

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0095c.6
(384 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
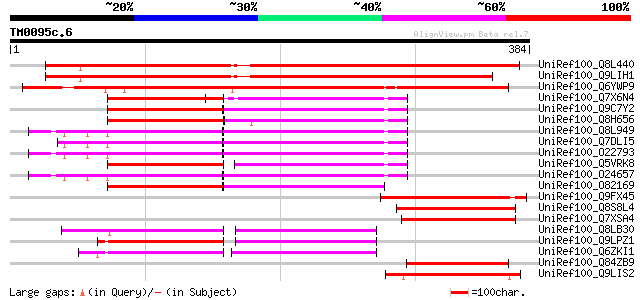
UniRef100_Q8L440 Hypothetical protein At3g20930 [Arabidopsis tha... 419 e-116
UniRef100_Q9LIH1 Arabidopsis thaliana genomic DNA, chromosome 3,... 387 e-106
UniRef100_Q6YWP9 RNA recognition motif (RRM)-containing protein-... 374 e-102
UniRef100_Q7X6N4 OSJNBa0041A02.8 protein [Oryza sativa] 104 4e-21
UniRef100_Q9C7Y2 Plastid protein, putative; 23108-24430 [Arabido... 103 1e-20
UniRef100_Q8H656 Putative plastid protein [Oryza sativa] 101 4e-20
UniRef100_Q8L949 Plastid protein [Arabidopsis thaliana] 99 2e-19
UniRef100_Q7DLI5 Plastid protein [Arabidopsis thaliana] 99 2e-19
UniRef100_O22793 Plastid protein [Arabidopsis thaliana] 99 2e-19
UniRef100_Q5VRK8 Putative DAL1 protein [Oryza sativa] 99 2e-19
UniRef100_O24657 DAL1 protein [Arabidopsis thaliana] 98 4e-19
UniRef100_O82169 Hypothetical protein At2g35240 [Arabidopsis tha... 94 6e-18
UniRef100_Q9FX45 RNA-binding glycine-rich protein, putative [Ara... 89 2e-16
UniRef100_Q8S8L4 Putative RNA-binding protein [Arabidopsis thali... 82 2e-14
UniRef100_Q7XSA4 OJ000126_13.13 protein [Oryza sativa] 81 6e-14
UniRef100_Q8LB30 DAG protein, putative [Arabidopsis thaliana] 77 6e-13
UniRef100_Q9LPZ1 T23J18.10 [Arabidopsis thaliana] 77 8e-13
UniRef100_Q6ZKI1 Putative DAG protein [Oryza sativa] 76 1e-12
UniRef100_Q84ZB9 RNA recognition motif (RRM)-containing protein-... 76 2e-12
UniRef100_Q9LIS2 Glycine-rich RNA binding protein-like [Arabidop... 75 2e-12
>UniRef100_Q8L440 Hypothetical protein At3g20930 [Arabidopsis thaliana]
Length = 374
Score = 419 bits (1078), Expect = e-116
Identities = 216/373 (57%), Positives = 276/373 (73%), Gaps = 32/373 (8%)
Query: 27 LPNKSYHSNLPSSSSSSISCNRFFT---------------------IIAAAAATTYPSFT 65
+ + S+ + +SSSS I NRFF ++ +++A + P +
Sbjct: 5 IASTSFFVPISNSSSSHIINNRFFPSFYSPNLNFGTFRKTSLSSSHLVFSSSAISAPPSS 64
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
T N +WMVL+DKPP V SKS ++DYYV+ L V+G+EK+AQ+ +YDAS+DTHFGFC
Sbjct: 65 TVLTRNSYWMVLLDKPPHWVSSKSAMVDYYVEILAKVLGNEKDAQVSIYDASFDTHFGFC 124
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSK 185
C IDE+ S QLASLPGV+S+RP+ D++S KK+Y + S + G+S LF G K
Sbjct: 125 CHIDEDASRQLASLPGVVSIRPEQDYSSEKKNYGIGSHK-GVS---------LFDHGTVK 174
Query: 186 HWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDC 245
HW+VR+DKPGV +VTKAQ+VD+ Q+L+KV+ NEKDAQMC+YHVSW+++FGFCC+LDE
Sbjct: 175 HWMVRIDKPGVGIVTKAQMVDHCVQLLSKVLWNEKDAQMCLYHVSWQSDFGFCCDLDERS 234
Query: 246 AQDLAGVPGVSSVQPDDNFESENKNYE-DSLSVPNSSEASQEASLRTKKLFVTGLSFYTS 304
A +LAGVPGV +V PD++FES NK+YE DS + S+ ++TKKLF+TGLSFYTS
Sbjct: 235 AVELAGVPGVLAVVPDNSFESLNKDYEGDSTQDSRDQDDSESPPVKTKKLFITGLSFYTS 294
Query: 305 EKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGKIINGWMIV 364
EKTLRAAFEGFGELV+VK+I+DKISKRSKGYAF+EYTTEEAAG ALKEMNGKIINGWMIV
