAF+ GR I DN L+A E FH ++ K A KL+++KAYDR++W FL+ +
Sbjct: 1268 SVEQSAFVQGRMITDNALLAFECFHAMQKNKKANHAACAYKLDLSKAYDRVDWRFLEMAM 1327
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
GF+ +WV+ IM+CVT + K NG + + P RG+RQG PL P+LF+ A+ +S
Sbjct: 1328 NKLGFARRWVNWIMKCVTSVRYMVKFNGTLLQSFAPTRGLRQGDPLLPFLFLFVADGLSL 1387
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
LL + + + ++ P I+HLLFADD+L F KA++ EA + +L+ Y + +GQ+
Sbjct: 1388 LLKEKVAQNSLTPFKVCRAAPGISHLLFADDTLLFFKAHQREAEVVKEVLSSYAMGTGQL 1447
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ K I+ + + IS IL++ +YLG P G+ ++ ++
Sbjct: 1448 INPAKCSILMGGASTPAVSEAISEILQVERDRFEDRYLGFPTPEGRMHKGRFQSLQAKIW 1507
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
++ W E L+ GKEVLIKAV+QA+ Y M I + P++ + FWW S +
Sbjct: 1508 KRVIQWGENHLSTGGKEVLIKAVIQAIPVYVMGIFKLPESVIDDLTKLTKNFWWDSMNGQ 1567
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R HWK W ++K K GGLGF+D+ N ALLA+QAWR+ T PD+L R LK YFP+
Sbjct: 1568 RKTHWKAWDSLTKPKSLGGLGFRDYRLFNQALLARQAWRLITYPDSLCARVLKAKYFPHG 1627
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAGDA 481
S + + + +S W+SI G + LK+ +W +G+G+ +++W+D W+ + P G A
Sbjct: 1628 SLIDTSFGSNSSPAWRSIEYGLDLLKKGIIWRVGNGNSIRIWRDSWLPRDHSRRPITGKA 1687
Query: 482 ELT---VNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYT 538
V+ LI D W++PK+ F A + I +S D + W K+G ++
Sbjct: 1688 NCRLKWVSDLITED-GSWDVPKIHQYFHNLDAEVILNICISSRSEEDFIAWHPDKNGMFS 1746
Query: 539 VKTGYQVAYSLK--EQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASL 596
V++ Y++A L E++ + +N W IW +VP K+K+F W+ + L +
Sbjct: 1747 VRSAYRLAAQLVNIEESSSSGTNNIN-KAWEMIWKCKVPQKVKIFAWRVASNCLATMVNK 1805
Query: 597 FRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVW--LSSQLHVQHLHQGSLTVKDWLTQ 654
+R + +C IC+++ E H L C +W + V + S+ + WL
Sbjct: 1806 KKRKLEQSDMCQICDRENEDDAHALCRCIQASQLWSCMHKSGSVSVDIKASVLGRFWLFD 1865
Query: 655 LVIRLQENSEDLSRGLTIVAYYLWGIWKERNE-CHFQQRPPDP---------LGTTIRIR 704
+ ++ E + + LW W RNE H + PP + +IR
Sbjct: 1866 CLEKIPEYEQ------AMFLMTLWRNWYVRNELIHGKSAPPTETSQRFIQSYVDLLFQIR 1919
Query: 705 ----ASMLNGMEDNTPSPI---PTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSI 757
A ++ G P+ P N+Q W+ P +KLNVD ++ + KG +
Sbjct: 1920 QAPQADLVKGKHVVRTVPLKGGPKYRVLNNHQPCWERPKDGWMKLNVDGSFDASSGKGGL 1979
Query: 758 AVVVFDEKGTNVLSHSKTIWAA-SPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSL 816
+++ + G + + K + +PL +E A E L LA++ L I + +DC ++V L
Sbjct: 1980 GMILRNSAGDIIFTSCKPLERCNNPLESELRACVEGLKLAIHWTLLPIQVETDCASVVQL 2039
Query: 817 IRERRTDWKVEA 828
++ D+ V A
Sbjct: 2040 LQGIGRDFSVLA 2051
>UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa]
Length = 1509
Score = 476 bits (1224), Expect = e-132
Identities = 275/781 (35%), Positives = 412/781 (52%), Gaps = 53/781 (6%)
Query: 44 VITKILVNKLKPHMNRLATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKL 103
+I K+L N+LK + + + AQSAF+ GR I DNILIA+E H+++N G+ + A KL
Sbjct: 717 LIPKVLANRLKKILPDVISPAQSAFVPGRLISDNILIAYEMTHYMRNKRSGQVGYAAFKL 776
Query: 104 EMNKAYDRLEWDFLKACLTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQ 163
+M+KAYDR+EW FL + GF WV+ IM+CV+ ++R +VNGE+S+ P+RG+RQ
Sbjct: 777 DMSKAYDRVEWSFLHDMMLKLGFHTDWVNLIMKCVSTVTYRIRVNGELSESFSPERGLRQ 836
Query: 164 GYPLSPYLFVLAAETMSHLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREE 223
G PLSPYLF+L AE S LL K +EEGR+ G+R+ P ++HLLFADDSL +AN E
Sbjct: 837 GDPLSPYLFLLCAEGFSALLSKTEEEGRLHGIRICQGAPSVSHLLFADDSLILCRANGGE 896
Query: 224 AYQLISILNDYTLASGQMISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPA 283
A QL +IL Y SGQ+I+ +KS ++FS NT S K + + L M + KYLGLP
Sbjct: 897 AQQLQTILQIYEECSGQVINKDKSAVMFSPNTSSLEKGAVMAALNMQRETTNEKYLGLPV 956
Query: 284 EWGKSRNNTLNWIKLRVLDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFC 343
G+SR +++K R+ +I GWKEKLL++AGKE+LIKAV Q + + M ++ C
