

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.15
(184 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
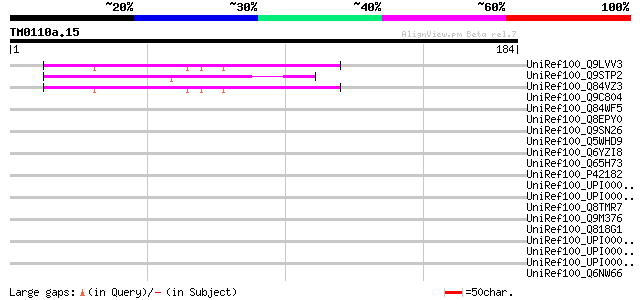
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LVV3 Arabidopsis thaliana genomic DNA, chromosome 5,... 58 1e-07
UniRef100_Q9STP2 Hypothetical protein T27E11.80 [Arabidopsis tha... 55 9e-07
UniRef100_Q84VZ3 At5g52990 [Arabidopsis thaliana] 55 9e-07
UniRef100_Q9C804 Hypothetical protein F10C21.23 [Arabidopsis tha... 44 0.003
UniRef100_Q84WF5 Hypothetical protein At1g33480 [Arabidopsis tha... 44 0.003
UniRef100_Q8EPY0 GTP-binding protein [Oceanobacillus iheyensis] 43 0.003
UniRef100_Q9SN26 Hypothetical protein F28M11.90 [Arabidopsis tha... 40 0.038
UniRef100_Q5WHD9 GTP-binding protein Era, Era/TrmE family [Bacil... 37 0.19
UniRef100_Q6YZI8 Putative vesicle-associated membrane protein 72... 37 0.19
UniRef100_Q65H73 Era [Bacillus licheniformis] 37 0.32
UniRef100_P42182 GTP-binding protein era homolog [Bacillus subti... 37 0.32
UniRef100_UPI000026BCAC UPI000026BCAC UniRef100 entry 35 0.93
UniRef100_UPI000034019D UPI000034019D UniRef100 entry 34 1.6
UniRef100_Q8TMR7 Predicted protein [Methanosarcina acetivorans] 34 1.6
UniRef100_Q9M376 Vesicle-associated membrane protein 727 [Arabid... 34 1.6
UniRef100_Q818G1 GTP-binding protein [Bacillus cereus] 34 2.1
UniRef100_UPI00003AA223 UPI00003AA223 UniRef100 entry 33 2.7
UniRef100_UPI00003CBD2B UPI00003CBD2B UniRef100 entry 33 2.7
UniRef100_UPI000026AA9E UPI000026AA9E UniRef100 entry 33 2.7
UniRef100_Q6NW66 Hypothetical protein zgc:85938 [Brachydanio rerio] 33 2.7
>UniRef100_Q9LVV3 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MNB8
[Arabidopsis thaliana]
Length = 272
Score = 57.8 bits (138), Expect = 1e-07
Identities = 39/113 (34%), Positives = 62/113 (54%), Gaps = 5/113 (4%)
Query: 13 HSLFSHTVNNRTYTFLI-DPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETL-DSVND-S 69
HS+FSHTV+ +TYTF I D F+YF+I D K ++ LNR+RS++++ + D +D
Sbjct: 46 HSMFSHTVHKKTYTFAIDDDSFVYFSISDESMEKPESFWVLNRLRSAIEDLIKDGGSDVE 105
Query: 70 TAPPPLS--LQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSA 120
T P+S LQ D + EI+ + D +VG + + +DS+
Sbjct: 106 TLINPVSHCLQLKLDPVFAEIVGVVDLELLDMDLVGSPRSVARESRNPSIDSS 158
>UniRef100_Q9STP2 Hypothetical protein T27E11.80 [Arabidopsis thaliana]
Length = 260
Score = 55.1 bits (131), Expect = 9e-07
Identities = 35/102 (34%), Positives = 52/102 (50%), Gaps = 14/102 (13%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRS---SLKETLDSVNDS 69
