

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.3
(169 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
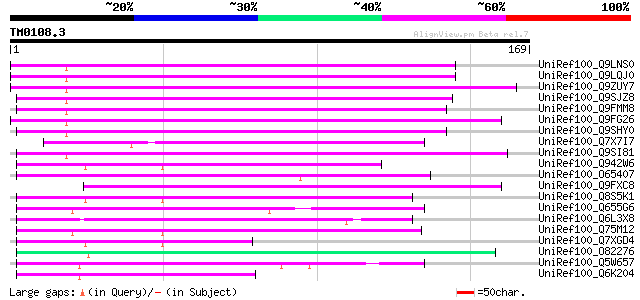
UniRef100_Q9LNS0 F1L3.4 [Arabidopsis thaliana] 74 2e-12
UniRef100_Q9LQJ0 F28G4.15 protein [Arabidopsis thaliana] 74 2e-12
UniRef100_Q9ZUY7 Putative reverse transcriptase [Arabidopsis tha... 71 9e-12
UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcrip... 69 6e-11
UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like... 64 2e-09
UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like... 64 2e-09
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 59 4e-08
UniRef100_Q7X7I7 OSJNBa0093F12.7 protein [Oryza sativa] 59 6e-08
UniRef100_Q9SI81 F23N19.5 [Arabidopsis thaliana] 57 1e-07
UniRef100_Q942W6 P0506E04.5 protein [Oryza sativa] 56 4e-07
UniRef100_O65407 Hypothetical protein F18E5.40 [Arabidopsis thal... 55 9e-07
UniRef100_Q9FXC8 F12A21.24 [Arabidopsis thaliana] 55 9e-07
UniRef100_Q8S5K1 Putative retroelement [Oryza sativa] 54 2e-06
UniRef100_Q655G6 Hypothetical protein P0009H10.11 [Oryza sativa] 53 3e-06
UniRef100_Q6L3X8 Hypothetical protein [Solanum demissum] 53 3e-06
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 53 3e-06
UniRef100_Q7XGD4 Putative reverse transcriptase [Oryza sativa] 53 3e-06
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 52 5e-06
UniRef100_Q5W657 Hypothetical protein OSJNBb0052F16.4 [Oryza sat... 51 1e-05
UniRef100_Q6K204 Hypothetical protein B1469H02.35 [Oryza sativa] 49 4e-05
>UniRef100_Q9LNS0 F1L3.4 [Arabidopsis thaliana]
Length = 253
Score = 73.6 bits (179), Expect = 2e-12
Identities = 50/146 (34%), Positives = 69/146 (47%), Gaps = 1/146 (0%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W P VG +K N DG S R A GGV+RD DGNW GF + + LAELW
Sbjct: 83 IAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLAELWGA 142
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
+++A +RG + + +E DS + + P S +VR + V +
Sbjct: 143 YYGLNIAWERGVTQLEMEIDSEMVVGFLRTGIDDSHPLSFLVRLCHGLLSKDWSVRISHV 202
Query: 120 PREANMVADCLAQNAHVFNFGVHLLH 145
REAN +AD LA A G HL +
Sbjct: 203 YREANRLADGLANYAFFLPLGFHLFN 228
>UniRef100_Q9LQJ0 F28G4.15 protein [Arabidopsis thaliana]
Length = 272
Score = 73.6 bits (179), Expect = 2e-12
Identities = 50/146 (34%), Positives = 69/146 (47%), Gaps = 1/146 (0%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W P VG +K N DG S R A GGV+RD DGNW GF + + LAELW
Sbjct: 102 IAWSPPRVGWFKLNTDGASRGNPRLATAGGVVRDGDGNWCYGFSLNIGICSAPLAELWGA 161
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
+++A +RG + + +E DS + + P S +VR + V +
Sbjct: 162 YYGLNIAWERGVTQLEMEIDSEMVVGFLRTGIDDSHPLSFLVRLCHGLLSKDWSVRISHV 221
Query: 120 PREANMVADCLAQNAHVFNFGVHLLH 145
REAN +AD LA A G HL +
Sbjct: 222 YREANRLADGLANYAFFLPLGFHLFN 247
>UniRef100_Q9ZUY7 Putative reverse transcriptase [Arabidopsis thaliana]
Length = 314
Score = 71.2 bits (173), Expect = 9e-12
Identities = 53/166 (31%), Positives = 72/166 (42%), Gaps = 1/166 (0%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W P G WK N DG S A GGVLRD +G W GF + + LAELW +
