

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0098b.3
(287 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
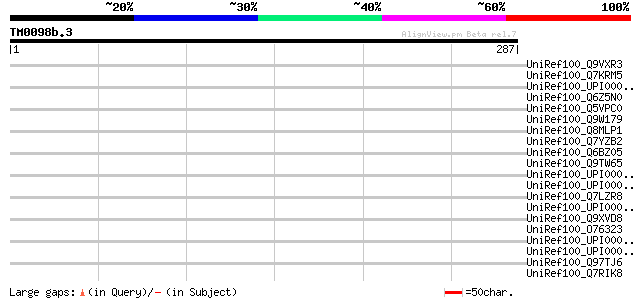
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9VXR3 CG8184-PB [Drosophila melanogaster] 42 0.024
UniRef100_Q7KRM5 CG33352-PA [Drosophila melanogaster] 37 0.78
UniRef100_UPI000021F26F UPI000021F26F UniRef100 entry 36 1.3
UniRef100_Q6Z5N0 Hypothetical protein P0684F11.29 [Oryza sativa] 35 1.7
UniRef100_Q5VPC0 Hypothetical protein OSJNBb0032K15.17 [Oryza sa... 35 1.7
UniRef100_Q9W179 CG4527-PB, isoform B [Drosophila melanogaster] 35 1.7
UniRef100_Q8MLP1 CG4527-PA, isoform A [Drosophila melanogaster] 35 1.7
UniRef100_Q7YZB2 Polo kinase kinase 1 [Drosophila melanogaster] 35 1.7
UniRef100_Q6BZ05 Debaryomyces hansenii chromosome A of strain CB... 35 1.7
UniRef100_Q9TW65 Dystrophin-1 [Caenorhabditis elegans] 35 1.7
UniRef100_UPI0000360560 UPI0000360560 UniRef100 entry 35 2.3
UniRef100_UPI0000342726 UPI0000342726 UniRef100 entry 35 2.3
UniRef100_Q7LZR8 RAD 23B protein [Ictalurus punctatus] 35 2.3
UniRef100_UPI000027C8D4 UPI000027C8D4 UniRef100 entry 35 3.0
UniRef100_Q9XVD8 Hypothetical protein C14A6.8 [Caenorhabditis el... 35 3.0
UniRef100_O76323 Synapsin s-syn-long [Loligo pealeii] 35 3.0
UniRef100_UPI000042F00D UPI000042F00D UniRef100 entry 34 3.9
UniRef100_UPI00002E750A UPI00002E750A UniRef100 entry 34 3.9
UniRef100_Q97TJ6 Arsenate reductase, arsC, protein-tyrosine-phos... 34 3.9
UniRef100_Q7RIK8 Hypothetical protein [Plasmodium yoelii yoelii] 34 3.9
>UniRef100_Q9VXR3 CG8184-PB [Drosophila melanogaster]
Length = 5146
Score = 41.6 bits (96), Expect = 0.024
Identities = 30/92 (32%), Positives = 44/92 (47%), Gaps = 4/92 (4%)
Query: 90 HLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPPPPPPSFTIATIVVV 149
H++EAL L + + N AS +L ++ S S + PPPPPPS + I V
Sbjct: 1485 HVIEALRTNASLEEATDYLLNNPEAS----SLSTTGGQSSSSAPPPPPPPSASTMDIDVD 1540
Query: 150 AIATGSITASNSTVVITNTHCHHHPPLMPLFI 181
A G T S S+ +++ + H LMP I
Sbjct: 1541 VPADGESTQSKSSTSPNSSYDYKHLKLMPSLI 1572
>UniRef100_Q7KRM5 CG33352-PA [Drosophila melanogaster]
Length = 1938
Score = 36.6 bits (83), Expect = 0.78
Identities = 24/95 (25%), Positives = 42/95 (43%), Gaps = 5/95 (5%)
Query: 86 TTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPPPPPPSFTIAT 145
TT + L + +L+K + +S +++S S S + PP PP+ +A
Sbjct: 1553 TTSFNALPHFPLSSSTSSLLSKVSSFSNSSSASPPTTAATSGSASSHYQPPQPPNAAVAN 1612
Query: 146 IVVVAIATGSIT-----ASNSTVVITNTHCHHHPP 175
