

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0108.13
(1262 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
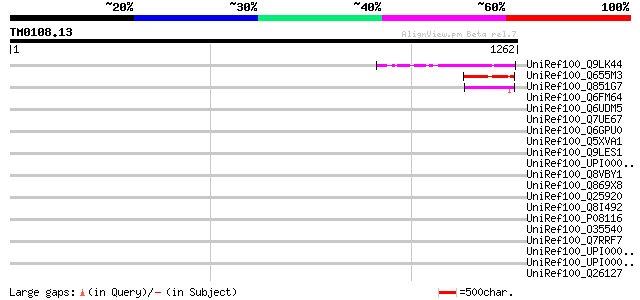
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LK44 Emb|CAB10191.1 [Arabidopsis thaliana] 152 8e-35
UniRef100_Q655M3 Hypothetical protein P0672D08.25 [Oryza sativa] 111 2e-22
UniRef100_Q851G7 Hypothetical protein OSJNBb0042N11.21 [Oryza sa... 67 3e-09
UniRef100_Q6FM64 Similar to sp|P43597 Saccharomyces cerevisiae Y... 46 0.008
UniRef100_Q6UDM5 Starmaker [Brachydanio rerio] 45 0.011
UniRef100_Q7UE67 Hypothetical protein [Rhodopirellula baltica] 43 0.054
UniRef100_Q6GPU0 LOC443607 protein [Xenopus laevis] 42 0.16
UniRef100_Q5XVA1 Hypothetical protein [Arabidopsis thaliana] 42 0.16
UniRef100_Q9LES1 Hypothetical protein T8M16_200 [Arabidopsis tha... 42 0.16
UniRef100_UPI000036634F UPI000036634F UniRef100 entry 41 0.20
UniRef100_Q8VBY1 Phosphophoryn precursor [Rattus norvegicus] 41 0.20
UniRef100_Q869X8 Similar to Homo sapiens (Human). dentin sialoph... 41 0.20
UniRef100_Q25920 Mature-parasite-infected erythrocyte surface an... 41 0.20
UniRef100_Q8I492 Phoshoprotein 300 [Plasmodium falciparum] 41 0.20
UniRef100_P08116 Processed variable antigen [Plasmodium falciparum] 41 0.20
UniRef100_O35540 Hepatoma-derived growth factor [Mus musculus] 40 0.35
UniRef100_Q7RRF7 Chloroquine resistance marker protein, putative... 40 0.35
UniRef100_UPI000032E5FC UPI000032E5FC UniRef100 entry 40 0.46
UniRef100_UPI000036F6AB UPI000036F6AB UniRef100 entry 40 0.59
UniRef100_Q26127 Microneme protein-1 [Plasmodium vivax] 40 0.59
>UniRef100_Q9LK44 Emb|CAB10191.1 [Arabidopsis thaliana]
Length = 566
Score = 152 bits (383), Expect = 8e-35
Identities = 117/350 (33%), Positives = 174/350 (49%), Gaps = 42/350 (12%)
Query: 914 RIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISDSDLRATS 973
R K K GS Y+R+LP L + ++ N P QK ++ IS L + S
Sbjct: 240 RGKLFKTPGSVNYRRMLPYLKDIQEDN---------PYPQKNTEEV----ISSPMLNSES 286
Query: 974 GGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGFQSSNSKCKVMQMCDEQ 1033
+ V + T SGT + ++ P R P + K + +
Sbjct: 287 DNEGTQEVVTSNVTRESGTSSDE--------NEEPLPCERVPVNLEQSDPDKEQETQIKH 338
Query: 1034 VLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVNEVMAREEATFALSESLINSKVKGNCS 1093
V+ + E++ G+ I + P+V + E + AL + +++ V G +
Sbjct: 339 VIPD----TENNLGSEIPLSS---------PLVGSRSSSEVNSSALHNTFVDNLV-GEEN 384
Query: 1094 FSISPDDKEKLS----EAHGCCLSLSQLELKEKLEVPA-VDFKKGILKRNPRGCRGLCTC 1148
+ + + K+S EAH + ++ L P+ KGILKR+ RGCRG+C+C
Sbjct: 385 MNGAEITEAKISAEELEAHSSDATAELVDPSVILATPSSFSPSKGILKRSMRGCRGICSC 444
Query: 1149 LNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPIDSVNDNPGFDRDQV 1208
LNC+SFRLHAERAFE+SRNQL D E + DL+ E+SHLRD+L + + + P + Q
Sbjct: 445 LNCSSFRLHAERAFEFSRNQLQDTEVMVLDLVGEISHLRDLLEKYNSADHSEP--YKSQA 502
Query: 1209 KEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFADHVQEKII 1258
EA ++A A +LAK RL M DDL IH RI N QR RV+FA ++ EK I
Sbjct: 503 GEASKRACEAAELAKSRLHQMNDDLQIHYRIPNEQRARVKFAHYIHEKTI 552
Score = 37.4 bits (85), Expect = 2.9
