

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0106.15
(120 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
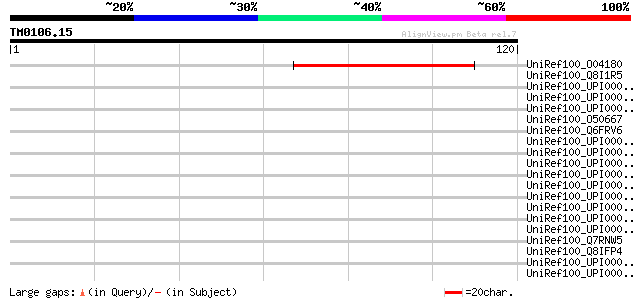
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O04180 Pge1 protein [Lotus japonicus] 61 7e-09
UniRef100_Q8I1R5 Hypothetical protein PFD0835c [Plasmodium falci... 35 0.52
UniRef100_UPI00002CE8D7 UPI00002CE8D7 UniRef100 entry 34 0.67
UniRef100_UPI00002C5783 UPI00002C5783 UniRef100 entry 33 1.1
UniRef100_UPI0000294384 UPI0000294384 UniRef100 entry 33 1.1
UniRef100_O50667 Hypothetical protein BBH09 [Borrelia burgdorferi] 33 1.1
UniRef100_Q6FRV6 Similar to sp|P48415 Saccharomyces cerevisiae Y... 33 1.1
UniRef100_UPI0000297E68 UPI0000297E68 UniRef100 entry 33 1.5
UniRef100_UPI000026051C UPI000026051C UniRef100 entry 33 1.5
UniRef100_UPI000030E215 UPI000030E215 UniRef100 entry 33 2.0
UniRef100_UPI0000306B01 UPI0000306B01 UniRef100 entry 33 2.0
UniRef100_UPI000025C73F UPI000025C73F UniRef100 entry 33 2.0
UniRef100_UPI00002FD798 UPI00002FD798 UniRef100 entry 32 2.6
UniRef100_UPI00002FC762 UPI00002FC762 UniRef100 entry 32 2.6
UniRef100_UPI00002FA3BB UPI00002FA3BB UniRef100 entry 32 2.6
UniRef100_UPI0000325E0D UPI0000325E0D UniRef100 entry 32 3.3
UniRef100_Q7RNW5 Hypothetical protein [Plasmodium yoelii yoelii] 32 3.3
UniRef100_Q8IFP4 Erythrocyte membrane-associated antigen, putati... 32 3.3
UniRef100_UPI00002C9EDF UPI00002C9EDF UniRef100 entry 32 4.4
UniRef100_UPI00002B9BF7 UPI00002B9BF7 UniRef100 entry 32 4.4
>UniRef100_O04180 Pge1 protein [Lotus japonicus]
Length = 210
Score = 60.8 bits (146), Expect = 7e-09
Identities = 26/43 (60%), Positives = 33/43 (76%)
Query: 68 VFDWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKFK 110
+ DWA+ VGK N +VVI+IRSDYG+ K +TLGCE GGK+K
Sbjct: 163 LIDWARCVGKENGYVVIVIRSDYGSAKRKPLITLGCERGGKYK 205
>UniRef100_Q8I1R5 Hypothetical protein PFD0835c [Plasmodium falciparum]
Length = 802
Score = 34.7 bits (78), Expect = 0.52
Identities = 21/101 (20%), Positives = 50/101 (48%), Gaps = 6/101 (5%)
Query: 10 DNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVF 69
+N E +H KV+++KE N+V+ +KE + + + ++ N ++ + + V + + +
Sbjct: 145 ENKIEEEHNKVNQLKEEHNKVNQLKEQHNKVNQLKEQHNKVNQLKEQHNKVNQLKEQHNI 204
Query: 70 DWAQLVGKNNDFVVIMIRSDYGTLNGKESLTLGCEHGGKFK 110
++ + +N I + G + + T+ +H GK K
Sbjct: 205 EFPATLKPSN------INNKKGVYHEEREETINKDHEGKIK 239
>UniRef100_UPI00002CE8D7 UPI00002CE8D7 UniRef100 entry
Length = 168
Score = 34.3 bits (77), Expect = 0.67
Identities = 26/94 (27%), Positives = 49/94 (51%), Gaps = 11/94 (11%)
Query: 3 HVLIEKVDNVAEGDHIKVDRVKEVDNEVDDM-------KEWGHILKEDEDLMNAIDDMNA 55
H IEK N++ D+I+ D++ +++ E D+ KE+ LKE +L + ++D N