HS+ SHTV+ RTY +ID F YFAI D KS+++ NR++S SL E + +
Sbjct: 46 HSMISHTVHKRTYALIIDGLFSYFAILDEVVAKSESVWLFNRLKSATESLMEDGSTADSL 105
Query: 70 TAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLK 111
P LQ+ D + EI A +G +H++ L+
Sbjct: 106 DNPTQHCLQSKLDPVFAEI-----------AAIGGNHNKDLE 136
>UniRef100_Q84VZ3 At5g52990 [Arabidopsis thaliana]
Length = 272
Score = 55.1 bits (131), Expect = 9e-07
Identities = 38/113 (33%), Positives = 61/113 (53%), Gaps = 5/113 (4%)
Query: 13 HSLFSHTVNNRTYTFLI-DPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETL-DSVND-S 69
HS+FSHTV+ +TYTF I D F+YF+I K ++ LNR+RS++++ + D +D
Sbjct: 46 HSMFSHTVHKKTYTFAIDDDSFVYFSISGESMEKPESFWVLNRLRSAIEDLIKDGGSDVE 105
Query: 70 TAPPPLS--LQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSA 120
T P+S LQ D + EI+ + D +VG + + +DS+
Sbjct: 106 TLINPVSHCLQLKLDPVFAEIVGVVDLELLDMDLVGSPRSVARESRNPSIDSS 158
>UniRef100_Q9C804 Hypothetical protein F10C21.23 [Arabidopsis thaliana]
Length = 508
Score = 43.5 bits (101), Expect = 0.003
Identities = 23/77 (29%), Positives = 37/77 (47%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAP 72
H + T +T+ FL+ F+YFAI D KS L FL ++R LKE + +
Sbjct: 300 HKWYFETRGKKTFGFLMKDDFVYFAIVDDVFKKSSVLDFLEKLRDELKEANKKNSRGSFS 359
Query: 73 PPLSLQTPFDLILQEIL 89
+S D I++ ++
Sbjct: 360 GSISFSNVQDQIVRRLI 376
>UniRef100_Q84WF5 Hypothetical protein At1g33480 [Arabidopsis thaliana]
Length = 255
Score = 43.5 bits (101), Expect = 0.003
Identities = 23/77 (29%), Positives = 37/77 (47%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDSVNDSTAP 72
H + T +T+ FL+ F+YFAI D KS L FL ++R LKE + +
Sbjct: 47 HKWYFETRGKKTFGFLMKDDFVYFAIVDDVFKKSSVLDFLEKLRDELKEANKKNSRGSFS 106
Query: 73 PPLSLQTPFDLILQEIL 89
+S D I++ ++
Sbjct: 107 GSISFSNVQDQIVRRLI 123
>UniRef100_Q8EPY0 GTP-binding protein [Oceanobacillus iheyensis]
Length = 300
Score = 43.1 bits (100), Expect = 0.003
Identities = 33/100 (33%), Positives = 54/100 (54%), Gaps = 4/100 (4%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKG-LAAAAATEL 140
+LI +++LHL V I + E + +K+++ AT++ R+S KG L T L
Sbjct: 194 ELIREKVLHLTKEEVPHSIAVVIENIEKNENEKLLIQ-ATIVTERSSQKGILIGKQGTML 252
Query: 141 VELEKDAVE--EALVGSAIQIEGFRRVWLDLRSRLSLLHE 178
+ K A EAL+GS + +E + +V D R+R + LHE
Sbjct: 253 KNIGKQARRDIEALLGSKVYLELWIKVKKDWRNRQNQLHE 292
>UniRef100_Q9SN26 Hypothetical protein F28M11.90 [Arabidopsis thaliana]
Length = 254
Score = 39.7 bits (91), Expect = 0.038
Identities = 18/49 (36%), Positives = 26/49 (52%)
Query: 13 HSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKE 61
