

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0110a.10
(1408 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
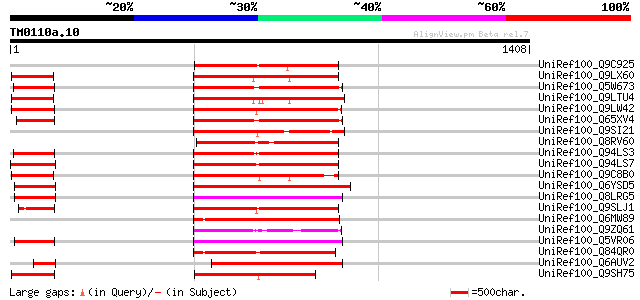
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 388 e-106
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 379 e-103
UniRef100_Q5W673 Putative helicase [Oryza sativa] 370 e-100
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 370 e-100
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 369 e-100
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 369 e-100
UniRef100_Q9SI21 Putative helicase [Arabidopsis thaliana] 365 5e-99
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 359 4e-97
UniRef100_Q94LS3 Putative helicase [Oryza sativa] 355 4e-96
UniRef100_Q94LS7 Putative helicase [Oryza sativa] 354 9e-96
UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thal... 351 1e-94
UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa] 351 1e-94
UniRef100_Q8LRG5 Putative helicase [Oryza sativa] 348 5e-94
UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana] 346 3e-93
UniRef100_Q6MW89 B1248C03.15 protein [Oryza sativa] 325 8e-87
UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana] 323 2e-86
UniRef100_Q5VR06 Helicase-like protein [Oryza sativa] 323 2e-86
UniRef100_Q84QR0 Helicase-like protein [Oryza sativa] 311 1e-82
UniRef100_Q6AUV2 Hypothetical protein OSJNBa0091B22.10 [Oryza sa... 305 5e-81
UniRef100_Q9SH75 Putative helicase [Arabidopsis thaliana] 300 2e-79
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 388 bits (996), Expect = e-106
Identities = 191/407 (46%), Positives = 260/407 (62%), Gaps = 17/407 (4%)
Query: 502 IEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPCG 561
IEFQKRGLPHAHILL++ KL++ D +I AE+PD K P L+ V + M+HGPCG
Sbjct: 57 IEFQKRGLPHAHILLFMHPTSKLSTAEDTDKIITAEIPDKKKKPGLYAVVKDCMIHGPCG 116
Query: 562 LSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPY 621
+ SPCM+NG+C K+FPK Y+D T D+DG+PVYRRR+TG+ + G DN +V+PY
Sbjct: 117 VGHPNSPCMENGKCKKYFPKSYSDTTKVDNDGFPVYRRRDTGIYVEKNGFQCDNRYVIPY 176
Query: 622 NPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEIKQY 681
N K+ ++YQAHIN+E CN+S SIKYLFKY++KG DRVT+++ K E DE+K Y
Sbjct: 177 NEKVSLRYQAHINVELCNQSGSIKYLFKYVHKGHDRVTVTVEPNDQDTAKK-EKDEVKDY 235
Query: 682 FDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIVKNS 741
FDCRY+S CEA+WRIF++ IH++ PV KL +H KQ V +K E E V+ R + +
Sbjct: 236 FDCRYVSACEAMWRIFKFPIHYRTTPVVKLFFHEEGKQPVYYKPGETTESVMDRLSSEAT 295
Query: 742 MFLAWMDANCK----------------YVHGRQLTYVEFPELFVYDPKSKSWHPRKQGES 785
FLAW N K +L + E P F ++ K K + R++G +
Sbjct: 296 QFLAWFQLNKKPPSRTIRANAKKLPKAAPDPTKLLFEEIPNHFTWNSKEKKFMIRERGFA 355
Query: 786 IGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREF 845
IGR++FVP + YYLR+LLN++RG T Y+DL++V VH +FR A ALGLL DD+E+
Sbjct: 356 IGRINFVPRTIEDAYYLRILLNIKRGVTSYKDLKTVKGVVHKSFRDAVFALGLLDDDKEY 415
Query: 846 IDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
I+GI D Y R LFV LL S+ P VW+ETW +L++ I
Sbjct: 416 INGIKDAKFWCSAKYVRRLFVIMLLSESLTKPEMVWDETWRILSEDI 462
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 379 bits (974), Expect = e-103
Identities = 189/412 (45%), Positives = 262/412 (62%), Gaps = 18/412 (4%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
M+T+EFQKRGLPHAHILL++ A+ KL + ID +I AE+PD P L+E + N M+HG
Sbjct: 778 MHTVEFQKRGLPHAHILLFMDAKSKLPTADDIDKIISAEIPDKDKEPELYEVIKNSMIHG 837
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM G+C+K +PKK+ D+T DGYP+YRRR T + G DNG+V
Sbjct: 838 PCGAANMNSPCMVEGKCSKQYPKKHQDITKVGKDGYPIYRRRMTEDYIEKGGFKCDNGYV 897
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRV------------TLSMSSGR 666
VPYN KL ++YQAHIN+E+CN+S SIKYLFKYINKG DRV T + +SG
Sbjct: 898 VPYNKKLSLRYQAHINVEWCNQSGSIKYLFKYINKGADRVVFIVEPVNQDKTTENATSGE 957
Query: 667 TTQDKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKEN 726
+ DEIK +FDCRY+S EAVWRI+++ + + VQ+L++H KQ V K +
Sbjct: 958 PPNSTEKKKDEIKDWFDCRYVSASEAVWRIYKFPLQDRSTAVQRLSFHDEGKQPVYAKPD 1017
Query: 727 EAIEDVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYDPKSKSWHPR 780
IEDVL R ++SMF+AW+ N G R+L Y + P F +D K+K W R
Sbjct: 1018 ADIEDVLERISNEDSMFMAWLTLNKNNDVGKNGKRARELLYSQIPAYFTWDGKNKQWVKR 1077
Query: 781 KQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLK 840
+G S+GR+++V YYLR+LLN+ +G Y+D+++ N V+ +F+ AC A G+L
Sbjct: 1078 IRGFSLGRINYVCRKMEVEYYLRVLLNIVKGPMSYDDIKTFNGVVYPSFKEACFARGILD 1137
Query: 841 DDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
DD+ +IDG+ + + G Y R F LL +S+ P HVW+ETWH+LA+ I
Sbjct: 1138 DDQVYIDGLHEASQFCFGDYLRNFFAMLLLSDSLARPEHVWSETWHLLAEDI 1189
Score = 132 bits (331), Expect = 1e-28
Identities = 62/116 (53%), Positives = 80/116 (68%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K Q++ SP F++CC KG + LP L D P L++NLL D SR+F +NIR YN +FA TS
Sbjct: 340 KKQQNKSPTFTLCCGKGNVKLPLLKDSPALINNLLTGDDALSRNFRENIRIYNMIFAMTS 399
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
+GG+VD + G+GP F + G NYH IGSL P GD K++QLYI DT+NEV NR