Identities = 44/155 (28%), Positives = 63/155 (40%), Gaps = 54/155 (34%)
Query: 492 NDDFILTTPPDAKI------YDNSAVNVDRS----KSV--------------PQDMKRLK 527
+DDF TTPPD+++ + S VN + KSV P KRL
Sbjct: 113 SDDFAQTTPPDSELLAISEEINGSVVNKSDTNLWRKSVLLPCSRPKIFKNTGPFSYKRLL 172
Query: 528 PMDLRTNTQESFSKGVCLKADS------------------------RKDSLLKSKSVRRP 563
P ++ + + S C K+ S R S LKS P
Sbjct: 173 PYLMQASDDGTSSSSRCSKSLSQNITKPVSQSMDSVYDKDSTGSFCRDTSPLKSVIASTP 232
Query: 564 H-----LPGKLFKTLGSVS-KRFLPFLMELAKDDP 592
+ GKLFKT GSV+ +R LP+L ++ +D+P
Sbjct: 233 NKNAAFSRGKLFKTPGSVNYRRMLPYLKDIQEDNP 267
>UniRef100_Q655M3 Hypothetical protein P0672D08.25 [Oryza sativa]
Length = 676
Score = 111 bits (277), Expect = 2e-22
Identities = 54/127 (42%), Positives = 87/127 (67%), Gaps = 11/127 (8%)
Query: 1131 KKGILKRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDML 1190
KKGILKR+ RGC+G+C CL+C++FRL A+RAFE+SR Q+ +A+++ +LLKE+S LR+++
Sbjct: 548 KKGILKRHTRGCKGICMCLDCSTFRLRADRAFEFSRKQMQEADDIIDNLLKEVSSLRNLM 607
Query: 1191 GRPIDSVNDNPGFDRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQRPRVRFA 1250
+ ++ + AC++A E +A+ER M +L+ HCRI PRV+FA
Sbjct: 608 --------EKSAGQQETKQTACQRASQVEVVARERRRQMLMELNSHCRIPG---PRVKFA 656
Query: 1251 DHVQEKI 1257
+V+E++
Sbjct: 657 QYVEERM 663
>UniRef100_Q851G7 Hypothetical protein OSJNBb0042N11.21 [Oryza sativa]
Length = 863
Score = 67.0 bits (162), Expect = 3e-09
Identities = 42/143 (29%), Positives = 74/143 (51%), Gaps = 17/143 (11%)
Query: 1132 KGILK--RNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDM 1189
KGILK +P + CTC+ AS L AE+A E+S+ Q+ D E +A L++ L+H+R +
Sbjct: 701 KGILKGTESPSPQKTTCTCMKAASVILDAEKAVEFSQRQMHDIENIASKLMRSLNHMRSI 760
Query: 1190 LGRPIDSVNDN--PGFDRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNL----- 1242
+ + S + + P F+ +++ A A E+ ++ L++M D + C+I L
Sbjct: 761 VDGNLLSESHSLLPTFNTAEIRAASEDALEVERTTRKWLTIMNKDCNRFCKILRLAGKKA 820
Query: 1243 --------QRPRVRFADHVQEKI 1257
+R ++ FAD K+
Sbjct: 821 VSHSEVPRKRKKITFADETGGKL 843
>UniRef100_Q6FM64 Similar to sp|P43597 Saccharomyces cerevisiae YFR016c [Candida
glabrata]
Length = 1012
Score = 45.8 bits (107), Expect = 0.008
Identities = 113/539 (20%), Positives = 195/539 (35%), Gaps = 76/539 (14%)
Query: 389 KSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSG----------------- 431
K+ES + + S S +L ++ N+ V GE S+ G
Sbjct: 256 KNESISNELATEKEQSPSTAVLSDDLKKNQDNVQGEKQSIEKGSKADADILIKPETNSKL 315
Query: 432 --DDKCFNESKCDPAADSD-GVKVVGNALHNDDFMQCSIHKNNLDYAKVT----NDSGCT 484
D NES D + SD +K +G + Q + ++ + +T N+
Sbjct: 316 VKDAVIANESDADKKSISDTTIKEIGESNAEIKEKQTTQSRSEKEIISLTAEENNNKVTI 375
Query: 485 SVQLGVLNDDFILTTPPDAKIYDNSAVNVDRSKSVPQDMKRLKPMDLRT---NTQESFSK 541
S + LNDD + P N+D KSV Q+ K + ++ +E +K
Sbjct: 376 SDTIPDLNDDTMKAASP----------NIDEIKSVQQESYDSKSKETKSEPEEQEEDMNK 425
Query: 542 GVCLKAD--SRKDSLLKSKSVRRPHLPGK------------------LFKTLGSVSKRFL 581
G + S S K ++++ P L + L + L +VS
Sbjct: 426 GQLEPENKVSSSPSSSKKQALKEPALESETEQEHEGNLLPTGNKTDVLEENLETVS-AVK 484
Query: 582 PFLMELAKDDPGISKFGHG-TPKKDEADMHGKRF-ELTLSSQPEEASIDGHKIE-----K 634