H+ + T+ R + FLI F+YFAI D +S L FL +R K+
Sbjct: 45 HTWYFETIGKRRFGFLIGDGFVYFAIVDEVLKRSSVLKFLEHLRDEFKK 93
>UniRef100_Q5WHD9 GTP-binding protein Era, Era/TrmE family [Bacillus clausii]
Length = 303
Score = 37.4 bits (85), Expect = 0.19
Identities = 31/101 (30%), Positives = 52/101 (50%), Gaps = 6/101 (5%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLA----AAAA 137
+LI +++LHL P +V + + V ATVI R+S KG+ A
Sbjct: 197 ELIREKVLHLTK-EEIPHSVAVVIEQIKKRDNGKVYVGATVIVERSSQKGIIIGKHGAML 255
Query: 138 TELVELEKDAVEEALVGSAIQIEGFRRVWLDLRSRLSLLHE 178
E+ +L + +E AL+GS++ +E + +V D R+R S L +
Sbjct: 256 KEVGQLARSDIE-ALLGSSVYLELWVKVQKDWRNRPSQLKD 295
>UniRef100_Q6YZI8 Putative vesicle-associated membrane protein 725 [Oryza sativa]
Length = 241
Score = 37.4 bits (85), Expect = 0.19
Identities = 49/184 (26%), Positives = 74/184 (39%), Gaps = 26/184 (14%)
Query: 4 PKKGSKGSTHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETL 63
P SK ST+S HT N FL+D F++ + D +S FL+R++ +
Sbjct: 42 PPNTSK-STYSCDGHTFN-----FLVDRGFVFLVVADEAVGRSVPFVFLDRVKEDFMQRY 95
Query: 64 DSVNDSTAPPPLSLQTPFDLILQE----ILHLDDGNRSP----PAVVGISHDEGLKKKKM 115
S D PL+ D L E I + D P + I+H E + K
Sbjct: 96 GSSIDEEGQHPLADDADDDDFLLEDRFSIAYNLDREFGPRLKDHMLYCINHPEEISK--- 152
Query: 116 VVDSATVIANRTSAKGLAAAAATELVELEKDAVEEALVGSA----IQIEGFRRVWLDLRS 171
+ V A+ T KG+ ++ LE+ E LVG Q + F R +LR
Sbjct: 153 ---LSKVKAHLTEVKGIMMDNIEKI--LERGEKIELLVGKTETLQSQADSFHRHGRELRR 207
Query: 172 RLSL 175
++ L
Sbjct: 208 KMWL 211
>UniRef100_Q65H73 Era [Bacillus licheniformis]
Length = 301
Score = 36.6 bits (83), Expect = 0.32
Identities = 31/106 (29%), Positives = 54/106 (50%), Gaps = 4/106 (3%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELV 141
+LI +++LHL V I + + + V SAT++ R S KG+ L+
Sbjct: 195 ELIREKVLHLTREEIPHSIAVSIESIQERENGSVYV-SATIVVERDSQKGIVIGKRGSLL 253
Query: 142 -ELEKDAVE--EALVGSAIQIEGFRRVWLDLRSRLSLLHESRSEKD 184
E+ K A EAL+GS++ +E + +V D R+++S L + +D
Sbjct: 254 KEIGKRARTDIEALLGSSVFLELWVKVQKDWRNKMSQLRDFGFRED 299
>UniRef100_P42182 GTP-binding protein era homolog [Bacillus subtilis]
Length = 301
Score = 36.6 bits (83), Expect = 0.32
Identities = 31/106 (29%), Positives = 54/106 (50%), Gaps = 4/106 (3%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELV 141
+LI +++LHL V I +G + V +AT++ R S KG+ L+
Sbjct: 195 ELIREKVLHLTREEIPHSIAVAIESIKGQDNGSVHV-AATIVVERDSQKGIVIGKKGSLL 253
Query: 142 -ELEKDAVE--EALVGSAIQIEGFRRVWLDLRSRLSLLHESRSEKD 184
E+ K A EAL+GS + +E + +V D R+++S L + ++D
Sbjct: 254 KEVGKRARADIEALLGSRVYLELWVKVQKDWRNKMSQLRDFGFKED 299
>UniRef100_UPI000026BCAC UPI000026BCAC UniRef100 entry
