

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0094b.3
(337 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
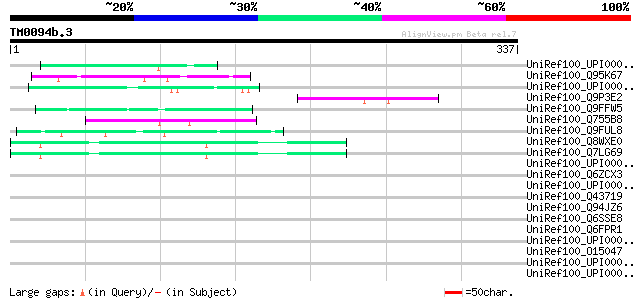
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI0000365031 UPI0000365031 UniRef100 entry 50 1e-04
UniRef100_Q95K67 Hypothetical protein [Macaca fascicularis] 49 2e-04
UniRef100_UPI0000437027 UPI0000437027 UniRef100 entry 49 3e-04
UniRef100_Q9P3E2 Related to transport protein USO1 [Neurospora c... 48 3e-04
UniRef100_Q9FFW5 Similarity to protein kinase [Arabidopsis thali... 48 4e-04
UniRef100_Q755B8 AFL095Wp [Ashbya gossypii] 48 4e-04
UniRef100_Q9FUL8 Arabinogalactan protein [Nicotiana alata] 47 7e-04
UniRef100_Q8WXE0 Cask-interacting protein 2 [Homo sapiens] 47 7e-04
UniRef100_Q7LG69 KIAA1139 protein [Homo sapiens] 47 7e-04
UniRef100_UPI0000437029 UPI0000437029 UniRef100 entry 47 0.001
UniRef100_Q6ZCX3 Putative diaphanous 1 [Oryza sativa] 47 0.001
UniRef100_UPI000034DD72 UPI000034DD72 UniRef100 entry 46 0.002
UniRef100_Q43719 Putative arabinogalactan-protein precursor [Lyc... 46 0.002
UniRef100_Q94JZ6 Protein kinase-like protein [Arabidopsis thaliana] 46 0.002
UniRef100_Q6SSE8 Minus agglutinin [Chlamydomonas reinhardtii] 46 0.002
UniRef100_Q6FPR1 Similarities with sp|P08640 Saccharomyces cerev... 46 0.002
UniRef100_UPI00001CE7BF UPI00001CE7BF UniRef100 entry 45 0.002
UniRef100_O15047 KIAA0339 protein [Homo sapiens] 45 0.002
UniRef100_UPI0000437028 UPI0000437028 UniRef100 entry 45 0.003
UniRef100_UPI0000248D73 UPI0000248D73 UniRef100 entry 45 0.003
>UniRef100_UPI0000365031 UPI0000365031 UniRef100 entry
Length = 762
Score = 49.7 bits (117), Expect = 1e-04
Identities = 37/123 (30%), Positives = 46/123 (37%), Gaps = 9/123 (7%)
Query: 21 ACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRP 80
A +P P P P A P S P AS PS + +P A P
Sbjct: 632 AAAPPSPAASPPSPAAPPSPAASPPSPAAPPSPAASPPSPAASPPPSPAASPPSPAASPP 691
Query: 81 VTLAEGQRSPSSIRPPI-----PSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
+ A SP++ PP PSP P+AP SP AA P P++S P
Sbjct: 692 PSPAASPPSPAASPPPAPAASPPSPAGAPPSPAAPPSPAAPPS----PAASPPSPAASPP 747
Query: 136 PYP 138
P P
Sbjct: 748 PSP 750
Score = 33.9 bits (76), Expect = 6.5
Identities = 29/110 (26%), Positives = 41/110 (36%), Gaps = 10/110 (9%)
Query: 39 LTGTGGGFVAPDPVVT--DSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRP- 95
L + G P P + A P + P A +P A P A SP++ P
Sbjct: 626 LRNSSGAAAPPSPAASPPSPAAPPSPAASPPSPAAPPSPAASPPSPAASPPPSPAASPPS 685
Query: 96 ----PIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPT---GPSSSSPPYP 138
P PSP P+A P + + A P+ PS ++PP P
Sbjct: 686 PAASPPPSPAASPPSPAASPPPAPAASPPSPAGAPPSPAAPPSPAAPPSP 735
>UniRef100_Q95K67 Hypothetical protein [Macaca fascicularis]
Length = 599
Score = 48.9 bits (115), Expect = 2e-04
Identities = 52/175 (29%), Positives = 77/175 (43%), Gaps = 37/175 (21%)
Query: 15 QSSPVSACSPKRSKVGE----PDPQTTPLTGTGGGFVAPDPVVTDSACDP-LASDHPSEV 69
+ P+ A S + K+G P+ TPLT P VV+ P L S PS+
Sbjct: 80 EPKPIPALSMETQKLGSVLSPESPKPTPLTPLEPQ--KPGSVVSPELQTPSLPSPEPSKP 137
Query: 70 NASQTPDAPRPVTLAEGQR-------SPSSIRPPIPSPVKV---------------TLGP 107
+ +P+ P+PV + E Q+ P + P P PVK TLGP
Sbjct: 138 ASVSSPEPPKPVPVCESQKPAPVPSPEPQKLAPVSPEPVKATLPNPKPQKHSHFPETLGP 197
Query: 108 SAPNSPGVGGEGENLSAA-QPTGPS-SSSPPYPFSFLLSHPYLEKLKGAPSQNPQ 160
+ +SP E L+A+ +P GPS ++SP S + P E K +PS +P+
Sbjct: 198 PSASSP----ESPVLAASPEPWGPSPAASPESRKSARTTSP--EPRKPSPSGSPE 246
>UniRef100_UPI0000437027 UPI0000437027 UniRef100 entry
Length = 877
Score = 48.5 bits (114), Expect = 3e-04
Identities = 49/173 (28%), Positives = 63/173 (36%), Gaps = 27/173 (15%)
Query: 13 KRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNAS 72
K PVS + + +K P + P G P + A P + PS S
Sbjct: 475 KPSPPPVSQPASQPAKPTPSQPASQPGQARPGQASQAQPTPSQPASQPASPGQPSPPPVS 534
Query: 73 QTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLG-------PSAP------NSPGVGGEG 119
Q P P GQ P P PSP + TL PS P + P +
Sbjct: 535 QQPARP-------GQARPGQPSPAQPSPAQPTLSQPARPAKPSPPPVSQPASQPAQASQA 587
Query: 120 ENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKG--APSQ----NPQDVAQEA 166
SA+QP P SPP P S S P + G P+Q +PQ +Q A
Sbjct: 588 HPQSASQPASPGQPSPP-PVSQPASQPASQARPGQARPAQPSPAHPQSASQPA 639
Score = 45.1 bits (105), Expect = 0.003
Identities = 49/166 (29%), Positives = 65/166 (38%), Gaps = 16/166 (9%)
Query: 4 KPRVAEVGGKRQSSPVSACSPKR-SKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLA 62
+P A + P SA P R + + P + +P PV + A P +
Sbjct: 618 RPGQARPAQPSPAHPQSASQPARPGQASQAQPTPSQPASQPASQPSPPPV-SQPASQPAS 676
Query: 63 SDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGE-- 120
HP +ASQ P PRP GQ P+ P PSP S P PG + +
Sbjct: 677 QAHPQ--SASQ-PARPRP-----GQARPAKPSPAQPSPAHPQ-SASQPARPGQASQAQPT 727
Query: 121 -NLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLK-GAPSQNPQDVAQ 164
+ A+QP P SPP P S S P + + G PS AQ
Sbjct: 728 PSQPASQPASPGQPSPP-PVSQSASQPASQPARPGQPSPAQPSPAQ 772
Score = 44.3 bits (103), Expect = 0.005
Identities = 38/145 (26%), Positives = 54/145 (37%), Gaps = 25/145 (17%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVA-PDPVVTDSACDPLASDHPSEVNASQ- 73
S P S P + +P P + P + G P P SA P + P++ SQ
Sbjct: 388 SQPASQARPGQPSPAQPTP-SQPASQPGQARPGQPSPAHPQSASQPASQPRPAKPTPSQP 446
Query: 74 -----TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAP---------NSPGVGGEG 119
+P P P +++ P RP PSP V+ S P + PG G
Sbjct: 447 ASQPASPGQPSPPPVSQPASQPGQARPAKPSPPPVSQPASQPAKPTPSQPASQPGQARPG 506
Query: 120 E--------NLSAAQPTGPSSSSPP 136
+ + A+QP P SPP
Sbjct: 507 QASQAQPTPSQPASQPASPGQPSPP 531
Score = 43.1 bits (100), Expect = 0.011
Identities = 40/156 (25%), Positives = 52/156 (32%), Gaps = 16/156 (10%)
Query: 15 QSSPVSACSPKRSKVGEP-------------DPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
Q+ P SA P R + G+ PQ+ G P + A P
Sbjct: 677 QAHPQSASQPARPRPGQARPAKPSPAQPSPAHPQSASQPARPGQASQAQPTPSQPASQPA 736
