

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0091a.6
(242 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
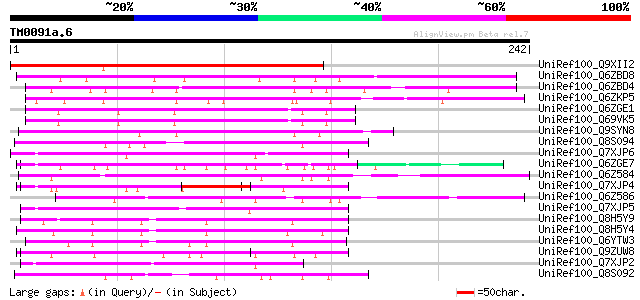
UniRef100_Q9XII2 Hypothetical protein At2g16050 [Arabidopsis tha... 259 4e-68
UniRef100_Q6ZBD8 Hypothetical protein P0685B10.17 [Oryza sativa] 162 5e-39
UniRef100_Q6ZBD4 Hypothetical protein P0685B10.21 [Oryza sativa] 119 5e-26
UniRef100_Q6ZKP5 Hypothetical protein OJ1118_A03.8 [Oryza sativa] 116 4e-25
UniRef100_Q6ZGE1 Putative disease-resistant protein [Oryza sativa] 112 1e-23
UniRef100_Q69VK5 Putative disease-resistant protein [Oryza sativa] 112 1e-23
UniRef100_Q9SYN8 F9H16.2 protein [Arabidopsis thaliana] 106 6e-22
UniRef100_Q8S094 P0470A12.25 protein [Oryza sativa] 106 6e-22
UniRef100_Q7XJP6 At2g37820 protein [Arabidopsis thaliana] 100 2e-20
UniRef100_Q6ZGE7 Hypothetical protein OJ1297_D02.2 [Oryza sativa] 100 4e-20
UniRef100_Q6Z584 Hypothetical protein OSJNBa0005L24.29 [Oryza sa... 97 5e-19
UniRef100_Q7XJP4 At2g37800 protein [Arabidopsis thaliana] 95 2e-18
UniRef100_Q6Z586 Hypothetical protein OSJNBa0005L24.22 [Oryza sa... 93 7e-18
UniRef100_Q7XJP5 At2g37810 protein [Arabidopsis thaliana] 89 1e-16
UniRef100_Q8H5Y9 CHP-rich zinc finger protein-like [Oryza sativa] 79 1e-13
UniRef100_Q8H5Y4 CHP-rich zinc finger protein-like [Oryza sativa] 77 5e-13
UniRef100_Q6YTW3 CHP-rich zinc finger protein-like [Oryza sativa] 76 9e-13
UniRef100_Q9ZUW8 Hypothetical protein At2g27660 [Arabidopsis tha... 75 1e-12
UniRef100_Q7XJP2 At2g37780 protein [Arabidopsis thaliana] 75 1e-12
UniRef100_Q8S092 P0470A12.27 protein [Oryza sativa] 72 9e-12
>UniRef100_Q9XII2 Hypothetical protein At2g16050 [Arabidopsis thaliana]
Length = 167
Score = 259 bits (662), Expect = 4e-68
Identities = 109/151 (72%), Positives = 128/151 (84%), Gaps = 5/151 (3%)
Query: 1 MKYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCS-----ICDYDLHMQCAIP 55
MKY++ISHFSHPQH L+F++ PFKCDGC E+GIGSRY+CS CD+DLH CA+P
Sbjct: 1 MKYADISHFSHPQHNLKFDYTEKPFKCDGCNEVGIGSRYRCSGDNHGSCDFDLHAHCALP 60
Query: 56 SPTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVL 115
S ++ HPFY KC+FQF++ PPGN RYCNACEKDV+GF+YHCK+CGFDLHPCCA LPMVL
Sbjct: 61 SASITHPFYKKCTFQFVARPPGNERRYCNACEKDVSGFLYHCKACGFDLHPCCAMLPMVL 120
