

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0105.5
(231 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
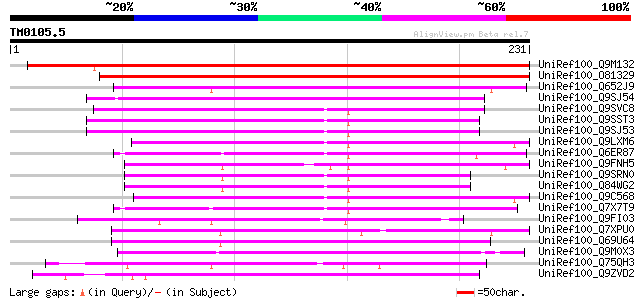
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M132 Putative hypoersensitive response protein [Arab... 221 2e-56
UniRef100_O81329 F3D13.5 protein [Arabidopsis thaliana] 209 4e-53
UniRef100_Q652J9 Harpin-induced family protein-like [Oryza sativa] 129 5e-29
UniRef100_Q9SJ54 Putative harpin-induced protein [Arabidopsis th... 128 1e-28
UniRef100_Q9SVC8 Hypothetical protein F22O6_150 [Arabidopsis tha... 124 1e-27
UniRef100_Q9SST3 Hypothetical protein AT4g09590 [Arabidopsis tha... 120 4e-26
UniRef100_Q9SJ53 Putative harpin-induced protein [Arabidopsis th... 118 1e-25
UniRef100_Q9LXM6 Hypothetical protein T10D17_10 [Arabidopsis tha... 115 1e-24
UniRef100_Q6ER87 Harpin-induced protein-like [Oryza sativa] 110 2e-23
UniRef100_Q9FNH5 Harpin-induced protein-like [Arabidopsis thaliana] 110 4e-23
UniRef100_Q9SRN0 T19F11.6 protein [Arabidopsis thaliana] 107 2e-22
UniRef100_Q84WG2 Putative VAMP protein SEC22 [Arabidopsis thaliana] 106 4e-22
UniRef100_Q9C568 NDR1/HIN1-like protein [Arabidopsis thaliana] 104 2e-21
UniRef100_Q7X7T9 OSJNBa0033G16.4 protein [Oryza sativa] 102 1e-20
UniRef100_Q9FI03 Similarity to harpin-induced protein [Arabidops... 101 2e-20
UniRef100_Q7XPU0 OSJNBa0088H09.16 protein [Oryza sativa] 92 1e-17
UniRef100_Q69U64 Harpin-induced protein 1 family (HIN1)-like [Or... 89 9e-17
UniRef100_Q9M0X3 Hypothetical protein AT4g05220 [Arabidopsis tha... 87 4e-16
UniRef100_Q75QH3 Hin1 like protein [Capsicum chinense] 82 8e-15
UniRef100_Q9ZVD2 Expressed protein [Arabidopsis thaliana] 79 9e-14
>UniRef100_Q9M132 Putative hypoersensitive response protein [Arabidopsis thaliana]
Length = 227
Score = 221 bits (562), Expect = 2e-56
Identities = 108/225 (48%), Positives = 151/225 (67%), Gaps = 2/225 (0%)
Query: 9 KGQNQHHHLPPPNKIQMNNHKASPSPYG--GGGGGSSPKRAACTFITVFLLAVGITLLVL 66
+G+ + H + + ++P Y GGGGG +RA C I L+ +GI L+L
Sbjct: 3 EGEAKAEHAAKADHKNAPSASSTPESYSKEGGGGGGDARRAICGAIFTILVILGIIALIL 62
Query: 67 WLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSYR 126
WLVYRPHKPR TVVGAA+Y LN T+PPL+S ++QF+VL RNPN+RVSI+YD+ S +V+Y+
Sbjct: 63 WLVYRPHKPRLTVVGAAIYDLNFTAPPLISTSVQFSVLARNPNRRVSIHYDKLSMYVTYK 122
Query: 127 NQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANGLAMDESYGVVGLKLVFLGR 186
+Q IT + LPPL L S V ++PV+GG +PV+ EVANGL DE+YGVV +++V GR
Sbjct: 123 DQIITPPLPLPPLRLGHKSTVVIAPVMGGNGIPVSPEVANGLKNDEAYGVVLMRVVIFGR 182
Query: 187 VKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQAQACDVDL 231
++WKAG ++T +G Y +CD+ + GQVPLL C VD+
Sbjct: 183 LRWKAGAIKTGRYGFYARCDVWLRFNPSSNGQVPLLAPSTCKVDV 227