P L ++ KD P HG T E K EL+ + P++ S+D K E
Sbjct: 485 PSLADVNKDLPLKEGVNHGKTVDVQELHTTSKPIEELSQAESPKKTSLDAMKKEDKLKDS 544
Query: 635 GPIHGTVESNALGNILFNSANELSNGNLPKLTSSQDLPELPMQSDVKEVVQECLSAPSVE 694
T+E++ I S E + P S+DL + P +S E V + +V
Sbjct: 545 RKTESTIEAHLDNRINEKSEAEEVGDDGPVDKKSEDLKDTPEKSLSTEAVTQSPKVHNVN 604
Query: 695 EPIEKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSLQERGPS 754
E V +D+ + D A + + + D + N +
Sbjct: 605 E--VSKVNEEEDKAAESLTVDKAGAENSHKPSNLKVLEAKDT-LSDTMNETKQDETMDKE 661
Query: 755 PKDQSMLYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQL 814
PK + L + ++++ E S+ S E ++ + ++ Q+ S L
Sbjct: 662 PKSTAKL---IKLNDEVDEEASTLADDVSDLTEEKLKVKDNVMHTTAEIPNNDAQTDSDL 718
Query: 815 DTDMQDLEE-NERSNG--AIRKSDDAVVLNHVGVSNEVPRPPPENIISKTQDMAAHGGN 870
+ +DL E NE N ++K D V+ + +NE P +IS D+ + N
Sbjct: 719 PKEKEDLNEFNEDKNNEDPVKKGDGDVLSKIMTENNETPE--KRVVISILDDLPSRHDN 775
>UniRef100_Q6UDM5 Starmaker [Brachydanio rerio]
Length = 613
Score = 45.4 bits (106), Expect = 0.011
Identities = 90/421 (21%), Positives = 145/421 (34%), Gaps = 53/421 (12%)
Query: 69 SSGNDEKREPPEED---DGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQSSA 125
S +K E P+ D DGD+ + D+D KD + + S
Sbjct: 184 SPDTTDKPEGPDSDSAPDGDSASAEKTDSDHSPDEDANKSSTEADKDDTSDKDSSQTDEK 243
Query: 126 RASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSSAE 185
DSD +D+ S E+ S +D + + +D+ +D G S A+
Sbjct: 244 H------DSDASDKDEKHEDKDEKSDEKDSSKDSEDK----SQEKSDK--SDDGSNSEAD 291
Query: 186 KGGLCVD--DATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDE 243
+ V+ D S + D D+ ++DS+D K +E D D S + + D
Sbjct: 292 EQKESVESKDHDSDSQDSDSAEKKEKHDDKDQDSSDSADSKDSDEDKDKDHSEQKDSEDH 351
Query: 244 EIVVTTPPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLK 303
E D E D DEGK +ED D + E S++ SK ++S +
Sbjct: 352 EHKEKHTKDKE----EHKDSDEGKDDEDKSKSD------EHDKDESESKEASKSDESEQE 401
Query: 304 SKYFSTAANHSFDLDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKD 363
K + + + D ++ + S + D+ + H D D
Sbjct: 402 EKKDDKSDSDNSSRDSHSDSDSD-------------------SHSDSDSDSHSDSHSDSD 442
Query: 364 EKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSG 423
D S S+ S+ + S +E K S S++ S + S
Sbjct: 443 SDSHSDSDSDSDSDSDSDSDSDSDSNSRD---KEEKKDKSSESRDEDSSDSDSKSNSESS 499
Query: 424 ECLSVPSGDDKCFNESKCDPAADSDGVKVVGNAL---HNDDFMQCSIHKNNLDYAKVTND 480
E + DDK + K D SD V N H+DD + + K D
Sbjct: 500 ETAEEDTNDDKDSSVEK-DKTDSSDSASVEANDSDDEHDDDSKDATPSSEDHTAEKTDED 558
Query: 481 S 481
S
Sbjct: 559 S 559
>UniRef100_Q7UE67 Hypothetical protein [Rhodopirellula baltica]
Length = 669
Score = 43.1 bits (100), Expect = 0.054
Identities = 87/471 (18%), Positives = 153/471 (32%), Gaps = 31/471 (6%)
Query: 5 PTLRESASAMEETLISRKRSNPSSSPPGALTRSRSQLFVHRNRSGQRRPD-SGPRQGGLY 63
P S+S+ ++ S S+PSSS + S S + SG S L
Sbjct: 11 PVAGSSSSSSSDSSSSSSSSSPSSSSSSGSSSSSSGSSSSSSMSGSSSSSMSSSSSSSLS 70
Query: 64 SGLGGSSGNDEKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQS 123
S SSG+ + + G + ++ S S
Sbjct: 71 SSSSSSSGSSSSSSGSSSSSSGSPSSSSSSGSSSSSSSG-------SSSSSSKSSSSSDS 123
Query: 124 SARASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSS 183