Sbjct: 295 EKTLRAAFEGFGELVEVKIIMDKISKRSKGYAFLEYTTEEAAGTALKEMNGKIINGWMIV 354
Query: 365 VDVAKTNPPRFNR 377
VDVAKT P R NR
Sbjct: 355 VDVAKTKPFRQNR 367
>UniRef100_Q9LIH1 Arabidopsis thaliana genomic DNA, chromosome 3, P1 clone:MFD22
[Arabidopsis thaliana]
Length = 396
Score = 387 bits (993), Expect = e-106
Identities = 198/353 (56%), Positives = 259/353 (73%), Gaps = 32/353 (9%)
Query: 27 LPNKSYHSNLPSSSSSSISCNRFFT---------------------IIAAAAATTYPSFT 65
+ + S+ + +SSSS I NRFF ++ +++A + P +
Sbjct: 5 IASTSFFVPISNSSSSHIINNRFFPSFYSPNLNFGTFRKTSLSSSHLVFSSSAISAPPSS 64
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
T N +WMVL+DKPP V SKS ++DYYV+ L V+G+EK+AQ+ +YDAS+DTHFGFC
Sbjct: 65 TVLTRNSYWMVLLDKPPHWVSSKSAMVDYYVEILAKVLGNEKDAQVSIYDASFDTHFGFC 124
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSK 185
C IDE+ S QLASLPGV+S+RP+ D++S KK+Y + S + G+S LF G K
Sbjct: 125 CHIDEDASRQLASLPGVVSIRPEQDYSSEKKNYGIGSHK-GVS---------LFDHGTVK 174
Query: 186 HWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDC 245
HW+VR+DKPGV +VTKAQ+VD+ Q+L+KV+ NEKDAQMC+YHVSW+++FGFCC+LDE
Sbjct: 175 HWMVRIDKPGVGIVTKAQMVDHCVQLLSKVLWNEKDAQMCLYHVSWQSDFGFCCDLDERS 234
Query: 246 AQDLAGVPGVSSVQPDDNFESENKNYE-DSLSVPNSSEASQEASLRTKKLFVTGLSFYTS 304
A +LAGVPGV +V PD++FES NK+YE DS + S+ ++TKKLF+TGLSFYTS
Sbjct: 235 AVELAGVPGVLAVVPDNSFESLNKDYEGDSTQDSRDQDDSESPPVKTKKLFITGLSFYTS 294
Query: 305 EKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGKI 357
EKTLRAAFEGFGELV+VK+I+DKISKRSKGYAF+EYTTEEAAG ALKEMNGK+
Sbjct: 295 EKTLRAAFEGFGELVEVKIIMDKISKRSKGYAFLEYTTEEAAGTALKEMNGKV 347
>UniRef100_Q6YWP9 RNA recognition motif (RRM)-containing protein-like [Oryza sativa]
Length = 399
Score = 374 bits (961), Expect = e-102
Identities = 199/368 (54%), Positives = 264/368 (71%), Gaps = 18/368 (4%)
Query: 10 SVNPNRSQVHQLPTNITLPNKSYHSNLPSSSSSSISCNRFFTIIAAAAATTYPSFTPSTT 69
+V+ + S V +++P+ S H+ R F +A+AAA++ P ++
Sbjct: 15 AVSTSTSAVATAAQTVSIPHLSPHTRRRRQ--------RRFLRLASAAASSPPPLPAASA 66
Query: 70 --HNRHWMVLMDKPPL----GVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFG 123
H W+V+M++PP G +S+++ +D+YV TL V+GS++EAQM +YDASWD +
Sbjct: 67 QPHCSRWVVVMERPPAPAGGGEVSRAEAVDHYVATLARVLGSQEEAQMRIYDASWDGSYE 126
Query: 124 FCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDYSLSSG--QAGLSSGLQTRTNMLFPA 181
F C+ID+E S LA +PGVL+V+PD D + + + SG A L + +N +
Sbjct: 127 FSCEIDDEASRDLAKMPGVLAVKPDTDKVDMSEKDNHGSGLSAANLGNFSDAVSNHSSSS 186
Query: 182 GNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCEL 241
G ++ WLVRM+KPGVEVVTKAQ+VD+Y Q L KV+GNEKDAQ+ IYH+SW+ ++GFCC +
Sbjct: 187 GENEFWLVRMEKPGVEVVTKAQMVDHYTQTLMKVLGNEKDAQVSIYHISWERDYGFCCHI 246
Query: 242 DEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVPNSSEASQEASLRTKKLFVTGLSF 301
DE+CA++LA V GV SVQPD NF S+NKNY+ S SSEA+Q A ++TK+LFVTGLSF
Sbjct: 247 DEECAKELADVSGVLSVQPDTNFGSDNKNYKGDDSF-KSSEATQ-AEVKTKRLFVTGLSF 304
Query: 302 YTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEMNGKIINGW 361