Query: 62 ASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGEN 121
+ PS SQ+ P GQ SP+ P P P K P + + P
Sbjct: 737 SPGQPSPPPVSQSASQPASQPARPGQPSPAQPSPAQPRPAK-PAHPQSASQPASQARPGQ 795
Query: 122 LSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQ 157
S AQPT +S P S +HP P Q
Sbjct: 796 PSPAQPTPSQPASQPAQAS--QAHPQSASQPARPGQ 829
Score = 41.2 bits (95), Expect = 0.041
Identities = 42/154 (27%), Positives = 59/154 (38%), Gaps = 9/154 (5%)
Query: 12 GKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA 71
G+ PVS + + +P P G +P PV + A P + P + +
Sbjct: 343 GQPSPPPVSQQPASQPRPAQPTPSQPASQPASPGQPSPPPV-SQPASQPASQARPGQPSP 401
Query: 72 SQ-TPDAP--RPVTLAEGQRSPS---SIRPPIPSPVKVTLGPSAPNS-PGVGGEGENLSA 124
+Q TP P +P GQ SP+ S P P PS P S P G+
Sbjct: 402 AQPTPSQPASQPGQARPGQPSPAHPQSASQPASQPRPAKPTPSQPASQPASPGQPSPPPV 461
Query: 125 AQPTG-PSSSSPPYPFSFLLSHPYLEKLKGAPSQ 157
+QP P + P P +S P + K PSQ
Sbjct: 462 SQPASQPGQARPAKPSPPPVSQPASQPAKPTPSQ 495
Score = 41.2 bits (95), Expect = 0.041
Identities = 44/176 (25%), Positives = 57/176 (32%), Gaps = 25/176 (14%)
Query: 15 QSSPVSACSPKR-SKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
Q SP P R + + P P++ P + A P +
Sbjct: 236 QPSPPPVSQPARPGQASQAQPSPPPVSQPASQPRPAKPTPSQPASQPAKPTPSQPASQPA 295
Query: 74 TPDAPRPVTLAEGQRSPSSI-------RPPIPSPVKVTLGPSAPNSPG------------ 114
P RP + Q SP S+ RP PSP V+ S P SPG
Sbjct: 296 RPGQARPAKPSPAQPSPPSVSQPASQARPGQPSPAPVSQPASQPASPGQPSPPPVSQQPA 355
Query: 115 ----VGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEA 166
+ A+QP P SPP P S S P + G PS +Q A
Sbjct: 356 SQPRPAQPTPSQPASQPASPGQPSPP-PVSQPASQPASQARPGQPSPAQPTPSQPA 410
Score = 37.4 bits (85), Expect = 0.59
Identities = 42/163 (25%), Positives = 60/163 (36%), Gaps = 28/163 (17%)
Query: 8 AEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPS 67
A G R + P S P V +P Q P G +P PV P+
Sbjct: 295 ARPGQARPAKP-SPAQPSPPSVSQPASQARP------GQPSPAPV-----------SQPA 336
Query: 68 EVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQP 127
AS +P PV+ Q+ S RP P+P + P++P P + A+QP
Sbjct: 337 SQPASPGQPSPPPVS----QQPASQPRPAQPTPSQPASQPASPGQPSPPPVSQ--PASQP 390
Query: 128 TGPSSSSPPYPFSFLLSHPYLEKLKGAPSQ----NPQDVAQEA 166
+ P P S P + + P Q +PQ +Q A
Sbjct: 391 ASQARPGQPSPAQPTPSQPASQPGQARPGQPSPAHPQSASQPA 433
Score = 34.7 bits (78), Expect = 3.8
Identities = 37/152 (24%), Positives = 49/152 (31%), Gaps = 9/152 (5%)
Query: 12 GKRQSSPVSACSPKRSKVGEPDP---QTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSE 68
G+ S P S P +++ G+ P + P + P AS HP
Sbjct: 141 GQPASQPASHPQPSQARPGQASPGRPASQPASHPQSSQARPGQASPGRPASQPAS-HPQS 199
Query: 69 VNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPT 128
A P P Q P S P P + + +P S AQP+
Sbjct: 200 SQAR--PGRPASQPAKSSQAHPQSASQPASQPGQASQAQPSPPPVSQPARPGQASQAQPS 257
Query: 129 GPSSSSP---PYPFSFLLSHPYLEKLKGAPSQ 157
P S P P P S P + K PSQ
Sbjct: 258 PPPVSQPASQPRPAKPTPSQPASQPAKPTPSQ 289
Score = 34.3 bits (77), Expect = 5.0
Identities = 36/131 (27%), Positives = 50/131 (37%), Gaps = 11/131 (8%)
Query: 15 QSSPVSACSPKRSKVGEPDPQTTPLTGTGG----GFVAPDPVVTDSACDPLASDHPSEVN 70
QS+ A P ++ +P P G +P PV ++ A PS+
Sbjct: 220 QSASQPASQPGQASQAQPSPPPVSQPARPGQASQAQPSPPPVSQPASQPRPAKPTPSQP- 278
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVT----LGPSAPNSPGVGGEGE-NLSAA 125
ASQ P P P A P RP PSP + + P++ PG + A+
Sbjct: 279 ASQ-PAKPTPSQPASQPARPGQARPAKPSPAQPSPPSVSQPASQARPGQPSPAPVSQPAS 337
Query: 126 QPTGPSSSSPP 136
QP P SPP
Sbjct: 338 QPASPGQPSPP 348
Score = 33.9 bits (76), Expect = 6.5
Identities = 37/128 (28%), Positives = 48/128 (36%), Gaps = 14/128 (10%)
Query: 15 QSSPVSACSPKRSKVGEPDP-QTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
QS+ A P R G+P P Q +P P SA P + P + + +Q
Sbjct: 748 QSASQPASQPARP--GQPSPAQPSPAQPRPA-----KPAHPQSASQPASQARPGQPSPAQ 800
Query: 74 -TPDAPRPVTLAEGQRSPSSI----RPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPT 128
TP P Q P S RP PSP + P A S G+ A+P
Sbjct: 801 PTPSQPASQPAQASQAHPQSASQPARPGQPSPAQPASQP-AQASQASPGQPNQPRPAKPA 859
Query: 129 GPSSSSPP 136
P S+S P
Sbjct: 860 HPQSASQP 867
>UniRef100_Q9P3E2 Related to transport protein USO1 [Neurospora crassa]
Length = 1171
Score = 48.1 bits (113), Expect = 3e-04
Identities = 38/105 (36%), Positives = 54/105 (51%), Gaps = 11/105 (10%)
Query: 192 GLRQEVEELQRDLDAKEITVASQEKFLAAKDKEIADLREELEG----QNEALAEAQSDLT 247
G+ +E EE +R L KE VA EK L K++E+ ++E LEG + EA+ +A ++
Sbjct: 952 GVIEEREEAERLLKEKEDAVAEVEKALKDKEREMVSMKERLEGMEKEKEEAVQKAVEEIR 1011
Query: 248 FKC-------KLLADYEAQAATLKEKIRSEAAHAAAKERVRGSLI 285
+ K A+ EA AAT EK + A AAAK S I
Sbjct: 1012 KEVETAKNNDKAEAEAEADAATGAEKAAAIEAEAAAKAAAAASEI 1056
>UniRef100_Q9FFW5 Similarity to protein kinase [Arabidopsis thaliana]
Length = 681
Score = 47.8 bits (112), Expect = 4e-04
Identities = 42/145 (28%), Positives = 56/145 (37%), Gaps = 7/145 (4%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
PV + SP V P P ++P + +P P V S P+ P + TP A
Sbjct: 50 PVVSSSPPPPVVSSPPPSSSP-PPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTPPA 108
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS-SSSPP 136
P P T++ +S PP P+ T P SP GE + P+ P S S P
Sbjct: 109 P-PQTVSPPPPPDASPSPPAPT----TTNPPPKPSPSPPGETPSPPGETPSPPKPSPSTP 163
Query: 137 YPFSFLLSHPYLEKLKGAPSQNPQD 161
P + P PS NP D
Sbjct: 164 TPTTTTSPPPPPATSASPPSSNPTD 188
Score = 34.3 bits (77), Expect = 5.0
Identities = 36/142 (25%), Positives = 50/142 (34%), Gaps = 15/142 (10%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
PV SP S P TTP P P + S P ++ P + + S +
Sbjct: 91 PVVIASPPPST-----PATTPPAPPQTVSPPPPPDASPSPPAPTTTNPPPKPSPSPPGET 145
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS-SSPP 136
P P PS P P+P T P P + ++ PT PS+ + PP
Sbjct: 146 PSPPGETPSPPKPS---PSTPTPTTTTSPPPPP------ATSASPPSSNPTDPSTLAPPP 196
Query: 137 YPFSFLLSHPYLEKLKGAPSQN 158
P + + K G S N
Sbjct: 197 TPLPVVPREKPIAKPTGPASNN 218
>UniRef100_Q755B8 AFL095Wp [Ashbya gossypii]
Length = 2535
Score = 47.8 bits (112), Expect = 4e-04