Sbjct: 15 HSEIEK--NLSNQDNIETDKLIKLNKEYSDLMPLANLIKEYNQYLKESNNLKDLLNDNND 72
Query: 56 LSHVVMKFQNRQVFDWAQLVGKNNDFVVIMIRSD 89
+ + + RQV + +L+ K D + +I D
Sbjct: 73 SIKELAEVELRQVDEKIKLLEK--DLIKSLIPKD 104
>UniRef100_UPI00002C5783 UPI00002C5783 UniRef100 entry
Length = 262
Score = 33.5 bits (75), Expect = 1.1
Identities = 20/75 (26%), Positives = 43/75 (56%), Gaps = 9/75 (12%)
Query: 18 IKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAI--DD----MNALSHVVMK---FQNRQV 68
+KV+RV +++N+++D K+ GH +K+ E++ + + DD M + H ++K +
Sbjct: 146 LKVNRVLDLENKMNDHKQMGHDIKDHENMNHNMKKDDKKMSMQGMKHSMVKPTWVIPHPI 205
Query: 69 FDWAQLVGKNNDFVV 83
+ A + G +D V+
Sbjct: 206 LNLAYIAGNGSDEVI 220
>UniRef100_UPI0000294384 UPI0000294384 UniRef100 entry
Length = 357
Score = 33.5 bits (75), Expect = 1.1
Identities = 19/76 (25%), Positives = 37/76 (48%), Gaps = 5/76 (6%)
Query: 18 IKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQLVGK 77
+K+ R + N + + WG + K+ ++++ + ++A SHV N + W +VG
Sbjct: 1 MKMKRGGSLSNPIAAYETWGKLSKQKDNVVVILTGLSASSHVASHSNNSKPGWWEGIVGP 60
Query: 78 N-----NDFVVIMIRS 88
+ N F VI + S
Sbjct: 61 DKAIDTNKFYVICVNS 76
>UniRef100_O50667 Hypothetical protein BBH09 [Borrelia burgdorferi]
Length = 1278
Score = 33.5 bits (75), Expect = 1.1
Identities = 21/62 (33%), Positives = 36/62 (57%), Gaps = 4/62 (6%)
Query: 3 HVLIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDE--DLMNAIDDMNALSHVV 60
H LI +D + E ++D+ ++V E+D KE+G ILKE E D+ ++I L ++
Sbjct: 519 HFLISCLDYLTEKVWYELDKFEDVKKELD--KEYGIILKESEEYDIQDSISKELVLKRML 576
Query: 61 MK 62
+K
Sbjct: 577 LK 578
>UniRef100_Q6FRV6 Similar to sp|P48415 Saccharomyces cerevisiae YPL085w SEC16
[Candida glabrata]
Length = 2238
Score = 33.5 bits (75), Expect = 1.1
Identities = 21/57 (36%), Positives = 33/57 (57%), Gaps = 5/57 (8%)
Query: 5 LIEKVDNVAEGDHI-KVDRVKEVD--NEVDDMKEWGHILKEDEDLMNAIDDMNALSH 58
L+E+VD+V E DH+ +VD +E D EVD ++E H E+ D +D + + H
Sbjct: 265 LVEEVDHVEEVDHVEEVDHSEEEDLVEEVDHVEEVDH--SEEVDHSEEVDHVEEVDH 319
>UniRef100_UPI0000297E68 UPI0000297E68 UniRef100 entry
Length = 219
Score = 33.1 bits (74), Expect = 1.5
Identities = 23/61 (37%), Positives = 33/61 (53%), Gaps = 3/61 (4%)
Query: 12 VAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDM-NALSHVVMKFQNRQVFD 70
+AEGD KVD+VK++ NE D+ G ++E L N I + +A H K N+ F
Sbjct: 150 LAEGDEGKVDKVKKIINENFDLSPRG--IREMLKLNNPIYQVTSAYGHFGRKPTNKGEFS 207
Query: 71 W 71
W
Sbjct: 208 W 208
>UniRef100_UPI000026051C UPI000026051C UniRef100 entry
Length = 197
Score = 33.1 bits (74), Expect = 1.5
Identities = 17/75 (22%), Positives = 37/75 (48%), Gaps = 1/75 (1%)
Query: 1 MSHVLIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVV 60