YTSEKTLRAAFE FGELV+VK+I+DKISKRSKGYAFIEYTTEEA GAALK MNG+IINGW
Sbjct: 305 YTSEKTLRAAFEPFGELVEVKIIMDKISKRSKGYAFIEYTTEEAGGAALKAMNGQIINGW 364
Query: 362 MIVVDVAK 369
MIVVDVAK
Sbjct: 365 MIVVDVAK 372
>UniRef100_Q7X6N4 OSJNBa0041A02.8 protein [Oryza sativa]
Length = 223
Score = 104 bits (260), Expect = 4e-21
Identities = 60/149 (40%), Positives = 88/149 (58%), Gaps = 10/149 (6%)
Query: 146 RPDPDFNSLKKDYSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIV 205
RP+ ++ L+ SGQ G T LFP + +HWL+ MDKPG E TK Q++
Sbjct: 55 RPESSYSPLR------SGQGG--DRAPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMI 106
Query: 206 DYYAQILTKVMGNEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFE 265
D Y Q L KV+G+E++A+ IY+VS + FGF CE+DE+ + L G+PGV V PD +
Sbjct: 107 DCYIQTLAKVVGSEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVD 166
Query: 266 SENKNYEDSLSVPNSSEASQEASLRTKKL 294
+ENK+Y L V + E Q + R +++
Sbjct: 167 AENKDYGAELFV--NGEIVQRSPERQRRV 193
Score = 89.4 bits (220), Expect = 2e-16
Identities = 42/86 (48%), Positives = 61/86 (70%)
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++MDKP +K Q+ID Y++TL V+GSE+EA+ +Y+ S + +FGF C+IDEE
Sbjct: 87 HWLIVMDKPGGEGATKQQMIDCYIQTLAKVVGSEEEAKKKIYNVSCERYFGFGCEIDEET 146
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL V PD ++ KDY
Sbjct: 147 SNKLEGLPGVLFVLPDSYVDAENKDY 172
>UniRef100_Q9C7Y2 Plastid protein, putative; 23108-24430 [Arabidopsis thaliana]
Length = 229
Score = 103 bits (256), Expect = 1e-20
Identities = 55/137 (40%), Positives = 79/137 (57%), Gaps = 2/137 (1%)
Query: 158 YSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
YS + S T LFP + +HWL+ MDKPG E TK Q++D Y Q L K++G
Sbjct: 67 YSPLKSGSNFSDRAPTEMAPLFPGCDYEHWLIVMDKPGGENATKQQMIDCYVQTLAKIIG 126
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + FGF CE+DE+ + L G+PGV + PD + ENK+Y L V
Sbjct: 127 SEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFILPDSYVDQENKDYGAELFV 186
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q R +K+
Sbjct: 187 --NGEIVQRPPERQRKI 201
Score = 88.6 bits (218), Expect = 3e-16
Identities = 41/86 (47%), Positives = 60/86 (69%)
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++MDKP +K Q+ID YV+TL ++GSE+EA+ +Y+ S + +FGF C+IDEE
Sbjct: 95 HWLIVMDKPGGENATKQQMIDCYVQTLAKIIGSEEEAKKKIYNVSCERYFGFGCEIDEET 154
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL + PD + KDY
Sbjct: 155 SNKLEGLPGVLFILPDSYVDQENKDY 180
>UniRef100_Q8H656 Putative plastid protein [Oryza sativa]
Length = 227
Score = 101 bits (251), Expect = 4e-20
Identities = 57/137 (41%), Positives = 82/137 (59%), Gaps = 4/137 (2%)
Query: 160 LSSGQAGLSSGLQTRTNM--LFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
+ SG G G + T M LFP + +HWL+ MDKPG E TK Q++D Y Q L KV+G
Sbjct: 66 MRSGGGGGGGGDRAPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVLG 125
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + FGF CE+DE+ + L G+PGV V PD + E K+Y L V