Sbjct: 400 LGGRVDNSMPKGKGPNMFRLQGGNYHLIGSLKPNPGDYAKYSQLYIVDTENEVDNR 455
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 370 bits (951), Expect = e-100
Identities = 180/404 (44%), Positives = 259/404 (63%), Gaps = 10/404 (2%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YTIEFQKRGLPHAHIL++L + K + ID +I AE+PD + FE V N+M+HG
Sbjct: 689 IYTIEFQKRGLPHAHILIFLDKKDKCPDASEIDRIISAEIPDKEEDREGFEAVENFMMHG 748
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG +K SPCM +C + FPKK+ TT D DG+P YRRR+ G + V LDN +V
Sbjct: 749 PCGEAKSNSPCMIENKCIRNFPKKFHSETTVDEDGFPTYRRRDNGRYIEKGNVKLDNRYV 808
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN LL+KYQAHIN+E CN+S SIKYLFKY++KG D+ T + S + DEI
Sbjct: 809 VPYNRDLLVKYQAHINVERCNRSKSIKYLFKYMHKGDDQATALIES---------DHDEI 859
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
K+Y +C Y+S +A WRIFQ+++H+++P V++L +HL +Q V+F ++ + ++R+ +
Sbjct: 860 KKYLECTYISGHDACWRIFQFEMHYRYPSVERLPFHLENEQQVIFPDSADLRKIVRKERI 919
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
+ F WM+ N R LTY EFP +V+ K K W+ RK+G+ IGR+ + P +G+
Sbjct: 920 GVTKFTQWMETNKINDEARDLTYAEFPSKWVWKNKLKQWNKRKKGKMIGRIYYAHPASGD 979
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGG 858
YYLRMLLN +G +E++R+V+ VH +F+ ACEALG L DDRE+++ I + + A G
Sbjct: 980 KYYLRMLLNTVKGPRTFEEIRTVDGVVHPSFKSACEALGFLDDDREWVECIREASNYASG 1039
Query: 859 AYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNN 902
R LF + L + +P +W W L++ I QRK+ N
Sbjct: 1040 NQLRHLFTTILCHCEVTDPKRIWESCWEDLSEDIEYK-QRKNLN 1082
Score = 141 bits (355), Expect = 2e-31
Identities = 67/111 (60%), Positives = 87/111 (78%), Gaps = 1/111 (0%)
Query: 11 PDFSICCMKGKIMLPYLADCPELLSNLL-RNTDTRSRHFLDNIRSYNSMFAFTSIGGKVD 69
P FS+CC +GK+ LP L P LSNL+ + RSR+++DNIR YNSMFAFTS+GGKVD
Sbjct: 265 PSFSLCCKQGKVDLPTLKKPPTYLSNLMCKEKGKRSRNYMDNIRVYNSMFAFTSMGGKVD 324
Query: 70 CGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRM 120
+N+G GP F ++GQNYHRI +L+P EGD P++AQLYIYDT+NEV+NR+
Sbjct: 325 REINNGSGPYVFRMNGQNYHRISTLLPEEGDKPRWAQLYIYDTENEVKNRI 375
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 370 bits (949), Expect = e-100
Identities = 189/436 (43%), Positives = 269/436 (61%), Gaps = 27/436 (6%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHILL+++A KL + ID +I AE+P+ P L+E + N M+HG
Sbjct: 446 MYTVEFQKRGLPHAHILLFMAANSKLPTADDIDKIISAEIPNKDKEPELYEVIKNSMMHG 505
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM +G+C+K +PKK+ ++T +DGYP+YRRR T + GV DN +V
Sbjct: 506 PCGSANTSSPCMVDGQCSKLYPKKHQEITKVGADGYPIYRRRLTDDYIEKGGVKCDNRYV 565
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTL------SMSSGRTTQDKP 672
VPYN KL ++YQAHIN+E+CN++ SIKYLFKYINKG DRV +S TT
Sbjct: 566 VPYNKKLSLRYQAHINVEWCNQNGSIKYLFKYINKGPDRVVFIVEPIKEATSSDTTAPVV 625
Query: 673 VED------DEIKQYFDC---------RYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPR 717
D DEIK +FDC RY+S EA+WRIF++ I H+ PVQKL++H
Sbjct: 626 ESDTTEKKKDEIKDWFDCSSYISFSPARYVSASEAIWRIFKFPIQHRSTPVQKLSFHDKG 685
Query: 718 KQVVLFKENEAIEDVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYD 771
KQ F + DVL R ++S FLAW+ N K G R Y E P F +D
Sbjct: 686 KQPAYFDAKAKMADVLERVSNEDSQFLAWLTLNRKNAVGKNGKRARDCLYAEIPAYFTWD 745
Query: 772 PKSKSWHPRKQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRG 831
++K + R +G S+GR+++V + YYLR+LLN+ RG Y+D+++VN V+ +++
Sbjct: 746 GENKQFKKRTRGFSLGRINYVSRKMEDEYYLRVLLNIVRGPQSYDDIKTVNGVVYPSYKL 805
Query: 832 ACEALGLLKDDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADG 891
AC A G+L DD+ +I+G+++ + G Y R F LL +S+ P HVW+ETWH+L++
Sbjct: 806 ACFARGILDDDQVYINGLIEASQFCFGDYLRNFFSMMLLSDSLARPEHVWSETWHLLSED 865
Query: 892 IACDLQRKHNNRGMCL 907
I + + N+ + L
Sbjct: 866 ILIKKRDEFKNQELTL 881
Score = 133 bits (334), Expect = 4e-29
Identities = 61/116 (52%), Positives = 80/116 (68%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K S +P FS+CC +G + LP+L + PEL+ LL+ D SRH+ IR YN +FA TS
Sbjct: 9 KKSNSENPKFSLCCGQGSVKLPFLKESPELIKKLLKGNDALSRHYRQFIRIYNMIFAMTS 68
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
+GGKVD + GRGP F + G NYH+IGSL P +GD K++QLYI DT+NEV+NR
Sbjct: 69 LGGKVDKSMPKGRGPAMFRLQGGNYHQIGSLKPKDGDYAKYSQLYIVDTENEVENR 124
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 369 bits (948), Expect = e-100
Identities = 181/412 (43%), Positives = 258/412 (61%), Gaps = 16/412 (3%)
Query: 500 YTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGP 559
YTIEFQKRGLPHAHI++W+ YK +D +I AE+PD + P L++ VS M+HGP
Sbjct: 769 YTIEFQKRGLPHAHIIVWMDPRYKFPIADHVDKIIFAEIPDKENDPKLYQAVSECMIHGP 828
Query: 560 CGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVV 619
C L SPCM+NG+C+K++PK + + T D++GYP+YRRR+TG + DN +VV
Sbjct: 829 CRLVNPNSPCMENGKCSKYYPKNHVENTFLDNEGYPIYRRRDTGRFIKKNKYPCDNRYVV 888
Query: 620 PYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGR-----------TT 668
PYN LL KY+AHIN+E+CN+S S+KYLFKY+NKG DRVT+S+ R T
Sbjct: 889 PYNDFLLRKYRAHINVEWCNQSVSVKYLFKYVNKGPDRVTVSLEPHRKEVVSEENNVGET 948
Query: 669 QDKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEA 728
+ P E ++++ YFDCRY+S CEA+WRI Y IH++ V KLT+H KQ + KE E