Sbjct: 144 IAWSKPEEGWWKLNTDGASRGNPGLASAGGVLRDEEGAWRGGFALNIGVCSAPLAELWGV 203
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
+ +A +R + + IE DS + + P S +VR I +V +
Sbjct: 204 YYGLYIAWERRVTRLEIEVDSEIVVGFLKIGINEVHPLSFLVRLCHDFISRDWRVRISHV 263
Query: 120 PREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTVVAS 165
REAN +AD LA A G H L R++L D+ S
Sbjct: 264 YREANRLADGLANYAFSLPLGFHSLSLVPDSLRFILLDDTSGATVS 309
>UniRef100_Q9SJZ8 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 321
Score = 68.6 bits (166), Expect = 6e-11
Identities = 49/143 (34%), Positives = 68/143 (47%), Gaps = 1/143 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W+ P G K N DG S A GGVLRD +G W+ GF + + LAELW +
Sbjct: 153 WVSPEDGWVKLNTDGASRGNPGFATAGGVLRDHNGAWIGGFAVNIGVCSAPLAELWGVYY 212
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A RG + +E DS +T+ P S ++R + G V + R
Sbjct: 213 GLFIAWGRGARRVELEVDSKMVVGFLTTGIADSHPLSFLLRLCYDFLSKGWIVRISHVYR 272
Query: 122 EANMVADCLAQNAHVFNFGVHLL 144
EAN +AD LA A + G+HLL
Sbjct: 273 EANRLADGLANYAFSLSLGLHLL 295
>UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 308
Score = 63.5 bits (153), Expect = 2e-09
Identities = 45/141 (31%), Positives = 63/141 (43%), Gaps = 1/141 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W+LP VG K N DG S A GGVLRDC+G W GF + + AELW +
Sbjct: 140 WVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGFSLNIGRCSAQHAELWGVYY 199
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ A ++ + +E DS + + P S +VR + V + R
Sbjct: 200 GLYFAWEKKVPRVELEVDSEAIVGFLKTGISDSHPLSFLVRLCHNFLQKDWLVRIVYVYR 259
Query: 122 EANMVADCLAQNAHVFNFGVH 142
EAN +AD LA + + G H
Sbjct: 260 EANCLADGLANYTILLSLGFH 280
>UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 63.5 bits (153), Expect = 2e-09
Identities = 51/161 (31%), Positives = 71/161 (43%), Gaps = 1/161 (0%)
Query: 1 LCWLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAI 59
+ W P G N DG S +A GGV+RD G+W+ GF + + LAELW +
Sbjct: 506 IAWRKPAEGWVTMNTDGASHGNPGQATAGGVIRDEHGSWLVGFALNIGVCSAPLAELWGV 565
Query: 60 RTAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLI 119
+ +A +RG + +E DSA + S P + +VR I V +
Sbjct: 566 YYGLVVAWERGWRRVRLEVDSALVVGFLQSGIGDSHPLAFLVRLCHGFISKDWIVRITHV 625
Query: 120 PREANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSM 160
REAN +AD LA A FG LL +L +D M
Sbjct: 626 YREANRLADGLANYAFTLPFGFLLLDSCPEHVSSILLEDVM 666
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 59.3 bits (142), Expect = 4e-08
Identities = 45/141 (31%), Positives = 61/141 (42%), Gaps = 1/141 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W+ P VG K N DG S A GGVLRDC G W GF + + AELW +
Sbjct: 561 WVSPCVGWVKVNTDGASRGNPGLASAGGVLRDCTGAWCGGFSLNIGRCSAPQAELWGVYY 620
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ A ++ + +E DS + + P S +VR + V + R
Sbjct: 621 GLYFAWEKKVPRVELEVDSEVIVGFLKTGISDSHPLSFLVRLCHGFLQKDWLVRIVHVYR 680
Query: 122 EANMVADCLAQNAHVFNFGVH 142
EAN +AD LA A + G H