Sbjct: 957 FVGRSRTKIFSYLKERIWQRIQGWKEKLLSRAGKEILIKAVAQVIPTFAMGCFELTKDLC 1016
Query: 344 QAICASIARFWWRSPGKERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYT 403
I IA++WW + K+ +HW W+ ++ K GGLGF+D NLA+LAKQ WR+
Sbjct: 1017 DQISKMIAKYWWSNQEKDNKMHWLSWNKLTLPKNMGGLGFRDIYIFNLAMLAKQGWRLIQ 1076
Query: 404 QPDALWVRTLKGLYFPNSSFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVW 463
PD+L R L+ YFP + +T+ S+ W+SI +G L+ +W +GDGS++ +W
Sbjct: 1077 DPDSLCSRVLRAKYFPLGDCFRPKQTSNVSYTWRSIQKGLRVLQNGMIWRVGDGSKINIW 1136
Query: 464 KDKWVS---DQVLQAPEAGDAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSL 520
D W+ + P + V LI P W+ L TF + + IP+ +
Sbjct: 1137 ADPWIPRGWSRKPMTPRGANLVTKVEELIDPYTGTWDEDLLSQTFWEEDVAAIKSIPVHV 1196
Query: 521 TQVPDKLCWPHTKDGEYTVKTGYQVAYSLKEQA-RGPQPHILN-----DPIWNQIWGIQV 574
++ D L W G +TVK+ Y+V ++ +A R P + N D W ++W + V
Sbjct: 1197 -EMEDVLAWHFDARGCFTVKSAYKVQREMERRASRNGCPGVSNWESGDDDFWKKLWKLGV 1255
Query: 575 P*KIKMFLWKACNEALPVKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSS 634
P KIK FLW+ C+ L ++A+L R + D C +C + E HL C + VW +
Sbjct: 1256 PGKIKHFLWRMCHNTLALRANLHHRGMDVDTRCVMCGRYNEDAGHLFFKCKPVKKVWQAL 1315
Query: 635 QL-HVQHLHQGSLTVKDWLTQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRP 693
L ++ + + + K+ L + R EN T LW WKERNE ++
Sbjct: 1316 NLEELRSMLEQQTSGKNVLQSIYCR-PENER------TSAIVCLWQWWKERNE----EKS 1364
Query: 694 PDPLGTTIRIRASMLNGMEDNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETC 753
P + A W PP +K+N D Y
Sbjct: 1365 P------------------------------RTGECAVWRRPPLNFVKINTDGAYSSNMK 1394
Query: 754 KGSIAVVVFDEKGTNVLSHS-KTIWAASPLAAEALAVREALLLAMNLELPKIYLVSDCLN 812
+G V+ D+ G + + + + AE +A A+ A + +I L +D +
Sbjct: 1395 QGGWGFVIRDQTGAVLQAGAGPAAYLQDAFHAEVVACAAAIKTASERGMSRIELETDSMM 1454
Query: 813 L 813
L
Sbjct: 1455 L 1455
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 472 bits (1214), Expect = e-131
Identities = 286/876 (32%), Positives = 459/876 (51%), Gaps = 41/876 (4%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ + IN+T + L+PK+ P+++ RPIS C YK+I+KIL+ +LK + +
Sbjct: 836 FFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRLKQCLGDVI 895
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ +Q+AF+ G+ I DN+L+AHE H LK+ + ++ ++A+K +++KAYDR+EW+FL+ +
Sbjct: 896 SDSQAAFVPGQNISDNVLVAHELLHSLKSRRECQSGYVAVKTDISKAYDRVEWNFLEKVM 955
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
GF+ +WV IM CVT S+ +NG ++ P RGIRQG PLSPYLF+ AE +S+
Sbjct: 956 IQLGFAPRWVKWIMTCVTSVSYEVLINGSPYGKIFPSRGIRQGDPLSPYLFLFCAEVLSN 1015
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
+L KA+ +I G+++ +C I+HLLFADDSLFF +A+ + QL I Y ASGQ
Sbjct: 1016 MLRKAEVNKQIHGMKITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQK 1075
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ KS IIF + + + ++ +L + GKYLGLP + G+ + +I +V
Sbjct: 1076 INYAKSSIIFGQKIPTMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTKVK 1135
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
++ +GW L+ AGKE++IKA+ A+ Y+M+ P C I + I FWW GKE
Sbjct: 1136 ERTEGWAYNYLSPAGKEIVIKAIAMALPVYSMNCFLLPTLICNEINSLITAFWW---GKE 1192
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
+G LGFKD N ALLAKQAWRI T P +L R KGLY+PN+
Sbjct: 1193 N---------------EGDLGFKDLHQFNRALLAKQAWRILTNPQSLLARLYKGLYYPNT 1237
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAG-- 479
++L++ + S+GW SI +GK L++ +GDG K+W+D W+ + P G
Sbjct: 1238 TYLRANKGGHASYGWNSIQEGKLLLQQGLRVRLGDGQTTKIWEDPWL-PTLPPRPARGPI 1296
Query: 480 -DAELTVNSLILPDVKQWN---LPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDG 535
D ++ V L + ++W+ +L+ +Q+A ++ LS D W +T++
Sbjct: 1297 LDEDMKVADLWRENKREWDPVIFEGVLNPEDQQLAKSLY---LSNYAARDSYKWAYTRNT 1353
Query: 536 EYTVKTGYQVAYSL---KEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPV 592