Sbjct: 949 NNDPQEQNQVEDYFDCRYVSACEAMWRIKGYPIHYRQTLVTKLTFHEKGKQPIYVKEGET 1008
Query: 729 IEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQ-GESIG 787
E VL R + F AW + N + +L Y + P + ++ K K++ RK G +G
Sbjct: 1009 AESVLYRVNDDETQFTAWFELNKRDPEAAKLLYEQIPNFYTWNGKDKNFRRRKMPGFVVG 1068
Query: 788 RMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFID 847
R++ VPP + Y+LR+L+N RG ++D++++ VH T+R AC ALGLL DD+++I+
Sbjct: 1069 RINHVPPKIDDAYHLRILINNIRGPKGFDDIKTIEGVVHKTYRDACYALGLLDDDKDYIN 1128
Query: 848 GILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRK 899
GI + Y R LFV L+ S+ +P VW TW +L++ D QRK
Sbjct: 1129 GIEEANFWCFHKYVRKLFVIMLIFESLSSPAVVWEHTWKILSE----DFQRK 1176
Score = 112 bits (281), Expect = 6e-23
Identities = 53/115 (46%), Positives = 74/115 (64%)
Query: 6 QKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIG 65
++ S S CCM+G+I+LP L + PE + L + +++F N+R YN +F+FTS+G
Sbjct: 333 KEHGSSTTSRCCMQGQIVLPMLKESPEYMWWLFTSDHPDAKNFRANVRPYNMLFSFTSLG 392
Query: 66 GKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRM 120
GKVD + GRGP F + G+NYH I +L P GD KF QLY+ DT NEV NR+
Sbjct: 393 GKVDRSVKKGRGPSMFALQGENYHLIDALKPKPGDYAKFQQLYVMDTDNEVDNRI 447
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 369 bits (946), Expect = e-100
Identities = 179/404 (44%), Positives = 257/404 (63%), Gaps = 10/404 (2%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YTIEFQKRGLPHAHIL++L + K + ID +I AE+PD + FE V N+M+HG
Sbjct: 580 IYTIEFQKRGLPHAHILIFLDKKDKCPDASEIDRIISAEIPDKEEDREGFEAVENFMMHG 639
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG +K SPCM +C + FPKK+ TT D DG+P YRRR+ G + V LDN +V
Sbjct: 640 PCGEAKSNSPCMIENKCIRNFPKKFHSETTVDEDGFPTYRRRDNGRYIEKGNVKLDNRYV 699
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN LL+KYQAHIN+E CN+S SIKYLFKY++KG D+ T + S + DEI
Sbjct: 700 VPYNRDLLVKYQAHINVERCNRSKSIKYLFKYMHKGDDQATALIES---------DHDEI 750
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
K+Y +C Y+S +A WRIFQ+++H+++P V++L +HL +Q V+F ++ + ++R+ +
Sbjct: 751 KKYLECTYISGHDACWRIFQFEMHYRYPSVERLPFHLENEQQVIFPDSADLRKIVRKERI 810
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
+ F WM+ N R TY EFP +V+ K K W+ RK+G+ IGR+ + P +G+
Sbjct: 811 GVTKFTQWMETNKINDEARDFTYAEFPSKWVWKNKLKQWNKRKKGKMIGRIYYAHPASGD 870
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGG 858
YYLRMLLN +G +E++R+V+ VH +F+ ACEALG L DDRE+++ I + + A G
Sbjct: 871 KYYLRMLLNTVKGPRTFEEIRTVDGVVHPSFKSACEALGFLDDDREWVECIREASNYASG 930
Query: 859 AYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNN 902
R LF + L + +P +W W L + I QRK+ N
Sbjct: 931 NQLRHLFTTILCHCEVTDPKRIWESCWEDLGEDIEYK-QRKNLN 973
Score = 130 bits (326), Expect = 4e-28
Identities = 63/103 (61%), Positives = 82/103 (79%), Gaps = 1/103 (0%)
Query: 19 KGKIMLPYLADCPELLSNLL-RNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRG 77
+GK+ LP L P LSNL+ + RSR+++DNIR YNSMFAFTS+GGKVD +N+G G
Sbjct: 164 RGKVDLPTLKKPPTYLSNLMCKEKGKRSRNYMDNIRVYNSMFAFTSMGGKVDREINNGSG 223
Query: 78 PPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRM 120
P F ++GQNYHRIG+L+P EGD P++AQLYIYDT+NEV+NR+
Sbjct: 224 PYVFRMNGQNYHRIGTLLPEEGDKPRWAQLYIYDTENEVKNRI 266
>UniRef100_Q9SI21 Putative helicase [Arabidopsis thaliana]
Length = 1219
Score = 365 bits (937), Expect = 5e-99
Identities = 180/424 (42%), Positives = 267/424 (62%), Gaps = 28/424 (6%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYTIEFQKRGLPHAHIL+WL ++ KL ID I AE+PD P LFE + MVHG
Sbjct: 264 MYTIEFQKRGLPHAHILIWLDSKCKLTRAEHIDKAISAEIPDKLKDPELFEVIKEMMVHG 323
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG+ + PCM+NG+C+KF+PK + T D +G+P+YRRR ++ DN +V
Sbjct: 324 PCGVVNPKCPCMENGKCSKFYPKDHVPKTIIDKEGFPIYRRRRIDDFVQKKDFKCDNRYV 383
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDK------- 671
+PYN L ++Y+AHIN+E+CN+S S+KY+FKYI+KG DRVT+ + S +++K
Sbjct: 384 IPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIHKGPDRVTVVVGSSLNSKNKEKGKQKV 443
Query: 672 --------PVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLF 723
P + +E++ +F+CRY+S CEA WRI +Y IH++ V KL++HLP +Q + F
Sbjct: 444 NADTDGSEPKKKNEVEDFFNCRYVSACEAAWRILKYPIHYRSTSVMKLSFHLPGEQYIYF 503
Query: 724 KENEAIEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQG 783
K +E +E VL + + S+ +A R+LTY P F YDPK K ++ RK+G
Sbjct: 504 KGDEEVETVLNKADLDGSIQIA-----------RKLTYPNIPTRFTYDPKEKKFNLRKKG 552
Query: 784 ESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDR 843
+IGR+++VP + YYLR+LLNV G +E+L++VN ++ ++ ACEALGLL +D+
Sbjct: 553 FAIGRINYVPRDIEDGYYLRILLNVVPGPRSFEELKTVNGVLYKEWKDACEALGLLDNDQ 612
Query: 844 EFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNNR 903
E+ID + + + G Y R LFV L +++ +P +VW TW L++ I + ++ N
Sbjct: 613 EYIDDLKRTSFWSSGWYLRQLFVIML--DALISPENVWAATWQHLSEDIQNEKKKYFNRP 670
Query: 904 GMCL 907
CL
Sbjct: 671 VTCL 674
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 359 bits (921), Expect = 4e-97
Identities = 172/388 (44%), Positives = 245/388 (62%), Gaps = 21/388 (5%)
Query: 506 KRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPCGLSKR 565