Identities = 33/120 (27%), Positives = 51/120 (42%), Gaps = 6/120 (5%)
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIP----SPVKVTLG 106
P T C P A+ P++ ++S + P P T Q S S+ +P P P + +
Sbjct: 2228 PPATSQPCPPPATSQPAQSSSSASQPCPPPATSQPAQSSSSASQPCPPPATSQPAQSSSS 2287
Query: 107 PSAPNSPGVGGE--GENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQ 164
S P P + + SA+QP PS++S P S S P P+Q+ +Q
Sbjct: 2288 ASQPCPPAATSQPAQSSSSASQPCPPSATSQPAQSSSSASQPCPPSATSQPAQSSSSASQ 2347
Score = 40.8 bits (94), Expect = 0.053
Identities = 28/95 (29%), Positives = 44/95 (45%), Gaps = 6/95 (6%)
Query: 58 CDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPS----PVKVTLGPSAPNSP 113
C P A+ P++ ++S + P P T Q S S+ +P P+ P + + S P P
Sbjct: 2254 CPPPATSQPAQSSSSASQPCPPPATSQPAQSSSSASQPCPPAATSQPAQSSSSASQPCPP 2313
Query: 114 GVGGEG--ENLSAAQPTGPSSSSPPYPFSFLLSHP 146
+ + SA+QP PS++S P S S P
Sbjct: 2314 SATSQPAQSSSSASQPCPPSATSQPAQSSSSASQP 2348
>UniRef100_Q9FUL8 Arabinogalactan protein [Nicotiana alata]
Length = 228
Score = 47.0 bits (110), Expect = 7e-04
Identities = 57/207 (27%), Positives = 79/207 (37%), Gaps = 38/207 (18%)
Query: 5 PRVAEVGGKRQSSPVSACSPKRSKVGEPD------------------PQTTPLTGTGGGF 46
P A VG K + P +A P + K P P T P T
Sbjct: 28 PAKAPVGAKATTPPAAA--PTKPKTPAPATAPASAPPTAVSTPPAAAPATAPTTPVVTSP 85
Query: 47 VAPDPVVTDSACDPLA---SDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPV-- 101
V+ P T ++ P A S P V Q+P AP PV P++ PP P PV
Sbjct: 86 VSAPPAKTPASSPPAAVPVSSPPLAVTPVQSPPAPAPVAAT----PPAASAPPAPVPVSA 141
Query: 102 ----KVTLGPSAPNSPGVGGEGENLSAAQPT-GPSSSSPPYPFSFLLSHPYLEKLKGAPS 156
+ P+ S G G + A+ P P SPP P P +E +PS
Sbjct: 142 PAVSETVPAPAPSKSKGKGKKKGKKHASSPAPSPDMMSPPAP-PTEAPGPSMESDSPSPS 200
Query: 157 QNPQDVAQE-AVLNLLRAGCLFAQVAW 182
N + A++ +L L AG +A ++W
Sbjct: 201 LNDESGAEKLKMLGSLVAG--WAVMSW 225
>UniRef100_Q8WXE0 Cask-interacting protein 2 [Homo sapiens]
Length = 1202
Score = 47.0 bits (110), Expect = 7e-04
Identities = 57/240 (23%), Positives = 85/240 (34%), Gaps = 41/240 (17%)
Query: 1 MSVKPRVAEVGGKRQSSPV----------SACSPKRSKVGEPDPQTTPLTGTGGGFVAPD 50
+++K R G + +PV S +R K E +P T L G P
Sbjct: 958 LTIKQRPKPAGPPPRETPVPPGLDFNLTESDTVKRRPKCREREPLQTALLAFGVASATPG 1017
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTL-AEGQRSPSSIRPPIPSPVKVTLGPSA 109
P PL S P E + + P P +L A+G +P + P + PV GP
Sbjct: 1018 PAA------PLPSPTPGESPPASSLPQPEPSSLPAQGVPTPLAPSPAMQPPVPPCPGPGL 1071
Query: 110 PNSPGVGGEGENLSAAQPTG----PSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE 165
+S GE A P P + + P P S + K AP P+ V
Sbjct: 1072 ESSAASRWNGETEPPAAPAALLKVPGAGTAPKPVSVACTQLAFSGPKLAPRLGPRPVPPP 1131
Query: 166 AVLNLLRAGCLFAQVAWQVRP-LTGIVGLRQEVEELQRDLDAKEITVASQEKFLAAKDKE 224
RP TG VG Q + L++ + + + EK + K++E
Sbjct: 1132 -------------------RPESTGTVGPGQAQQRLEQTSSSLAAALRAAEKSIGTKEQE 1172
>UniRef100_Q7LG69 KIAA1139 protein [Homo sapiens]
Length = 1124
Score = 47.0 bits (110), Expect = 7e-04
Identities = 57/240 (23%), Positives = 85/240 (34%), Gaps = 41/240 (17%)
Query: 1 MSVKPRVAEVGGKRQSSPV----------SACSPKRSKVGEPDPQTTPLTGTGGGFVAPD 50
+++K R G + +PV S +R K E +P T L G P
Sbjct: 880 LTIKQRPKPAGPPPRETPVPPGLDFNLTESDTVKRRPKCREREPLQTALLAFGVASATPG 939
Query: 51 PVVTDSACDPLASDHPSEVNASQTPDAPRPVTL-AEGQRSPSSIRPPIPSPVKVTLGPSA 109
P PL S P E + + P P +L A+G +P + P + PV GP
Sbjct: 940 PAA------PLPSPTPGESPPASSLPQPEPSSLPAQGVPTPLAPSPAMQPPVPPCPGPGL 993
Query: 110 PNSPGVGGEGENLSAAQPTG----PSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE 165
+S GE A P P + + P P S + K AP P+ V
Sbjct: 994 ESSAASRWNGETEPPAAPAALLKVPGAGTAPKPVSVACTQLAFSGPKLAPRLGPRPVPPP 1053
Query: 166 AVLNLLRAGCLFAQVAWQVRP-LTGIVGLRQEVEELQRDLDAKEITVASQEKFLAAKDKE 224
RP TG VG Q + L++ + + + EK + K++E
Sbjct: 1054 -------------------RPESTGTVGPGQAQQRLEQTSSSLAAALRAAEKSIGTKEQE 1094
>UniRef100_UPI0000437029 UPI0000437029 UniRef100 entry
Length = 1010
Score = 46.6 bits (109), Expect = 0.001
Identities = 52/168 (30%), Positives = 67/168 (38%), Gaps = 21/168 (12%)
Query: 16 SSPVSA-CSPKRSKVGEPDP-QTTPLT-----GTGGGFVAPDPVVTDSACDPLASDHPSE 68
S P S P +++ G+P P Q TP G P V+ A P + HP
Sbjct: 782 SQPASQPARPGQARPGQPSPAQPTPSQPASQPGQARPAKPSPPPVSQPASQPASQAHPQS 841
Query: 69 VN--ASQTPDAPRPVTLAE--GQRSPSSIRPPIPSPVKVT----LGPSAPNSPGVGGEGE 120
+ ASQ P P P A GQ P RP PSP + + S P PG + +
Sbjct: 842 ASQPASQ-PAKPTPSQPASQPGQARPGQARPAKPSPAQPSPAHPQSASQPARPGQASQAQ 900
Query: 121 ---NLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLK-GAPSQNPQDVAQ 164
+ A+QP P SPP P S S P + + G PS AQ
Sbjct: 901 PTPSQPASQPASPGQPSPP-PVSQSASQPASQPARPGQPSPAQPSPAQ 947
Score = 43.5 bits (101), Expect = 0.008
Identities = 45/167 (26%), Positives = 64/167 (37%), Gaps = 22/167 (13%)
Query: 15 QSSPVSACSP-KRSKVGEPDPQ--TTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA 71
Q SP S P +++ G+P P + P + P V+ A P + PS
Sbjct: 515 QPSPPSVSQPASQARPGQPSPAPVSQPASQPASPGQPSPPPVSQPASQPASPGQPSPPPV 574
Query: 72 SQTPDAPRPVTLAEGQRS---------PSSIRPPIPSPVKVTLGPSAPNSPGVGGEGE-- 120
SQ P + A Q + P P PSP V+ P++ PG + +
Sbjct: 575 SQPASQPAQASQAHPQSASQPASQPARPGQASPAQPSPPPVSQ-PASQARPGQASQAQPT 633
Query: 121 -NLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEA 166
+ A+QP P SPP P S S P A +PQ +Q A
Sbjct: 634 PSQPASQPASPGQPSPP-PVSQPASQP-----AQASQAHPQSASQPA 674
Score = 41.6 bits (96), Expect = 0.031
Identities = 35/146 (23%), Positives = 48/146 (31%), Gaps = 3/146 (2%)
Query: 12 GKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA 71
G+ + P ++ PQ+ G P + A P + PS
Sbjct: 862 GQARPGQARPAKPSPAQPSPAHPQSASQPARPGQASQAQPTPSQPASQPASPGQPSPPPV 921
Query: 72 SQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS 131
SQ+ P GQ SP+ P P P K P + + P S AQPT
Sbjct: 922 SQSASQPASQPARPGQPSPAQPSPAQPRPAK-PAHPQSASQPASQARPGQPSPAQPTPSQ 980
Query: 132 SSSPPYPFSFLLSHPYLEKLKGAPSQ 157
+S P S +HP P Q
Sbjct: 981 PASQPAQAS--QAHPQSASQPARPGQ 1004
Score = 38.1 bits (87), Expect = 0.34
Identities = 40/146 (27%), Positives = 54/146 (36%), Gaps = 19/146 (13%)