Query: 116 DDGEVKLYLYRKVSSPCHRCGRKGRSWSYRS 146
DDGE L+LYRKVSS CHRCG+KGRSWSYRS
Sbjct: 121 DDGETNLFLYRKVSSSCHRCGKKGRSWSYRS 151
>UniRef100_Q6ZBD8 Hypothetical protein P0685B10.17 [Oryza sativa]
Length = 269
Score = 162 bits (411), Expect = 5e-39
Identities = 91/255 (35%), Positives = 135/255 (52%), Gaps = 23/255 (9%)
Query: 4 SEISHFSHPQHKLRFEHNA-FPFKCDGCKEIG-IGSRYKCSICDYDLHMQCAIPSPTLVH 61
++++H++HP+H+L A PF+CDGC+E G G RY+C+ C++DLH CA+P TL H
Sbjct: 9 AKLTHWAHPEHELTLAATAGAPFRCDGCQEPGGDGPRYRCAPCNFDLHTDCALPPATLQH 68
Query: 62 PFYTK---CSFQFMSSPPGNIP--RYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVL- 115
P K C+F F+ PP R C+AC DV GFV+HC DLHPCCA L +
Sbjct: 69 PLLFKGGGCTFVFLREPPAPAAASRQCDACGDDVRGFVFHCADRDLDLHPCCASLEDRIV 128
Query: 116 ----DDGEVKLYLYRKVSSP------CHRCGRKGRS--WSYRSKCKSYN--LHVACVREM 161
DG+ +++ K +S C CG K R W YR + + +HVACV+E+
Sbjct: 129 TGGGGDGDGRVFELTKAASSSSSRRRCGVCGDKSRRTFWFYRGRFDGEDVFIHVACVKEL 188
Query: 162 VVENWDHVYVAGGKGRNKVVETSVPGLKNTLYAAHNSRRSKGKVKKCCEIAGLAVQFVIS 221
V W+ Y G ++ P ++ L + R G ++ +I G+ V +I+
Sbjct: 189 AVRRWEASY-RRRSGAGQIALAGAPLMEGALQSLPRRTRRSGGFERFSKIVGVIVSAIIA 247
Query: 222 AVLGDPTALIAGVVG 236
+ G+P LIA V G
Sbjct: 248 VIFGNPMGLIAAVAG 262
>UniRef100_Q6ZBD4 Hypothetical protein P0685B10.21 [Oryza sativa]
Length = 839
Score = 119 bits (299), Expect = 5e-26
Identities = 92/270 (34%), Positives = 125/270 (46%), Gaps = 48/270 (17%)
Query: 8 HFSHPQHKLRF----EHNAFPFKCDGCKEIGIG--SRYKCSI--CDYDLHMQCAIPSPTL 59
H HP HKL+ + A F CDGCKE+G +RY+C CD+DLH CA+ L
Sbjct: 18 HPFHPAHKLKLITADDAGAGRFVCDGCKELGGAGCARYECEEAGCDFDLHAPCALAPDVL 77
Query: 60 VHPFYT----KCSFQFMSSPP-------GNIPRYCNACEKDVNGFVYHCKSCGFDLHPCC 108
SF + PP G++ R C+AC DV GFVYHC DLHPCC
Sbjct: 78 PAGRALFKGGAASFVLLHEPPPTAAPDDGDV-RVCDACGDDVRGFVYHCFDRDLDLHPCC 136
Query: 109 AKLPMVLDDGEVKLYLYRKVSSP--CHRCGRKG-------RSWSYRS---KCKSYNLHVA 156
A LP + G L ++P C C +G W+Y S ++ +LHVA
Sbjct: 137 AHLPGRVALGGAAFELSSGGTAPRRCLLCTEEGSRPHLRRNYWTYSSDDLDGEAVHLHVA 196
Query: 157 CVREMVVE------NWDHVYVAGGKGRNKVVETSVPGLKNTLYAAHNSRRSKG----KVK 206
CV+ M E + H GG GRN +P ++ + AA R+ G K+K
Sbjct: 197 CVKRMAYESSRAGSSSSHRTDGGGGGRN------MPVIRAPVQAAVALRKKNGRPRSKLK 250
Query: 207 KCCEIAGLAVQFVISAVLGDPTALIAGVVG 236
K +I ++ + + GDPTA+ VVG
Sbjct: 251 KLLKIVVFVLRVIAGVLFGDPTAMAVAVVG 280
