

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0102b.4
(278 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
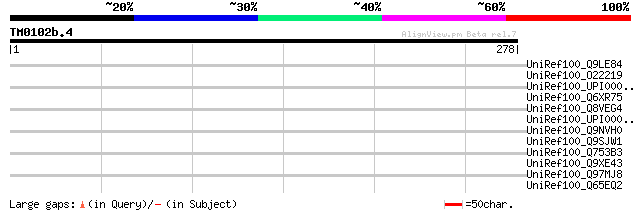
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LE84 Hypothetical protein AT4g08820 [Arabidopsis tha... 42 0.013
UniRef100_O22219 Putative Ta11-like non-LTR retroelement protein... 36 0.96
UniRef100_UPI00001CFA61 UPI00001CFA61 UniRef100 entry 35 1.6
UniRef100_Q6XR75 Acetylcholinesterase [Rhipicephalus sanguineus] 35 1.6
UniRef100_Q8VEG4 Protein C14orf114 homolog [Mus musculus] 35 1.6
UniRef100_UPI0000369D47 UPI0000369D47 UniRef100 entry 35 2.8
UniRef100_Q9NVH0 Protein C14orf114 [Homo sapiens] 35 2.8
UniRef100_Q9SJW1 Hypothetical protein At2g01050 [Arabidopsis tha... 34 3.6
UniRef100_Q753B3 AFR411Cp [Ashbya gossypii] 34 3.6
UniRef100_Q9XE43 Putative non-LTR retrolelement reverse transcri... 33 6.2
UniRef100_Q97MJ8 Uncharacterized membrane protein, ortholog YYAS... 33 8.1
UniRef100_Q65EQ2 Rnr [Bacillus licheniformis] 33 8.1
>UniRef100_Q9LE84 Hypothetical protein AT4g08820 [Arabidopsis thaliana]
Length = 404
Score = 42.4 bits (98), Expect = 0.013
Identities = 44/195 (22%), Positives = 83/195 (42%), Gaps = 21/195 (10%)
Query: 100 LKPWSSSFAP-----INRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WS +F P + WVR T IP++ + + + +G ++K+D +T F
Sbjct: 94 VQDWSPNFDPLRDDIVTTPVWVRLTNIPVNYYHRCLLEEIARGLGKLLKVDLNTITFGRG 153
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISI--IEEGEALLHPNCCCREESSVEDDVFS 212
F+R+ + L +LI G Y ++ +EEG ++ + R E + VF+
Sbjct: 154 RFARVCIEVNLAK--PLKGTVLINGDRYFVAYEEVEEGFTVVRQS-GIRVEKPAQKMVFA 210
Query: 213 GWSLNGVTVTP----------ELSESGGEDDDGLWPVETKSQHDLQGSPNHSLGDNKNPR 262
S G + E+S G ++ + E L+GS + KN R
Sbjct: 211 AGSSGGRSNRSLRELPKNQGVEISNRFGGLEEDMVSAEIGEVAILEGSNKENKYKGKNMR 270
Query: 263 SNTGAVNRLLSSRLN 277
+G V ++ +++N
Sbjct: 271 KESG-VTQVKEAQVN 284
>UniRef100_O22219 Putative Ta11-like non-LTR retroelement protein [Arabidopsis
thaliana]
Length = 418
Score = 36.2 bits (82), Expect = 0.96
Identities = 33/154 (21%), Positives = 66/154 (42%), Gaps = 12/154 (7%)
Query: 67 LRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKPWSSS-------FAPINRVTWVRCT 119
L +R E +L+E+L+ + E+ ++ W F P+ W+R
Sbjct: 72 LSQERFQFFFKSEDDLLEILKTGVWTQDEWCVVMERWIEKSTEEYLMFLPV----WMRLR 127
Query: 120 GIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARKILIKG 179
IP++ +T D K + VG V+K++ D + ++ R+ V N + ++I +
Sbjct: 128 NIPVNYYTQDTIKKIASCVGKVLKVELDLEKSQAQDYIRVQVIIDVRNPLRNFKEIQLP- 186
Query: 180 TTYTISIIEEGEALLHPNCCCREESSVEDDVFSG 213
T +S+ + E + C+ + + D SG
Sbjct: 187 TGEIVSVTFDYERIRKRCFLCQRLTHEKGDCDSG 220
>UniRef100_UPI00001CFA61 UPI00001CFA61 UniRef100 entry
Length = 570
Score = 35.4 bits (80), Expect = 1.6
Identities = 29/98 (29%), Positives = 45/98 (45%), Gaps = 5/98 (5%)