+AI + S T A + + ++H HH PP
Sbjct: 1613 SKDMAIYSSSFTKNPAAAQSPNMRQAHSHQHHQPP 1647
>UniRef100_UPI000021F26F UPI000021F26F UniRef100 entry
Length = 220
Score = 35.8 bits (81), Expect = 1.3
Identities = 25/111 (22%), Positives = 45/111 (40%), Gaps = 4/111 (3%)
Query: 81 CESDSTTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPPPPPPS 140
C S +T V+H + P P + + +S + SS+++ L S PPPP
Sbjct: 112 CSSSTTAVLHH----SSPPPQQSSSSTAAVLHHSSPPPQQCSSSTTAVLHCSSAPPPPQQ 167
Query: 141 FTIATIVVVAIATGSITASNSTVVITNTHCHHHPPLMPLFITEVTHTNIAP 191
+ ++++ ++ +S+ H PPL T V H + P
Sbjct: 168 SSTTAVLLLHYSSPPPQQCSSSTTAVLHHSSPPPPLQQSSTTAVLHRSSPP 218
>UniRef100_Q6Z5N0 Hypothetical protein P0684F11.29 [Oryza sativa]
Length = 207
Score = 35.4 bits (80), Expect = 1.7
Identities = 26/78 (33%), Positives = 38/78 (48%), Gaps = 16/78 (20%)
Query: 114 ASQVDKALLSSSSSSLS-------------LSHPPPPPPSFTIATIVVVAIATGSITASN 160
+S A L+S++SS+S H PPPPP+ T A + + S ++S+
Sbjct: 42 SSSSTTARLTSATSSVSRHRHPQPRRRPSGCCHCPPPPPAPTSA---ASSSLSSSSSSSS 98
Query: 161 STVVITNTHCHHHPPLMP 178
S+ V H HHHPP P
Sbjct: 99 SSSVRHCRHRHHHPPPPP 116
>UniRef100_Q5VPC0 Hypothetical protein OSJNBb0032K15.17 [Oryza sativa]
Length = 342
Score = 35.4 bits (80), Expect = 1.7
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 7/51 (13%)
Query: 128 SLSLSHPPPPPPSFTIATIVVVAIATGSITASNSTVVITNTHCHHHPPLMP 178
S LS PPPPPP ++ T + A A S +A + CH H +P
Sbjct: 220 SRPLSQPPPPPPPSSLPTAIPTATAAASFSA-------PDRRCHRHCSRLP 263
>UniRef100_Q9W179 CG4527-PB, isoform B [Drosophila melanogaster]
Length = 1703
Score = 35.4 bits (80), Expect = 1.7
Identities = 32/119 (26%), Positives = 50/119 (41%), Gaps = 22/119 (18%)
Query: 82 ESDSTTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLS--------- 132
ESD + +A P PL KP +D + K +S ++ +
Sbjct: 465 ESDKKHFVKKGKAPPPPSPLGLANAKPAASDSQTSPKKLATPEPTSPVTTAIEVAIGQEA 524
Query: 133 -HPPPPPPSFTIATIV-VVAIATGSITASNSTVVITNTHC-----------HHHPPLMP 178
P P PPS T ++IV V ++A+ S + S S V++++ HHH PL P
Sbjct: 525 MEPKPQPPSPTASSIVSVQSVASSSSSGSVSNAVLSSSTSLITINSDPPTPHHHQPLPP 583
>UniRef100_Q8MLP1 CG4527-PA, isoform A [Drosophila melanogaster]
Length = 1300
Score = 35.4 bits (80), Expect = 1.7
Identities = 32/119 (26%), Positives = 50/119 (41%), Gaps = 22/119 (18%)
Query: 82 ESDSTTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLS--------- 132
ESD + +A P PL KP +D + K +S ++ +
Sbjct: 465 ESDKKHFVKKGKAPPPPSPLGLANAKPAASDSQTSPKKLATPEPTSPVTTAIEVAIGQEA 524