>UniRef100_Q6ZKP5 Hypothetical protein OJ1118_A03.8 [Oryza sativa]
Length = 275
Score = 116 bits (291), Expect = 4e-25
Identities = 88/266 (33%), Positives = 126/266 (47%), Gaps = 39/266 (14%)
Query: 8 HFSH-PQHKLRFEHNAFP-FKCDGCKEIGIGSRYKCS--ICDYDLHMQCAIPSPTLVHPF 63
H +H P HKL+ FKCDGC E G G RY+C C++DLH CA+ T H
Sbjct: 15 HPAHRPGHKLKLVRTGGQKFKCDGCMEHGDGPRYRCERETCNFDLHTCCALAPATREHRL 74
Query: 64 YTKCSFQFMSSPP----GNIPRYCNACEKDVN--GFVYHCK-----SCGFDLHPCCAKLP 112
+ C+F + PP R C+AC + V+ G VYHC G DLHP CA LP
Sbjct: 75 FPGCTFVLLPEPPPPTAAGERRICDACGEGVHARGLVYHCSGRGDGGLGLDLHPTCASLP 134
Query: 113 MVLDDGEVKLYLYRKVSS----PC--HRCGRKGRSWSYRSKC------KSYNLHVACVRE 160
G +++ RK +S C RCG R W YRS ++ LHVAC++
Sbjct: 135 ARFAVGGGRVFELRKEASRRCAECGEMRCGGGRRFWFYRSYSYADGDGEALYLHVACLKR 194
Query: 161 MVVENWDHVYVAGGKGRNKVVETSVPGLKNTLYAAHNSRR------SKGKVKKCCEIAGL 214
M + Y A R+ V +S P ++ L + +RR G +++ I
Sbjct: 195 MQTQ-----YGAAADVRSVQVMSS-PVMEGVLRSLPPARRRATAAGGGGGLERFLTIVAG 248
Query: 215 AVQFVISAVLGDPTALIAGVVGSLMS 240
++ +I + GDPT LI VG++++
Sbjct: 249 VIRAIIGVIFGDPTFLIELAVGAILN 274
>UniRef100_Q6ZGE1 Putative disease-resistant protein [Oryza sativa]
Length = 846
Score = 112 bits (279), Expect = 1e-23
Identities = 62/169 (36%), Positives = 95/169 (55%), Gaps = 16/169 (9%)
Query: 8 HFSHPQHKLRFEHN--AFPFKCDGCKEIGIGSRYKCSICDYDLH--MQCAIPSPTLVHPF 63
H H LR + + ++P +CDGCKE+G G R+KC C+ ++ M CA TL HP
Sbjct: 597 HTDHMHENLRLDKSVRSWPTQCDGCKELGAGHRFKCEQCNSKVYYDMCCATAPHTLKHPL 656
Query: 64 YTKCSFQFMSSP-PGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKL 122
+ F+F+ P R C+AC ++GFVYHC G DLHP CA+LP+ + + +
Sbjct: 657 FPGSVFRFLRKPLASECGRACDACGDLMHGFVYHCFERGLDLHPRCARLPVRTANVKGYV 716
Query: 123 YLYRKVSSPCHRC--------GRKGRSWSYRS--KCKSYNLHVACVREM 161
R+VS+ C RC G + + WSYRS + + N+H+AC++++
Sbjct: 717 MELRRVSA-CSRCCICMCGKEGYRNKFWSYRSSQEGQDINVHMACLKDL 764
>UniRef100_Q69VK5 Putative disease-resistant protein [Oryza sativa]
Length = 809
Score = 112 bits (279), Expect = 1e-23
Identities = 62/169 (36%), Positives = 95/169 (55%), Gaps = 16/169 (9%)
Query: 8 HFSHPQHKLRFEHN--AFPFKCDGCKEIGIGSRYKCSICDYDLH--MQCAIPSPTLVHPF 63
H H LR + + ++P +CDGCKE+G G R+KC C+ ++ M CA TL HP
Sbjct: 597 HTDHMHENLRLDKSVRSWPTQCDGCKELGAGHRFKCEQCNSKVYYDMCCATAPHTLKHPL 656