>UniRef100_O81329 F3D13.5 protein [Arabidopsis thaliana]
Length = 202
Score = 209 bits (533), Expect = 4e-53
Identities = 100/191 (52%), Positives = 136/191 (70%)
Query: 41 GSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQ 100
G +RA C I L+ +GI L+LWLVYRPHKPR TVVGAA+Y LN T+PPL+S ++Q
Sbjct: 12 GGDARRAICGAIFTILVILGIIALILWLVYRPHKPRLTVVGAAIYDLNFTAPPLISTSVQ 71
Query: 101 FNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPV 160
F+VL RNPN+RVSI+YD+ S +V+Y++Q IT + LPPL L S V ++PV+GG +PV
Sbjct: 72 FSVLARNPNRRVSIHYDKLSMYVTYKDQIITPPLPLPPLRLGHKSTVVIAPVMGGNGIPV 131
Query: 161 TVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVP 220
+ EVANGL DE+YGVV +++V GR++WKAG ++T +G Y +CD+ + GQVP
Sbjct: 132 SPEVANGLKNDEAYGVVLMRVVIFGRLRWKAGAIKTGRYGFYARCDVWLRFNPSSNGQVP 191
Query: 221 LLQAQACDVDL 231
LL C VD+
Sbjct: 192 LLAPSTCKVDV 202
>UniRef100_Q652J9 Harpin-induced family protein-like [Oryza sativa]
Length = 218
Score = 129 bits (325), Expect = 5e-29
Identities = 74/195 (37%), Positives = 113/195 (57%), Gaps = 11/195 (5%)
Query: 47 AACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALN----TTSPPLLSAALQFN 102
A L A LVL+LVYRP P+ +V AAVY L ++S L+A++QF
Sbjct: 23 ATAALAVAALAAAAAAALVLYLVYRPVMPQASVPRAAVYRLALANASSSAHALAASVQFT 82
Query: 103 VLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTV 162
+++ NP+ R S+ YD A+ SYR +P+ LPP+ ++ + V++SP++GG +PV+
Sbjct: 83 LVLHNPSDRASLLYDGLVAYASYRGEPVMPPAPLPPVAQDRGADVAMSPLLGGAAVPVSP 142
Query: 163 EVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKK-------GF 215
+ A LA D + V L+LV +GRVK+++G R+ LYV+C+++VGL G
Sbjct: 143 DAARALAADCAARRVQLRLVVMGRVKYRSGPFRSGWRDLYVRCNVVVGLSTEAAVAGGGG 202
Query: 216 VGQVPLLQAQACDVD 230
G VPLL+ C VD
Sbjct: 203 GGDVPLLEYPRCAVD 217
>UniRef100_Q9SJ54 Putative harpin-induced protein [Arabidopsis thaliana]
Length = 210
Score = 128 bits (322), Expect = 1e-28
Identities = 66/177 (37%), Positives = 99/177 (55%), Gaps = 1/177 (0%)
Query: 35 YGGGGGGSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPL 94
+GGGGGG + R C I F++ V IT+ ++W++ +P KPRF + A VYA N + P L
Sbjct: 9 HGGGGGGGTASRI-CGVIIGFIIIVLITIFLVWIILQPTKPRFILQDATVYAFNLSQPNL 67
Query: 95 LSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIG 154
L++ Q + RN N R+ IYYDR + +YRNQ IT + +PP + SP +
Sbjct: 68 LTSNFQITIASRNRNSRIGIYYDRLHVYATYRNQQITLRTAIPPTYQGHKEDNVWSPFVY 127
Query: 155 GTPMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGL 211
G +P+ A L +++ G V L + GRV+WK G L T + L+V+C + L
Sbjct: 128 GNSVPIAPFNAVALGDEQNRGFVTLIIRADGRVRWKVGTLITGKYHLHVRCQAFINL 184
>UniRef100_Q9SVC8 Hypothetical protein F22O6_150 [Arabidopsis thaliana]
Length = 208
Score = 124 bits (312), Expect = 1e-27
Identities = 63/175 (36%), Positives = 99/175 (56%), Gaps = 2/175 (1%)
