

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0096b.3
(223 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
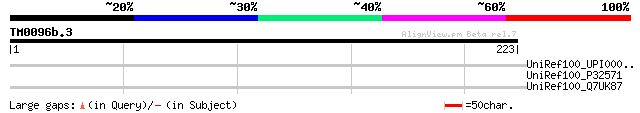
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI0000298CD9 UPI0000298CD9 UniRef100 entry 35 1.4
UniRef100_P32571 Ubiquitin carboxyl-terminal hydrolase 4 [Saccha... 34 3.2
UniRef100_Q7UK87 Hypothetical protein [Rhodopirellula baltica] 32 9.3
>UniRef100_UPI0000298CD9 UPI0000298CD9 UniRef100 entry
Length = 152
Score = 35.0 bits (79), Expect = 1.4
Identities = 26/74 (35%), Positives = 37/74 (49%), Gaps = 3/74 (4%)
Query: 45 GLSIQTRGMLSPHIHHRMQDVGALVELDPGTPVPEVLALDSSLILPDALKFVSVMTKAIR 104
G SI G++S + +G L+E+ P VLA +S+ PDAL +SV
Sbjct: 23 GTSIAWIGVISHFMVEWCARIGCLLEIPPVVMGTTVLAAGTSI--PDALSSISVAKDGYA 80
Query: 105 DLIIAKQVGV-VFD 117
D+ +A VG VFD
Sbjct: 81 DMAVANAVGSNVFD 94
>UniRef100_P32571 Ubiquitin carboxyl-terminal hydrolase 4 [Saccharomyces cerevisiae]
Length = 926
Score = 33.9 bits (76), Expect = 3.2
Identities = 17/37 (45%), Positives = 23/37 (61%), Gaps = 4/37 (10%)
Query: 28 VVDDVVYHLQLLSGNTNGLSIQTRGMLSPHIHHRMQD 64
+ DD VY ++GNT+GLS+Q +SP I H M D
Sbjct: 338 IKDDSVY----INGNTSGLSLQHLPKMSPSIRHSMDD 370
>UniRef100_Q7UK87 Hypothetical protein [Rhodopirellula baltica]
Length = 487
Score = 32.3 bits (72), Expect = 9.3
Identities = 19/54 (35%), Positives = 29/54 (53%), Gaps = 3/54 (5%)
Query: 16 VLVRMARVVSLVVVDDVVYHLQLLSGNT---NGLSIQTRGMLSPHIHHRMQDVG 66
VL AR++ L + DD L +LSG T +G S GM+ +H R+ ++G
Sbjct: 384 VLAAYARLLELNLTDDQWSRLAILSGGTDGEDGPSDAAGGMIDGDVHRRIMELG 437
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.142 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 373,227,365
Number of Sequences: 2790947
Number of extensions: 15283091
Number of successful extensions: 33774
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 33774
Number of HSP's gapped (non-prelim): 3
length of query: 223
length of database: 848,049,833
effective HSP length: 123
effective length of query: 100
effective length of database: 504,763,352
effective search space: 50476335200
effective search space used: 50476335200
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0096b.3