Query: 7 EGSKVDRGRMTALELWSGPEIPKEEWMLRSLVREMKNLELIPS--LREAFMVEGHITIQV 64
E +V G L S P KEE L +RE N +++ L+EA +E I +
Sbjct: 447 ERRQVRSGARALLNAESLPAHRKEE--LLHALREFYNTDIVTEEMLQEAASLETRIYNE- 503
Query: 65 RYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKP 102
Y+ ++ H E L L+Q WR F+D+++P
Sbjct: 504 SYIPHGLKVVQRHTEGGLRSLMQLESRWRQHFLDSMQP 541
>UniRef100_Q6XR75 Acetylcholinesterase [Rhipicephalus sanguineus]
Length = 593
Score = 35.4 bits (80), Expect = 1.6
Identities = 20/67 (29%), Positives = 28/67 (40%)
Query: 190 GEALLHPNCCCREESSVEDDVFSGWSLNGVTVTPELSESGGEDDDGLWPVETKSQHDLQG 249
GE L +C E+ + + W+ T P LSE G + WP T S+ L
Sbjct: 502 GEPLNDTHCYSEEDKVLSRRIMRYWANFAKTGNPNLSEGGSSESITNWPERTDSKRHLVL 561
Query: 250 SPNHSLG 256
N S+G
Sbjct: 562 DVNESVG 568
>UniRef100_Q8VEG4 Protein C14orf114 homolog [Mus musculus]
Length = 496
Score = 35.4 bits (80), Expect = 1.6
Identities = 30/98 (30%), Positives = 44/98 (44%), Gaps = 5/98 (5%)
Query: 7 EGSKVDRGRMTALELWSGPEIPKEEWMLRSLVREMKNLELIPS--LREAFMVEGHITIQV 64
E +V G L S P KEE L +RE N ++I L EA +E I +
Sbjct: 373 ERRQVRSGARALLNAESLPAHRKEE--LLHALREFYNTDIITEEMLHEAASLETRIYNE- 429
Query: 65 RYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKP 102
Y+ ++ H E L L+Q WR F+D+++P
Sbjct: 430 SYIPHGLKVVQRHTEGGLRSLMQLESRWRQHFLDSMQP 467
>UniRef100_UPI0000369D47 UPI0000369D47 UniRef100 entry
Length = 496
Score = 34.7 bits (78), Expect = 2.8
Identities = 28/98 (28%), Positives = 46/98 (46%), Gaps = 5/98 (5%)
Query: 7 EGSKVDRGRMTALELWSGPEIPKEEWMLRSLVREMKNLELIPS--LREAFMVEGHITIQV 64
E +V G L S P KEE L +RE N +++ L+EA +E I+ +
Sbjct: 373 ERRQVRSGARALLNAESLPAHRKEE--LLQALREFYNTDVVTEEMLQEAASLETRISNE- 429
Query: 65 RYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKP 102
Y+ ++ H + L L+Q WR F+D+++P
Sbjct: 430 NYVPHGLKVVQCHSQGGLRSLMQLESRWRQHFLDSMQP 467
>UniRef100_Q9NVH0 Protein C14orf114 [Homo sapiens]
Length = 496
Score = 34.7 bits (78), Expect = 2.8
Identities = 28/98 (28%), Positives = 46/98 (46%), Gaps = 5/98 (5%)
Query: 7 EGSKVDRGRMTALELWSGPEIPKEEWMLRSLVREMKNLELIPS--LREAFMVEGHITIQV 64
E +V G L S P KEE L +RE N +++ L+EA +E I+ +
Sbjct: 373 ERRQVRSGARALLNAESLPTQRKEE--LLQALREFYNTDVVTEEMLQEAASLETRISNE- 429
Query: 65 RYLRDDRVLLTNHQESNLVELLQGSKGWRSEFVDNLKP 102
Y+ ++ H + L L+Q WR F+D+++P
Sbjct: 430 NYVPHGLKVVQCHSQGGLRSLMQLESRWRQHFLDSMQP 467
>UniRef100_Q9SJW1 Hypothetical protein At2g01050 [Arabidopsis thaliana]
Length = 515
Score = 34.3 bits (77), Expect = 3.6
Identities = 25/104 (24%), Positives = 45/104 (43%), Gaps = 9/104 (8%)
Query: 100 LKPWSSSFAP-----INRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WSS F P + WVR + IP + + + +G +K+D +T F
Sbjct: 147 VQDWSSRFDPLRDDIVTTPVWVRLSNIPYNYYHRCLLMEIARGLGRPLKVDMNTINFDKG 206
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNC 198
F+R+ + L +LI G Y ++ EG + + +C
Sbjct: 207 RFARVCIEVNLAK--PLKGTVLINGDRYFVAY--EGLSKICSSC 246