Query: 38 GGGGSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSA 97
GGG R C I F++ V IT+ ++W++ RP KPRF + A VYA N + P LL++
Sbjct: 9 GGGKEVVVRKLCAAIIAFIVIVLITIFLVWVILRPTKPRFVLQDATVYAFNLSQPNLLTS 68
Query: 98 ALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGT 156
Q + RNPN ++ IYYDR + +Y NQ IT + +PP + + H +V++ SP + GT
Sbjct: 69 NFQVTIASRNPNSKIGIYYDRLHVYATYMNQQITLRTAIPPTY-QGHKEVNVWSPFVYGT 127
Query: 157 PMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGL 211
+P+ + L ++ G VGL + G V+WK L T + ++V+C + L
Sbjct: 128 AVPIAPYNSVALGEEKDRGFVGLMIRADGTVRWKVRTLITGKYHIHVRCQAFINL 182
>UniRef100_Q9SST3 Hypothetical protein AT4g09590 [Arabidopsis thaliana]
Length = 211
Score = 120 bits (300), Expect = 4e-26
Identities = 61/176 (34%), Positives = 98/176 (55%), Gaps = 2/176 (1%)
Query: 35 YGGGGGGSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPL 94
+GGGGGG C I F++ V +T+ ++W++ +P P F + VYA N + P L
Sbjct: 9 HGGGGGGGGTACRICGAIIGFIIIVLMTIFLVWIILQPKNPEFILQDTTVYAFNLSQPNL 68
Query: 95 LSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVI 153
L++ Q + RN N + IYYD A+ SYRNQ IT LPP + ++H + S+ SP++
Sbjct: 69 LTSKFQITIASRNRNSNIGIYYDHLHAYASYRNQQITLASDLPPTY-QRHKEDSVWSPLL 127
Query: 154 GGTPMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV 209
G +P+ A L +++ GV L + G+V+WK G L ++ L+V+C +
Sbjct: 128 YGNQVPIAPFNAVALGDEQNSGVFTLTICVDGQVRWKVGTLTIGNYHLHVRCQAFI 183
>UniRef100_Q9SJ53 Putative harpin-induced protein [Arabidopsis thaliana]
Length = 211
Score = 118 bits (295), Expect = 1e-25
Identities = 62/176 (35%), Positives = 98/176 (55%), Gaps = 2/176 (1%)
Query: 35 YGGGGGGSSPKRAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPL 94
+GGGGGG C I F++ V +T+ ++ ++ +P KP F + VYA N + P L
Sbjct: 9 HGGGGGGGGTACRICGAIIGFIIIVLMTIFLVSIILQPKKPEFILQDTTVYAFNLSQPNL 68
Query: 95 LSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVI 153
L++ Q + RN N + IYYD A+ SYRNQ IT LPP + ++H + S+ SP++
Sbjct: 69 LTSKFQITIASRNRNSNIGIYYDHLHAYASYRNQQITLASDLPPTY-QRHKENSVWSPLL 127
Query: 154 GGTPMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLV 209
G +P+ A L +++ GV L + GRV+WK G L ++ L+V+C +
Sbjct: 128 YGNQVPIAPFNAVALGDEQNSGVFTLTICVDGRVRWKVGTLTIGNYHLHVRCQAFI 183
>UniRef100_Q9LXM6 Hypothetical protein T10D17_10 [Arabidopsis thaliana]
Length = 206
Score = 115 bits (287), Expect = 1e-24
Identities = 64/181 (35%), Positives = 98/181 (53%), Gaps = 5/181 (2%)
Query: 55 FLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSI 114
FL AV + ++W + PH PRF + A +YA N + P L++ LQ + RNPN ++ I
Sbjct: 27 FLAAVLFVVFLVWAILHPHGPRFVLQDATIYAFNVSQPNYLTSNLQVTLSSRNPNDKIGI 86
Query: 115 YYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGLAMDES 173
+YDR + SYRNQ +T LLP + + H V++ SP + GT +PV + L+ D +
Sbjct: 87 FYDRLDIYASYRNQQVTLATLLPATY-QGHLDVTIWSPFLYGTTVPVAPYFSPALSQDLT 145