Sbjct: 126 SEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVDPEYKDYGAELFV 185
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 186 --NGEIVQRSPERQRRV 200
Score = 88.2 bits (217), Expect = 4e-16
Identities = 42/86 (48%), Positives = 60/86 (68%)
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++MDKP +K Q+ID Y++TL V+GSE+EA+ +Y+ S + +FGF C+IDEE
Sbjct: 94 HWLIVMDKPGGEGATKQQMIDCYIQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDEET 153
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL V PD + KDY
Sbjct: 154 SNKLEGLPGVLFVLPDSYVDPEYKDY 179
>UniRef100_Q8L949 Plastid protein [Arabidopsis thaliana]
Length = 219
Score = 99.4 bits (246), Expect = 2e-19
Identities = 55/137 (40%), Positives = 80/137 (58%), Gaps = 2/137 (1%)
Query: 158 YSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
YS + + S T LFP + +HWL+ MDKPG E TK Q++D Y Q L KV+G
Sbjct: 58 YSPLNSGSNFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVVG 117
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + GF CE+DE+ + L G+PGV V PD + ENK+Y L V
Sbjct: 118 SEEEAKKRIYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDYGAELFV 177
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 178 --NGEIVQRSPERQRRV 192
Score = 87.0 bits (214), Expect = 8e-16
Identities = 58/166 (34%), Positives = 86/166 (50%), Gaps = 25/166 (15%)
Query: 15 RSQVHQLPTNITLPNKSYHSNLPSS---SSSSISCN-RFFTIIAAA--AATTYPSFTPST 68
R L +N+TL +PSS S + C+ RFF+I A + +TY +
Sbjct: 9 RHLTRALLSNVTLMAPP---RIPSSVHYGGSRLGCSTRFFSIRCGANRSGSTYSPLNSGS 65
Query: 69 THN----------------RHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMC 112
+ HW+++MDKP +K Q+ID Y++TL V+GSE+EA+
Sbjct: 66 NFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVVGSEEEAKKR 125
Query: 113 LYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
+Y+ S + + GF C+IDEE S++L LPGVL V PD + KDY
Sbjct: 126 IYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDY 171
>UniRef100_Q7DLI5 Plastid protein [Arabidopsis thaliana]
Length = 198
Score = 99.4 bits (246), Expect = 2e-19
Identities = 55/137 (40%), Positives = 80/137 (58%), Gaps = 2/137 (1%)
Query: 158 YSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
YS + + S T LFP + +HWL+ MDKPG E TK Q++D Y Q L KV+G
Sbjct: 37 YSPLNSGSNFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVVG 96
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + GF CE+DE+ + L G+PGV V PD + ENK+Y L V
Sbjct: 97 SEEEAKKRIYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDYGAELFV 156
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 157 --NGEIVQRSPERQRRV 171
Score = 86.7 bits (213), Expect = 1e-15
Identities = 53/145 (36%), Positives = 79/145 (53%), Gaps = 22/145 (15%)
Query: 36 LPSS---SSSSISCN-RFFTIIAAA--AATTYPSFTPSTTHN----------------RH 73
+PSS S + C+ RFF+I A + +TY + + H
Sbjct: 6 IPSSVHYGGSRLGCSTRFFSIRCGANRSGSTYSPLNSGSNFSDRPPTEMAPLFPGCDYEH 65