Query: 133 -HPPPPPPSFTIATIV-VVAIATGSITASNSTVVITNTHC-----------HHHPPLMP 178
P P PPS T ++IV V ++A+ S + S S V++++ HHH PL P
Sbjct: 525 MEPKPQPPSPTASSIVSVQSVASSSSSGSVSNAVLSSSTSLITINSDPPTPHHHQPLPP 583
>UniRef100_Q7YZB2 Polo kinase kinase 1 [Drosophila melanogaster]
Length = 1342
Score = 35.4 bits (80), Expect = 1.7
Identities = 32/119 (26%), Positives = 50/119 (41%), Gaps = 22/119 (18%)
Query: 82 ESDSTTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLS--------- 132
ESD + +A P PL KP +D + K +S ++ +
Sbjct: 465 ESDKKHFVKKGKAPPPPSPLGLANAKPAASDSQTSPKKLATPEPTSPVTTAIEVAIGQEA 524
Query: 133 -HPPPPPPSFTIATIV-VVAIATGSITASNSTVVITNTHC-----------HHHPPLMP 178
P P PPS T ++IV V ++A+ S + S S V++++ HHH PL P
Sbjct: 525 MEPKPQPPSPTASSIVSVQSVASSSSSGSVSNAVLSSSTSLITINSDPPTPHHHQPLPP 583
>UniRef100_Q6BZ05 Debaryomyces hansenii chromosome A of strain CBS767 of Debaryomyces
hansenii [Debaryomyces hansenii]
Length = 1152
Score = 35.4 bits (80), Expect = 1.7
Identities = 24/85 (28%), Positives = 37/85 (43%), Gaps = 15/85 (17%)
Query: 114 ASQVDKALLSSSSSSLSLSHPPPPPPSFTIATIV----------VVAIATGSITASNSTV 163
+ + LL S ++S S S PPPPPPS + V +++ + S T S +
Sbjct: 1016 SKSISNPLLRSPTASTSASTPPPPPPSRKVGNPVKPPIGFSSTPLISSRSNSATPSRGSP 1075
Query: 164 VIT-----NTHCHHHPPLMPLFITE 183
+ T N HP L P+ T+
Sbjct: 1076 ISTPSTTGNEQNQEHPKLNPIVPTK 1100
>UniRef100_Q9TW65 Dystrophin-1 [Caenorhabditis elegans]
Length = 3674
Score = 35.4 bits (80), Expect = 1.7
Identities = 17/65 (26%), Positives = 37/65 (56%), Gaps = 3/65 (4%)
Query: 214 KVSLDDVVRWSVPFLMVERTMVQGLTDREGKPEQEFIGRNKFVKEKVQEKSEDRIEVDIS 273
K+ LD+VVRW M E+ Q + +G ++ GR +++QE+ +D ++++++
Sbjct: 2181 KLELDEVVRWCE---MAEKEAAQNVNSLDGDGLEKLDGRLAQFTKELQERKDDMVQLEMA 2237
Query: 274 YSTIV 278
+ I+
Sbjct: 2238 KNMII 2242
>UniRef100_UPI0000360560 UPI0000360560 UniRef100 entry
Length = 167
Score = 35.0 bits (79), Expect = 2.3
Identities = 35/119 (29%), Positives = 44/119 (36%), Gaps = 23/119 (19%)
Query: 91 LLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPPPPPPSF--------- 141
LL A P PL L+ P + L SSSSS S + PPPPPPS
Sbjct: 58 LLPPWAHPSPLLLSLSLPPLPS------PPLRPSSSSSSSTTPPPPPPPSLRHKYTHTHT 111
Query: 142 ---TIATIVVVAIATGSITASNSTVVITNTHCHHHPPLMPLFITEVTHTNIAPRKHVTS 197
T ++ A T T+TH H H TH+N ++H S
Sbjct: 112 HTHTHTHTHILTQPRTHAPAHTHTHTHTHTHTHTH-----THTATHTHSNTRTQQHTHS 165
>UniRef100_UPI0000342726 UPI0000342726 UniRef100 entry
Length = 297
Score = 35.0 bits (79), Expect = 2.3
Identities = 28/98 (28%), Positives = 45/98 (45%), Gaps = 2/98 (2%)