Query: 24 PKRSKVGEPDP-QTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVT 82
P ++ +P P Q +P + + G P P SA P + P A TP P
Sbjct: 717 PGQASQAQPSPAQPSPPSVSQPG--QPSPAHPQSASQPASQPRP----AKPTPSQPASQP 770
Query: 83 LAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQP--TGPSSSSPPYPFS 140
Q P S P P + G + P P + A+QP P+ SPP P S
Sbjct: 771 AQASQAHPQSASQPASQPARP--GQARPGQPSPAQPTPSQPASQPGQARPAKPSPP-PVS 827
Query: 141 FLLSHPYLEKLKGAPSQNPQDVAQEA 166
S P A +PQ +Q A
Sbjct: 828 QPASQP-------ASQAHPQSASQPA 846
Score = 38.1 bits (87), Expect = 0.34
Identities = 37/154 (24%), Positives = 52/154 (33%), Gaps = 8/154 (5%)
Query: 12 GKRQSSPVSACSPKRSKVGEPDP---QTTPLTGTGGGFVAPDPVVTDS-ACDPLASDHPS 67
G+ S P S P +++ G+ P + P + P A P + PS
Sbjct: 342 GRPASQPASHPQPSQARPGQASPGRPASQPASHPQSSQARPGQASPGRPASQPASHPQPS 401
Query: 68 EVNASQ-TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQ 126
+ Q +P P Q P S P P + + +P S AQ
Sbjct: 402 QARPGQASPGRPASQPAKSSQAHPQSASQPASQPGQASQAQPSPPPVSQPARPGQASQAQ 461
Query: 127 PTGPSSSSP---PYPFSFLLSHPYLEKLKGAPSQ 157
P+ P S P P P S P + K PSQ
Sbjct: 462 PSPPPVSQPASQPRPAKPTPSQPASQPAKPTPSQ 495
Score = 37.0 bits (84), Expect = 0.76
Identities = 42/183 (22%), Positives = 69/183 (36%), Gaps = 19/183 (10%)
Query: 3 VKPRVAEVGGKRQSSPVSA----CSPKRSKVGEPDPQTTPLT----GTGGGFVAPDPVVT 54
++P A G R +S ++ P +++ G+P P + P + G + P
Sbjct: 31 IQPGQARSGKPRPASQPASQPPPIQPGQARSGKPRPASQPASQPPPAQPGQARSGKPRPA 90
Query: 55 DSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSA---PN 111
A+ +P+ + A +P + PS RP SP + P++ P
Sbjct: 91 SQPASQPATPNPARPGQVRQAQAGQPASQPASHPQPSQARPGQASPGRPASQPASQPPPA 150
Query: 112 SPGVGGEGENLSAAQPT--------GPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVA 163
PG G+ A+QP G + S P P S S P + A S P+ +
Sbjct: 151 QPGQARSGKPRPASQPASQPPPIQPGQARSGKPRPASQPASQPPPIQPGQARSGKPRPAS 210
Query: 164 QEA 166
Q A
Sbjct: 211 QPA 213
Score = 36.2 bits (82), Expect = 1.3
Identities = 46/162 (28%), Positives = 63/162 (38%), Gaps = 19/162 (11%)
Query: 15 QSSPVSACSPKRSKVGEPDPQTTPLTGTGG----GFVAPDPVVTDSACDPLASDHPSEVN 70
QS+ A P ++ +P P G +P PV ++ A PS+
Sbjct: 426 QSASQPASQPGQASQAQPSPPPVSQPARPGQASQAQPSPPPVSQPASQPRPAKPTPSQP- 484
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVT----LGPSAPNSPGVGGEGE-NLSAA 125
ASQ P P P A P RP PSP + + P++ PG + A+
Sbjct: 485 ASQ-PAKPTPSQPASQPARPGQARPAKPSPAQPSPPSVSQPASQARPGQPSPAPVSQPAS 543
Query: 126 QPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQ-NPQDVAQEA 166
QP P SPP P S S P +P Q +P V+Q A
Sbjct: 544 QPASPGQPSPP-PVSQPASQP------ASPGQPSPPPVSQPA 578
Score = 36.2 bits (82), Expect = 1.3
Identities = 41/171 (23%), Positives = 58/171 (32%), Gaps = 20/171 (11%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN--ASQ 73
S P S P + +P Q T + P A A P V+ ASQ
Sbjct: 468 SQPASQPRPAKPTPSQPASQPAKPTPSQPASQPARPGQARPAKPSPAQPSPPSVSQPASQ 527
Query: 74 T-PDAPRPVTLAEGQRSPSS----IRPPIPSPVKVTLGPSAPNSPGVG------------ 116
P P P +++ P+S PP+ P P P+ P V
Sbjct: 528 ARPGQPSPAPVSQPASQPASPGQPSPPPVSQPASQPASPGQPSPPPVSQPASQPAQASQA 587
Query: 117 -GEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEA 166
+ + A+QP P +SP P +S P + G SQ +Q A
Sbjct: 588 HPQSASQPASQPARPGQASPAQPSPPPVSQPASQARPGQASQAQPTPSQPA 638
Score = 35.0 bits (79), Expect = 2.9
Identities = 42/184 (22%), Positives = 68/184 (36%), Gaps = 34/184 (18%)
Query: 12 GKRQSSPVSACSPKRSKVGEPDPQ--TTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEV 69
G+ +PVS + + + G+P P + P + P V+ A P +
Sbjct: 531 GQPSPAPVSQPASQPASPGQPSPPPVSQPASQPASPGQPSPPPVSQPASQPAQASQAHPQ 590
Query: 70 NASQTPDAP-RPVTLAEGQRSPSSIRPPI------------PSPVKVTLGPSAP------ 110
+ASQ P RP + Q SP + P P+P + P++P
Sbjct: 591 SASQPASQPARPGQASPAQPSPPPVSQPASQARPGQASQAQPTPSQPASQPASPGQPSPP 650
Query: 111 ------NSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGA-PSQ-NPQDV 162
+ P + SA+QP + P P +HP+ + A P Q +P V
Sbjct: 651 PVSQPASQPAQASQAHPQSASQPASQARPGQPSP-----AHPHQPASQPASPGQPSPPPV 705
Query: 163 AQEA 166
+Q A
Sbjct: 706 SQPA 709
Score = 34.7 bits (78), Expect = 3.8
Identities = 42/168 (25%), Positives = 59/168 (35%), Gaps = 26/168 (15%)
Query: 12 GKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN- 70
G+ S P S P +++ G+ P G A P + A AS S+
Sbjct: 388 GRPASQPASHPQPSQARPGQASP----------GRPASQPAKSSQAHPQSASQPASQPGQ 437
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
ASQ +P PV+ + +S P P PV P P + P+ P
Sbjct: 438 ASQAQPSPPPVS-QPARPGQASQAQPSPPPVSQPASQPRPAKPTPSQPASQPAKPTPSQP 496
Query: 131 SSS------------SPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEA 166
+S SP P +S P + G PS P V+Q A
Sbjct: 497 ASQPARPGQARPAKPSPAQPSPPSVSQPASQARPGQPS--PAPVSQPA 542
>UniRef100_Q6ZCX3 Putative diaphanous 1 [Oryza sativa]
Length = 893
Score = 46.6 bits (109), Expect = 0.001
Identities = 44/140 (31%), Positives = 53/140 (37%), Gaps = 14/140 (10%)
Query: 15 QSSPVSACSPKRSKVGEPDPQ---TTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA 71
QS CSP R+ P P ++P+ +G P P P P A
Sbjct: 272 QSPSTPRCSPVRTLASPPPPPAPTSSPVRMSGPPPPPPPPAPNSCPSRPAPPPPPPPPLA 331
Query: 72 SQT----PDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQP 127
S + P AP P L SP+ PP P P T+ SAP P + G S P
Sbjct: 332 STSSPPRPAAPSPCQLHTSTSSPARPVPP-PPPTLSTIRSSAPTPPLLPGATSAPSPPPP 390
Query: 128 TGPSSSS------PPYPFSF 141
P SSS PP P SF
Sbjct: 391 PPPCSSSNQLSAPPPPPPSF 410
>UniRef100_UPI000034DD72 UPI000034DD72 UniRef100 entry
Length = 4006
Score = 45.8 bits (107), Expect = 0.002
Identities = 43/134 (32%), Positives = 54/134 (40%), Gaps = 17/134 (12%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P+S+C+ + E +PQ L G+G A L S P A P A
Sbjct: 3864 PLSSCASPPPPISE-EPQQEQLPGSG------------QARGRLDSGPPCRYPAQAVPQA 3910
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLS-AAQPTGPSSSSPP 136
P LA R PSS I S + + + P V G G N S AA P P SPP
Sbjct: 3911 PGSNPLA---RQPSSEHLDILSSILASFNSTVLTPPAVSGPGPNGSPAALPPPPPHLSPP 3967
Query: 137 YPFSFLLSHPYLEK 150