Query: 174 YGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQ---AQACDVD 230
G+V L + G V+WK G + + L+V C + L F G P ++ Q C VD
Sbjct: 146 AGMVLLNIKIDGWVRWKVGTWVSGRYRLHVNCPAYITLAGHFSGDGPAVKYQLVQRCAVD 205
Query: 231 L 231
+
Sbjct: 206 V 206
>UniRef100_Q6ER87 Harpin-induced protein-like [Oryza sativa]
Length = 224
Score = 110 bits (276), Expect = 2e-23
Identities = 71/186 (38%), Positives = 102/186 (54%), Gaps = 6/186 (3%)
Query: 47 AACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIR 106
AAC I +L VG LV++L RP KP F + + +++ P L SA Q + R
Sbjct: 21 AAC--ILALVLVVGFIALVVYLALRPSKPSFYLQDLQLRSVDLGDPSL-SATAQVTLASR 77
Query: 107 NPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVA 165
NPN V ++Y R FV+YR++P+T V LPP + + H V++ SPV+ G +PV VA
Sbjct: 78 NPNDHVGVHYRRLDVFVTYRDEPVTVPVSLPPTY-QGHRDVTIWSPVLSGESVPVAGFVA 136
Query: 166 NGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCD-LLVGLKKGFVGQVPLLQA 224
+ L D + G V L++ GRVKWK G + + L+V C +L G VG +PL A
Sbjct: 137 DALRQDVAAGYVALQVKVDGRVKWKVGSWVSGSYHLFVSCPAMLASAGPGGVGPMPLGGA 196
Query: 225 QACDVD 230
A V+
Sbjct: 197 SAAVVN 202
>UniRef100_Q9FNH5 Harpin-induced protein-like [Arabidopsis thaliana]
Length = 207
Score = 110 bits (274), Expect = 4e-23
Identities = 65/188 (34%), Positives = 100/188 (52%), Gaps = 12/188 (6%)
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPP--LLSAALQFNVLIRNPN 109
I++ LL + + +L++W + +P KPRF + A V+ N + P LL++ QF + RNPN
Sbjct: 24 ISIILLLILVVILLVWAILQPSKPRFVLQDATVFNFNVSGNPPNLLTSNFQFTLSSRNPN 83
Query: 110 KRVSIYYDRFSAFVSYRNQPITQQVLLPPLFL---EKHSQVSL-SPVIGGTPMPVTVEVA 165
++ IYYDR + SYR+Q IT LP L + H +V++ SP +GG +PV A
Sbjct: 84 DKIGIYYDRLDVYASYRSQQIT----LPSPMLTTYQGHKEVNVWSPFVGGYSVPVAPYNA 139
Query: 166 NGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQV--PLLQ 223
L D S G + L L GRV+WK G T + L+V+C L+ G + +
Sbjct: 140 FYLDQDHSSGAIMLMLHLDGRVRWKVGSFITGKYHLHVRCHALINFGSSAAGVIVGKYML 199
Query: 224 AQACDVDL 231
+ C V +
Sbjct: 200 TETCSVSV 207
>UniRef100_Q9SRN0 T19F11.6 protein [Arabidopsis thaliana]
Length = 209
Score = 107 bits (267), Expect = 2e-22
Identities = 59/157 (37%), Positives = 88/157 (55%), Gaps = 4/157 (2%)
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPP--LLSAALQFNVLIRNPN 109
I L + +T+L++W + +P KPRF + A VYA N + P LL++ Q + RNPN
Sbjct: 22 IIFVLFIIFLTILLIWAILQPSKPRFILQDATVYAFNVSGNPPNLLTSNFQITLSSRNPN 81
Query: 110 KRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGL 168
++ IYYDR + +YR+Q IT +PP + + H V + SP + GT +P+ L
Sbjct: 82 NKIGIYYDRLDVYATYRSQQITFPTSIPPTY-QGHKDVDIWSPFVYGTSVPIAPFNGVSL 140
Query: 169 AMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKC 205
D+ GVV L + GRV+WK G T + L+VKC
Sbjct: 141 DTDKDNGVVLLIIRADGRVRWKVGTFITGKYHLHVKC 177