Sbjct: 681 EANRLADGLANYAFSLSLGFH 701
>UniRef100_Q7X7I7 OSJNBa0093F12.7 protein [Oryza sativa]
Length = 127
Score = 58.5 bits (140), Expect = 6e-08
Identities = 40/127 (31%), Positives = 64/127 (49%), Gaps = 5/127 (3%)
Query: 12 KFNVDGSVWQTREAGCGGVLRDCDGNW---VQGFCRRL*SSNPLLAELWAIRTAVDLAVQ 68
K NVDGS + G G +LR+ G+ V GF + S + L AE+ A + + LA+Q
Sbjct: 2 KLNVDGSYQENNTGGIGAILRNSTGDVIFAVYGFVEQ--SQSALEAEILACKEGIRLALQ 59
Query: 69 RGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVAD 128
PIIIE+D AEA + S L ++ +VR I+ + ++ + R N ++
Sbjct: 60 WTLLPIIIESDCAEAIQMILSEGKLRSAHTFLVREIQGMVKGAREMKLKKVNRNCNRISH 119
Query: 129 CLAQNAH 135
LA ++
Sbjct: 120 VLANKSY 126
>UniRef100_Q9SI81 F23N19.5 [Arabidopsis thaliana]
Length = 233
Score = 57.4 bits (137), Expect = 1e-07
Identities = 47/161 (29%), Positives = 65/161 (40%), Gaps = 1/161 (0%)
Query: 3 WLLPPVGSWKFNVDG-SVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P +G K N DG S A GG LR+ G W GF + LAELW +
Sbjct: 65 WSKPSLGWCKLNTDGASHGNPGLATAGGALRNEYGEWCFGFALNIGRCLAPLAELWGVYY 124
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A RG + + +E DS + + P S +VR + V + R
Sbjct: 125 GLFMAWDRGITRLELEVDSEMVVGFLRTGIGSSHPLSFLVRMCHGFLSRDWIVRIGHVYR 184
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQDSMTV 162
EAN +AD LA A G H P +L D + +
Sbjct: 185 EANRLADGLANYAFDLPLGYHAFASPPNSLDSILRDDELGI 225
>UniRef100_Q942W6 P0506E04.5 protein [Oryza sativa]
Length = 325
Score = 55.8 bits (133), Expect = 4e-07
Identities = 36/121 (29%), Positives = 60/121 (48%), Gaps = 2/121 (1%)
Query: 3 WLLPPVGSWKFNVDGSVWQTR-EAGCGGVLRDCDGNWVQGFCRRL*S-SNPLLAELWAIR 60
W+ P G K NVDGS + + + G G VLRD GN + C L ++ L AE+ R
Sbjct: 153 WIRPQEGWMKLNVDGSYYPSDGKGGTGAVLRDSSGNLIFAACGVLHRPASALEAEMVDCR 212
Query: 61 TAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIP 120
+ +A+Q PII+ETD E + S + + + ++R ++ + ++V +
Sbjct: 213 EGISMALQWTLLPIIVETDCLEMVQLIHSDEKAMSDLAFLIREVKLLLKGNREIVIKKVS 272
Query: 121 R 121
R
Sbjct: 273 R 273
>UniRef100_O65407 Hypothetical protein F18E5.40 [Arabidopsis thaliana]
Length = 229
Score = 54.7 bits (130), Expect = 9e-07
Identities = 37/136 (27%), Positives = 65/136 (47%), Gaps = 1/136 (0%)
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTA 62
W P G K N DGS + +A GGV R+ ++ G+ + + +AE A++
Sbjct: 63 WKKPENGRIKLNFDGSRGREGQASIGGVFRNHKAEFLLGYSESIGEATSTMAEFAALKRG 122
Query: 63 VDLAVQRGCSPIIIETDSAEAHAAVTSAQPL-IPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
++LA++ G + + +E D+ ++ L ++ V +I+ +P V + R
Sbjct: 123 LELALENGLTDLWLEGDAKIIMDIISRRGRLRCEKTNKHVNYIKVVMPELNNCVLSHVYR 182
Query: 122 EANMVADCLAQNAHVF 137
E N VAD LA+ H F
Sbjct: 183 EGNRVADKLAKLGHQF 198
>UniRef100_Q9FXC8 F12A21.24 [Arabidopsis thaliana]
Length = 803
Score = 54.7 bits (130), Expect = 9e-07
Identities = 41/136 (30%), Positives = 60/136 (43%)
Query: 25 AGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTAVDLAVQRGCSPIIIETDSAEAH 84
A GGV+RD GNW GF + + LAELW + LA + + + +E DS
Sbjct: 658 ATAGGVIRDGAGNWCGGFALNIGRCSAPLAELWGVYYGFYLAWTKALTRVELEVDSELVV 717