P ++ S P L K
Sbjct: 3968 PPTAYPRSPPTLSK 3981
>UniRef100_Q43719 Putative arabinogalactan-protein precursor [Lycopersicon
esculentum]
Length = 215
Score = 45.8 bits (107), Expect = 0.002
Identities = 49/192 (25%), Positives = 73/192 (37%), Gaps = 17/192 (8%)
Query: 5 PRVAEVGGKRQSSPVSACSPKRSKVGEP-------DPQTTPLTGTGGGFVAPDPVVTDSA 57
P A VG K ++P +A +P + K P P P+ AP V +
Sbjct: 24 PAAAPVGAKAGTTPPAA-APTKPKTPAPATAPASAPPTAVPVAPVTAPVTAPTTPVVAAP 82
Query: 58 CDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPV------KVTLGPSAPN 111
AS P + AS P P E P+ PP +PV + T P+
Sbjct: 83 VSAPASSPPLKAPASSPPVQSPPAPAPEVATPPAVSTPPAAAPVAAPVASETTPAPAPSK 142
Query: 112 SPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE-AVLNL 170
G +G+ +A+ P SPP P S +PS N + A++ +L
Sbjct: 143 GKVKGKKGKKHNASPAPSPDMMSPPAPPSEAPGPSMDSDSAPSPSLNDESGAEKLKMLGS 202
Query: 171 LRAGCLFAQVAW 182
L AG +A ++W
Sbjct: 203 LVAG--WAVMSW 212
>UniRef100_Q94JZ6 Protein kinase-like protein [Arabidopsis thaliana]
Length = 652
Score = 45.8 bits (107), Expect = 0.002
Identities = 37/134 (27%), Positives = 53/134 (38%), Gaps = 10/134 (7%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
++P +P S P TT +P P T S+ P PS + S P
Sbjct: 3 TAPSPGTTPSPSPPSPPTNSTTTTPPPAAS--SPPPTTTPSSPPP----SPSTNSTSPPP 56
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSP---VKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS 132
+P P +L P S+ PPIP P +T P +P +P + + P PS
Sbjct: 57 SSPLPPSLPPPS-PPGSLTPPIPQPSPSAPITPSPPSPTTPSNPRSPPSPNQGPPNTPSG 115
Query: 133 SSPPYPFSFLLSHP 146
S+P P + S P
Sbjct: 116 STPRTPSNTKPSPP 129
>UniRef100_Q6SSE8 Minus agglutinin [Chlamydomonas reinhardtii]
Length = 3889
Score = 45.8 bits (107), Expect = 0.002
Identities = 35/118 (29%), Positives = 49/118 (40%), Gaps = 16/118 (13%)
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIP------SPV 101
+P+P + P + + PS V S P +P P + A +P PP P SPV
Sbjct: 1148 SPEPPSPAPSPPPPSPEPPSPVPPSPPPPSPEPPSPAPPSPAPPLPLPPSPHTQSPPSPV 1207
Query: 102 KVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
+ PSAP+ P + QP P + SPP P S P+ AP+ P
Sbjct: 1208 PPSPAPSAPSPP----------SPQPPSPLAPSPPSPAPQAPSSPFPPPQPTAPTAPP 1255
Score = 44.7 bits (104), Expect = 0.004
Identities = 33/122 (27%), Positives = 45/122 (36%), Gaps = 9/122 (7%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
+P SA P P P + V P P+ P + + PS V S P
Sbjct: 797 APPSAAPPSPMPPSPAPPSPDPPSPKPPSPVPPSPL-------PPSPEPPSPVPPSPPPA 849
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
+P P + A P S PP P+P P +P P L + +P P+ PP
Sbjct: 850 SPEPTSPAPPSPPPPSPEPPSPAPPSPP--PPSPEPPSPAPPSPPLPSPEPPSPAPPLPP 907
Query: 137 YP 138
P
Sbjct: 908 PP 909
Score = 44.3 bits (103), Expect = 0.005
Identities = 27/90 (30%), Positives = 40/90 (44%), Gaps = 2/90 (2%)
Query: 49 PDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPS 108
P P + + P + + PS S P +P P + A P S +PP SPV + P
Sbjct: 777 PSPSIPPPSPGPPSPEPPSPAPPSAAPPSPMPPSPAPPSPDPPSPKPP--SPVPPSPLPP 834
Query: 109 APNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+P P ++ +PT P+ SPP P
Sbjct: 835 SPEPPSPVPPSPPPASPEPTSPAPPSPPPP 864
Score = 42.4 bits (98), Expect = 0.018
Identities = 35/125 (28%), Positives = 49/125 (39%), Gaps = 10/125 (8%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNA-SQT 74
S P + P P P P + P P A A P + +Q+
Sbjct: 1143 SPPPPSPEPPSPAPSPPPPSPEPPSPVPPSPPPPSPEPPSPAPPSPAPPLPLPPSPHTQS 1202
Query: 75 PDAPRPVTLAEGQRSPSSIRPPIP-SPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSS 133
P +P P + A SP S +PP P +P + P AP+SP QPT P++
Sbjct: 1203 PPSPVPPSPAPSAPSPPSPQPPSPLAPSPPSPAPQAPSSP--------FPPPQPTAPTAP 1254
Query: 134 SPPYP 138
PP+P
Sbjct: 1255 PPPFP 1259
Score = 40.8 bits (94), Expect = 0.053
Identities = 38/137 (27%), Positives = 55/137 (39%), Gaps = 16/137 (11%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL--ASDHPSE---VNA 71
+P S P + + P TP + +P+P V PL + PS V
Sbjct: 1326 APPSPMPPSPAPLAPQPPSPTPPSPAPPVPPSPEPPVPPGPDPPLPPSPTPPSPQPPVPP 1385
Query: 72 SQTPDAPRPVTLAEGQRSPS----------SIRPPIPSPVKVTLGPSAPNSPGVGGEGEN 121
S TP +P+P + A +PS S +PP P+P PS P++P
Sbjct: 1386 SPTPPSPQPPSPAPPSPAPSAPLQPSPDPPSPQPPSPAPGPPPSPPSPPSTPSPPSPAP- 1444
Query: 122 LSAAQPTGPSSSSPPYP 138
L+ A P P + PP P
Sbjct: 1445 LAPAPPVPPMAPQPPSP 1461
Score = 37.7 bits (86), Expect = 0.45
Identities = 31/121 (25%), Positives = 41/121 (33%), Gaps = 5/121 (4%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P + P P + P P P + + PS V S P +
Sbjct: 1061 PPSPAPPSPAPPSPEPPSPAPPSPAPPSPEPPSPAPPSPR--PPSPEPPSPVPPSPPPPS 1118
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPY 137
P P + A P S+ PP P+P P SP + +P P SPP
Sbjct: 1119 PEPPSPAPPSPPPPSLEPPSPAPPSPPPPSPEPPSP---APSPPPPSPEPPSPVPPSPPP 1175
Query: 138 P 138
P
Sbjct: 1176 P 1176
Score = 37.7 bits (86), Expect = 0.45
Identities = 38/143 (26%), Positives = 46/143 (31%), Gaps = 1/143 (0%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
SP P + P P + P AP P + +P + PS S P
Sbjct: 989 SPAPLPPPPSPEPPSPAPPSPPPPSPEPPSPAP-PSPPPPSPEPPSPAPPSPPPPSPEPP 1047
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
+P P + A P S PP P+P AP SP A P S PP
Sbjct: 1048 SPAPPSPAPPSPEPPSPAPPSPAPPSPEPPSPAPPSPAPPSPEPPSPAPPSPRPPSPEPP 1107
Query: 137 YPFSFLLSHPYLEKLKGAPSQNP 159
P P E AP P
Sbjct: 1108 SPVPPSPPPPSPEPPSPAPPSPP 1130
Score = 37.0 bits (84), Expect = 0.76
Identities = 32/126 (25%), Positives = 46/126 (36%), Gaps = 5/126 (3%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
SP + P + P P + P AP P+ + +P + PS S P
Sbjct: 930 SPAPSPPPPSPEPPSPAPPSPPPPSPEPPSPAP-PLPPPPSPEPPSPAPPSPPPPSPEPP 988
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPS----S 132
+P P+ PS P P P P+ P+ P E + + P PS S
Sbjct: 989 SPAPLPPPPSPEPPSPAPPSPPPPSPEPPSPAPPSPPPPSPEPPSPAPPSPPPPSPEPPS 1048
Query: 133 SSPPYP 138
+PP P
Sbjct: 1049 PAPPSP 1054
Score = 36.6 bits (83), Expect = 1.00
Identities = 27/122 (22%), Positives = 38/122 (31%)
Query: 17 SPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPD 76