Query: 74 WMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEIS 133
W+++MDKP +K Q+ID Y++TL V+GSE+EA+ +Y+ S + + GF C+IDEE S
Sbjct: 66 WLIVMDKPGGEGATKQQMIDCYIQTLAKVVGSEEEAKKRIYNVSCERYLGFGCEIDEETS 125
Query: 134 SQLASLPGVLSVRPDPDFNSLKKDY 158
++L LPGVL V PD + KDY
Sbjct: 126 TKLEGLPGVLFVLPDSYVDPENKDY 150
>UniRef100_O22793 Plastid protein [Arabidopsis thaliana]
Length = 219
Score = 99.4 bits (246), Expect = 2e-19
Identities = 55/137 (40%), Positives = 80/137 (58%), Gaps = 2/137 (1%)
Query: 158 YSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
YS + + S T LFP + +HWL+ MDKPG E TK Q++D Y Q L KV+G
Sbjct: 58 YSPLNSGSNFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVVG 117
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + GF CE+DE+ + L G+PGV V PD + ENK+Y L V
Sbjct: 118 SEEEAKKRIYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDYGAELFV 177
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 178 --NGEIVQRSPERQRRV 192
Score = 87.0 bits (214), Expect = 8e-16
Identities = 58/166 (34%), Positives = 86/166 (50%), Gaps = 25/166 (15%)
Query: 15 RSQVHQLPTNITLPNKSYHSNLPSS---SSSSISCN-RFFTIIAAA--AATTYPSFTPST 68
R L +N+TL +PSS S + C+ RFF+I A + +TY +
Sbjct: 9 RHLTRALLSNVTLMAPP---RIPSSVHYGGSRLGCSTRFFSIRCGANRSGSTYSPLNSGS 65
Query: 69 THN----------------RHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMC 112
+ HW+++MDKP +K Q+ID Y++TL V+GSE+EA+
Sbjct: 66 NFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVVGSEEEAKKR 125
Query: 113 LYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
+Y+ S + + GF C+IDEE S++L LPGVL V PD + KDY
Sbjct: 126 IYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDY 171
>UniRef100_Q5VRK8 Putative DAL1 protein [Oryza sativa]
Length = 165
Score = 99.0 bits (245), Expect = 2e-19
Identities = 54/128 (42%), Positives = 77/128 (59%), Gaps = 2/128 (1%)
Query: 167 LSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCI 226
LS T LFP + +HWL+ MDKPG E TK Q++D Y Q L KV+G+E++A+ I
Sbjct: 13 LSRRAPTEMAPLFPGCDYEHWLIVMDKPGGEGATKQQMIDCYIQTLAKVLGSEEEAKKKI 72
Query: 227 YHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSVPNSSEASQE 286
Y+VS + FGF CE+DE+ + L G+PGV V PD + E K+Y L V + E Q
Sbjct: 73 YNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVDPEYKDYGAELFV--NGEIVQR 130
Query: 287 ASLRTKKL 294
+ R +++
Sbjct: 131 SPERQRRV 138
Score = 88.2 bits (217), Expect = 4e-16
Identities = 42/86 (48%), Positives = 60/86 (68%)
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++MDKP +K Q+ID Y++TL V+GSE+EA+ +Y+ S + +FGF C+IDEE
Sbjct: 32 HWLIVMDKPGGEGATKQQMIDCYIQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDEET 91
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL V PD + KDY
Sbjct: 92 SNKLEGLPGVLFVLPDSYVDPEYKDY 117
>UniRef100_O24657 DAL1 protein [Arabidopsis thaliana]
Length = 219
Score = 97.8 bits (242), Expect = 4e-19
Identities = 54/137 (39%), Positives = 80/137 (57%), Gaps = 2/137 (1%)