+ GLPHAHILL+L E K+ + A +D I AE+PD +P L E V M+HGPCGL+ +
Sbjct: 392 RAGLPHAHILLFLEKEAKIPTTADVDKFIWAEIPDKYTHPILHEVVKEMMIHGPCGLANK 451
Query: 566 ESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVPYNPKL 625
SPCM+ G+C+K FPK + T + DG+P+Y+RR+ G + G+ LDN FV+PYNP L
Sbjct: 452 NSPCMEKGKCSKKFPKDFTSSTHINKDGFPIYKRRDDGRFVEKNGILLDNRFVIPYNPTL 511
Query: 626 LMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEIKQYFDCR 685
L+KYQAHIN+E+ ++S +KYLFKYINKG D+VT + G +EIK+++DCR
Sbjct: 512 LLKYQAHINVEWVSQSKIVKYLFKYINKGNDKVTARIKEG-------TGPNEIKRWYDCR 564
Query: 686 YLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIVKNSMFLA 745
Y+SPCEA WRIF +D+HH Q +++ ++E +EDVL + + S FLA
Sbjct: 565 YISPCEAAWRIFGFDLHHS-----------STSQNIIYDDDEDVEDVLNKEENQTSQFLA 613
Query: 746 WMDANCKYV---HGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGELYYL 802
++ N K R+ TY+EFP FV+ K W R++ S+GR+ V P G+ +YL
Sbjct: 614 FLKTNKKIAEDPEARKFTYIEFPSHFVWKKKQMEWTLRQRSVSVGRVYHVTPSAGQRFYL 673
Query: 803 RMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGGAYTR 862
R+LLN +G T YED+++V ++ TF+ AC AL LL+DD+E+ID I++ + G Y R
Sbjct: 674 RILLNKVKGPTCYEDIKTVGGILYPTFKDACYALSLLEDDQEYIDTIIEASQWGSGRYMR 733
Query: 863 ALFVSFLLPNSMCNPLHVWNETWHVLAD 890
LF L S+ P HVW TW+ L+D
Sbjct: 734 RLFAQLLTSESLSTPYHVWLNTWNQLSD 761
>UniRef100_Q94LS3 Putative helicase [Oryza sativa]
Length = 1501
Score = 355 bits (912), Expect = 4e-96
Identities = 170/394 (43%), Positives = 251/394 (63%), Gaps = 5/394 (1%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
++TIEFQKRGLPHAHIL++L K + ID +ICAE+PD P FE V N+M+HG
Sbjct: 607 VFTIEFQKRGLPHAHILIFLDKRGKSLEPSQIDELICAEIPDRDKDPETFEAVKNFMMHG 666
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + +SPCM + +C +FFP+ ++D T D +P+YRRR+ G + V+L+NGFV
Sbjct: 667 PCGEANPKSPCMVDHKCNRFFPRGFSDETIIDEVNFPIYRRRDDGRQIKKGRVNLNNGFV 726
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN LL K+Q H+N+E+ N+S SIKYLFK I D+ T + T +K +DEI
Sbjct: 727 VPYNKDLLAKFQVHMNVEWFNRSRSIKYLFKSICNEDDQATAVVEE---TDEK--NNDEI 781
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
K+Y C+Y + EA WRIF++ +H++ PPV++L++H +Q V+F ++ +++++RR
Sbjct: 782 KRYLGCKYTTATEACWRIFKFPLHYQEPPVERLSFHEENEQHVIFPDSTDLQEIVRRPRS 841
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
+MF WM+ N ++ R+LTY EFP + +D K W RK G IGR+ P +GE
Sbjct: 842 GVTMFTEWMETNKRHEDARELTYSEFPTKWTWDKNVKKWVRRKGGMKIGRIYNAHPASGE 901
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGG 858
YYLR++LN +GCT +ED+R+VN VH +++ AC ALG L DD E+I+ I + + A G
Sbjct: 902 RYYLRVILNTAKGCTTFEDIRTVNGTVHSSYKSACHALGFLNDDSEWIECIKEASCWASG 961
Query: 859 AYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
R LF + L + +P +W +W L+ I
Sbjct: 962 MKLRQLFATVLCHCEVTDPKRLWESSWEKLSKDI 995
Score = 127 bits (319), Expect = 2e-27
Identities = 57/110 (51%), Positives = 80/110 (71%)
Query: 11 PDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDC 70
P F +CC +GK+ LP L + P L++LL RS ++ NIRSYNSMFAFTS+GG VD
Sbjct: 184 PSFGLCCKQGKVALPPLKEQPPYLTSLLTRDGGRSTNYQQNIRSYNSMFAFTSMGGTVDR 243
Query: 71 GLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRM 120
+N+G GP F ++GQN+H IG+L+P + P+F QLY+YDT+NE++NR+
Sbjct: 244 KINNGHGPYIFRLNGQNHHHIGTLLPEGSNKPRFQQLYVYDTENEIENRI 293
>UniRef100_Q94LS7 Putative helicase [Oryza sativa]
Length = 1573
Score = 354 bits (909), Expect = 9e-96
Identities = 167/394 (42%), Positives = 251/394 (63%), Gaps = 4/394 (1%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
++TIEFQK+GLPHAHIL++L + K + ID +ICAE+PD P FE V N+M+HG
Sbjct: 622 IFTIEFQKKGLPHAHILIFLDKKEKCLKPSQIDKMICAEIPDSNKDPETFEAVKNFMMHG 681
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + +SPCM + +C ++FPK ++D T D +P+Y+RR+ G + ++L+NGFV
Sbjct: 682 PCGETNPKSPCMVDHKCDRYFPKGFSDETIIDEVNFPIYKRRDDGRQIKKGRINLNNGFV 741
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN LL+K+QAHIN+E+ N+S SI+YLFK I G D+ T + T +D +DEI
Sbjct: 742 VPYNKDLLVKFQAHINVEWFNRSKSIRYLFKSIYNGDDQATAVVEETDTAKD----NDEI 797
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
K+Y C Y++ EA WRIF + +H++ P VQ+L +H+ +Q V+F ++ +++++R
Sbjct: 798 KRYLGCSYMTATEACWRIFTFPLHYQEPSVQRLFFHVENEQQVIFPDSTDLQEIIRHPRS 857
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
+MF WM+ N K+ R+LTY EFP + + K K W RK + IGR+ P +GE
Sbjct: 858 GVTMFTEWMETNKKHEDARELTYSEFPTKWTWVNKVKKWVRRKGRKKIGRIYNAHPASGE 917
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGG 858
YYLR++LN +GCT +ED+R+VN VH +++ AC ALG L DD E+I+ I + + A G
Sbjct: 918 RYYLRVILNTAKGCTTFEDIRTVNGFVHSSYKSACHALGFLNDDNEWIECIKEASCWASG 977
Query: 859 AYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
+ LF + L + +P +W W L+ I
Sbjct: 978 IELQQLFATILCHCEVTDPKSLWESIWEELSKDI 1011
Score = 134 bits (336), Expect = 3e-29
Identities = 63/123 (51%), Positives = 83/123 (67%)
Query: 2 SVKAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAF 61
S Q+ P F +CC +GK+ LP L + P LS+LL S ++ NIRSYNSMFAF
Sbjct: 190 SFSRQRKNQPSFGLCCKQGKVALPPLKEPPHFLSSLLARDGGTSENYQQNIRSYNSMFAF 249
Query: 62 TSIGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMS 121