S+ +S SD +D + S S + S V + +SD +D +
Sbjct: 124 SSNSS----SSDSSDSSHSSSASSSSDSSDSSTSSSSSVSSSQSSDSSDSSDSSTSTSLP 179
Query: 184 AEKGGLCVDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKINDFDE 243
+ D ++ + S D SS ++ D +N+DSSD + + SG + D
Sbjct: 180 SVSSSSSSDGSSSSSSGSSD--SSSNSQSSDSSNSDSSDSS--DSTSLPSISGS-SSSDG 234
Query: 244 EIVVTTPPDAELLGDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLK 303
+ D+ S + + + +G + ++G + DS + S +
Sbjct: 235 SSASSDSSDSSSNSQSSDSSNSESSDSSDSISLPSISGSSSSAGSSSASSDSSNSSSNSQ 294
Query: 304 SKYFSTAANH----SFDLDLYAG----------ARSQTAFPGQIVQGPGFEACGGTNWPD 349
S ST+ ++ S L +G + SQ++ G ++ + P
Sbjct: 295 SSDSSTSESNDSSASVSLPSVSGSSSSASDSSSSESQSSDSGSSSDSSTSQSSESASTPS 354
Query: 350 NFVGTKKLGHCDKDEKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNL 409
+ SS SE S E SES N S S S +
Sbjct: 355 VSSSQSSTSESASSTSESSNSESSQSESSTSESRSRSESDSESLPSYSWNSSTSSDSDSE 414
Query: 410 LESPMELNEREVSGECLSVPSGDDKCFNESKCDPAADSDGVKVVGNALHND 460
++ + SV +ES D ++DS +A ++D
Sbjct: 415 SSDSESESQSQSDSTSQSVSDSGSDSQSESSIDRSSDSQSYSASASASYSD 465
>UniRef100_Q6GPU0 LOC443607 protein [Xenopus laevis]
Length = 325
Score = 41.6 bits (96), Expect = 0.16
Identities = 57/254 (22%), Positives = 93/254 (36%), Gaps = 30/254 (11%)
Query: 203 DLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKIN-----DFDEEIVVTTPPDAELLG 257
DL S + ++ +DS E GNDG S + D D+E P + EL G
Sbjct: 21 DLFGSDAESENEQKESDSGSGSDSEPGNDGSGSNRSGSDSDRDEDDEHSAVKPSNKELFG 80
Query: 258 DSKVD--GDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSF 315
D D G + G + + ++ N+ E + D + ND ++ + A H
Sbjct: 81 DDSEDEVGSQHSGSYNR-------SERSYNASE-TAHSDQEVNDQSDAEQHSGSEALHDE 132
Query: 316 DLDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLP 375
D D G RS + P G +A N D G + +E+ M D +L
Sbjct: 133 DNDEDVGQRSDHSSPRSEADGSDKDA----NSDDEKWGREDKSDQSDEEEKMQNSDDELQ 188
Query: 376 LSSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSGDDKC 435
S + E E G + SS E+ +E + + + + DD+
Sbjct: 189 HSDRDEHRQSDE-----------EGQRPSSDAEEQETHQSDDEEQDNHHSDDLHNSDDEE 237
Query: 436 FNESKCDPAADSDG 449
++ + SDG
Sbjct: 238 NDQQHYNDRPHSDG 251
>UniRef100_Q5XVA1 Hypothetical protein [Arabidopsis thaliana]
Length = 700
Score = 41.6 bits (96), Expect = 0.16
Identities = 30/128 (23%), Positives = 61/128 (47%), Gaps = 6/128 (4%)
Query: 1136 KRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPID 1195
+RN + + S + +++A +S+ Q+ D + VA L KEL +R + R +
Sbjct: 540 RRNTQAAKAQPFSTGGTSIQGCSQKAIAFSQGQMRDFQNVAARLTKELKSMRQITKRCLQ 599
Query: 1196 SVNDNPGF---DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQR---PRVRF 1249
+ ++ + D+VK A E+ K+ LS+++ D + C++ ++ R P
Sbjct: 600 AESNTSNMSDCNLDEVKTVIGNAEKTEESCKKWLSIIERDCNRFCKLMSMVREDSPATEN 659
Query: 1250 ADHVQEKI 1257
H ++KI
Sbjct: 660 IVHKKKKI 667
>UniRef100_Q9LES1 Hypothetical protein T8M16_200 [Arabidopsis thaliana]
Length = 670
Score = 41.6 bits (96), Expect = 0.16
Identities = 30/128 (23%), Positives = 61/128 (47%), Gaps = 6/128 (4%)
Query: 1136 KRNPRGCRGLCTCLNCASFRLHAERAFEYSRNQLLDAEEVAQDLLKELSHLRDMLGRPID 1195