+YTV++GY VA + +E+ P + P+ +IW +++ KIK F+W+ + AL
Sbjct: 1354 QYTVRSGYWVATHVNLTEEEIINPLEG--DVPLKQEIWRLKITPKIKHFIWRCLSGALST 1411
Query: 593 KASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWL 652
L R+I DP C C +E++ H++ C + + VW S+ + + +++ +
Sbjct: 1412 TTQLRNRNIPADPTCQRCCNADETINHIIFTCSYAQVVWRSANFSGSNRLCFTDNLEENI 1471
Query: 653 TQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNGME 712
++ + + + GL + + +W +WK RNE FQQ P + +E
Sbjct: 1472 RLILQGKKNQNLPILNGL-MPFWIMWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATEWVE 1530
Query: 713 ----DNTPSPIPTQG--RKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKG 766
D S Q R + +W PP LK N D+ Y + S ++ D G
Sbjct: 1531 TMVNDTAISHNTAQSNDRPLSRSKQWSSPPEGFLKCNFDSGYVQGRDYTSTGWILRDCNG 1590
Query: 767 TNVLSH-SKTIWAASPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWK 825
+ S +K + S L AEAL AL + ++ D L L +LI +
Sbjct: 1591 RVLHSGCAKLQQSYSALQAEALGFLHALQMVWIRGYCYVWFEGDNLELTNLINKTEDHHL 1650
Query: 826 VEAVLRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
+E +L DI+ +V R RNL A + K
Sbjct: 1651 LETLLYDIRFWMTKLPFSSIGYVNRERNLAADKLTK 1686
>UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1229
Score = 471 bits (1212), Expect = e-131
Identities = 303/883 (34%), Positives = 451/883 (50%), Gaps = 65/883 (7%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF P +NET + L+PK P + RPI+ CN YK++ KI+ +++ + +L
Sbjct: 369 FFTSKNFPRRMNETHIRLIPKDLGPRKVADYRPIALCNIFYKIVAKIMTKRMQLILPKLI 428
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
++ QSAF+ GR I DN+LI HE HFL+ + K MA+K +M+KAYDR+EWDFLK L
Sbjct: 429 SENQSAFVPGRVISDNVLITHEVLHFLRTSSAKKHCSMAVKTDMSKAYDRVEWDFLKKVL 488
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
FGF W+D +++CVT S+ F +NG +VVP RG+RQG PLSP LF+L E +S
Sbjct: 489 QRFGFHSIWIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSG 548
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
L +AQ ++ G+R++ P + HLLFADD++FF+K++ E +L IL+ Y ASGQ
Sbjct: 549 LCTRAQRLRQLPGVRVSINGPRVNHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQS 608
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ KS + FS T S K Q+ IL++ GKYLGLP +G+ + + I ++
Sbjct: 609 INFHKSSVTFSSKTPRSVKGQVKRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIR 668
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
K W + L+QAGK+V++KAVL +M Y+MS + P C+ I + + RFWW +
Sbjct: 669 QKSHSWASRFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRFWWDTKPDV 728
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R W W ++ K GGLGF+D N +LLAK WR+ P++L R L G Y +S
Sbjct: 729 RKTSWVAWSKLTNPKNAGGLGFRDIERCNDSLLAKLGWRLLNSPESLLSRILLGKYCHSS 788
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAGDA 481
SF++ ++ S GW+SI+ G+E LKE W+I +G +V +W D W+S P G A
Sbjct: 789 SFMECKLPSQPSHGWRSIIAGREILKEGLGWLITNGEKVSIWNDPWLSISKPLVP-IGPA 847
Query: 482 -----ELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGE 536
+L V++LI + QW+ K+ P + Q+P ++ DKL W K G+
Sbjct: 848 LREHQDLRVSALINQNTLQWDWNKIAVILPNYENL-IKQLPAPSSRGVDKLAWLPVKSGQ 906
Query: 537 YTVKTGYQVAYSLKEQARGPQPHILNDPIW-NQIWGIQVP*KIKMFLWKACNEALPVKAS 595
YT ++GY +A A P P + W + +W +Q KIK +WKA EALPV
Sbjct: 907 YTSRSGYGIA----SVASIPIPQTQFN--WQSNLWKLQTLPKIKHLMWKAAMEALPVGIQ 960
Query: 596 LFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGS--LTVKDWLT 653
L RRHI+P C C ES HL C + VW + L + GS L L
Sbjct: 961 LVRRHISPSAACHRCGA-PESTTHLFFHCEFAAQVWELAPLQETTVPPGSSMLDALSLLK 1019
Query: 654 QLVIRLQENSEDLSRGLTIVAYYLW--GIWKERNECHFQQRPPDPLGTTIRIRASMLNGM 711
+ +I G+T A + W GI+ + LGT R L
Sbjct: 1020 KAIIL-------PPTGVTSAALFPWICGIYGK-------------LGTMTRAILDALAWQ 1059
Query: 712 EDNTPSPIPTQGRKPNYQARWDVPPPWTLKL-------NVDATYFKETCKGSIAVVVFDE 764
P + R V P L + VDA + ++ S+A +
Sbjct: 1060 SAQRCLP----------KTRNVVHPISQLPVLRSGYFCFVDAAWIAQS---SLAGSGWVF 1106