Query: 35 IIRYSFSEWVASFSSFFGGSCSLYVELLAKKHGLLLDINLGYRWVICESDSTTVIHLLEA 94
II F + +ASF S G + ++A +G +IN +RW E I L
Sbjct: 107 IINIIFLKEIASFISILGCFIVFFGVVIAIAYGDRKNIN--HRWETIEGSLKIGIFLAIC 164
Query: 95 LAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLS 132
A+ + V+ KP +N A + A + ++SS + LS
Sbjct: 165 AALCQAIGVVMMKPILNQGADPIASAAIRTASSCVLLS 202
>UniRef100_Q7LZR8 RAD 23B protein [Ictalurus punctatus]
Length = 385
Score = 35.0 bits (79), Expect = 2.3
Identities = 24/83 (28%), Positives = 35/83 (41%)
Query: 107 KPTINDMASQVDKALLSSSSSSLSLSHPPPPPPSFTIATIVVVAIATGSITASNSTVVIT 166
KP A+Q SSSSS+ S + P PP + + AT T + + S S+V+
Sbjct: 76 KPKAATAAAQSSTTAASSSSSTSSTTTPTVPPVAASAATTTTTTTTTTTDSTSESSVIEE 135
Query: 167 NTHCHHHPPLMPLFITEVTHTNI 189
P P +T+ NI
Sbjct: 136 KAAEEKPPSSTPASSGSLTNVNI 158
>UniRef100_UPI000027C8D4 UPI000027C8D4 UniRef100 entry
Length = 580
Score = 34.7 bits (78), Expect = 3.0
Identities = 23/68 (33%), Positives = 31/68 (44%), Gaps = 5/68 (7%)
Query: 77 RWVICESDSTTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSS-----SSLSL 131
RW ++HLLE L + P +LN + + + V +L SS S L
Sbjct: 55 RWGSRNGRVRDLLHLLEGLELLRPRDLILNGQSSPPLRTPVTSSLRPDSSPFEGVSCLKP 114
Query: 132 SHPPPPPP 139
S PPPPPP
Sbjct: 115 SPPPPPPP 122
>UniRef100_Q9XVD8 Hypothetical protein C14A6.8 [Caenorhabditis elegans]
Length = 129
Score = 34.7 bits (78), Expect = 3.0
Identities = 16/64 (25%), Positives = 32/64 (50%)
Query: 97 MPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPPPPPPSFTIATIVVVAIATGSI 156
M P +D N +D + A+L ++ ++ + PPPPPP T+A +++ +
Sbjct: 1 MGDPFFDPTNNLLKSDGNEYQNLAILDPNAPAVGGNEPPPPPPPLTVAQDALISDNGAKV 60
Query: 157 TASN 160
T+ +
Sbjct: 61 TSGD 64
>UniRef100_O76323 Synapsin s-syn-long [Loligo pealeii]
Length = 503
Score = 34.7 bits (78), Expect = 3.0
Identities = 31/125 (24%), Positives = 53/125 (41%), Gaps = 8/125 (6%)
Query: 18 SGLWMKMQSSNLMSVDDIIRYSFSEWVASFSSFFGGSCSLYVELLAKKHG--LLLDINLG 75
SG W K + + M + + WV S FGG + VE L K G ++++N
Sbjct: 309 SGNW-KANTGSAMLEQIQMNEKYKLWVDECSQLFGGLDIVAVEALQGKDGREYIIEVNDS 367
Query: 76 YRWVICESDSTTVIHLLEALAMPHPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPP 135
++ E+ + E + +Y KP N M+ + + S++ S + PP
Sbjct: 368 SMALLGETQEEDRRLIAEMVLQKMHMYC---KP--NTMSQAMSSGTIQSAADSTATPPPP 422
Query: 136 PPPPS 140
PP P+
Sbjct: 423 PPRPA 427
>UniRef100_UPI000042F00D UPI000042F00D UniRef100 entry
Length = 893
Score = 34.3 bits (77), Expect = 3.9
Identities = 21/43 (48%), Positives = 25/43 (57%), Gaps = 2/43 (4%)