>UniRef100_Q84WG2 Putative VAMP protein SEC22 [Arabidopsis thaliana]
Length = 209
Score = 106 bits (265), Expect = 4e-22
Identities = 58/157 (36%), Positives = 88/157 (55%), Gaps = 4/157 (2%)
Query: 52 ITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPP--LLSAALQFNVLIRNPN 109
I L + +T+L++W + +P KPRF + A VYA N + P +L++ Q + RNPN
Sbjct: 22 IIFVLFIIFLTILLIWAILQPSKPRFILQDATVYAFNVSGNPPNILTSNFQITLSSRNPN 81
Query: 110 KRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGL 168
++ IYYDR + +YR+Q IT +PP + + H V + SP + GT +P+ L
Sbjct: 82 NKIGIYYDRLDVYATYRSQQITFPTSIPPTY-QGHKDVDIWSPFVYGTSVPIAPFNGVSL 140
Query: 169 AMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKC 205
D+ GVV L + GRV+WK G T + L+VKC
Sbjct: 141 DTDKDNGVVLLIIRADGRVRWKVGTFITGKYHLHVKC 177
>UniRef100_Q9C568 NDR1/HIN1-like protein [Arabidopsis thaliana]
Length = 210
Score = 104 bits (259), Expect = 2e-21
Identities = 58/180 (32%), Positives = 96/180 (53%), Gaps = 5/180 (2%)
Query: 56 LLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIY 115
++AV + ++W + PH PRF + + N + P LS+ LQ V RNPN ++ I+
Sbjct: 32 IVAVAFVVFLVWAILHPHGPRFVLQDVTINDFNVSQPNFLSSNLQVTVSSRNPNDKIGIF 91
Query: 116 YDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVANGLAMDESY 174
YDR +V+YRNQ +T LLP + + H +V++ SP + G+ +PV +++ L D
Sbjct: 92 YDRLDIYVTYRNQEVTLARLLPSTY-QGHLEVTVWSPFLIGSAVPVAPYLSSALNEDLFA 150
Query: 175 GVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQ---AQACDVDL 231
G+V L + G V+WK G + + L+V C + + G P ++ Q C VD+
Sbjct: 151 GLVLLNIKIDGWVRWKVGSWVSGSYRLHVNCPAFITVTGKLTGTGPAIKYQLVQRCAVDV 210
>UniRef100_Q7X7T9 OSJNBa0033G16.4 protein [Oryza sativa]
Length = 220
Score = 102 bits (253), Expect = 1e-20
Identities = 66/181 (36%), Positives = 94/181 (51%), Gaps = 5/181 (2%)
Query: 47 AACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIR 106
AAC I + VG +LV++L P KP F + + ++ S P +S LQ + R
Sbjct: 20 AAC--ILTLAVLVGFIVLVIYLAIHPSKPSFYLQDVQLRNIDL-SDPAISLNLQVTIASR 76
Query: 107 NPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSL-SPVIGGTPMPVTVEVA 165
NPN RV +YY F +YR +PIT V LP ++ + H VS+ SPV+ G +PV VA
Sbjct: 77 NPNDRVGVYYKTLHVFTTYREEPITVPVELPAIY-QGHKDVSVWSPVMSGESVPVGQYVA 135
Query: 166 NGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQAQ 225
+ + D + G V L + GRVKWK G + + L+V C L+ G VG + A
Sbjct: 136 DAMRQDIAAGYVLLHVKVDGRVKWKVGSWVSGGYHLFVTCPALLAASGGNVGGAFAMSAT 195
Query: 226 A 226
A
Sbjct: 196 A 196
>UniRef100_Q9FI03 Similarity to harpin-induced protein [Arabidopsis thaliana]
Length = 213
Score = 101 bits (251), Expect = 2e-20
Identities = 57/177 (32%), Positives = 96/177 (54%), Gaps = 9/177 (5%)
Query: 31 SPSPYGGGGGGSSPKRAACTFITVFLLAVGITLLV--LWLVYRPHKPRFTVVGAAVYALN 88
SP GG + R F T G+ L++ +WL+ P +P F++ A +Y+LN