Query: 765 KGTNVLSHSKTIWAA------SPLAAEALAVREALLLAMNLELPKIYLVSDCLNLVSLIR 818
+ L ++A S L+AEA A++ ALL A+ L + ++SD ++V +
Sbjct: 1107 QSATALEKETATYSAGCRRLPSALSAEAWAIKSALLHALQLGRTDLMVLSDSKSVVDALT 1166
Query: 819 ERRTDWKVEAVLRDIQD*RNLFRVCGFFWVPRTRNLCAHYVAK 861
+ ++ +L +I+ R F F ++ R+ N A AK
Sbjct: 1167 SNISINEIYGLLMEIRALRVSFHSLCFNFISRSANAIADATAK 1209
>UniRef100_O22220 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1094
Score = 466 bits (1199), Expect = e-129
Identities = 278/785 (35%), Positives = 421/785 (53%), Gaps = 26/785 (3%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FFQ G LP N T + L+PK+ KP + +RPIS C+ +YK+I+KIL +LK ++ +
Sbjct: 203 FFQTGILPEDWNHTHLCLIPKITKPARMADIRPISLCSVMYKIISKILSARLKKYLPVIV 262
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ QSAF++ R + DNI++AHE H L+ K +FM K +M+KAYDR+EW FLK L
Sbjct: 263 SPTQSAFVAERLVSDNIILAHEIVHNLRTNEKISKDFMVFKTDMSKAYDRVEWPFLKGIL 322
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
+ GF+ W++ +M CV+ S+ +NG+ + P RG+RQG PLSP+LFVL E + H
Sbjct: 323 LALGFNSTWINWMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEALIH 382
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
+L +A++ G+I G++ P + HLLFADD+L KA++ E +++ L+ Y SGQM
Sbjct: 383 ILNQAEKIGKISGIQFNGTGPSVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQM 442
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ EKS I F TK I + + GKYLGLP + S+ +IK ++
Sbjct: 443 INSEKSAITFGAKVNEETKQWIMNRSGIQTEGGTGKYLGLPECFQGSKQVLFGFIKEKLQ 502
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
++ GW K L+Q GK++L+K++ A Y M+ R + C + + + FWW S +
Sbjct: 503 SRLSGWYAKTLSQGGKDILLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFWWNSVQDK 562
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
+ IHW + K GG GFKD N ALLAKQA R++T D+L + LK Y+ NS
Sbjct: 563 KKIHWIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYMNS 622
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEA--- 478
FL + + R S+ WQSIL G+E L IIG+G VW D W+ D + PE+
Sbjct: 623 DFLSATKGTRPSYAWQSILYGRELLVSGLKKIIGNGENTYVWMDNWIFDDKPRRPESLQI 682
Query: 479 -GDAELTVNSLILPDVKQWNLPKLLSTFP-RQIAFKVFQIPLSLTQVPDKLCWPHTKDGE 536
D +L V+ LI P + WNL L FP ++I Q P++ Q D CW T G
Sbjct: 683 MVDIQLKVSQLIDPFSRNWNLNMLRDLFPWKEIQIICQQRPMASRQ--DSFCWFGTNHGL 740
Query: 537 YTVKTGY-----QVAYSLKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALP 591
YTVK+ Y QV + ++A QP + +P++ +IW + KIK+FLWK A+
Sbjct: 741 YTVKSEYDLCSRQVHKQMFKEAE-EQPSL--NPLFGKIWNLNSAPKIKVFLWKVLKGAVA 797
Query: 592 VKASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDW 651
V+ L R + + C +C + E++ H+L CP R VW + + + G +
Sbjct: 798 VEDRLRTRGVLIEDGCSMCPEKNETLNHILFQCPLARQVWALTPMQSPNHGFGDSIFTN- 856
Query: 652 LTQLVIRLQENSEDLSRGLTIVA-YYLWGIWKERNECHFQQRPPDPLGT-TIRIRASMLN 709
VI N+E LS L V+ + +W +WK RN+ F + +G+ ++ I L
Sbjct: 857 -VNHVIGNCHNTE-LSPHLRYVSPWIIWILWKNRNKRLF-----EGIGSVSLSIVGKALE 909
Query: 710 GMEDNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNV 769
++ + ++P W P LK N+ + K+ ++ VV + KG V
Sbjct: 910 DCKEWLKAHELICSKEPTKDLTWIPPLMNELKCNIGIAWSKKHQMAGVSWVVRNWKG-RV 968
Query: 770 LSHSK 774
L HS+
Sbjct: 969 LLHSR 973
>UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 459 bits (1180), Expect = e-127
Identities = 268/719 (37%), Positives = 386/719 (53%), Gaps = 36/719 (5%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF +G +P N T + L+PK P + LRPIS C+ LYK+I+KI+ +L+P + +
Sbjct: 437 FFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIV 496
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ QSAF+S R I DNIL+AHE H LK + +EFMA+K +M+KAYDR+EW +L++ L
Sbjct: 497 SDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMAVKSDMSKAYDRVEWSYLRSLL 556
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
S GF KWV+ IM CV+ ++ +N ++ QRG+RQG PLSP+LFVL E ++H