Query: 85 AAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPREANMVADCLAQNAHVFNFGVHLL 144
+ + P S +VR + V + REAN +AD LA A G H L
Sbjct: 718 GFLKTGIGDQHPLSFLVRLCHGLLSKDWIVRITHVYREANRLADGLANYAFSLPLGFHSL 777
Query: 145 HHPLMECRYLLYQDSM 160
+ +L++DS+
Sbjct: 778 IDVPDDLEVILHEDSL 793
>UniRef100_Q8S5K1 Putative retroelement [Oryza sativa]
Length = 1888
Score = 53.9 bits (128), Expect = 2e-06
Identities = 42/131 (32%), Positives = 59/131 (44%), Gaps = 2/131 (1%)
Query: 3 WLLPPVGSWKFNVDGSVWQTR-EAGCGGVLRDCDGNWVQGFCRRL*S-SNPLLAELWAIR 60
WL P GS K NVDGS ++ + G G VLR+C G + C + S+ L EL A R
Sbjct: 1728 WLKPLPGSMKLNVDGSFQESDGKGGIGAVLRNCTGEVIFAACGHVDCCSSALETELLACR 1787
Query: 61 TAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIP 120
+ LA+Q PI+IETD A + ++R I + + F
Sbjct: 1788 DGLALALQWTLLPIVIETDCLAMIHLFRDATGAKSELAFLIREIDSLLVGNRDISFSKCL 1847
Query: 121 REANMVADCLA 131
R N ++ LA
Sbjct: 1848 RSQNHISHYLA 1858
>UniRef100_Q655G6 Hypothetical protein P0009H10.11 [Oryza sativa]
Length = 240
Score = 53.1 bits (126), Expect = 3e-06
Identities = 44/140 (31%), Positives = 62/140 (43%), Gaps = 12/140 (8%)
Query: 3 WLLPPVGSWKFNVDGSV-WQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P G K N DGS T A GGV RD DG ++ G+ R+ ++ +AEL A+R
Sbjct: 79 WRKPEEGWMKLNFDGSSKHSTGIASIGGVYRDHDGAFLLGYAERIGTATSSVAELAALRR 138
Query: 62 AVDLAVQRGCSPIIIETDSAEA------HAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVV 115
++LAV+ G + E DS A V S + L + R I +P +
Sbjct: 139 GLELAVRNGWRRVWAEGDSKAVVDVVCDRADVQSEEDL-----RLCREIAALLPQLDDMA 193
Query: 116 FDLIPREANMVADCLAQNAH 135
+ R N VA A+ H
Sbjct: 194 VSHVRRGGNKVAHGFAELGH 213
>UniRef100_Q6L3X8 Hypothetical protein [Solanum demissum]
Length = 1155
Score = 53.1 bits (126), Expect = 3e-06
Identities = 43/130 (33%), Positives = 61/130 (46%), Gaps = 4/130 (3%)
Query: 3 WLLPPVGSWKFNVDGSVWQTREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRTA 62
W+ PP K N DGS R G GG+LR+ G + F +L AE ++
Sbjct: 992 WIKPPTMFAKLNTDGSCVNGR-CGGGGILRNALGQVIMAFTIKLGEGTSSWAEAMSMLHG 1050
Query: 63 VDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSI-PHGMKVVFDLIPR 121
+ L +QRG + II ETDS A+T + V+ I++ + HG + + R
Sbjct: 1051 MQLCIQRGVNMIIGETDSILLAKAITENWSIPWRMYIPVKKIQKMVEEHGF--IINHCLR 1108
Query: 122 EANMVADCLA 131
EAN AD LA
Sbjct: 1109 EANQPADKLA 1118
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 52.8 bits (125), Expect = 3e-06
Identities = 40/134 (29%), Positives = 60/134 (43%), Gaps = 2/134 (1%)
Query: 3 WLLPPVGSWKFNVDGSV-WQTREAGCGGVLRDCDGNWVQGFCRRL*S-SNPLLAELWAIR 60
W P G K NVDGS + + G G +LR+C GN + CR L S S PL AEL A
Sbjct: 1777 WERPRNGWMKLNVDGSFDINSEKGGIGMILRNCLGNVIFSSCRSLDSCSGPLEAELHACV 1836
Query: 61 TAVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIP 120
+ LA+ PI +ETD + + + I + + + ++ +
Sbjct: 1837 EGLHLALHWTLLPIQVETDCSSVIQLLNHPDKDRSVLANIAQEAKSLMAGDRQIAISKVQ 1896
Query: 121 REANMVADCLAQNA 134
R N+++ LA A
Sbjct: 1897 RSQNVISHFLANKA 1910
>UniRef100_Q7XGD4 Putative reverse transcriptase [Oryza sativa]