Sbjct: 8 SPKHCAKKGGININNRHKKLFFTFSTFFSGLLLIIFLVWLILHPERPEFSLTEADIYSLN 67
Query: 89 --TTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQ 146
T+S LL++++Q + +NPNK+V IYYD+ + +YR Q IT + LPP F + H +
Sbjct: 68 LTTSSTHLLNSSVQLTLFSKNPNKKVGIYYDKLLVYAAYRGQQITSEASLPP-FYQSHEE 126
Query: 147 VS-LSPVIGGTPMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLY 202
++ L+ + GT +PV ++ + S G + + + G+++WK G TW G Y
Sbjct: 127 INLLTAFLQGTELPVAQSFGYQISRERSTGKIIIGMKMDGKLRWKIG---TWVSGAY 180
>UniRef100_Q7XPU0 OSJNBa0088H09.16 protein [Oryza sativa]
Length = 220
Score = 91.7 bits (226), Expect = 1e-17
Identities = 65/201 (32%), Positives = 91/201 (44%), Gaps = 17/201 (8%)
Query: 46 RAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSP--PLLSAALQFNV 103
R C + +L +LV+WL RPHKPRF + +V LN T P L +Q V
Sbjct: 22 RRLCWALVALVLLTLFIVLVVWLALRPHKPRFYLQDLSVLCLNVTPPASAYLFTTMQATV 81
Query: 104 LIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGG---TPMPV 160
RN N RV +YYD+ + Y++ IT LP + + Q SP + +P
Sbjct: 82 AARNDNGRVGVYYDKVDVYAQYKDVAITVPTRLPVEYQGHYDQSVWSPFLQSLDHVVLPP 141
Query: 161 TVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKK------- 213
+ VA LA DE+ G V + + G V+WK G + H+ L V C L+ +
Sbjct: 142 NLAVA--LAQDETAGYVLVDIRLDGWVRWKVGTWISGHYHLRVNCPALLTVNDGKGSYGV 199
Query: 214 ---GFVGQVPLLQAQACDVDL 231
G G QA AC VD+
Sbjct: 200 NYGGGDGYFRFQQAAACAVDV 220
>UniRef100_Q69U64 Harpin-induced protein 1 family (HIN1)-like [Oryza sativa]
Length = 227
Score = 89.0 bits (219), Expect = 9e-17
Identities = 54/182 (29%), Positives = 86/182 (46%), Gaps = 13/182 (7%)
Query: 46 RAACTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSP------------- 92
R C I LL + + +L++WL+ RP KPRF + V LN T+
Sbjct: 19 RRCCAAIFGILLLLLLIVLIVWLILRPTKPRFYLNDLTVVCLNVTTGGSYAGATASSGYF 78
Query: 93 PLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPV 152
L+ +Q + RN N+RV IYYDR + Y+ IT LPP++ SP
Sbjct: 79 SFLTVTMQTTLAARNGNERVGIYYDRADVYAEYKGLRITVPTSLPPVYQGHPDLTVWSPF 138
Query: 153 IGGTPMPVTVEVANGLAMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLK 212
+ G + + +A + DE+ G + + + G +++KAG T H+ L V+C L+ +
Sbjct: 139 LSGNNVQLPPYLAVSITQDETAGYLLVTIRVDGWIRYKAGAFITGHYHLRVRCPALLIVN 198
Query: 213 KG 214
G
Sbjct: 199 DG 200
>UniRef100_Q9M0X3 Hypothetical protein AT4g05220 [Arabidopsis thaliana]
Length = 226
Score = 86.7 bits (213), Expect = 4e-16
Identities = 48/181 (26%), Positives = 89/181 (48%), Gaps = 4/181 (2%)
Query: 49 CTFITVFLLAVGITLLVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNP 108
C + L VG+ +LWL RPH+PRF + V L+ + + +A + FNV I NP
Sbjct: 47 CAMFLLVLFFVGVIAFILWLSLRPHRPRFHIQDFVVQGLDQPTG-VENARIAFNVTILNP 105
Query: 109 NKRVSIYYDRFSAFVSYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANGL 168
N+ + +Y+D + Y++Q + LL P F + + ++ + G + V