Sbjct: 557 LSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTH 616
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
LL KAQ EG ++G++ + P + HLLFADDSLF KA+RE++ L IL Y A+GQ
Sbjct: 617 LLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQT 676
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I++ KS I F K I + L + G YLGLP + S+ + L+++K R+
Sbjct: 677 INLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLK 736
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
+K+D W + L+Q GKEVL+K+V AM + MS + P C+ + +++A FWW S
Sbjct: 737 EKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHS 796
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R IHW+ W + K GGLGF+D + N ALLAKQAWR+ PD L R LK YF +
Sbjct: 797 RKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDAT 856
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAG-- 479
FL + + R S+GW+SIL G+E L + +GDG+ + VW D W+ D +AP
Sbjct: 857 DFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGDGASLFVWIDPWIDDNGFRAPWRKNL 916
Query: 480 --DAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEY 537
D L V +L+ P W+ L F + ++ I ++Q D W K G++
Sbjct: 917 IYDVTLKVKALLNPRTGFWDEEVLHDLFLPEDILRIKAIKPVISQA-DFFVWKLNKSGDF 975
Query: 538 TVKTGYQVAYSLKEQ----ARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVK 593
+VK+ Y +AY K Q QP L + Q+W +Q KIK+FLWK C E
Sbjct: 976 SVKSAYWLAYQTKSQNLRSEVSMQPSTLG--LKTQVWNLQTDPKIKIFLWKVCGEL---- 1029
Query: 594 ASLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKDWLT 653
ES H L CP +R +W S + ++ +
Sbjct: 1030 --------------------GESTNHTLFLCPLSRQIWALSDYPFPPDGFSNGSIYSNIN 1069
Query: 654 QLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNGME 712
L + ++N E I + LW IWK RN F+ T I+IR ++ E
Sbjct: 1070 HL-LENKDNKEWPINLRKIFPWILWRIWKNRNSFIFEGISYPATDTVIKIRDDVVEWFE 1127
>UniRef100_Q7XLZ5 OSJNBa0086O06.14 protein [Oryza sativa]
Length = 1205
Score = 450 bits (1157), Expect = e-124
Identities = 249/774 (32%), Positives = 400/774 (51%), Gaps = 39/774 (5%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF+ G +N+TV+ ++PK + P + RP+S CN +YKV+ K LVN+L+P + +
Sbjct: 462 FFETGEWKEGMNDTVIVMIPKTNAPVEMKDFRPVSLCNVIYKVVAKCLVNRLRPLLQEII 521
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
++ QSAF+ GR I DN L+A E FH + + +F A+KL+++KAYDR++W FL L
Sbjct: 522 SETQSAFVPGRMITDNALVAFECFHSIHKCTRESQDFCALKLDLSKAYDRVDWGFLDGAL 581
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
GF W IM CVT + ++NG + + P RG+R+G PL+PYLF+ A+ +S+
Sbjct: 582 QKLGFGNIWRKWIMSCVTSVRYSVRLNGNMLEPFYPTRGLREGDPLNPYLFLFIADGLSN 641
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
+L + ++E +IQ L++ P ++HLLFADDSL F KA +A ++ L+ Y +GQ+
Sbjct: 642 ILQRRRDERQIQPLKVCRSAPGVSHLLFADDSLLFFKAEVIQATRIKEALDLYERCTGQL 701
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ ++ ++FS + I ++L++ K LGLP G+ + IK R
Sbjct: 702 INPKECSLLFSALCPQERQDGIKAVLQVERTCFDDKCLGLPTPDGRMKAEQFQPIKERFE 761
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
++ W E+ L+ AGKE LIK+V QA+ YTM + + P+ FC+ + FWW E
Sbjct: 762 KRLTDWSERFLSLAGKEALIKSVAQALPTYTMGVFKMPERFCEEYEQLVRNFWWGHEKGE 821
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
+ +HW W ++ K GGLGF+D N ALLA+QAWR+ PD+L R LK Y+PN
Sbjct: 822 KKVHWIAWEKLTSPKLLGGLGFRDIRCFNQALLARQAWRLIESPDSLCARVLKAKYYPNG 881
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVS---DQVLQAPEA 478
+ + + +S W+ I+ G E LK+ +W IGDGS+ K+W++ WV+ + + +
Sbjct: 882 TITDTAFPSVSSPTWKGIVHGLELLKKGLIWRIGDGSKTKIWRNHWVAHGENLKILEKKT 941
Query: 479 GDAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYT 538
+ + V LI+ D K WN P + + A ++ +I + + D W + K G ++
Sbjct: 942 WNRVIYVRELIVTDTKTWNEPLIRHIIREEDADEILKIRIPQREEEDFPAWHYEKTGIFS 1001
Query: 539 VKTGYQVAYSL----KEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKA 594
V++ Y++A++L EQA IW+ +W V K+++F WK + L