Query: 158 YSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
YS + + S T LFP + +HWL+ MDKPG E TK +++D Y Q L KV+G
Sbjct: 58 YSPLNSGSNFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKHEMIDCYIQTLAKVVG 117
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + GF CE+DE+ + L G+PGV V PD + ENK+Y L V
Sbjct: 118 SEEEAKKRIYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDYGAELFV 177
Query: 278 PNSSEASQEASLRTKKL 294
+ E Q + R +++
Sbjct: 178 --NGEIVQRSPERQRRV 192
Score = 85.5 bits (210), Expect = 2e-15
Identities = 57/166 (34%), Positives = 86/166 (51%), Gaps = 25/166 (15%)
Query: 15 RSQVHQLPTNITLPNKSYHSNLPSS---SSSSISCN-RFFTIIAAA--AATTYPSFTPST 68
R L +N+TL +PSS S + C+ RFF+I A + +TY +
Sbjct: 9 RHLTRALLSNVTLMAPP---RIPSSVHYGGSRLGCSTRFFSIRCGANRSGSTYSPLNSGS 65
Query: 69 THN----------------RHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMC 112
+ HW+++MDKP +K ++ID Y++TL V+GSE+EA+
Sbjct: 66 NFSDRPPTEMAPLFPGCDYEHWLIVMDKPGGEGATKHEMIDCYIQTLAKVVGSEEEAKKR 125
Query: 113 LYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
+Y+ S + + GF C+IDEE S++L LPGVL V PD + KDY
Sbjct: 126 IYNVSCERYLGFGCEIDEETSTKLEGLPGVLFVLPDSYVDPENKDY 171
>UniRef100_O82169 Hypothetical protein At2g35240 [Arabidopsis thaliana]
Length = 232
Score = 94.0 bits (232), Expect = 6e-18
Identities = 49/120 (40%), Positives = 70/120 (57%)
Query: 158 YSLSSGQAGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMG 217
YS + S T LFP + +HWL+ M+KPG E K Q++D Y Q L K++G
Sbjct: 70 YSPLKSGSNFSDRPPTEMAPLFPGCDYEHWLIVMEKPGGENAQKQQMIDCYVQTLAKIVG 129
Query: 218 NEKDAQMCIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNYEDSLSV 277
+E++A+ IY+VS + FGF CE+DE+ + L G+PGV V PD + E K+Y L V
Sbjct: 130 SEEEARKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVDPEFKDYGAELFV 189
Score = 85.5 bits (210), Expect = 2e-15
Identities = 41/86 (47%), Positives = 59/86 (67%)
Query: 73 HWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEI 132
HW+++M+KP K Q+ID YV+TL ++GSE+EA+ +Y+ S + +FGF C+IDEE
Sbjct: 98 HWLIVMEKPGGENAQKQQMIDCYVQTLAKIVGSEEEARKKIYNVSCERYFGFGCEIDEET 157
Query: 133 SSQLASLPGVLSVRPDPDFNSLKKDY 158
S++L LPGVL V PD + KDY
Sbjct: 158 SNKLEGLPGVLFVLPDSYVDPEFKDY 183
>UniRef100_Q9FX45 RNA-binding glycine-rich protein, putative [Arabidopsis thaliana]
Length = 181
Score = 89.0 bits (219), Expect = 2e-16
Identities = 47/108 (43%), Positives = 70/108 (64%), Gaps = 3/108 (2%)
Query: 275 LSVPNSSEASQEASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKG 334
LS +SS +S KL+V+GLSF T+E TLR FE FG L+ + +++DK++ R KG
Sbjct: 60 LSTDSSSSPPSSSSGPKTKLYVSGLSFRTTEDTLRDTFEQFGNLIHMNMVMDKVANRPKG 119
Query: 335 YAFIEYTTEEAAGAALKEMNGKIINGWMIVVDVAKTNPPRFNRGHARP 382
+AF+ Y TEE A A++ M+GK ++G +I V+ AKT R + A+P
Sbjct: 120 FAFLRYETEEEAMKAIQGMHGKFLDGRVIFVEEAKT---RSDMSRAKP 164
>UniRef100_Q8S8L4 Putative RNA-binding protein [Arabidopsis thaliana]
Length = 142
Score = 82.4 bits (202), Expect = 2e-14