Length = 142
Score = 35.0 bits (79), Expect = 0.93
Identities = 25/102 (24%), Positives = 46/102 (44%), Gaps = 7/102 (6%)
Query: 58 SLKETLDSVNDSTAPPPLSLQTPFDLILQEILHLDDGNRSPPAVVGISHDEGLKKKKMVV 117
++ + ++ ND T L+ P ++ + E +SPPA V L KK +
Sbjct: 34 NIMDFCNAFNDKTKEMEKGLKVPVEITVFEDRTFTFITKSPPAAV-------LLKKAAGI 86
Query: 118 DSATVIANRTSAKGLAAAAATELVELEKDAVEEALVGSAIQI 159
DS + NRT + + E+ E++ D + + +AI+I
Sbjct: 87 DSGSGEPNRTKVAKIPVSKVQEIAEMKMDDLNANDIDAAIKI 128
>UniRef100_UPI000034019D UPI000034019D UniRef100 entry
Length = 203
Score = 34.3 bits (77), Expect = 1.6
Identities = 18/42 (42%), Positives = 28/42 (65%), Gaps = 4/42 (9%)
Query: 108 EGLKKKKMVVDSATVIANRTSAKGLAAA----AATELVELEK 145
+ +KKKK+ ++ A VI+N+ +AKGLA A TE++E K
Sbjct: 17 KAIKKKKIPINPAVVISNKPNAKGLAIAKKMGIQTEVIESSK 58
>UniRef100_Q8TMR7 Predicted protein [Methanosarcina acetivorans]
Length = 678
Score = 34.3 bits (77), Expect = 1.6
Identities = 22/69 (31%), Positives = 35/69 (49%), Gaps = 6/69 (8%)
Query: 21 NNRTYTFLIDPPFIYFAI--FDHRHTKSQTLTFLNRIRSSLKETLDSVN----DSTAPPP 74
N+ Y ++ D PF I F +TKS T+T+ + SL T D+ N D+T P
Sbjct: 163 NDNGYAYIEDVPFSEVVIGGFVGTYTKSGTVTYPDSTIFSLGSTFDAENVNFIDATVTPT 222
Query: 75 LSLQTPFDL 83
+ +P+D+
Sbjct: 223 YTKDSPYDI 231
>UniRef100_Q9M376 Vesicle-associated membrane protein 727 [Arabidopsis thaliana]
Length = 240
Score = 34.3 bits (77), Expect = 1.6
Identities = 15/61 (24%), Positives = 31/61 (50%), Gaps = 2/61 (3%)
Query: 14 SLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSSLKETLDS--VNDSTA 71
S ++++ + T+ FL+D F++ + D +S FL R++ K+ ++ ND
Sbjct: 44 SKYTYSCDGHTFNFLVDNGFVFLVVADESTGRSVPFVFLERVKEDFKKRYEASIKNDERH 103
Query: 72 P 72
P
Sbjct: 104 P 104
>UniRef100_Q818G1 GTP-binding protein [Bacillus cereus]
Length = 301
Score = 33.9 bits (76), Expect = 2.1
Identities = 31/107 (28%), Positives = 56/107 (51%), Gaps = 6/107 (5%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKK-MVVDSATVIANRTSAKGLAAAAATEL 140
+LI +++LHL P V + D K++ V +AT++ R S KG+ ++
Sbjct: 195 ELIREKVLHLT--REEVPHSVAVVIDAIQKREGGAVYINATIVVERASQKGIIIGKQGKM 252
Query: 141 V-ELEKDAVE--EALVGSAIQIEGFRRVWLDLRSRLSLLHESRSEKD 184
+ E+ K A EAL+GS + +E + +V D R+++S L + +D
Sbjct: 253 LKEVGKRARFDIEALLGSKVFLEVWVKVQKDWRNKMSQLRDLGFRED 299
>UniRef100_UPI00003AA223 UPI00003AA223 UniRef100 entry
Length = 1981
Score = 33.5 bits (75), Expect = 2.7
Identities = 30/110 (27%), Positives = 51/110 (46%), Gaps = 12/110 (10%)
Query: 43 HTKSQTLTF---LNRIRSSLKETLDSVNDSTAPPPLSLQTPFDLILQE------ILHLDD 93