Sbjct: 106 NQHMGVYFDSMEGSIYYKDQRVGLIPLLNPFFQQPTNTTIVTGTLTGASLTVNSNRWTEF 165
Query: 169 AMDESYGVVGLKLVFLGRVKWKAGGLRTWHHGLYVKCDLLVGLKKGFVGQVPLLQAQACD 228
+ D + G VG +L + +++K + HH ++ C+++VG + G + +P + C
Sbjct: 166 SNDRAQGTVGFRLDIVSTIRFKLHRWISKHHRMHANCNIVVG-RDGLI--LPKFNHKRCP 222
Query: 229 V 229
V
Sbjct: 223 V 223
>UniRef100_Q75QH3 Hin1 like protein [Capsicum chinense]
Length = 228
Score = 82.4 bits (202), Expect = 8e-15
Identities = 63/209 (30%), Positives = 99/209 (47%), Gaps = 26/209 (12%)
Query: 17 LPPPNKIQMNNHKASPSPYGGGGGGSSPKRAACTF----------ITVFLLAVGITLLVL 66
+PPP+K Y G GSS +C F I L+ +G+ LVL
Sbjct: 15 IPPPSK-----------SYHRHGRGSSCNPCSCLFGCLCNCIFQIIFTILIVLGVIALVL 63
Query: 67 WLVYRPHKPRFTVVGAAVYALN-TTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFVSY 125
WLV RP+K +F V A + + +T+ L L N+ IRNPNKR+ IYYD A Y
Sbjct: 64 WLVLRPNKVKFYVTDATLTQFDLSTANNTLYYDLALNMTIRNPNKRIGIYYDSIEARGMY 123
Query: 126 RNQPITQQVLLPPLFLEKHSQV-SLSPVIGGTPMPVTVE-VANGLAMDESYGVVGLKLVF 183
+ + Q L P F + H SL PV+ G + + + + +++ GV +++
Sbjct: 124 KGERFASQNLEP--FYQGHKNTSSLHPVLKGQSLVLLGDREKSNYNNEKNSGVYDIEMKL 181
Query: 184 LGRVKWKAGGLRTWHHGLYVKCDLLVGLK 212
R++ K G ++T ++CD+ V L+
Sbjct: 182 YMRIRLKIGWIKTHKIKPKIECDIKVPLE 210
>UniRef100_Q9ZVD2 Expressed protein [Arabidopsis thaliana]
Length = 260
Score = 79.0 bits (193), Expect = 9e-14
Identities = 54/206 (26%), Positives = 99/206 (47%), Gaps = 16/206 (7%)
Query: 11 QNQHHHLPPPNKI----QMNNHKASPSPYGGGGGGSSPKRAACTFIT-VFLLAV--GITL 63
++Q + +PPP Q++ K + S + + C+F+ VF+L V GI+
Sbjct: 43 KDQIYRIPPPENAHRFEQLSRKKTNRS---------NCRCCFCSFLAAVFILIVLAGISF 93
Query: 64 LVLWLVYRPHKPRFTVVGAAVYALNTTSPPLLSAALQFNVLIRNPNKRVSIYYDRFSAFV 123
VL+L+YRP P++++ G +V +N S +S + V RN N ++ +YY++ S+
Sbjct: 94 AVLYLIYRPEAPKYSIEGFSVSGINLNSTSPISPSFNVTVRSRNGNGKIGVYYEKESSVD 153
Query: 124 SYRNQPITQQVLLPPLFLEKHSQVSLSPVIGGTPMPVTVEVANGLAMDESYGVVGLKLVF 183
Y N ++P + + + V+ G+ + +T + + + S V KL
Sbjct: 154 VYYNDVDISNGVMPVFYQPAKNVTVVKLVLSGSKIQLTSGMRKEMRNEVSKKTVPFKLKI 213
Query: 184 LGRVKWKAGGLRTWHHGLYVKCDLLV 209
VK K G ++TW + V CD+ V
Sbjct: 214 KAPVKIKFGSVKTWTMIVNVDCDVTV 239
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 432,984,749
Number of Sequences: 2790947
Number of extensions: 19007037
Number of successful extensions: 60300
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 48
Number of HSP's that attempted gapping in prelim test: 60144
Number of HSP's gapped (non-prelim): 128
length of query: 231
length of database: 848,049,833
effective HSP length: 123
effective length of query: 108
effective length of database: 504,763,352
effective search space: 54514442016
effective search space used: 54514442016
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0105.5