Query: 105 LNKPTINDM--ASQVDKALLSSSSSSLSLSHPPPPPPSFTIAT 145
LNKP+ +S + LLS SS S SLS PPP PP+ AT
Sbjct: 816 LNKPSSRSPRESSILSHPLLSGSSQSSSLSPPPPTPPNTQRAT 858
>UniRef100_UPI00002E750A UPI00002E750A UniRef100 entry
Length = 289
Score = 34.3 bits (77), Expect = 3.9
Identities = 30/113 (26%), Positives = 47/113 (41%), Gaps = 21/113 (18%)
Query: 53 GSCSLYVELLAKKHGLLLDINLGYRWVICESDSTTVIHLLEALAMPHPLYDVLNKPTIND 112
G C+ + ++ G +L + L E D TV + + P P L P ++
Sbjct: 146 GKCNSEMPVIGSFIGAVLPLKL------VELDGLTVADICPSPPPPFPPPPPLPPPPVSS 199
Query: 113 MASQVDKALLSSSSSSLSLSHPPPPPPSFTIATIVVVAIATGSITASNSTVVI 165
SSSSSS S S P PPPP + + A+GS++ + T +
Sbjct: 200 S---------SSSSSSSSSSSPSPPPP------VTLTLRASGSVSDYSDTTAL 237
>UniRef100_Q97TJ6 Arsenate reductase, arsC, protein-tyrosine-phosphatase family
enzyme [Clostridium acetobutylicum]
Length = 136
Score = 34.3 bits (77), Expect = 3.9
Identities = 17/47 (36%), Positives = 29/47 (61%)
Query: 226 PFLMVERTMVQGLTDREGKPEQEFIGRNKFVKEKVQEKSEDRIEVDI 272
PF++ + T GL D GK ++EFI K ++EKV++ ++ I +I
Sbjct: 88 PFVLSKHTEDWGLDDPSGKSDEEFIRTAKTIEEKVKDLAKRIINKEI 134
>UniRef100_Q7RIK8 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 847
Score = 34.3 bits (77), Expect = 3.9
Identities = 43/191 (22%), Positives = 73/191 (37%), Gaps = 24/191 (12%)
Query: 99 HPLYDVLNKPTINDMASQVDKALLSSSSSSLSLSHPPPPPPSFTIATIVV-----VAIAT 153
HP+ L + N + KAL P PP AT A T
Sbjct: 333 HPIQPKLQLESHNKENETLQKALSEIK---------PQSPPEAAAATTTTPSPSTAATTT 383
Query: 154 GSITASNSTVVITNT-----HCHHHPPLMP--LFITEVTHTNIAPRKHVTSGADKERLVR 206
S + + +T +I +T H HH P P + I +V H I P+ + +G D ++
Sbjct: 384 PSPSTATTTPIIPSTEVSSQHPHHQQPKPPQQVQINQVIHEPIQPQHTMKNGIDSYFILN 443
Query: 207 GTEAGVPKVSLDDVVRWSVPFLMVERTMVQGLTDREGKPEQEFIGRNKFVKEKVQEKSED 266
E+ P S D + P + E ++ + ++ K + +N ++ V + S
Sbjct: 444 PDESN-PNGSKDQLKHEKKPEIPKENSIQE--VEKYPKTNENDNEQNTNLQNSVDQTSIQ 500
Query: 267 RIEVDISYSTI 277
E +S S +
Sbjct: 501 TKESSVSGSAL 511
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 477,093,089
Number of Sequences: 2790947
Number of extensions: 19650704
Number of successful extensions: 108059
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 107872
Number of HSP's gapped (non-prelim): 133
length of query: 287
length of database: 848,049,833
effective HSP length: 126
effective length of query: 161
effective length of database: 496,390,511
effective search space: 79918872271
effective search space used: 79918872271
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0098b.3