+RN + + S + +++A +S+ Q+ D + VA L KEL +R + R +
Sbjct: 510 RRNTQAAKAQPFSTGGTSIQGCSQKAIAFSQGQMRDFQNVAARLTKELKSMRQITKRCLQ 569
Query: 1196 SVNDNPGF---DRDQVKEACRKAYAAEQLAKERLSMMKDDLSIHCRITNLQR---PRVRF 1249
+ ++ + D+VK A E+ K+ LS+++ D + C++ ++ R P
Sbjct: 570 AESNTSNMSDCNLDEVKTVIGNAEKTEESCKKWLSIIERDCNRFCKLMSMVREDSPATEN 629
Query: 1250 ADHVQEKI 1257
H ++KI
Sbjct: 630 IVHKKKKI 637
>UniRef100_UPI000036634F UPI000036634F UniRef100 entry
Length = 369
Score = 41.2 bits (95), Expect = 0.20
Identities = 80/410 (19%), Positives = 144/410 (34%), Gaps = 60/410 (14%)
Query: 660 GNLPKLTSSQDLPELPMQSDVKEVVQECLSAPSVEEPIEKAVVGSKDECSREPNFDLCSA 719
G+L TS Q L + + + + + + + P V +P + G K EC N D S
Sbjct: 13 GSLEVCTSDQQLDQ-QLDQQLDQQLDQPVCGPPVPQPAPLSA-GIKRECVINSNSDNNSG 70
Query: 720 IVDFHSTKIVANDGHDVDIKHMQNSISSLQERGPSPKDQSMLYVNVGVSNQTLEHHSSEV 779
+ S V N ++ + K +S SS + S N ++N++ +HS+
Sbjct: 71 NNNSSSNNSVNNSNNNNNSKKSSSSNSSSSNNSNNSSSNSSS--NNSINNRSNNNHSNSN 128
Query: 780 KGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQLDTDMQDLEENERSNGAIRKSDDAVV 839
S NN S + S++ + + + N SN + S++
Sbjct: 129 NNRSNNNH--------------SNSNNNSSSNNHSNNNSSNNSNNNSSNNSNNNSNNNSS 174
Query: 840 LNHVGVSNEVPRPPPENIISKTQDMAAHGGNKEGSVQKGIAYGSKGN--NGRKGCYAYEI 897
NH N K+ H N + + S N N R
Sbjct: 175 NNH-----------SNNNSKKSSSSNNHSNNNSRNNNNSSSNNSNNNSNNNRSNNNNNSS 223
Query: 898 ENASESKITPVLNRCPRIKSLKHAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLD 957
N+S +K T N + + ++G+ +N+KK ++ S + + K +
Sbjct: 224 NNSSSNKNT---NNSGKSNNSNNSGNNNTS------NNSKKSSNNNSSSNNSKKSSNNSN 274
Query: 958 QTPLLPISDSDLRATSGGDSNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGF 1017
S S+ + + +N++ + NS + N N D+SS S S
Sbjct: 275 NN-----SSSNNHSNNNSSNNNNNSSSN---NSNNNNRSNNNNSSNNRDNSSNNSNSNNN 326
Query: 1018 QSSNSKCKVMQMCDEQVLLNGLCKPESSSGTPISVRRIDSPSTPLRPIVN 1067
S+NS + P S SGT ++ P P + ++
Sbjct: 327 SSNNSNNNIFP------------PPCSISGTAAAMETAAPPPLPRQQAIS 364
>UniRef100_Q8VBY1 Phosphophoryn precursor [Rattus norvegicus]
Length = 970
Score = 41.2 bits (95), Expect = 0.20
Identities = 83/444 (18%), Positives = 145/444 (31%), Gaps = 38/444 (8%)
Query: 70 SGNDEKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSPQSSARASL 129
+G+D E+D D D +G D KD + S + +
Sbjct: 489 NGSDSDSHAGEDDSSDDT-----SDTDDSDSNGDDDSESKDKDESDNSNHDNDSDSESKS 543
Query: 130 DLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGSSAEKGGL 189
D DSD + S SSE D +SD +D + SS
Sbjct: 544 DSSDSDSDSSDSSDSSDSSDSSESSDSSDSSD-----SSDSSDSSDSSDSSDSSDSSDSD 598
Query: 190 CVDDATMKVTVSPDLGISSQARDF----------DRANTDSSDHKQFEEGNDGDCSGKIN 239
D + + S D SS + D D +N+DSSD + +D S +
Sbjct: 599 SSDSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSDSSDSSDSSDSSDSSDSSDSS 658
Query: 240 DFDEEIVVTTPPDAELLGDS-----KVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKD 294
D + + D+ DS D D + D+ + + +S + D
Sbjct: 659 DSSDSSDSSDSSDSSESSDSSDSSDSSDSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD 718
Query: 295 SKRNDSVLKSKYFSTAANHSFDLDLYAGA-----RSQTAFPGQIVQGPGFEACGGTNWPD 349
S +DS S+ ++ S D + + S ++ ++ ++ D