Sbjct: 1002 VRSVYRLAWNLARKTSEQASSSSGGADGRKIWDNVWKANVQPKVRVFAWKLAQDRLATWE 1061
Query: 595 SLFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQG--SLTVKDWL 652
+ +R I C IC Q EE+ +H + C +A+ S + H + S+T DWL
Sbjct: 1062 NKKKRKIEMFGTCPICGQKEETGFHATVECTLAKALRASLREHWTLPDESLFSMTGPDWL 1121
Query: 653 TQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNECHFQQRPPDPLGTTIRIRASMLNGME 712
L+ RL + Q P I+ + M
Sbjct: 1122 LVLLDRLSSEKK-------------------------AQLPARKHMEDIKGKGPMF---- 1152
Query: 713 DNTPSPIPTQGRKPNYQARWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKG 766
P + + +W PP + KLNVDA Y ET + S +++ D +G
Sbjct: 1153 -QDPCQKEQTCQLNAEKEKWSCPPDGSAKLNVDAAYRTETGEASAGIIIRDCRG 1205
>UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 444 bits (1142), Expect = e-123
Identities = 246/612 (40%), Positives = 354/612 (57%), Gaps = 11/612 (1%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF +G +P N T + L+PK P + LRPIS C+ LYK+I+KI+ +L+P + +
Sbjct: 440 FFADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIV 499
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+ QSAF+S R I DNIL+AHE H LK + +EFMA+K +M+KAYDR+EW +L++ L
Sbjct: 500 SDTQSAFVSERLITDNILVAHELVHSLKVHPRISSEFMAVKSDMSKAYDRVEWSYLRSLL 559
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
S GF KWV+ IM CV+ ++ +N ++ QRG+RQG PLSP+LFVL E ++H
Sbjct: 560 LSLGFHLKWVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTH 619
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
LL KAQ EG ++G++ + P + HLLFADDSLF KA+RE++ L IL Y A+GQ
Sbjct: 620 LLNKAQWEGALEGIQFSENGPMVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQT 679
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I++ KS I F K I + L + G YLGLP + S+ + L+++K R+
Sbjct: 680 INLNKSSITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLK 739
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
+K+D W + L+Q GKEVL+K+V AM + MS + P C+ + +++A FWW S
Sbjct: 740 EKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCENLESAMASFWWDSCDHS 799
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R IHW+ W + K GGLGF+D + N ALLAKQAWR+ PD L R LK YF +
Sbjct: 800 RKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDAT 859
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAG-- 479
FL + + R S+GW+SIL G+E L + +GDG+ + VW D W+ D +AP
Sbjct: 860 DFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGDGASLFVWIDPWIDDNGFRAPWRKNL 919
Query: 480 --DAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEY 537
D L V +L+ P W+ L F + ++ I ++Q D W K G++
Sbjct: 920 IYDVTLKVKALLNPRTGFWDEEVLHDLFLPEDILRIKAIKPVISQA-DFFVWKLNKSGDF 978
Query: 538 TVKTGYQVAYSLKEQ----ARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVK 593
+VK+ Y +AY K Q QP L + Q+W +Q KIK+FLWK + LPV
Sbjct: 979 SVKSAYWLAYQTKSQNLRSEVSMQPSTLG--LKTQVWNLQTDPKIKIFLWKVLSGILPVA 1036
Query: 594 ASLFRRHIAPDP 605
+L R ++ P
Sbjct: 1037 ENLNGRGMSLIP 1048
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 436 bits (1120), Expect = e-120
Identities = 221/638 (34%), Positives = 357/638 (55%), Gaps = 7/638 (1%)
Query: 1 SFFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRL 60
+FFQ G +P +N+T + L+PK D+P ++ RPIS CN +YKV++K LVN+L+P ++ L
Sbjct: 1113 NFFQSGLMPKGVNDTAIVLIPKKDQPIDLKDYRPISLCNVVYKVVSKCLVNRLRPILDDL 1172
Query: 61 ATQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKAC 120
++ QSAFI GR I DN L+A E FH ++ K + A KL+++KAYDR++W FL+
Sbjct: 1173 VSKEQSAFIQGRMITDNALLAFECFHSIQKNKKANSAACAYKLDLSKAYDRVDWRFLELA 1232
Query: 121 LTSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMS 180
L GF+ +WV IM CVT + K NG + + P RG+RQG PLSP+LF+ A+ +S
Sbjct: 1233 LNKLGFAHRWVSWIMLCVTTVRYSVKFNGTLLRSFAPTRGLRQGEPLSPFLFLFVADGLS 1292
Query: 181 HLLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQ 240
LL + + + L++ + P I++LLFADD+L F KA ++EA + +L +Y +GQ