TS+GG VD +N GRGP F ++GQNYH IG+L+P + P+F QLYIYDT+NE++NR+
Sbjct: 250 TSMGGAVDRKINKGRGPYVFRLNGQNYHHIGTLLPKGSNKPRFQQLYIYDTENEIKNRIE 309
Query: 122 HFR 124
R
Sbjct: 310 ASR 312
>UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thaliana]
Length = 1678
Score = 351 bits (900), Expect = 1e-94
Identities = 180/412 (43%), Positives = 252/412 (60%), Gaps = 43/412 (10%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHILL++ A+ KL + ID +I AE+PD + P L+E + N M+HG
Sbjct: 788 MYTVEFQKRGLPHAHILLFMHAKSKLPTADDIDKLISAEIPDKEKEPDLYEVIKNSMIHG 847
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM +G C+K +PKK+ D+T SDGYP+YRRR T + G+ DN +V
Sbjct: 848 PCGSANVNSPCMVDGECSKLYPKKHQDITKIGSDGYPIYRRRKTDDYVEKGGIKCDNRYV 907
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTT----QDKPVE 674
VPYN KL ++Y AHIN+E+CN++ SIKYLFKYINKG D+V + + T + P +
Sbjct: 908 VPYNKKLSLRYNAHINVEWCNQNGSIKYLFKYINKGPDKVVFIVEPTQQTTAGDSETPQQ 967
Query: 675 D--------DEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKEN 726
+ +EIK +FDCRY+S EAVWRIF+Y I H PVQKL++H+ KQ F
Sbjct: 968 EQGSAEKKKNEIKDWFDCRYVSASEAVWRIFKYPIQHISTPVQKLSFHVEGKQPAYFDPK 1027
Query: 727 EAIEDVLRRNIVKNSMFLAWMDANCKYVHG------RQLTYVEFPELFVYDPKSKSWHPR 780
IEDVL R +S F+AW+ N + G R+ Y E P F +D ++KS+ R
Sbjct: 1028 SNIEDVLERVANVDSQFMAWLTLNRRNAVGKNGKRARECLYAEIPAYFTWDGENKSFKKR 1087
Query: 781 KQGESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLK 840
+G SIGR+ +V + Y+LR+LLN+ RG T Y ++++ + V+ TF+ AC A G+L
Sbjct: 1088 TRGFSIGRIHYVSRKMEDEYFLRVLLNIVRGPTSYAEIKTYDGVVYKTFKEACFARGILD 1147
Query: 841 DDREFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
DD+ FIDG+++ HV ++TWH+LA+ I
Sbjct: 1148 DDQVFIDGLVEAT-------------------------HVRSQTWHLLAEDI 1174
Score = 120 bits (301), Expect = 3e-25
Identities = 57/116 (49%), Positives = 75/116 (64%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K + + F++CC +G + LP+L + P LL NLL S+H+ DN R +N +FA TS
Sbjct: 350 KKETNKESVFTLCCGEGSVKLPFLKESPHLLKNLLSGNHPLSKHYRDNARIFNMVFAMTS 409
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNR 119
GGKVD + GRGP F + G NYH IGSL T GD K++QLYI DT+NEV+NR
Sbjct: 410 FGGKVDKSMPKGRGPAMFRLQGGNYHLIGSLKLTPGDYAKYSQLYIIDTENEVENR 465
>UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa]
Length = 1516
Score = 351 bits (900), Expect = 1e-94
Identities = 183/430 (42%), Positives = 261/430 (60%), Gaps = 5/430 (1%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YT+EFQKRGLPH H L+WL+A S ++ID ICAE+PD +E VS +M+HG
Sbjct: 511 LYTVEFQKRGLPHIHCLVWLAAATADVSASIIDGFICAEIPDYDTDRLGYELVSEFMMHG 570
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PC + ++ PCMKN +C+K +PK + D T D G+ +YRRRN G S + G+ LDN V
Sbjct: 571 PCSDANKKCPCMKNDKCSKHYPKDFQDETIVDESGFTIYRRRNDGRSIMKNGILLDNRSV 630
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSM-SSGRTTQDKPVED-- 675
VPYN LL KY+AHIN+E+CNKSN IKYLFKYI KG DR + ++G+T P D
Sbjct: 631 VPYNMALLKKYEAHINVEWCNKSNLIKYLFKYITKGHDRARIYFETTGKTQNASPNHDLA 690
Query: 676 --DEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVL 733
+EI +Y D R+LS EA+ R+F++DIH++ PPV++L HLP K V +++ + VL
Sbjct: 691 PRNEILEYMDARFLSTYEALHRLFEFDIHYRVPPVERLVVHLPGKNFVRYEKGADLRAVL 750
Query: 734 RRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVP 793
K SM W + N K LTY EFP+ + ++P SK+WH R IGR+ +V
Sbjct: 751 ESPGAKRSMLTEWFETNKKNSKAHSLTYCEFPKEWTWEPSSKTWHERTPAPKIGRIYYVH 810
Query: 794 PGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVA 853
P GELYYLRMLL + +G Y D+R+ + V+ T+R ACEA GLL+ D E+ +
Sbjct: 811 PTAGELYYLRMLLMIVKGAQSYADVRTYDGVVYGTYREACEARGLLEGDNEWHLLFDEAI 870
Query: 854 TLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNNRGMCL*C*H** 913
A A R LFV+ LL S+ + ++++ W + D I L++ +N + H
Sbjct: 871 VTASSAQLRQLFVTVLLYCSVGDVRSLFDKYWLYMTDDIHNRLKKALDNPHCVIPHDHLL 930
Query: 914 DFIRYHIIAI 923
+ + + +IA+
Sbjct: 931 NMLLHELIAV 940
Score = 121 bits (304), Expect = 1e-25
Identities = 60/112 (53%), Positives = 81/112 (71%), Gaps = 1/112 (0%)
Query: 13 FSICCMKGKIMLPYLADCPELLSNLLR-NTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCG 71
+++CC G+I LP L PE L++LL N D RS++FL IRSYNSMFAF+S+G +D
Sbjct: 90 YNLCCKGGRIQLPKLRAPPEPLASLLNYNGDARSKNFLRQIRSYNSMFAFSSMGAAIDKS 149
Query: 72 LNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMSHF 123
+N G P F I+G +HRIG+LVP+ G PKFAQLY+YD +NE+QNR++ F
Sbjct: 150 INTGNAPYVFKINGVVHHRIGTLVPSCGSPPKFAQLYVYDPENELQNRLNIF 201
>UniRef100_Q8LRG5 Putative helicase [Oryza sativa]
Length = 1453
Score = 348 bits (894), Expect = 5e-94
Identities = 175/409 (42%), Positives = 245/409 (59%), Gaps = 5/409 (1%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YT+EFQKRGLPH H ++W +A S +DS+ICAE+PD P + V +M+HG
Sbjct: 475 LYTVEFQKRGLPHIHCIVWRAAADAEFSATAVDSLICAEIPDVFSDPLGYALVDEFMIHG 534
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + CMKNG C+K FPK + + TT D G+ VYRRRN G + G+ LDN +V
Sbjct: 535 PCGDKNKSCVCMKNGHCSKHFPKGFQEETTMDEFGFTVYRRRNDGRYVVKNGIKLDNRWV 594
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVE---- 674
VPYN KLL KYQAHIN+E CNKSN IKYLFKYI KG DR L + T +K V+