+ + ++ ++ ++ D V E+ N + +K IL E +DL + N L H V
Sbjct: 100 LDEMTLDLQKRISHAENFSTDLVHEIRNPLASLKSASEILDETKDLDQRLKLTNILKHDV 159
Query: 61 MKFQNRQVFDWAQLV 75
+ + R + D++Q++
Sbjct: 160 QRIE-RLITDYSQIL 173
>UniRef100_UPI000030E215 UPI000030E215 UniRef100 entry
Length = 185
Score = 32.7 bits (73), Expect = 2.0
Identities = 16/58 (27%), Positives = 34/58 (58%), Gaps = 1/58 (1%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMK 62
L EK N+ + K + +++ +++DM+ I K+DE + N ++D N + H+++K
Sbjct: 120 LQEKFVNIVVQVNGKKKTLVKIEKDLEDMEVVNRI-KKDEKISNILNDKNLIKHIIVK 176
>UniRef100_UPI0000306B01 UPI0000306B01 UniRef100 entry
Length = 178
Score = 32.7 bits (73), Expect = 2.0
Identities = 19/76 (25%), Positives = 37/76 (48%), Gaps = 5/76 (6%)
Query: 18 IKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFDWAQLVGK 77
+K+ R + N + + WG + K+ ++++ + ++A SHV N + W +VG
Sbjct: 28 MKMKRGGSLSNPILAYETWGKLSKQKDNVVVILTGLSASSHVASHSNNSKPGWWEGIVGP 87
Query: 78 N-----NDFVVIMIRS 88
+ N F VI + S
Sbjct: 88 DKAIDTNKFFVICVNS 103
>UniRef100_UPI000025C73F UPI000025C73F UniRef100 entry
Length = 141
Score = 32.7 bits (73), Expect = 2.0
Identities = 17/68 (25%), Positives = 39/68 (57%), Gaps = 1/68 (1%)
Query: 11 NVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMKFQNRQVFD 70
+VA ++ + ++VDNE+ + E G+ + + L + IDD++ L+ ++ ++ F+
Sbjct: 44 SVARQQNMSEETARKVDNEIRKIVEKGYE-RAKKVLTDKIDDLHKLAKALLVYETLTGFE 102
Query: 71 WAQLVGKN 78
+L+ KN
Sbjct: 103 IEELINKN 110
>UniRef100_UPI00002FD798 UPI00002FD798 UniRef100 entry
Length = 265
Score = 32.3 bits (72), Expect = 2.6
Identities = 17/75 (22%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Query: 1 MSHVLIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVV 60
M + E ++ ++ D V E+ N + +K ILK+ +D+ ++ L+H V
Sbjct: 28 MEDMTNELQKRISHAENFSTDLVHEIRNPLASLKSASEILKDTKDINQRTKLLDILTHDV 87
Query: 61 MKFQNRQVFDWAQLV 75
+ + R + D++Q++
Sbjct: 88 QRIE-RLITDYSQML 101
>UniRef100_UPI00002FC762 UPI00002FC762 UniRef100 entry
Length = 194
Score = 32.3 bits (72), Expect = 2.6
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 3/56 (5%)
Query: 6 IEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGH---ILKEDEDLMNAIDDMNALSH 58
++K +A GD IKVD K EV D+K+W + ++ ED+D +SH
Sbjct: 46 VKKNLKLAYGDIIKVDTSKFESLEVPDLKKWDYPLEVIYEDDDFAIINKPFGIISH 101
>UniRef100_UPI00002FA3BB UPI00002FA3BB UniRef100 entry
Length = 286
Score = 32.3 bits (72), Expect = 2.6
Identities = 17/75 (22%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Query: 1 MSHVLIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVV 60
M + E ++ ++ D V E+ N + +K ILK+ +D+ ++ L+H V
Sbjct: 106 MEDMTNELQKRISHAENFSTDLVHEIRNPLASLKSASEILKDTKDINQRTKLLDILTHDV 165
Query: 61 MKFQNRQVFDWAQLV 75