Sbjct: 1293 LLLKEKVAQNSLTPLKICRQAPGISYLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQ 1352
Query: 241 MISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRV 300
+I+ K I+F + SS I + L++ + +YLG P G+ ++ ++
Sbjct: 1353 LINPAKCSILFGEASPSSVSEDIRNTLQVERDNFEDRYLGFPTPEGRMHKGRFQSLQAKI 1412
Query: 301 LDKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGK 360
++ W E L+ GKE+LIKAV+QA+ Y M + +FP + + FWW +
Sbjct: 1413 AKRVIQWGENFLSSGGKEILIKAVIQAIPVYVMGLFKFPDSVYDELTKMTRNFWWGADNG 1472
Query: 361 ERGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPN 420
R HW+ W ++KAK GGLGF+D+ N ALL +QAWR+ P++L + LK YFP+
Sbjct: 1473 RRRTHWRAWDSLTKAKINGGLGFRDYKLFNQALLTRQAWRLIEFPNSLCAQVLKAKYFPH 1532
Query: 421 SSFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAGD 480
S + +A S W I G + LK+ +W IG+G+ V++W+D W+ + + P +
Sbjct: 1533 GSLTDTTFSANASPTWHGIEYGLDLLKKGIIWRIGNGNSVRIWRDPWIPRDLSRRPVSSK 1592
Query: 481 AELT---VNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEY 537
A V+ LI D W+ K+ F + A + +I +S D + W K G +
Sbjct: 1593 ANCRLKWVSDLIAED-GTWDSAKINQYFLKIDADIIQKICISARLEEDFIAWHPDKTGRF 1651
Query: 538 TVKTGYQVAYSLKE--QARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKAS 595
+V++ Y++A L + LN W IW VP K+++F W+ + +L +
Sbjct: 1652 SVRSAYKLALQLADMNNCSSSSSSRLNKS-WELIWKCNVPQKVRIFAWRVASNSLATMEN 1710
Query: 596 LFRRHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLS 633
+R++ VC IC++++E H L C ++W++
Sbjct: 1711 KKKRNLERFDVCGICDREKEDAGHALCRCVHANSLWVN 1748
Score = 51.2 bits (121), Expect = 1e-04
Identities = 39/136 (28%), Positives = 72/136 (52%), Gaps = 6/136 (4%)
Query: 731 RWDVPPPWTLKLNVDATYFKETCKGSIAVVVFDEKGTNVLSHSKTIWAAS-PLAAEALAV 789
RW+ P +KLNVD ++ + KG I +++ + G + S +++ + S PL AE A
Sbjct: 1776 RWERPRNGWMKLNVDGSFDINSEKGGIGMILRNCLGNVIFSSCRSLDSCSGPLEAELHAC 1835
Query: 790 REALLLAMNLELPKIYLVSDCLNLVSLIRERRTDWKVEAVLRDIQD*RNLF---RVCGFF 846
E L LA++ L I + +DC +++ L+ D V A + Q+ ++L R
Sbjct: 1836 VEGLHLALHWTLLPIQVETDCSSVIQLLNHPDKDRSVLANI--AQEAKSLMAGDRQIAIS 1893
Query: 847 WVPRTRNLCAHYVAKQ 862
V R++N+ +H++A +
Sbjct: 1894 KVQRSQNVISHFLANK 1909
>UniRef100_Q7XGD3 Putative retroelement [Oryza sativa]
Length = 1694
Score = 430 bits (1105), Expect = e-118
Identities = 216/603 (35%), Positives = 339/603 (55%), Gaps = 4/603 (0%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF G +P +NET + L+PK ++P+ + RPIS CN +YK+++K LVN+L+P ++ L
Sbjct: 1088 FFSLGTMPSGVNETAIVLIPKTEQPQELKDFRPISLCNVVYKIVSKCLVNRLRPILDDLV 1147
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
+Q QSAF+ GR I DN LIA E FH ++ + + A KL+++KAYDR++W+FL+ +
Sbjct: 1148 SQNQSAFVPGRLITDNALIAFEYFHHIQRNKNPENAYSAYKLDLSKAYDRVDWEFLEQAM 1207
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
GF+ WV IM C+T + K+NG + P RG+RQG PLSP+LF+ A+ +S
Sbjct: 1208 VKLGFAHCWVKWIMACITLVRYAVKLNGTLLNTFAPSRGLRQGDPLSPFLFLFVADGLSF 1267
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
LL + ++G + + + P + HLLFADD+ F KA ++A + +L Y +GQ+
Sbjct: 1268 LLEEKVDQGAMNPIHICRHAPAVLHLLFADDTPLFFKAANDQAVVMKEVLETYASCTGQL 1327
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ K I+F +++ ++ ++QI L++S KYLG P G+ ++ R+
Sbjct: 1328 INPSKCSIMFGQSSPTAVQNQIKQTLQISN-SFEDKYLGFPTPEGRMTKGKFQSLQERLW 1386
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
+I W EK L+ GKE+LIKAVLQAM Y M + + P++ C+ + + FWW +
Sbjct: 1387 KRIMLWGEKFLSTGGKEILIKAVLQAMPVYVMGLFKLPESVCEELTKLVRNFWWGAEKGM 1446
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R HWK W + K KGG+GF+DF N ALLA+QAWR+ +PD+L R LK +Y+PN
Sbjct: 1447 RRTHWKSWDCLIAQKLKGGMGFRDFRLFNQALLARQAWRLLDRPDSLCARILKAIYYPNG 1506
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAPEAG-- 479
S + + S GW++I G E LK+ +W IG+G V++W+D W+ + P +
Sbjct: 1507 SLIDTSFGGNASPGWRAIEYGLELLKKGIIWRIGNGKSVRLWRDPWIPRSYSRRPISAKR 1566