S S SP+ P P + F P P + P +P+
Sbjct: 1214 SAPSPPSPQPPSPLAPSPPSPAPQAPSSPFPPPQPTAPTAPPPPFPPSPAPPSPTPPSPE 1273
Query: 77 APRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPP 136
P P + +P S PP P+P P SP ++ + P S PP
Sbjct: 1274 PPAPQPPSPTPHAPPSPEPPSPTPPSPLPPSPEPPSPSPPSPAPSVPSPPSPAPPSPMPP 1333
Query: 137 YP 138
P
Sbjct: 1334 SP 1335
Score = 35.4 bits (80), Expect = 2.2
Identities = 26/91 (28%), Positives = 37/91 (40%), Gaps = 3/91 (3%)
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
+P+P + P + + PS S P +P P + A P S PP P+P P
Sbjct: 925 SPEPPSPAPSPPPPSPEPPSPAPPSPPPPSPEPPSPAPPLPPPPSPEPPSPAPPSPP--P 982
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+P P S +P P+ SPP P
Sbjct: 983 PSPEPPSPAPLPPPPS-PEPPSPAPPSPPPP 1012
Score = 35.4 bits (80), Expect = 2.2
Identities = 40/144 (27%), Positives = 49/144 (33%), Gaps = 3/144 (2%)
Query: 17 SPVSACSPKRS-KVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTP 75
SPV P S + P P + P AP P + +P + PS S P
Sbjct: 840 SPVPPSPPPASPEPTSPAPPSPPPPSPEPPSPAP-PSPPPPSPEPPSPAPPSPPLPSPEP 898
Query: 76 DAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
+P P P S PP P P AP+ P E + + P P S P
Sbjct: 899 PSPAPPLPPPPSPEPPSPAPPSPPPPSPEPPSPAPSPPPPSPEPPSPAPPSPP-PPSPEP 957
Query: 136 PYPFSFLLSHPYLEKLKGAPSQNP 159
P P L P E AP P
Sbjct: 958 PSPAPPLPPPPSPEPPSPAPPSPP 981
Score = 35.0 bits (79), Expect = 2.9
Identities = 25/83 (30%), Positives = 33/83 (39%), Gaps = 13/83 (15%)
Query: 59 DPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPI---PSPVKVTLGPSAPNSPGV 115
+P + PS + S P +P P + A SP S PP PSP + P +P P
Sbjct: 1291 EPPSPTPPSPLPPSPEPPSPSPPSPAPSVPSPPSPAPPSPMPPSPAPLAPQPPSPTPP-- 1348
Query: 116 GGEGENLSAAQPTGPSSSSPPYP 138
+ P P S PP P
Sbjct: 1349 --------SPAPPVPPSPEPPVP 1363
Score = 35.0 bits (79), Expect = 2.9
Identities = 35/126 (27%), Positives = 46/126 (35%), Gaps = 23/126 (18%)
Query: 23 SPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSE-----VNASQTPDA 77
SP+ PDP P P T + P + PS + S P +
Sbjct: 1357 SPEPPVPPGPDPPLPPSPTPPSPQPPVPPSPTPPSPQPPSPAPPSPAPSAPLQPSPDPPS 1416
Query: 78 PRPVTLAEG-----QRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSS 132
P+P + A G PS+ PP P+P L P+ P P A QP P
Sbjct: 1417 PQPPSPAPGPPPSPPSPPSTPSPPSPAP----LAPAPPVPP---------MAPQPPSPPL 1463
Query: 133 SSPPYP 138
SPP+P
Sbjct: 1464 PSPPFP 1469
Score = 34.7 bits (78), Expect = 3.8
Identities = 34/131 (25%), Positives = 46/131 (34%), Gaps = 19/131 (14%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
P S P + P P P + P P +P + PS A +P+
Sbjct: 1026 PPSPEPPSPAPPSPPPPSPEPPSPAPPSPAPPSP-------EPPSPAPPSP--APPSPEP 1076
Query: 78 PRPVTLAEGQRSPS-------SIRPPI---PSPVKVTLGPSAPNSPGVGGEGENLSAAQP 127
P P + SP S RPP PSPV + P +P P + +P
Sbjct: 1077 PSPAPPSPAPPSPEPPSPAPPSPRPPSPEPPSPVPPSPPPPSPEPPSPAPPSPPPPSLEP 1136
Query: 128 TGPSSSSPPYP 138
P+ SPP P
Sbjct: 1137 PSPAPPSPPPP 1147
Score = 34.3 bits (77), Expect = 5.0
Identities = 31/117 (26%), Positives = 42/117 (35%), Gaps = 13/117 (11%)
Query: 24 PKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTL 83
P + P P + PL AP P+ + +P + PS S P +P P
Sbjct: 878 PPSPEPPSPAPPSPPLPSPEPPSPAP-PLPPPPSPEPPSPAPPSPPPPSPEPPSPAPSPP 936
Query: 84 AEGQRSPSSIRP--PIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
PS P P PSP + P P P + +P P+ SPP P
Sbjct: 937 PPSPEPPSPAPPSPPPPSPEPPSPAPPLPPPP----------SPEPPSPAPPSPPPP 983
Score = 33.9 bits (76), Expect = 6.5
Identities = 22/73 (30%), Positives = 28/73 (38%), Gaps = 2/73 (2%)
Query: 66 PSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAA 125
PS S TP +P P + SP S P +PSP P +P P +
Sbjct: 1288 PSPEPPSPTPPSPLPPSPEPPSPSPPSPAPSVPSPPSPA--PPSPMPPSPAPLAPQPPSP 1345
Query: 126 QPTGPSSSSPPYP 138
P P+ PP P
Sbjct: 1346 TPPSPAPPVPPSP 1358
Score = 33.9 bits (76), Expect = 6.5
Identities = 23/91 (25%), Positives = 32/91 (34%), Gaps = 2/91 (2%)
Query: 48 APDPVVTDSACDPLASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGP 107
+P+P P S P P +P P + A P S PP P+P P
Sbjct: 969 SPEPPSPAPPSPPPPSPEPPSPAPLPPPPSPEPPSPAPPSPPPPSPEPPSPAPPSPP--P 1026
Query: 108 SAPNSPGVGGEGENLSAAQPTGPSSSSPPYP 138
+P P + +P P+ SP P
Sbjct: 1027 PSPEPPSPAPPSPPPPSPEPPSPAPPSPAPP 1057
>UniRef100_Q6FPR1 Similarities with sp|P08640 Saccharomyces cerevisiae YIR019c STA1
[Candida glabrata]
Length = 979
Score = 45.8 bits (107), Expect = 0.002
Identities = 31/114 (27%), Positives = 50/114 (43%), Gaps = 9/114 (7%)
Query: 54 TDSACDPLASDHPSEVN-ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNS 112
TDS C+ + PS V +S P + P ++ +PSS+ P +P + P+S
Sbjct: 320 TDSKCEDIPPIEPSSVKPSSMNPSSVNPSSMNPSSMNPSSMNPSSMNPSSMNPSSMNPSS 379
Query: 113 PGVGGEGENLSAAQPTGPSSSSPPYPFSFLLS----HPYLEKLKGAPSQNPQDV 162
P N S+ P+ PSS +PP+ S ++ +P S NP +
Sbjct: 380 P----SSINPSSMNPSSPSSMNPPFANSSSMNPSSMNPSSMNPSSPSSMNPSSI 429
Score = 43.1 bits (100), Expect = 0.011
Identities = 43/169 (25%), Positives = 69/169 (40%), Gaps = 14/169 (8%)
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
S+ P SSP S+ +P +P + + + P V S+ +P
Sbjct: 405 SMNPSSMNPSSMNPSSP-SSMNPSSINPSSMNPSSINPSSMNPSSMNPSSV-NPSSMNP- 461
Query: 62 ASDHPSEVN------ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVT-LGPSAPNSPG 114
+S +PS VN +S P + P ++ +PSS+ P PS + + + PS+P+S
Sbjct: 462 SSVNPSSVNPSSMNPSSMNPSSMNPSSMNPSSVNPSSMNPSSPSSMNPSSMNPSSPSSMN 521
Query: 115 VGGEGE----NLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNP 159
+ S+ P+ PSS +P P S S P S NP
Sbjct: 522 PSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNP 570
Score = 35.8 bits (81), Expect = 1.7
Identities = 36/169 (21%), Positives = 66/169 (38%), Gaps = 6/169 (3%)
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQT-TPLTGTGGGFVAPDPVVTDSACDP 60
S+ P +P S S S +P + + + + + P + + P
Sbjct: 506 SMNPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSPSSMNPSSP 565
Query: 61 LASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGE 120
+ + P ++S P + P ++ +PSS+ P +P ++ PS+ N +