HT++ L+ L+R S + L V D APP + T + +L E LH+
Sbjct: 558 HTQTGELSTKHSLDRELQSSYQLLVIVQDGGAPPRSATGTIYVTVLDENDNAPVFLHITG 617
Query: 94 GNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVEL 143
G P V +S L ++++V + TV+A + G +AT+L+ +
Sbjct: 618 GQELPVQVGALSVVRALNREEVVQHNLTVVA---ADHGFPRRSATQLLSI 664
>UniRef100_UPI00003CBD2B UPI00003CBD2B UniRef100 entry
Length = 301
Score = 33.5 bits (75), Expect = 2.7
Identities = 31/107 (28%), Positives = 56/107 (51%), Gaps = 6/107 (5%)
Query: 82 DLILQEILHLDDGNRSPPAVVGISHDEGLKKKK-MVVDSATVIANRTSAKGLAAAAATEL 140
+LI +++LHL P V + D K++ V +AT++ R S KG+ ++
Sbjct: 195 ELIREKVLHLT--REEVPHSVAVVIDAIQKREGGAVYINATIVVERPSQKGIIIGKQGKM 252
Query: 141 V-ELEKDAVE--EALVGSAIQIEGFRRVWLDLRSRLSLLHESRSEKD 184
+ E+ K A EAL+GS + +E + +V D R+++S L + +D
Sbjct: 253 LKEVGKRARFDIEALLGSKVFLEVWVKVQKDWRNKMSQLRDLGFRED 299
>UniRef100_UPI000026AA9E UPI000026AA9E UniRef100 entry
Length = 241
Score = 33.5 bits (75), Expect = 2.7
Identities = 18/58 (31%), Positives = 33/58 (56%)
Query: 94 GNRSPPAVVGISHDEGLKKKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVEEA 151
G P ++V I D G+ K +++ SAT +A+ S +AA + + V+L + +E+A
Sbjct: 75 GGLVPGSLVLIGGDPGIGKSTLILQSATEMAHHRSVLYVAAEESAQQVKLRWNRIEDA 132
>UniRef100_Q6NW66 Hypothetical protein zgc:85938 [Brachydanio rerio]
Length = 632
Score = 33.5 bits (75), Expect = 2.7
Identities = 39/158 (24%), Positives = 64/158 (39%), Gaps = 16/158 (10%)
Query: 6 KGSKGSTHSLFSHTVNNRTYTFLIDPPFIYFAIFDHRHTKSQTLTFLNRIRSS-----LK 60
+GS ST V+NR +I PP + +H+ T+ +R S L+
Sbjct: 183 RGSISSTKGPLQ--VSNRNSGTVIPPPLVRGGAQVSKHSTHNTVVMPPLVRGSQQIATLR 240
Query: 61 ETLDSVNDSTAPPPLSLQTPFDL---------ILQEILHLDDGNRSPPAVVGISHDEGLK 111
+ + + PPPL L T + I+Q+ L + +V + GLK
Sbjct: 241 QQQQQQHSGSGPPPLLLGTRTSVPTVQVQGQRIIQQGLIRVASVGNANVLVSQASSTGLK 300
Query: 112 KKKMVVDSATVIANRTSAKGLAAAAATELVELEKDAVE 149
+ + S++ AN + A AAA +LEK +E
Sbjct: 301 TSQSGLGSSSSNANDSPASRQAAAKLALRKQLEKTLLE 338
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.132 0.368
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 290,346,321
Number of Sequences: 2790947
Number of extensions: 10823863
Number of successful extensions: 56167
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 56143
Number of HSP's gapped (non-prelim): 43
length of query: 184
length of database: 848,049,833
effective HSP length: 120
effective length of query: 64
effective length of database: 513,136,193
effective search space: 32840716352
effective search space used: 32840716352
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0110a.15