Sbjct: 719 SSDSDSSDSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSDSSDSSDSSDSSDSSD 778
Query: 350 NFVGTKKLGHCDKDEKGMGGQDFQLPLSSQSEKPSEPELKSESFTMRETNGSKLSSSQNL 409
+ + D + SS S S+ S+S +++ S SS N
Sbjct: 779 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNS 838
Query: 410 LESPMELNEREVSGECLSVPSGD--------DKCFNESKCDPAADSDGVKVVGNALHNDD 461
+S + + S S S D D + D + SDG G++ +D
Sbjct: 839 SDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDGDSSDGDSSDSDS 898
Query: 462 FMQCSIHKNNLDYAKVTNDSGCTS 485
S + ++ D + ++ S S
Sbjct: 899 SDSDSSNSSDSDSSDSSDSSSSDS 922
>UniRef100_Q869X8 Similar to Homo sapiens (Human). dentin sialophosphoprotein
[Dictyostelium discoideum]
Length = 1206
Score = 41.2 bits (95), Expect = 0.20
Identities = 78/372 (20%), Positives = 151/372 (39%), Gaps = 39/372 (10%)
Query: 698 EKAVVGSKDECSREPNFDLCSAIVDFHSTKIVANDGHDVDIKHMQNSISSLQERGPSPKD 757
E + GS +E S + + S H+ +N G + + H N SS S +
Sbjct: 666 ESSNQGSNNESSNNESSNQGSNNESSHNE--TSNQGSNNESSH--NESSSQGSNNESAHN 721
Query: 758 QSMLYVNVGVSNQTLEHHSSEVKGFSVNNGEVKQLLNMEKRKYVSRYPSEGQSSSQLDTD 817
+S N G +N++ H+ S +G S N N S S QSS+ L ++
Sbjct: 722 ESS---NQGSNNES-SHNESSSQG-SNNESSNNDSNNQNSNNEPSNNESSNQSSNNLSSN 776
Query: 818 MQDLEENERSNGAIRKSDDAVVLNHVG----VSNEVPRPPPENIISKTQ----------- 862
+ + G+ +S + N+ +++E P + S++
Sbjct: 777 NESSNNESSNQGSNNESSHSESSNNESSNQSLNSEYNNEPSQGSNSESSNESSNQSLNSE 836
Query: 863 ---DMAAHGGNKEGSVQKGIAYGSKGNNGRKGCYAYEIENASESKITPVLNRCPRIKSLK 919
D ++HG N E S + ++GS + + + N S + + + + +
Sbjct: 837 SNNDSSSHGSNSESSNNESSSHGSNNESSNNESSSQDSSNYSSNNGSSNNESSSQGSNNE 896
Query: 920 HAGSPTYKRLLPLLSNAKKYNSCASVNRHDPKHQKFLDQTPLLPISD---SDLRATSGGD 976
+ + S+++ N+ +S N +P +Q +++ S+ S+ ++S G
Sbjct: 897 SSNQGSNNESSNNESSSQGSNNESSHN--EPSNQSSNNESSNNESSNNESSNNESSSKGS 954
Query: 977 SNDSVPMEHFTANSGTKQQKELNACELNDDSSSPRSRSPGF--QSSNSKCKVMQMCDEQV 1034
+N+S E ++N G+ + N E N++SS+ S S G +SSN++ + Q
Sbjct: 955 NNESSNNE--SSNQGSNNESSNN--ESNNESSNNESSSQGSNNESSNNESSNNE-SSSQS 1009
Query: 1035 LLNGLCKPESSS 1046
NG ESS+
Sbjct: 1010 SNNGSSNNESSN 1021
>UniRef100_Q25920 Mature-parasite-infected erythrocyte surface antigen [Plasmodium
falciparum]
Length = 1510
Score = 41.2 bits (95), Expect = 0.20
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 268 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 327
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 328 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 385
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 386 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 425
>UniRef100_Q8I492 Phoshoprotein 300 [Plasmodium falciparum]
Length = 1434
Score = 41.2 bits (95), Expect = 0.20
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 265 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 324
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 325 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 382
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 383 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 422
>UniRef100_P08116 Processed variable antigen [Plasmodium falciparum]
Length = 224