Sbjct: 595 VPYNMKLLKKYQAHINVESCNKSNMIKYLFKYITKGGDRTKLYFETTGNTPNKTVDGTVL 654
Query: 675 -DDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVL 733
+EI +Y + R+LS CEA WR F++DIH++ P V++L HLP V +K+ ++ +L
Sbjct: 655 PPNEIDEYINARFLSTCEAFWRAFEFDIHYRVPAVERLPIHLPNMNFVQYKKGTDLKKLL 714
Query: 734 RRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVP 793
K +M W + N K+ + R LTY +FP+ + +D ++ W PR E IGR+ +V
Sbjct: 715 DSPAAKKTMLTEWFECNKKHPNARTLTYCDFPKQWTWDNSARCWRPRTPVEKIGRIYYVS 774
Query: 794 PGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVA 853
P GELYYLRMLL +G Y D+R+ V+ TFR ACE+ GLL++D ++ +
Sbjct: 775 PAAGELYYLRMLLMTVKGAKSYADVRTFEGTVYPTFRQACESRGLLENDNDWHLLFDEAI 834
Query: 854 TLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNN 902
A R LFV+ ++ S+ N ++++ W D I L+ +N
Sbjct: 835 VSASSLQLRQLFVTVVMFCSVGNVRSLFDKYWLYFTDDIQHRLRTALSN 883
Score = 119 bits (299), Expect = 5e-25
Identities = 59/112 (52%), Positives = 80/112 (70%), Gaps = 1/112 (0%)
Query: 13 FSICCMKGKIMLPYLADCPELLSNLLR-NTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCG 71
+++CC GKI LP L P++L+ LL+ + D +S+ FL IRSYNS+FAFTS+G V+
Sbjct: 52 YNLCCRGGKISLPELKYPPDMLAKLLKFDGDAQSKRFLRQIRSYNSLFAFTSLGADVEKS 111
Query: 72 LNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMSHF 123
+N+G P F I+G +HRIGSL+P G PKFAQLYIYDT+NE NR++ F
Sbjct: 112 INNGTAPYVFKINGVVHHRIGSLLPQRGAKPKFAQLYIYDTENETANRINIF 163
>UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana]
Length = 1250
Score = 346 bits (887), Expect = 3e-93
Identities = 175/403 (43%), Positives = 251/403 (61%), Gaps = 11/403 (2%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YTIEFQKRGLPHAHILLWL + K + ID I AE+P P V +M+HG
Sbjct: 357 VYTIEFQKRGLPHAHILLWLQGDLKKPTPNDIDKYISAEIPVKDKDPEGHTLVEQHMMHG 416
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG + SPCM+ G C+K FP+++ + T + G+ +YRRRN + LDN FV
Sbjct: 417 PCGKDRPSSPCMEKGICSKKFPREFVNHTKMNESGFILYRRRNDQRYVLKGQTRLDNRFV 476
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTT---------Q 669
VP+N ++L KY+AHIN+E+CNKS++IKYLFKYI KGVD+ T + G + +
Sbjct: 477 VPHNLEILKKYKAHINVEWCNKSSAIKYLFKYITKGVDKATFIIQKGNSVNGQGSGNGFE 536
Query: 670 DKPVEDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAI 729
+KP +EI +Y DCRYLS CEA+WRIF ++IHH PPVQ+L HLP +Q +F+E E +
Sbjct: 537 EKP--RNEINEYLDCRYLSACEAMWRIFMFNIHHHNPPVQRLPLHLPGEQSTIFEEEENL 594
Query: 730 EDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRM 789
E+V R + +M + + N R+L YV+ P +FV+D +K + RKQ E+IGR+
Sbjct: 595 ENVEYRYGHERTMLTEYFELNKICEDARKLKYVQVPTMFVWDSTNKMYTRRKQRENIGRI 654
Query: 790 SFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGI 849
+ P G+LYYLR+LLN +G T ++ L++V VH++F+ AC GLL D+E+ D +
Sbjct: 655 VNILPTAGDLYYLRILLNKVKGATSFDYLKTVGGVVHESFKAACHTRGLLDGDKEWHDAM 714
Query: 850 LDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGI 892
+ A + R+LFV L+ + PL +W+ W +AD +
Sbjct: 715 DEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWSHCWESMADDV 757
Score = 95.9 bits (237), Expect = 8e-18
Identities = 50/99 (50%), Positives = 65/99 (65%), Gaps = 4/99 (4%)
Query: 23 MLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTSIGGKVDCGLNDGRGPPQFV 82
+LP P++L LL T S H+ NIR+YNS+ AFTS+G ++D + GP F
Sbjct: 27 VLPPEPQPPQMLKKLL----TESPHYQRNIRTYNSILAFTSMGAQIDKNVMHKGGPFTFR 82
Query: 83 ISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMS 121
I GQN+H++GSLVP EG PK QLYI+DT NEV NR+S
Sbjct: 83 IHGQNHHKLGSLVPEEGKPPKILQLYIFDTANEVHNRIS 121
>UniRef100_Q6MW89 B1248C03.15 protein [Oryza sativa]
Length = 1550
Score = 325 bits (832), Expect = 8e-87
Identities = 164/397 (41%), Positives = 245/397 (61%), Gaps = 3/397 (0%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YTIEFQKRGLPH HI++WL+ + L++ + S I A+ PDP + ++ V N+MVHG
Sbjct: 486 IYTIEFQKRGLPHVHIIIWLAKKEPLDA-KKVGSYISAQFPDPAVDKIGYDAVCNFMVHG 544
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG S CM G+C KF+PK++ + TT +G+ Y R N G++ R V++DN FV
Sbjct: 545 PCGPHNPSSVCMSEGKCTKFYPKEFCEETTILENGFTQYARPNNGITFKRNEVEIDNRFV 604
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVED-DE 677
VP+N L++K+QAHIN+E N KYLF+Y+ KG D + S + + E +E
Sbjct: 605 VPHNVDLVVKFQAHINVEKVNYDGMHKYLFRYVTKGFDCSRVGFHSNSSNSESSSETINE 664
Query: 678 IKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNI 737
I Y +CR ++P +A WR+ Q+DIHH P V++L +LP + V+F E++ +E+V+
Sbjct: 665 INNYLECRCVTPNDAAWRLLQFDIHHTDPSVERLPVYLPFENNVVFTEDDDLEEVIDDPN 724
Query: 738 VKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRK-QGESIGRMSFVPPGT 796
S AW++AN + R+LTY+EFP+ + + K K W R+ IGR++ V P
Sbjct: 725 NSKSKLTAWLEANMENPSARELTYIEFPKYWTWHNKEKYWDGRRGASRRIGRIAHVSPSQ 784
Query: 797 GELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLA 856
GE YYLRMLL++ RG + ++R+V++ + TFR ACEALGLL DD+E+ + I D A A
Sbjct: 785 GEAYYLRMLLHIVRGPRSFAEIRTVSSVEYPTFRAACEALGLLGDDQEWSNAIKDAAQWA 844
Query: 857 GGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIA 893
R LFV+ LL + NP ++NE +++ IA
Sbjct: 845 LPFQLRQLFVTMLLFCEVTNPTRLFNEHMSCMSEDIA 881
Score = 38.1 bits (87), Expect = 1.9