Sbjct: 566 SSMNPPFANSSSMNPSSVNPSSMNPSSMNPSSMNPSSMNPS--SMNPSSMNPSSMNPSSM 623
Query: 121 NLSAAQPTG--PSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEAV 167
N S+ P+ PSS +P P S S + + S NP V +V
Sbjct: 624 NPSSMNPSSMNPSSMNPSSPSSMNPSSINPSSMNPS-SMNPSSVNPSSV 671
>UniRef100_UPI00001CE7BF UPI00001CE7BF UniRef100 entry
Length = 2759
Score = 45.4 bits (106), Expect = 0.002
Identities = 40/143 (27%), Positives = 59/143 (40%), Gaps = 29/143 (20%)
Query: 14 RQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
RQ SP+ P +P P + PV+ + PL S P EV A+
Sbjct: 939 RQPSPLLLLPPPAGLTSDPGPSVRRVPAVQ----RDSPVIVRNPDVPLPSKFPGEVGAAG 994
Query: 74 TPDA----------PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLS 123
A P P +A G+ S+ PP+P+PV +T+ P+AP + ++
Sbjct: 995 EARAGGPGRGCRETPVPPGVASGK---PSLPPPLPAPVPITVPPAAPTA---------VA 1042
Query: 124 AAQPTGPSSSSPPYPFSFLLSHP 146
PT +SSP P +F HP
Sbjct: 1043 QPMPTLGLASSPFQPVAF---HP 1062
Score = 33.5 bits (75), Expect = 8.5
Identities = 34/99 (34%), Positives = 41/99 (41%), Gaps = 9/99 (9%)
Query: 23 SPKRSKVGE-PDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDAPRPV 81
SP R K P P T +G GGG APD +DS + D S TP R V
Sbjct: 872 SPGRRKTELLPHPGTLGASGAGGGGAAPDFPKSDSLDSGV--DSVSHTPTPSTPAGFRAV 929
Query: 82 TLA-----EGQRSPSSIRPPIPSPVKVTLGPSAPNSPGV 115
+ A Q SP + PP P+ + GPS P V
Sbjct: 930 SPAVPFSRSRQPSPLLLLPP-PAGLTSDPGPSVRRVPAV 967
>UniRef100_O15047 KIAA0339 protein [Homo sapiens]
Length = 1709
Score = 45.4 bits (106), Expect = 0.002
Identities = 31/126 (24%), Positives = 49/126 (38%), Gaps = 4/126 (3%)
Query: 11 GGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN 70
G +S S CS GE D + + + + + S+ +S SE +
Sbjct: 1008 GEDEESDSSSKCSLYADSDGENDSTSDSESSSSSSSSSS----SSSSSSSSSSSSSSESS 1063
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
+ + RP L P + P P+PV+V + SP + S A+P GP
Sbjct: 1064 SEDEEEEERPAALPSASPPPREVPVPTPAPVEVPVPERVAGSPVTPLPEQEASPARPAGP 1123
Query: 131 SSSSPP 136
+ SPP
Sbjct: 1124 TEESPP 1129
>UniRef100_UPI0000437028 UPI0000437028 UniRef100 entry
Length = 728
Score = 45.1 bits (105), Expect = 0.003
Identities = 42/155 (27%), Positives = 58/155 (37%), Gaps = 26/155 (16%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVA-PDPVVTDSACDPLASDHPSEVNASQ- 73
S P S P + +P P + P + G P P SA P + P++ SQ
Sbjct: 248 SQPASQARPGQPSPAQPTP-SQPASQPGQARPGQPSPAHPQSASQPASQPRPAKPTPSQP 306
Query: 74 -----TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAP---------NSPGVGGEG 119
+P P P +++ P RP PSP V+ S P + PG G
Sbjct: 307 ASQPASPGQPSPPPVSQPASQPGQARPAKPSPPPVSQPASQPAKPTPSQPASQPGQARPG 366
Query: 120 E--------NLSAAQPTGPSSSSPPYPFSFLLSHP 146
+ + A+QP P SPP P S S P
Sbjct: 367 QASQAQPTPSQPASQPASPGQPSPP-PVSQPASQP 400
Score = 44.3 bits (103), Expect = 0.005
Identities = 45/165 (27%), Positives = 64/165 (38%), Gaps = 22/165 (13%)
Query: 16 SSPVSACSPKRSKVGEPDP-QTTPLTGTGG------GFVAP----DPVVTDSACDPLASD 64
S P S P +++ G+P P Q +P T G P P V+ A P +
Sbjct: 540 SQPASQARPGQARPGQPSPAQPSPAQPTPSQPASQPGQARPAKPSPPPVSQPASQPAQAS 599
Query: 65 --HPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENL 122
HP + + A +P + Q SP+ P PS + S P PG
Sbjct: 600 QAHPQSASQPASQPASQPGQASPAQPSPAQPSPGQPSQPTPSQPASQPARPG------QA 653
Query: 123 SAAQPTGPSSSSP---PYPFSFLLSHPYLEKLKGAPSQNPQDVAQ 164
S AQP+ P S P P P S P + + +P+Q Q +Q
Sbjct: 654 SQAQPSPPPVSQPASQPRPAKPTPSQPASQPGQASPAQPSQPASQ 698
Score = 42.4 bits (98), Expect = 0.018
Identities = 41/157 (26%), Positives = 61/157 (38%), Gaps = 17/157 (10%)
Query: 2 SVKPRVAEVGGKR---QSSPVSACSPKRSKVGEPDPQTTPLT-----GTGGGFVAPDPVV 53
S KPR A + PVS + + ++ G+P P P++ G P V
Sbjct: 67 SGKPRPASQPASQVQPSPPPVSQPASQPARPGQPSPAPPPVSQPARPGQASQAQPSPPPV 126
Query: 54 TDSACDPLAS----DHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSA 109
+ A P + P+ A TP P GQ P+ P PSP V+ P+
Sbjct: 127 SQPASQPRPAKPTPSQPASQPAKPTPSQPASQPARPGQARPAKPSPAQPSPPSVSQPPAK 186
Query: 110 PNSPGVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHP 146
P+ + + A+QP P+ +P P S S P
Sbjct: 187 PSP----SQPASQPASQPR-PAKPTPSQPASQPASQP 218
Score = 41.6 bits (96), Expect = 0.031
Identities = 45/166 (27%), Positives = 61/166 (36%), Gaps = 19/166 (11%)
Query: 12 GKRQSSPVS--ACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEV 69
G+ PVS A P +++ +P P P++ P P + A P +
Sbjct: 314 GQPSPPPVSQPASQPGQARPAKPSPP--PVSQPASQPAKPTP--SQPASQPGQARPGQAS 369
Query: 70 NASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLG--------PSAPNSPGVGGEGEN 121
A TP P + GQ SP + P P + G P+ P+ P V G+
Sbjct: 370 QAQPTPSQPASQPASPGQPSPPPVSQPASQPGQARPGQASQAQPSPAQPSPPSVSQPGQP 429
Query: 122 LSAA--QPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQE 165
A QP P SPP P S S P K PSQ A +
Sbjct: 430 SPAHPHQPASPGQPSPP-PVSQPASQP--RPAKPTPSQPASQPASQ 472
Score = 40.0 bits (92), Expect = 0.090
Identities = 45/159 (28%), Positives = 59/159 (36%), Gaps = 28/159 (17%)
Query: 2 SVKPRVAEVGGKRQSSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPL 61
S KPR A + +S P +++ G+P P + P + P P A
Sbjct: 21 SGKPRPAS----QPASQPPPIQPGQARSGKPRPASQPAS-------QPPPAQPGQARS-- 67
Query: 62 ASDHPSEVNASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGEN 121
P+ ASQ +P PV+ Q + P P PV S P PG
Sbjct: 68 GKPRPASQPASQVQPSPPPVSQPASQPARPGQPSPAPPPV------SQPARPG------Q 115
Query: 122 LSAAQPTGPSSSSP---PYPFSFLLSHPYLEKLKGAPSQ 157
S AQP+ P S P P P S P + K PSQ
Sbjct: 116 ASQAQPSPPPVSQPASQPRPAKPTPSQPASQPAKPTPSQ 154
Score = 38.5 bits (88), Expect = 0.26
Identities = 33/128 (25%), Positives = 44/128 (33%), Gaps = 11/128 (8%)
Query: 15 QSSPVSACSPKRSKVGEPDP----QTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN 70
Q+ P SA P +P Q +P + G P P AS
Sbjct: 600 QAHPQSASQPASQPASQPGQASPAQPSPAQPSPGQPSQPTP-------SQPASQPARPGQ 652
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
ASQ +P PV+ Q P+ P P+ P+ P+ P + A PT P
Sbjct: 653 ASQAQPSPPPVSQPASQPRPAKPTPSQPASQPGQASPAQPSQPASQPRPAKPAQANPTSP 712
Query: 131 SSSSPPYP 138
S P P