Score = 41.2 bits (95), Expect = 0.20
Identities = 41/168 (24%), Positives = 72/168 (42%), Gaps = 10/168 (5%)
Query: 257 GDSKVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFD 316
G+SK G+ + E ++ TG+++ +GE +SK ++KY +
Sbjct: 63 GESKETGESKETGESKETGESKETGESKETGESKETGESKETRIYEETKYNKITSEFRET 122
Query: 317 LDLYAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPL 376
++ S+ G V GP +E +N TKKL +++E G++
Sbjct: 123 ENVKITEESKDR-EGNKVSGP-YENSENSNVTSESEETKKLAEKEENEGEKLGENVNDGA 180
Query: 377 SSQSEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGE 424
S SE P + T +E NG+K SS + + P E NE++ +
Sbjct: 181 SENSEDP-------KKLTEQEENGTKESSEETKDDKPEE-NEKKADNK 220
>UniRef100_O35540 Hepatoma-derived growth factor [Mus musculus]
Length = 669
Score = 40.4 bits (93), Expect = 0.35
Identities = 47/183 (25%), Positives = 74/183 (39%), Gaps = 23/183 (12%)
Query: 120 SPQSSARASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLG 179
+P +S A + SD ++ A LG GS ++K V V + +DR +D
Sbjct: 87 NPHASYSAPPPVSSSD-SEAPEADLGCGSDVDKDKESRRVMTVTAVTTTATSDRMESDSD 145
Query: 180 FGSSAEKGGLCVDDATMKVTVSPDLGISSQARDFDRANTDSSDHKQFEEGNDGDCSGKIN 239
S++ GL +KV+VS +S D D+A+ S+ E+ S K +
Sbjct: 146 SDKSSDHSGLKRKTPVLKVSVSKRARRASS--DLDQASVSPSE----EDSESPSESEKTS 199
Query: 240 DFD---EEIVVTTPPDAELLG-------------DSKVDGDEGKGEEDLLVRDAPFTGKA 283
D D E+ PP LG DSK D D K E + + +P + +
Sbjct: 200 DQDFTPEKKTAARPPRRGPLGGRKKKKVPSASDSDSKADSDGAKEEPVVTAQPSPSSSSS 259
Query: 284 ENS 286
+S
Sbjct: 260 SSS 262
>UniRef100_Q7RRF7 Chloroquine resistance marker protein, putative [Plasmodium yoelii
yoelii]
Length = 3604
Score = 40.4 bits (93), Expect = 0.35
Identities = 48/242 (19%), Positives = 95/242 (38%), Gaps = 17/242 (7%)
Query: 84 GDTLLVKYLRKKAKRDDDGGDLPRLLTKDLRARRVYSP-QSSARASLDLIDSDCADRKIA 142
G+ LL+ + K + + + P + K+ + + +++ +L +D +D K
Sbjct: 205 GNNLLLNTMHNNLKIEKNESNYPDDMNKNTNISSISNTNETNDIKQTNLNHADSSDNKNI 264
Query: 143 GLGFGSPSSEEK---SGVHVDDVMIAVNSDCTDRKIADLGFGSSAEKGGLCVDDATM--K 197
FGS ++ S + D I +N+ TD + + + ++D T +
Sbjct: 265 NTQFGSTDTDANMFNSSIDNSDKNIVINNLNTDNSNDLIQHEHTEKSNSPNINDQTNVNE 324
Query: 198 VTVSPDLGISSQARDFDRAN-TDSSDHKQFEEGNDGDCSGKINDFDEEIVVTTPPDAELL 256
+ S + D + N TD +++ + + ND +CSG ND D V T
Sbjct: 325 IDNSENKNHDINVNDSNTENNTDVTNNNELPKSNDNECSGNSNDNDNNNVDTL------- 377
Query: 257 GDSKVDGDEGKGEE--DLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHS 314
D + D KG E DL N + + S+ + V+++ ++ +N
Sbjct: 378 -DKSENDDNAKGSEINDLQNGSEKNNTPVSNRSQNETFLSSENKNDVIENVVMTSNSNED 436
Query: 315 FD 316
D
Sbjct: 437 ND 438
>UniRef100_UPI000032E5FC UPI000032E5FC UniRef100 entry
Length = 291
Score = 40.0 bits (92), Expect = 0.46
Identities = 26/76 (34%), Positives = 37/76 (48%), Gaps = 2/76 (2%)
Query: 24 SNPSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYSGLGGSSGNDEKREPPEEDD 83
S P S PP LTR+ S F+ R ++G P+ G R G ++ G G E+R P
Sbjct: 75 STPPSRPPPGLTRAESPSFIWRAKAGGWGPEQGRRAGAVHLAQGTRGG--EQRTAPSVLI 132
Query: 84 GDTLLVKYLRKKAKRD 99
D +L + R +K D
Sbjct: 133 YDEVLQQQRRLLSKLD 148
>UniRef100_UPI000036F6AB UPI000036F6AB UniRef100 entry
Length = 620