Identities = 18/29 (62%), Positives = 21/29 (72%)
Query: 58 MFAFTSIGGKVDCGLNDGRGPPQFVISGQ 86
MFA TS+G KV+ +NDGR P F ISGQ
Sbjct: 324 MFAMTSMGAKVNESINDGRAPYVFKISGQ 352
>UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana]
Length = 1230
Score = 323 bits (829), Expect = 2e-86
Identities = 166/401 (41%), Positives = 235/401 (58%), Gaps = 37/401 (9%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
MYT+EFQKRGLPHAHI++W+ YK ++ +D +I AE+PD + +P L++ VS M+HG
Sbjct: 405 MYTVEFQKRGLPHAHIIVWMDPRYKFHTADHVDKIIFAEIPDKEKHPELYQAVSECMIHG 464
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PC L SPCM+NG+C+K++PK + + T+ D GYP+YRRR++G + DN +V
Sbjct: 465 PCRLVNPNSPCMENGKCSKYYPKNHVENTSLDYKGYPIYRRRDSGRFIEKNKYQCDNWYV 524
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPVEDDEI 678
VPYN LL KY+AHIN+E+CN+S SIKYLFKY+NKG DRVT + + G D P E +++
Sbjct: 525 VPYNDVLLRKYRAHINVEWCNQSVSIKYLFKYVNKGPDRVTQN-NVGEINND-PQERNQV 582
Query: 679 KQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRNIV 738
+ YFDCR Y IH++ V KLT+H KQ V KE E E VL R
Sbjct: 583 QDYFDCR------------GYPIHYRQTSVTKLTFHEKGKQSVYVKEGETAESVLYRVNN 630
Query: 739 KNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVPPGTGE 798
+ F+AW + N + +L Y + P + ++ VPP +
Sbjct: 631 DETQFIAWFELNKRDPEAAKLLYEQIPNFYT-------------------INHVPPKIDD 671
Query: 799 LYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATLAGG 858
Y+LR+L+N R ++D+++V VH T+R AC ALGLL DD+E+I GI +
Sbjct: 672 AYHLRILINNIRAPKGFDDIKTVEGVVHKTYRDACYALGLLDDDKEYIHGIEEANFWCSP 731
Query: 859 AYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRK 899
Y R FV L+ S+ +P+ VW TW +L + D QRK
Sbjct: 732 KYVRKSFVIMLISESLSSPVVVWEHTWKILFE----DFQRK 768
Score = 37.0 bits (84), Expect = 4.3
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLL 38
+ +KSA+P ++ CCM+G+I+LP L + PE + LL
Sbjct: 83 RRKKSANPVYTGCCMQGQIVLPMLKESPEYMWWLL 117
>UniRef100_Q5VR06 Helicase-like protein [Oryza sativa]
Length = 1427
Score = 323 bits (828), Expect = 2e-86
Identities = 152/408 (37%), Positives = 240/408 (58%), Gaps = 5/408 (1%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+Y++EFQKRGLPH HIL+WL + +I MID I E+PDP+ P + ++ +M+HG
Sbjct: 456 LYSVEFQKRGLPHVHILVWLDKKPSEITIEMIDKWISTEIPDPREDPLGYILIAEHMMHG 515
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG PCMK G+C+KF+PK++ D T F +G+ Y+RRNT + + +LDN +V
Sbjct: 516 PCGAKNENCPCMKKGKCSKFYPKEFNDQTNFTENGFAQYKRRNTNIYIRKDNHNLDNRWV 575
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRV-----TLSMSSGRTTQDKPV 673
VP+N LL +YQAH+N+E+ N+S +KYL KY+NKG D+ + + ++
Sbjct: 576 VPHNLFLLKRYQAHLNVEFVNQSRMLKYLCKYVNKGGDKAKIIFQRIKQGIDHSENEQTE 635
Query: 674 EDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVL 733
+ DEI++Y +CRY+ + +WR+ Y+IH+ WPPV+++ HLP +V ++ +++++
Sbjct: 636 KIDEIEEYLECRYICDQDGMWRLLGYEIHYHWPPVERMPVHLPLMNMVKLTKDTKLKNII 695
Query: 734 RRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIGRMSFVP 793
+ +M W AN + R LTY EFP+ + +D K + W R+ G I R+ +V
Sbjct: 696 ENPDNQRTMLTEWFMANQLHEEARNLTYCEFPQKWKWDKKERKWVKRQHGFKIARLYYVK 755
Query: 794 PGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVA 853
P GE +YLRMLL + +G YED+R+ N + TF+ C A GLL DD E+ + A
Sbjct: 756 PTEGERFYLRMLLMIVKGAKNYEDIRTYNGITYKTFKETCAARGLLMDDNEWYKTFDEAA 815
Query: 854 TLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHN 901
+ A R+LF+ LL ++ + + W + D I L K++
Sbjct: 816 SWATSPQLRSLFIILLLYCNLEDERKFFETNWEKMVDDIKFQLISKYH 863
Score = 119 bits (298), Expect = 7e-25
Identities = 55/109 (50%), Positives = 77/109 (70%), Gaps = 1/109 (0%)
Query: 13 FSICCMKGKIMLPYLADCPELLSNLLRNT-DTRSRHFLDNIRSYNSMFAFTSIGGKVDCG 71
+S CC KI +P + PE L+ LL N D S+HF+ IR YNS+F+FTS+GG +D
Sbjct: 32 YSNCCKNNKIKIPPFQNPPETLARLLNNKEDNLSKHFMQKIRQYNSLFSFTSMGGTIDKN 91
Query: 72 LNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRM 120
+N+G GP F ++GQ +HRIG L+P + PKFA+LYI+DTKNE++NR+
Sbjct: 92 INNGDGPYVFRVNGQIHHRIGCLLPKPNEIPKFAELYIFDTKNEIENRI 140
>UniRef100_Q84QR0 Helicase-like protein [Oryza sativa]
Length = 1330
Score = 311 bits (796), Expect = 1e-82
Identities = 160/388 (41%), Positives = 236/388 (60%), Gaps = 11/388 (2%)
Query: 499 MYTIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHG 558
+YT+EFQKRGLPH HI++WLS E L++ +D I A++P+P L P FE V+++M+HG
Sbjct: 428 VYTVEFQKRGLPHVHIIIWLSKEEPLDA-EKVDLRISAQLPNPTLDPIGFEAVTSFMIHG 486
Query: 559 PCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFV 618
PCG SPCM GRC+KF+PK++ + T +G+ Y R N + T + GVD+DN +
Sbjct: 487 PCGPGISYSPCMSEGRCSKFYPKEFCEHTYILQNGFTQYARPNNQIVT-KNGVDIDNRVI 545
Query: 619 VPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVD--RVTLSMSSGRTTQDKPVEDD 676
VP+N L++KYQ HIN+E N KYLFKY+ K D R + +S T +
Sbjct: 546 VPHNVDLVVKYQDHINLESVNHDGMHKYLFKYVTKVYDCSRAGIRRNSANETIN------ 599
Query: 677 EIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENEAIEDVLRRN 736
EI Y +CR ++P + WR+ Q+DIHH P V++L HLP + V++ E++ +E+V+
Sbjct: 600 EIDNYLECRCVTPNDVAWRLLQFDIHHTDPSVERLPVHLPLENNVVYTEDDDLEEVIENP 659
Query: 737 IVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQ-GESIGRMSFVPPG 795