>UniRef100_Q753B3 AFR411Cp [Ashbya gossypii]
Length = 283
Score = 34.3 bits (77), Expect = 3.6
Identities = 30/121 (24%), Positives = 54/121 (43%), Gaps = 13/121 (10%)
Query: 159 LLVRTTTLNFITLARK----ILIKGTTYTISIIE-EGEALLHPNCCCREESS---VEDDV 210
L+ ++ L+ + LAR+ I++ G T+++ EG+ ++P C S+ ++DD
Sbjct: 101 LVPKSDPLSLLILARQLDVDIMLWGGTHSVEAYTLEGKFFINPGSCTGAFSTDWPLQDDA 160
Query: 211 FSGWS-----LNGVTVTPELSESGGEDDDGLWPVETKSQHDLQGSPNHSLGDNKNPRSNT 265
G LNG P+ +E D++GL + S + + LG N R
Sbjct: 161 LQGEGSPSAKLNGAATAPQPTEGEAGDENGLQQELSTSSRTPREAEARVLGSPNNRRELE 220
Query: 266 G 266
G
Sbjct: 221 G 221
>UniRef100_Q9XE43 Putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 855
Score = 33.5 bits (75), Expect = 6.2
Identities = 26/104 (25%), Positives = 44/104 (42%), Gaps = 9/104 (8%)
Query: 100 LKPWSSSFAPI-NRVT----WVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVL 154
++ WS F P+ + +T WVR IPL ++ + +G +K+D T
Sbjct: 122 VQAWSPEFDPLRDEITTTPIWVRLMNIPLSLYHTSILMGIAGSLGKPVKVDMTTLHVERA 181
Query: 155 EFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALLHPNC 198
F+R+ + L +L+ G Y +S EG A + C
Sbjct: 182 RFARMCIEVDLAK--PLKGTLLLNGERYFVSY--EGLANICSRC 221
>UniRef100_Q97MJ8 Uncharacterized membrane protein, ortholog YYAS B.subtilis
[Clostridium acetobutylicum]
Length = 227
Score = 33.1 bits (74), Expect = 8.1
Identities = 21/80 (26%), Positives = 40/80 (49%), Gaps = 1/80 (1%)
Query: 108 APINRVTWVRCTGIPLHMWTMDCFKHLLLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLN 167
+PI+ V + G PL M T ++ L VG VI + +D + +++ LV ++
Sbjct: 35 SPISSVPYTLSMGFPLTMGTFTFILNMFLIVGQVIFLGKDFKKVQLMQIPISLVFGYFID 94
Query: 168 FITLARKILIKGTTYTISII 187
F T++ ++ TTY ++
Sbjct: 95 F-TMSLLSVLNPTTYVFKLV 113
>UniRef100_Q65EQ2 Rnr [Bacillus licheniformis]
Length = 767
Score = 33.1 bits (74), Expect = 8.1
Identities = 36/107 (33%), Positives = 52/107 (47%), Gaps = 15/107 (14%)
Query: 135 LLQVGDVIKIDEDTSEFVVLEFSRLLVRTTTLNFITLARKILIKGTTYTISIIEEGEALL 194
+L++ D + E V LE L+VRT + + + LIKG +S +G A +
Sbjct: 30 MLEITDSDEYKELVKALVTLEEKGLVVRTRSNRYGLPEKMNLIKG---KVSAHAKGFAFV 86
Query: 195 HPNCCCREESSVEDDVF-----SGWSLNGVTVTPELS-ESGGEDDDG 235
P E+SS+ DDVF ++NG TV LS ESGG +G
Sbjct: 87 LP-----EDSSL-DDVFIPPSELNTAMNGDTVLVRLSTESGGTKKEG 127
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 469,471,973
Number of Sequences: 2790947
Number of extensions: 19056948
Number of successful extensions: 48537
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 48536
Number of HSP's gapped (non-prelim): 12
length of query: 278
length of database: 848,049,833
effective HSP length: 126
effective length of query: 152
effective length of database: 496,390,511
effective search space: 75451357672
effective search space used: 75451357672
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0102b.4