Query: 480 DAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTV 539
+ L S ++ W+ K+ F A + I LS D + W K G ++V
Sbjct: 1567 NCRLRWVSDLIDQSGSWDTDKISQHFLPMDAEAILNIRLSSRLEDDFIAWHPDKLGRFSV 1626
Query: 540 KTGYQVAYSLKEQARGPQPH-ILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFR 598
++ Y +A +L G N WN +W P K+K+F W+A +LP + +
Sbjct: 1627 RSAYHLAVALAHVDEGSSSSGNGNSRAWNALWKCNAPQKVKIFAWRAITNSLPTLENKKK 1686
Query: 599 RHI 601
R +
Sbjct: 1687 RKL 1689
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 423 bits (1087), Expect = e-116
Identities = 226/694 (32%), Positives = 371/694 (52%), Gaps = 38/694 (5%)
Query: 2 FFQEGYLPPFINETVVALVPKVDKPENIGQLRPISCCNFLYKVITKILVNKLKPHMNRLA 61
FF G + +N+T + L+PK+++P ++ RPIS CN +YKV++K LVN+L+P ++ L
Sbjct: 862 FFHSGIMLEGVNDTAIVLIPKIEQPRDLRDFRPISLCNVIYKVLSKCLVNRLRPFLDELI 921
Query: 62 TQAQSAFISGRQIQDNILIAHEAFHFLKNTCKGKTEFMAIKLEMNKAYDRLEWDFLKACL 121
++ QSAF+ R I DN L+A E FHF++ ++ A KL+++KAYDR++W FL+ +
Sbjct: 922 SKNQSAFVPRRMITDNALLAFECFHFIQRNKNPRSAACAYKLDLSKAYDRVDWRFLEQSM 981
Query: 122 TSFGFSEKWVDRIMQCVTGASFRFKVNGEVSKRVVPQRGIRQGYPLSPYLFVLAAETMSH 181
+GFS WV +M C++ V +G R P+LF+ A+ +S
Sbjct: 982 YKWGFSHCWVSWVMTCIS---------------TVCDKGTR----CDPFLFLFVADGLSL 1022
Query: 182 LLLKAQEEGRIQGLRMAPECPPITHLLFADDSLFFAKANREEAYQLISILNDYTLASGQM 241
LL + E+G I +R+ + P I+HLLFADD+L F KA+ ++ + + Y ++GQ+
Sbjct: 1023 LLEEKVEQGAISPIRVCHQAPGISHLLFADDTLLFFKADLSQSQAIKEVFGSYATSTGQL 1082
Query: 242 ISIEKSGIIFSRNTLSSTKHQISSILRMSAWDSPGKYLGLPAEWGKSRNNTLNWIKLRVL 301
I+ K I+F + +++ I++ L++++ + KYLGLP G+ ++ R+
Sbjct: 1083 INPTKCSILFGNSLPIASRDAITNCLQIASTEFEDKYLGLPTPGGRMHKGRFQSLRERIW 1142
Query: 302 DKIDGWKEKLLNQAGKEVLIKAVLQAMSAYTMSIIRFPQNFCQAICASIARFWWRSPGKE 361
KI W E L+ GKEVLIKAV+QA+ Y M I + P++ C+ + + FWW + +
Sbjct: 1143 KKILQWGENYLSSGGKEVLIKAVIQAIPVYVMGIFKLPESVCEDLTSLTRNFWWGAEKGK 1202
Query: 362 RGIHWKDWHFISKAKCKGGLGFKDFSTMNLALLAKQAWRIYTQPDALWVRTLKGLYFPNS 421
R HWK W ++K+K GGLGFKD N ALLA+QAWR+ PD+L R LK Y+PN
Sbjct: 1203 RKTHWKAWKSLTKSKSLGGLGFKDIRLFNQALLARQAWRLIDNPDSLCARVLKAKYYPNG 1262
Query: 422 SFLQSGRTARNSWGWQSILQGKEFLKEAGLWIIGDGSEVKVWKDKWVSDQVLQAP--EAG 479
S + + S GWQ+I G E +K+ +W IG+G V+VW+D W+ + + P
Sbjct: 1263 SIVDTSFGGNASPGWQAIEHGLELVKKGIIWRIGNGRSVRVWQDPWLPRDLSRRPITPKN 1322
Query: 480 DAELTVNSLILPDVKQWNLPKLLSTFPRQIAFKVFQIPLSLTQVPDKLCWPHTKDGEYTV 539
+ + + ++ D W+ K+ F + ++ S D + W K G ++V
Sbjct: 1323 NCRIKWVADLMLDNGMWDANKINQIFLPVDVEIILKLRTSSRDEEDFIAWHPDKLGNFSV 1382
Query: 540 KTGYQVAYS-LKEQARGPQPHILNDPIWNQIWGIQVP*KIKMFLWKACNEALPVKASLFR 598
+T Y++A + KE+A + W +W VP K+K+F W+A + LP + +
Sbjct: 1383 RTAYRLAENWAKEEASSSSSDVNIRKAWELLWKCNVPSKVKIFTWRATSNCLPTWDNKKK 1442
Query: 599 RHIAPDPVCFICNQDEESVYHLLLGCPWTRAVWLSSQLHVQHLHQGSLTVKD------WL 652
R++ C IC ++E H L CP + +WL+ ++ + SL + D WL
Sbjct: 1443 RNLEISDTCVICGMEKEDTMHALCRCPQAKHLWLA----MKESNDLSLRMDDHLLGPSWL 1498
Query: 653 TQLVIRLQENSEDLSRGLTIVAYYLWGIWKERNE 686
+ L ++ + + LW IW NE
Sbjct: 1499 FNRLALLPDHEQPM------FLMVLWRIWFVHNE 1526
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.323 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,464,064,292
Number of Sequences: 2790947
Number of extensions: 60298633
Number of successful extensions: 121464
Number of sequences better than 10.0: 1045
Number of HSP's better than 10.0 without gapping: 634
Number of HSP's successfully gapped in prelim test: 411
Number of HSP's that attempted gapping in prelim test: 119040
Number of HSP's gapped (non-prelim): 1540
length of query: 863
length of database: 848,049,833
effective HSP length: 136
effective length of query: 727
effective length of database: 468,481,041
effective search space: 340585716807
effective search space used: 340585716807
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0111a.1