Sbjct: 713 GQPSQPTP 720
Score = 37.4 bits (85), Expect = 0.59
Identities = 37/149 (24%), Positives = 56/149 (36%), Gaps = 15/149 (10%)
Query: 15 QSSPVSACSP-KRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
Q SP P +++ G+P P P + + P + A P + P V SQ
Sbjct: 485 QPSPPPVSQPASQARPGQPSP-AHPQSASQPASQPAKPTPSQPASQPASQPSPPPV--SQ 541
Query: 74 TPDAPRPVTLAEGQRSPSSIRPPIPSPVKVT-----LGPSAPNSPGVGGEGENLSAAQPT 128
RP GQ SP+ P P+P + P+ P+ P V + A
Sbjct: 542 PASQARPGQARPGQPSPAQPSPAQPTPSQPASQPGQARPAKPSPPPVSQPASQPAQASQA 601
Query: 129 GPSSSSPPYPFSFLLSHPYLEKLKGAPSQ 157
P S+S P S P + + +P+Q
Sbjct: 602 HPQSASQP------ASQPASQPGQASPAQ 624
Score = 37.0 bits (84), Expect = 0.76
Identities = 42/150 (28%), Positives = 56/150 (37%), Gaps = 14/150 (9%)
Query: 16 SSPVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ-T 74
S P S P +P P G +P PV + A P + P + + +Q T
Sbjct: 212 SQPASQPRP-----AQPTPSQPASQPASPGQPSPPPV-SQPASQPASQARPGQPSPAQPT 265
Query: 75 PDAP--RPVTLAEGQRSPS---SIRPPIPSPVKVTLGPSAPNS-PGVGGEGENLSAAQPT 128
P P +P GQ SP+ S P P PS P S P G+ +QP
Sbjct: 266 PSQPASQPGQARPGQPSPAHPQSASQPASQPRPAKPTPSQPASQPASPGQPSPPPVSQPA 325
Query: 129 G-PSSSSPPYPFSFLLSHPYLEKLKGAPSQ 157
P + P P +S P + K PSQ
Sbjct: 326 SQPGQARPAKPSPPPVSQPASQPAKPTPSQ 355
Score = 37.0 bits (84), Expect = 0.76
Identities = 35/113 (30%), Positives = 40/113 (34%), Gaps = 16/113 (14%)
Query: 66 PSEVNASQTPDA---------PRPVTLAEGQR---SPSSIRPPIPSPVKVTLGPSAPNSP 113
P+ ASQ P A PRP + Q P R P P P P
Sbjct: 3 PASQPASQPPPAQPGQARSGKPRPASQPASQPPPIQPGQARSGKPRPASQPASQPPPAQP 62
Query: 114 GVGGEGENLSAAQPTGPSSSSPPYPFSFLLSHPYLEKLKGAPSQNPQDVAQEA 166
G G+ A+QP SPP P S S P G PS P V+Q A
Sbjct: 63 GQARSGKPRPASQPASQVQPSPP-PVSQPASQP---ARPGQPSPAPPPVSQPA 111
Score = 36.6 bits (83), Expect = 1.00
Identities = 42/161 (26%), Positives = 54/161 (33%), Gaps = 11/161 (6%)
Query: 16 SSPVS--ACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQ 73
S P S A P+ +K P + P + P + A S P ASQ
Sbjct: 190 SQPASQPASQPRPAKPTPSQPASQPASQPRPAQPTPSQPASQPASPGQPSPPPVSQPASQ 249
Query: 74 TPDAPRPVTLAEGQRSPS-------SIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQ 126
RP + Q +PS RP PSP + P + A+Q
Sbjct: 250 PASQARPGQPSPAQPTPSQPASQPGQARPGQPSPAHPQSASQPASQPRPAKPTPSQPASQ 309
Query: 127 PTGPSSSSPPYPFSFLLSHP-YLEKLKGAPSQNPQDVAQEA 166
P P SPP P S S P K +P Q +Q A
Sbjct: 310 PASPGQPSPP-PVSQPASQPGQARPAKPSPPPVSQPASQPA 349
Score = 34.7 bits (78), Expect = 3.8
Identities = 34/126 (26%), Positives = 46/126 (35%), Gaps = 10/126 (7%)
Query: 16 SSPVSA-CSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVN---- 70
S P S P +++ +P P P P + A P + P++
Sbjct: 153 SQPASQPARPGQARPAKPSPAQPSPPSVSQPPAKPSP--SQPASQPASQPRPAKPTPSQP 210
Query: 71 ASQTPDAPRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGP 130
ASQ PRP Q + P PSP V+ S P S G+ S AQPT
Sbjct: 211 ASQPASQPRPAQPTPSQPASQPASPGQPSPPPVSQPASQPASQARPGQP---SPAQPTPS 267
Query: 131 SSSSPP 136
+S P
Sbjct: 268 QPASQP 273
>UniRef100_UPI0000248D73 UPI0000248D73 UniRef100 entry
Length = 428
Score = 45.1 bits (105), Expect = 0.003
Identities = 39/140 (27%), Positives = 53/140 (37%), Gaps = 11/140 (7%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
PVSA +P+R V P P+ P++ AP P P P A + P
Sbjct: 173 PVSAPAPERPPVSAPAPERPPVS-------APAP-ERPPVSAPAPERPPVSAPAPERPPV 224
Query: 78 PRPVTLAEGQRSPSSIRPPI--PSPVKVTLGPSAPNSPGVGGEGENLSAAQPTGPSSSSP 135
P +P+ RPP+ P+P + + AP P V P SS
Sbjct: 225 SAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERSSM 284
Query: 136 PYPFSFLLSHPYLEKLKGAP 155
P P +LL+ P KL P
Sbjct: 285 PVPV-WLLALPAPLKLLALP 303
Score = 41.6 bits (96), Expect = 0.031
Identities = 39/130 (30%), Positives = 58/130 (44%), Gaps = 17/130 (13%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPD-PVVTDSACD--PLASDHPSEVNASQT 74
PVSA +P+R V P P+ P++ AP+ P V+ A + P+++ P S
Sbjct: 113 PVSAPAPERPPVSAPAPERPPVSAP-----APERPPVSAPAPERPPVSAPAPERPPVSAP 167
Query: 75 PDAPRPVTLAEGQRSPSSI----RPPI--PSPVKVTLGPSAPNSPGVGG---EGENLSAA 125
PV+ +R P S RPP+ P+P + + AP P V E +SA
Sbjct: 168 APERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAP 227
Query: 126 QPTGPSSSSP 135
P P S+P
Sbjct: 228 APERPPVSAP 237
Score = 41.6 bits (96), Expect = 0.031
Identities = 39/130 (30%), Positives = 58/130 (44%), Gaps = 17/130 (13%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPD-PVVTDSACD--PLASDHPSEVNASQT 74
PVSA +P+R V P P+ P++ AP+ P V+ A + P+++ P S
Sbjct: 133 PVSAPAPERPPVSAPAPERPPVSAP-----APERPPVSAPAPERPPVSAPAPERPPVSAP 187
Query: 75 PDAPRPVTLAEGQRSPSSI----RPPI--PSPVKVTLGPSAPNSPGVGG---EGENLSAA 125
PV+ +R P S RPP+ P+P + + AP P V E +SA
Sbjct: 188 APERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAPAPERPPVSAP 247
Query: 126 QPTGPSSSSP 135
P P S+P
Sbjct: 248 APERPPVSAP 257
Score = 37.0 bits (84), Expect = 0.76
Identities = 26/93 (27%), Positives = 39/93 (40%), Gaps = 7/93 (7%)
Query: 18 PVSACSPKRSKVGEPDPQTTPLTGTGGGFVAPDPVVTDSACDPLASDHPSEVNASQTPDA 77
PVSA +P+R V P P+ P++ AP P + P+ + P
Sbjct: 253 PVSAPAPERPPVSAPAPERPPVS-------APAPERSSMPVPVWLLALPAPLKLLALPVP 305
Query: 78 PRPVTLAEGQRSPSSIRPPIPSPVKVTLGPSAP 110
P PV L +P + P P+P ++ P AP
Sbjct: 306 PAPVQLPPVPSAPVQLPPVPPAPAQLPPVPPAP 338
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.310 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 593,911,530
Number of Sequences: 2790947
Number of extensions: 27647277
Number of successful extensions: 141165
Number of sequences better than 10.0: 3203
Number of HSP's better than 10.0 without gapping: 653
Number of HSP's successfully gapped in prelim test: 2671
Number of HSP's that attempted gapping in prelim test: 127851
Number of HSP's gapped (non-prelim): 11378
length of query: 337
length of database: 848,049,833
effective HSP length: 128
effective length of query: 209
effective length of database: 490,808,617
effective search space: 102579000953
effective search space used: 102579000953
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0094b.3