Identities = 41/88 (46%), Positives = 58/88 (65%)
Query: 287 ASLRTKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAA 346
++L + +LFV+GLS T+ + L+ AF FG+LV +VI D+ S RSKG+ F+ Y T E A
Sbjct: 29 STLTSPRLFVSGLSRLTTNEKLQDAFASFGQLVDARVITDRDSGRSKGFGFVTYATIEDA 88
Query: 347 GAALKEMNGKIINGWMIVVDVAKTNPPR 374
A EMN K ++GW+I VD A+ PR
Sbjct: 89 EKAKAEMNAKFLDGWVIFVDPARPREPR 116
>UniRef100_Q7XSA4 OJ000126_13.13 protein [Oryza sativa]
Length = 137
Score = 80.9 bits (198), Expect = 6e-14
Identities = 35/84 (41%), Positives = 59/84 (69%)
Query: 291 TKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAAL 350
T +LF+ GLS + +E +L AF +G++++ ++ DK++ R KG+ F+++ +EEAA A
Sbjct: 36 TYRLFIGGLSQFATEDSLAEAFSQYGQVLEATIVTDKMTNRPKGFGFVKFASEEAANKAK 95
Query: 351 KEMNGKIINGWMIVVDVAKTNPPR 374
+EMNGK++NG +I VD+AK R
Sbjct: 96 EEMNGKVLNGRVIYVDIAKAKMNR 119
>UniRef100_Q8LB30 DAG protein, putative [Arabidopsis thaliana]
Length = 232
Score = 77.4 bits (189), Expect = 6e-13
Identities = 48/131 (36%), Positives = 68/131 (51%), Gaps = 11/131 (8%)
Query: 39 SSSSSISCNRFFTIIAAAAATTYPSFTPSTTHNR-----------HWMVLMDKPPLGVIS 87
S S++ S R + AA + Y S ++ R HW+++M+ P S
Sbjct: 44 SISTAGSRRRVAIVKAATVDSDYSSKRSNSNEQRETIMLPGCDYNHWLIVMEFPKDPAPS 103
Query: 88 KSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRP 147
+ Q+ID Y+ TL TV+GS +EA+ +Y S T+ GF C IDEE S + LPGVL V P
Sbjct: 104 RDQMIDTYLNTLATVLGSMEEAKKNMYAFSTTTYTGFQCTIDEETSEKFKGLPGVLWVLP 163
Query: 148 DPDFNSLKKDY 158
D + KDY
Sbjct: 164 DSYIDVKNKDY 174
Score = 68.9 bits (167), Expect = 2e-10
Identities = 33/104 (31%), Positives = 58/104 (55%)
Query: 168 SSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIY 227
S+ + R ++ P + HWL+ M+ P ++ Q++D Y L V+G+ ++A+ +Y
Sbjct: 71 SNSNEQRETIMLPGCDYNHWLIVMEFPKDPAPSRDQMIDTYLNTLATVLGSMEEAKKNMY 130
Query: 228 HVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
S T GF C +DE+ ++ G+PGV V PD + +NK+Y
Sbjct: 131 AFSTTTYTGFQCTIDEETSEKFKGLPGVLWVLPDSYIDVKNKDY 174
>UniRef100_Q9LPZ1 T23J18.10 [Arabidopsis thaliana]
Length = 232
Score = 77.0 bits (188), Expect = 8e-13
Identities = 41/93 (44%), Positives = 56/93 (60%), Gaps = 1/93 (1%)
Query: 66 PSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGSEKEAQMCLYDASWDTHFGFC 125
P +N HW+++M+ P S+ Q+ID Y+ TL TV+GS +EA+ +Y S T+ GF
Sbjct: 83 PGCDYN-HWLIVMEFPKDPAPSRDQMIDTYLNTLATVLGSMEEAKKNMYAFSTTTYTGFQ 141
Query: 126 CDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
C IDEE S + LPGVL V PD + KDY
Sbjct: 142 CTIDEETSEKFKGLPGVLWVLPDSYIDVKNKDY 174
Score = 68.9 bits (167), Expect = 2e-10
Identities = 33/104 (31%), Positives = 58/104 (55%)
Query: 168 SSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQMCIY 227
S+ + R ++ P + HWL+ M+ P ++ Q++D Y L V+G+ ++A+ +Y
Sbjct: 71 SNSNEQRETIMLPGCDYNHWLIVMEFPKDPAPSRDQMIDTYLNTLATVLGSMEEAKKNMY 130
Query: 228 HVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
S T GF C +DE+ ++ G+PGV V PD + +NK+Y