+ S AW++AN ++ R+ TY+EFPE F + K W R+ IGR++ V P
Sbjct: 660 GNQKSKLTAWLEANSQFPQAREHTYIEFPEYFTWHASEKYWDIRRGCYNKIGRIAHVDPT 719
Query: 796 TGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDDREFIDGILDVATL 855
GE YYLRMLL++ +G + ++R+++ H TFR ACEALGLL DD+E+ + D
Sbjct: 720 KGEKYYLRMLLHIVKGPKTFSEIRNISGQQHPTFRAACEALGLLGDDQEWSHALNDAVHW 779
Query: 856 AGGAYTRALFVSFLLPNSMCNPLHVWNE 883
A R LFV+ LL + NP ++ E
Sbjct: 780 ALPYQLRQLFVTILLFCEVTNPQRLFTE 807
Score = 42.7 bits (99), Expect = 0.079
Identities = 17/35 (48%), Positives = 26/35 (73%)
Query: 84 SGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQN 118
S Q + SL+P++G P++AQLYI+DT+NE+ N
Sbjct: 102 SNQVFRHHRSLIPSQGRRPEYAQLYIFDTENEISN 136
>UniRef100_Q6AUV2 Hypothetical protein OSJNBa0091B22.10 [Oryza sativa]
Length = 1500
Score = 305 bits (782), Expect = 5e-81
Identities = 156/360 (43%), Positives = 218/360 (60%), Gaps = 5/360 (1%)
Query: 548 FECVSNYMVHGPCGLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTN 607
+E VS +M+HGPCG + + PCMKN +C+K +PK + D T D + +YR+RN G
Sbjct: 370 YELVSEFMMHGPCGDANKSCPCMKNDKCSKNYPKDFQDETIIDEAAFTIYRQRNDGRCII 429
Query: 608 RRGVDLDNGFVVPYNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSM-SSGR 666
+ G+ LDN VVPYN KLL KY+AHIN+E+CNKSN IKYLFKYI KG DR + ++ +
Sbjct: 430 KNGIPLDNRSVVPYNMKLLKKYEAHINVEWCNKSNLIKYLFKYITKGQDRAKIYFETTAK 489
Query: 667 TTQDKPVED----DEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVL 722
TT P D DEI +Y D R+LS CEA+ R+F++DIH++ P V++L HLP K V
Sbjct: 490 TTNASPNHDLAPRDEILEYMDARFLSTCEALHRLFEFDIHYRVPLVERLVVHLPGKNYVR 549
Query: 723 FKENEAIEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQ 782
+++ + VL K SM W +AN KY LTY EFP+ + ++ S+ W R
Sbjct: 550 YEKGSDLRAVLESPAAKRSMLTEWFEANKKYSKAHSLTYCEFPKEWSWESSSRVWRQRTP 609
Query: 783 GESIGRMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTFRGACEALGLLKDD 842
IGR+ +V P GELYYLRMLL + +G Y D+R+ NN V+ TFR ACEA GLL+ D
Sbjct: 610 AAKIGRIYYVHPTAGELYYLRMLLMIVKGAQSYVDVRTFNNIVYATFREACEARGLLEGD 669
Query: 843 REFIDGILDVATLAGGAYTRALFVSFLLPNSMCNPLHVWNETWHVLADGIACDLQRKHNN 902
E+ + A R LFV+ LL S+ + ++++ W + D I L++ +N
Sbjct: 670 NEWYLLFDEAIVSASSQQLRQLFVTILLYCSVNDVRSLFDKYWLYMTDDIHSKLKKALDN 729
Score = 68.6 bits (166), Expect = 1e-09
Identities = 32/60 (53%), Positives = 42/60 (69%)
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMSHF 123
+G VD +N G P F I+ +HRIG+LVP+ G PKFAQLYIYD +NE+QNR++ F
Sbjct: 1 MGAIVDKSINTGNAPYVFKINEVVHHRIGTLVPSRGSPPKFAQLYIYDAENEIQNRLNIF 60
>UniRef100_Q9SH75 Putative helicase [Arabidopsis thaliana]
Length = 1241
Score = 300 bits (768), Expect = 2e-79
Identities = 145/342 (42%), Positives = 214/342 (62%), Gaps = 13/342 (3%)
Query: 501 TIEFQKRGLPHAHILLWLSAEYKLNSIAMIDSVICAEVPDPKLYPSLFECVSNYMVHGPC 560
T+ FQKRGLPHAHILL++ K + ID+++ AE+PD P L+E V++ M+HGPC
Sbjct: 406 TVMFQKRGLPHAHILLFMDKSCKFPTSDDIDNILSAEIPDKAKDPKLYEVVNDCMIHGPC 465
Query: 561 GLSKRESPCMKNGRCAKFFPKKYADVTTFDSDGYPVYRRRNTGVSTNRRGVDLDNGFVVP 620
G + +ESPC+ +G+C+KFFPKK + TT DGYP+YRRR + + G+ DN +VVP
Sbjct: 466 GAANKESPCIVDGKCSKFFPKKLVEQTTVGKDGYPIYRRRESEHFVEKGGIKCDNTYVVP 525
Query: 621 YNPKLLMKYQAHINIEYCNKSNSIKYLFKYINKGVDRVTLSMSSGRTTQDKPV------- 673
YN L ++Y+AHIN+E+C +S SIKYLFKYINKG DRV + + T + +
Sbjct: 526 YNRMLSLRYRAHINVEWCKQSGSIKYLFKYINKGQDRVAIVVEPKDKTSNMVLFSGSQKL 585
Query: 674 ------EDDEIKQYFDCRYLSPCEAVWRIFQYDIHHKWPPVQKLTYHLPRKQVVLFKENE 727
+D EIK YFDCRY+S EAVWRIF++ I ++ PV KL+YHLP KQ + F++ +
Sbjct: 586 LVAVIDDDKEIKDYFDCRYVSASEAVWRIFKFPIQYRTTPVMKLSYHLPGKQPLCFEDTQ 645
Query: 728 AIEDVLRRNIVKNSMFLAWMDANCKYVHGRQLTYVEFPELFVYDPKSKSWHPRKQGESIG 787
I+++ + ++ MF+ ++ N + RQ Y E P F +D ++K W R++G IG
Sbjct: 646 NIDELSEKKANEDFMFIGFLKLNQECEFARQFIYTEIPPYFTWDGQNKQWKLRERGFYIG 705
Query: 788 RMSFVPPGTGELYYLRMLLNVQRGCTKYEDLRSVNNNVHDTF 829
RM++ YY+++LL + G ED+R+ + V F
Sbjct: 706 RMNYASIKMEPEYYMKILLGIVCGPKSDEDIRTYKDVVRRKF 747
Score = 137 bits (346), Expect = 2e-30
Identities = 63/119 (52%), Positives = 84/119 (69%)
Query: 4 KAQKSASPDFSICCMKGKIMLPYLADCPELLSNLLRNTDTRSRHFLDNIRSYNSMFAFTS 63
K +KS P FS+CC++G + LP+L + PEL+ LL D RHF +NIR+YN +F+ TS
Sbjct: 10 KRRKSKKPKFSLCCLQGSVKLPFLTESPELIRELLSCDDALRRHFRENIRAYNMLFSMTS 69
Query: 64 IGGKVDCGLNDGRGPPQFVISGQNYHRIGSLVPTEGDNPKFAQLYIYDTKNEVQNRMSH 122
+GGKVD G+GP QF + G NYH IGS++P EGD KF+QLYI DT NEV+NR ++
Sbjct: 70 LGGKVDRSNPQGKGPNQFQLHGANYHLIGSMLPGEGDYAKFSQLYIVDTGNEVENRSTN 128
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.353 0.158 0.566
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,066,885,280
Number of Sequences: 2790947
Number of extensions: 80470078
Number of successful extensions: 374000
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 87
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 373580
Number of HSP's gapped (non-prelim): 249
length of query: 1408
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1268
effective length of database: 457,317,253
effective search space: 579878276804
effective search space used: 579878276804
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0110a.10