Score = 39.7 bits (91), Expect = 0.59
Identities = 47/176 (26%), Positives = 68/176 (37%), Gaps = 20/176 (11%)
Query: 10 SASAMEETLISRKRSNPSSSPPGALTRSRSQLFVHRNRSGQRRPDSGPRQGGLYSGLGGS 69
S+S+ + S S+PSSS G + + S LF G +G SG G+
Sbjct: 457 SSSSSSSSSSSSSSSSPSSSSGGESSETSSDLF------------EGSNEGSSSSGSSGA 504
Query: 70 SGNDEKREPPEEDDGDTLLVKYLRKKAKRDDDGGDLPRLLTK---DLRARRVYSPQSSAR 126
R P D+ +L L + ++D D PR LTK D A R + + +
Sbjct: 505 RREGRHRAPVTFDESGSLPFLSLAQFFLLNEDDDDQPRGLTKEQIDNLAMRSFGENDALK 564
Query: 127 ASLDLIDSDCADRKIAGLGFGSPSSEEKSGVHVDDVMIAVNSDCTDRKIADLGFGS 182
I K+ L P S E VH D ++ NS C + A L G+
Sbjct: 565 TCSVCITEYTEGNKLRKL----PCSHEYH-VHCIDRWLSENSTCPICRRAVLASGN 615
>UniRef100_Q26127 Microneme protein-1 [Plasmodium vivax]
Length = 757
Score = 39.7 bits (91), Expect = 0.59
Identities = 104/477 (21%), Positives = 169/477 (34%), Gaps = 57/477 (11%)
Query: 260 KVDGDEGKGEEDLLVRDAPFTGKAENSGEGLSRKDSKRNDSVLKSKYFSTAANHSFDLDL 319
KV GD G + +D+ TGKA +G+G + + V +S + + + D
Sbjct: 305 KVPGDSTHGNVNS-GQDSSTTGKAV-TGDGQNGNQTPAESDVQRSDIAESVSAKNVDPQK 362
Query: 320 YAGARSQTAFPGQIVQGPGFEACGGTNWPDNFVGTKKLGHCDKDEKGMGGQDFQLPLSSQ 379
RS + G E G +N + P
Sbjct: 363 SVSERSDDTASVTGIAEAGKENLGASN----------------SRPSESTVEANSPGDDT 406
Query: 380 SEKPSEPELKSESFTMRETNGSKLSSSQNLLESPMELNEREVSGECLSVPSGDDKCFNES 439
S P + E+ + NG S + + P E E S E ++ P + K
Sbjct: 407 VNSASIPVVSGENPLVTPYNGLGHSKDNSDSDGPAEFAESTKSAESMANPDSNSK----G 462
Query: 440 KCDPAADSDGVKVVGNALHNDDFMQCSIHKNNLDYAKVTNDSGCTSVQLGVLNDDFILTT 499
+ D+D K ++ ++ D + T D+ + GV D
Sbjct: 463 ETGKGQDNDMAKATKDSSNSSDGTSSATGD--------TTDAVDREINKGVPEDRDKTVG 514
Query: 500 PPDAKIYDNSAVNVDRSKSVPQDMKRLKPMDLRTN--TQESFSKGVCLKADSRKD----S 553
D DNSA N D + V +D R TN ++ K DS++ +
Sbjct: 515 SKDGGGEDNSA-NKDAATVVGEDRIRENSAGGSTNDRSKNDTEKNGASTPDSKQSEDATA 573
Query: 554 LLKSKSVRRPHLPGKLFK-TLGSVSKRFLPFLMELAKDDPGISKFGHGTPKKDEA-DMHG 611
L K++S+ + T S+ + +L K D + + P D+ D+ G
Sbjct: 574 LSKTESLESTESGDRTTNDTTNSLENKNGGKEKDLQKHDFKSNDTPNEEPNSDQTTDVEG 633
Query: 612 KRFELTLSSQPEEASIDGHKIEKGPIHG--TVESNALGNILFNSANELSNGNLP-KLTSS 668
+ SI K E+G T N + L NS N LSNG L K
Sbjct: 634 H----------DRDSIKNDKAERGKHMNKDTFTKNTNSHHL-NSNNNLSNGKLDIKEYKY 682
Query: 669 QDLPELPMQSDVKEVVQECLSAPSVE--EPIEKAVVGSKDECSREPNFDLCSAIVDF 723
+D+ + V++C + S+E +E + S + CSRE + +LC +I DF
Sbjct: 683 RDVKATRKNIILMSSVRKCNNNISLEYCNSVEDKI--SSNTCSREKSKNLCCSISDF 737
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.131 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,186,783,819
Number of Sequences: 2790947
Number of extensions: 100043537
Number of successful extensions: 210081
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 103
Number of HSP's that attempted gapping in prelim test: 209753
Number of HSP's gapped (non-prelim): 347
length of query: 1262
length of database: 848,049,833
effective HSP length: 139
effective length of query: 1123
effective length of database: 460,108,200
effective search space: 516701508600
effective search space used: 516701508600
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0108.13