Length = 230
Score = 52.8 bits (125), Expect = 3e-06
Identities = 34/79 (43%), Positives = 44/79 (55%), Gaps = 2/79 (2%)
Query: 3 WLLPPVGSWKFNVDGSVWQTR-EAGCGGVLRDCDGNWVQGFCRRL*S-SNPLLAELWAIR 60
WL P GS K NVDGS ++ + G G VLR+C G + C + S+ L EL A R
Sbjct: 112 WLKPLPGSMKLNVDGSFQESDGKGGIGAVLRNCTGEVIFAACGHVDCCSSALETELLACR 171
Query: 61 TAVDLAVQRGCSPIIIETD 79
+ LA+Q PI+IETD
Sbjct: 172 DGLALALQWTLLPIVIETD 190
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 52.4 bits (124), Expect = 5e-06
Identities = 43/157 (27%), Positives = 63/157 (39%), Gaps = 1/157 (0%)
Query: 3 WLLPPVGSWKFNVDGSVWQTRE-AGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W +P G K DG+ A GG +R+ G W+ GF + S LAELW
Sbjct: 1063 WQVPSDGWVKITTDGASRGNHGLAAAGGAIRNGQGEWLGGFALNIGSCAAPLAELWGAYY 1122
Query: 62 AVDLAVQRGCSPIIIETDSAEAHAAVTSAQPLIPPYSEIVRHIRRSIPHGMKVVFDLIPR 121
+ +A +G + ++ D +++ P S +VR + V + R
Sbjct: 1123 GLLIAWDKGFRRVELDLDCKLVVGFLSTGVSNAHPLSFLVRLCQGFFTRDWLVRVSHVYR 1182
Query: 122 EANMVADCLAQNAHVFNFGVHLLHHPLMECRYLLYQD 158
EAN +AD LA A G+H R LL D
Sbjct: 1183 EANRLADGLANYAFTLPLGLHCFDACPEGVRLLLLAD 1219
>UniRef100_Q5W657 Hypothetical protein OSJNBb0052F16.4 [Oryza sativa]
Length = 220
Score = 50.8 bits (120), Expect = 1e-05
Identities = 43/139 (30%), Positives = 62/139 (43%), Gaps = 10/139 (7%)
Query: 3 WLLPPVGSWKFNVDGSVWQ--TREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIR 60
W PPVG K N DGSV+ +R A GGV+R CDG V F + E A+
Sbjct: 49 WAPPPVGWCKLNFDGSVFNDGSRRASIGGVIRGCDGGVVLAFAETTEHWTVGVVEARALI 108
Query: 61 TAVDLAVQRGCSPIIIETDSAEAHAAV--TSAQPLIPP--YSEIVRHIRRSIPHGMKVVF 116
+ LA+ I++E D + Q IP + EI+ +RR ++ ++
Sbjct: 109 KGLKLALACFVERIVVEGDDLVLVQLLRGEETQTRIPAAMHDEILSLLRRFTEFEVRHIY 168
Query: 117 DLIPREANMVADCLAQNAH 135
RE N VA L + A+
Sbjct: 169 ----REGNSVAHTLCRQAY 183
>UniRef100_Q6K204 Hypothetical protein B1469H02.35 [Oryza sativa]
Length = 212
Score = 49.3 bits (116), Expect = 4e-05
Identities = 30/79 (37%), Positives = 42/79 (52%), Gaps = 1/79 (1%)
Query: 3 WLLPPVGSWKFNVDGSVWQ-TREAGCGGVLRDCDGNWVQGFCRRL*SSNPLLAELWAIRT 61
W P G K N DGS T+ A GGV RD +G +V G+ R+ + +AEL A+R
Sbjct: 51 WRKPEAGWIKLNFDGSSKHATKIASIGGVYRDHEGAFVLGYAERIGRATSSVAELAALRR 110
Query: 62 AVDLAVQRGCSPIIIETDS 80
++L V+ G + E DS
Sbjct: 111 GLELVVRNGWRRVWAEGDS 129
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.328 0.141 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 283,744,761
Number of Sequences: 2790947
Number of extensions: 10235052
Number of successful extensions: 24850
Number of sequences better than 10.0: 166
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 137
Number of HSP's that attempted gapping in prelim test: 24760
Number of HSP's gapped (non-prelim): 169
length of query: 169
length of database: 848,049,833
effective HSP length: 118
effective length of query: 51
effective length of database: 518,718,087
effective search space: 26454622437
effective search space used: 26454622437
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0108.3