+ + R + D++Q++
Sbjct: 166 QRIE-RLITDYSQML 179
>UniRef100_UPI0000325E0D UPI0000325E0D UniRef100 entry
Length = 70
Score = 32.0 bits (71), Expect = 3.3
Identities = 17/58 (29%), Positives = 35/58 (60%), Gaps = 1/58 (1%)
Query: 5 LIEKVDNVAEGDHIKVDRVKEVDNEVDDMKEWGHILKEDEDLMNAIDDMNALSHVVMK 62
L EK N+ + K + +++ +++D KE + +K+DE + N ++D N L H+++K
Sbjct: 5 LHEKFVNIVVQINGKKKSLIKIEKDLED-KEVINKVKKDEKISNILNDKNLLKHIIVK 61
>UniRef100_Q7RNW5 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 407
Score = 32.0 bits (71), Expect = 3.3
Identities = 15/46 (32%), Positives = 27/46 (58%), Gaps = 1/46 (2%)
Query: 22 RVKEVDNEVDDMKEWG-HILKEDEDLMNAIDDMNALSHVVMKFQNR 66
R+KE DN V +KE H+L+ ++++ N + LS ++ F N+
Sbjct: 53 RIKEEDNIVSKIKELNLHLLQNEDEINNMRQENETLSSQIINFSNK 98
>UniRef100_Q8IFP4 Erythrocyte membrane-associated antigen, putative [Plasmodium
falciparum]
Length = 4261
Score = 32.0 bits (71), Expect = 3.3
Identities = 22/61 (36%), Positives = 35/61 (57%), Gaps = 7/61 (11%)
Query: 6 IEKVDNVAEGDHIK-VDRVKEVDN--EVDDMKEWGHILKEDE----DLMNAIDDMNALSH 58
++ VDN+ D +K VD +K VD VD MK + D+ D MN+ID+MN++ +
Sbjct: 600 MKNVDNMKNVDKMKNVDDMKNVDKMKNVDKMKNVDKMKNVDKMNSVDKMNSIDNMNSMDN 659
Query: 59 V 59
+
Sbjct: 660 M 660
>UniRef100_UPI00002C9EDF UPI00002C9EDF UniRef100 entry
Length = 320
Score = 31.6 bits (70), Expect = 4.4
Identities = 24/91 (26%), Positives = 45/91 (49%), Gaps = 11/91 (12%)
Query: 3 HVLIEKVDNVAEGDHIKVDRVKEVDNEVDDM-------KEWGHILKEDEDLMNAIDDMNA 55
H+ +EK + E D D+ E+ E D+ KE+ + KE++DL N I D N+
Sbjct: 6 HLSLEKELSSGELDK---DKYAEISKEYSDLNGIIKQAKEYLNFKKENDDLNNIIKDKNS 62
Query: 56 LSHVVMKFQNRQVFDWAQLVGKNNDFVVIMI 86
S ++ +F N ++ + + KN + + +
Sbjct: 63 DSEMI-EFANIELVNLKKNYFKNEKIIKLFL 92
>UniRef100_UPI00002B9BF7 UPI00002B9BF7 UniRef100 entry
Length = 302
Score = 31.6 bits (70), Expect = 4.4
Identities = 15/55 (27%), Positives = 29/55 (52%), Gaps = 4/55 (7%)
Query: 21 DRVKEVDNEVDDMKEWGHILKEDEDLMNAID----DMNALSHVVMKFQNRQVFDW 71
D+++ + N++ K + +LK D + N ID + ++ V+ F + VFDW
Sbjct: 126 DKIRRIQNKLGQKKFYLKLLKNDPLVKNKIDPTDVQRSIRAYEVISFTKKSVFDW 180
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.139 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 212,141,576
Number of Sequences: 2790947
Number of extensions: 8347343
Number of successful extensions: 23719
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 23674
Number of HSP's gapped (non-prelim): 99
length of query: 120
length of database: 848,049,833
effective HSP length: 96
effective length of query: 24
effective length of database: 580,118,921
effective search space: 13922854104
effective search space used: 13922854104
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0106.15