Sbjct: 131 AFSTTTYTGFQCTIDEETSEKFKGLPGVLWVLPDSYIDVKNKDY 174
>UniRef100_Q6ZKI1 Putative DAG protein [Oryza sativa]
Length = 229
Score = 76.3 bits (186), Expect = 1e-12
Identities = 44/113 (38%), Positives = 63/113 (54%), Gaps = 7/113 (6%)
Query: 52 IIAAAAATTYPS------FTPSTTHNRHWMVLMDKPPLGVISKSQVIDYYVKTLQTVMGS 105
++AAAA S P +N HW+++M+ P ++ Q+ID Y+ TL TV+GS
Sbjct: 59 VVAAAAGAGTGSEQRETILLPGCDYN-HWLIVMEFPKDPAPTREQMIDTYLNTLATVLGS 117
Query: 106 EKEAQMCLYDASWDTHFGFCCDIDEEISSQLASLPGVLSVRPDPDFNSLKKDY 158
+EA+ +Y S T+ GF C +DEE S + LPGVL V PD + KDY
Sbjct: 118 MEEAKKNMYAFSTTTYTGFQCTVDEETSEKFKGLPGVLWVLPDSYIDVKNKDY 170
Score = 75.9 bits (185), Expect = 2e-12
Identities = 37/107 (34%), Positives = 60/107 (55%)
Query: 165 AGLSSGLQTRTNMLFPAGNSKHWLVRMDKPGVEVVTKAQIVDYYAQILTKVMGNEKDAQM 224
AG +G + R +L P + HWL+ M+ P T+ Q++D Y L V+G+ ++A+
Sbjct: 64 AGAGTGSEQRETILLPGCDYNHWLIVMEFPKDPAPTREQMIDTYLNTLATVLGSMEEAKK 123
Query: 225 CIYHVSWKTNFGFCCELDEDCAQDLAGVPGVSSVQPDDNFESENKNY 271
+Y S T GF C +DE+ ++ G+PGV V PD + +NK+Y
Sbjct: 124 NMYAFSTTTYTGFQCTVDEETSEKFKGLPGVLWVLPDSYIDVKNKDY 170
>UniRef100_Q84ZB9 RNA recognition motif (RRM)-containing protein-like [Oryza sativa]
Length = 133
Score = 75.9 bits (185), Expect = 2e-12
Identities = 34/76 (44%), Positives = 53/76 (69%)
Query: 294 LFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYAFIEYTTEEAAGAALKEM 353
LFV+GL+ T+ LR AF FG++++ +VI D+IS S+G+ F++Y T E AG +K M
Sbjct: 37 LFVSGLNKRTTSDGLREAFSKFGQVIEARVITDRISGYSRGFGFVKYATVEEAGEGIKGM 96
Query: 354 NGKIINGWMIVVDVAK 369
+GK ++GW+I + AK
Sbjct: 97 DGKFLDGWVIFAEYAK 112
>UniRef100_Q9LIS2 Glycine-rich RNA binding protein-like [Arabidopsis thaliana]
Length = 136
Score = 75.5 bits (184), Expect = 2e-12
Identities = 42/104 (40%), Positives = 67/104 (64%), Gaps = 4/104 (3%)
Query: 279 NSSEASQEASLR--TKKLFVTGLSFYTSEKTLRAAFEGFGELVQVKVIIDKISKRSKGYA 336
N S SLR + KLFV GLS+ T + +L+ AF FGE+ + VI D+ + RS+G+
Sbjct: 20 NGPVTSMLGSLRYMSSKLFVGGLSWGTDDSSLKQAFTSFGEVTEATVIADRETGRSRGFG 79
Query: 337 FIEYTTEEAAGAALKEMNGKIINGWMIVVDVA--KTNPPRFNRG 378
F+ ++ E++A A+KEM+GK +NG I V++A +++ PR + G
Sbjct: 80 FVSFSCEDSANNAIKEMDGKELNGRQIRVNLATERSSAPRSSFG 123
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.130 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 653,136,458
Number of Sequences: 2790947
Number of extensions: 26466184
Number of successful extensions: 79349
Number of sequences better than 10.0: 3960
Number of HSP's better than 10.0 without gapping: 2949
Number of HSP's successfully gapped in prelim test: 1011
Number of HSP's that attempted gapping in prelim test: 72705
Number of HSP's gapped (non-prelim): 6603
length of query: 384
length of database: 848,049,833
effective HSP length: 129
effective length of query: 255
effective length of database: 488,017,670
effective search space: 124444505850
effective search space used: 124444505850
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0095c.6