Query: 64 YTKCSFQFMSSP-PGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKL 122
+ F+F+ P R C+AC ++GFVYHC G DLHP CA+LP+ + + +
Sbjct: 657 FPGSVFRFLRKPLASECGRACDACGDLMHGFVYHCFERGLDLHPRCARLPVRTANVKGYV 716
Query: 123 YLYRKVSSPCHRC--------GRKGRSWSYRS--KCKSYNLHVACVREM 161
R+VS+ C RC G + + WSYRS + + N+H+AC++++
Sbjct: 717 MELRRVSA-CSRCCICMCGKEGYRNKFWSYRSSQEGQDINVHMACLKDL 764
>UniRef100_Q9SYN8 F9H16.2 protein [Arabidopsis thaliana]
Length = 318
Score = 106 bits (264), Expect = 6e-22
Identities = 64/193 (33%), Positives = 86/193 (44%), Gaps = 21/193 (10%)
Query: 5 EISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL-VHPF 63
E+ HF HPQH L + C GCKE G G RY C CDY LH CA+ P L HPF
Sbjct: 66 EMVHFGHPQHVLVKVELPDIYTCAGCKEEGAGVRYVCQECDYQLHEFCALAPPQLKSHPF 125
Query: 64 YTKCSFQFMSSPP--GNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVK 121
+ + F + P G + C+ C + G+ + CK+C F +HP CA L L +
Sbjct: 126 HYQHQLLFFAKPAKGGIVKSKCDVCGRSPKGYTFRCKACSFQMHPGCAMLSPSLSSSSLH 185
Query: 122 LYLYRKVSSP--------------CHRCGRKGRSWS-YRSKCKSYNLHVACVREMVVENW 166
+ R + S C C R R+ YR Y+LH C ++ V
Sbjct: 186 HHPLRLLPSSSAGSTTGGDSGGFLCGECKRGKRTGRVYRCTVCDYHLHAVCAKDAAVNG- 244
Query: 167 DHVYVAGGKGRNK 179
+ G KGR+K
Sbjct: 245 --LRANGHKGRDK 255
>UniRef100_Q8S094 P0470A12.25 protein [Oryza sativa]
Length = 260
Score = 106 bits (264), Expect = 6e-22
Identities = 64/174 (36%), Positives = 83/174 (46%), Gaps = 17/174 (9%)
Query: 3 YSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSI--CDYDLHMQCAI-PSPTL 59
Y+EI +H +HK+ PF C GCKE+G RY C C++ H C + P T
Sbjct: 4 YTEILDTNHHEHKVCLVRKDEPFICSGCKELGFELRYACHTAGCNHQYHRSCTLQPLNTR 63
Query: 60 VH-PFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDG 118
V PFY F F S A V G+VY+C LHPCCA LP V+
Sbjct: 64 VPAPFYKHDFFFFKS--------VSTAGSSQVRGYVYYCPDKKVSLHPCCADLPRVITTE 115
Query: 119 EVKLYLYRKVSSPCHRC-----GRKGRSWSYRSKCKSYNLHVACVREMVVENWD 167
V+L L RK++ C C G W+Y S K LHVACVR+ +V ++
Sbjct: 116 TVQLKLERKITKKCGMCHERNQGSFSNPWAYASSEKMIQLHVACVRKALVSQFE 169
>UniRef100_Q7XJP6 At2g37820 protein [Arabidopsis thaliana]
Length = 290
Score = 100 bits (250), Expect = 2e-20
Identities = 55/165 (33%), Positives = 78/165 (46%), Gaps = 9/165 (5%)
Query: 1 MKYSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCA---IPSP 57
+K + HF+H H L ++ F CDGCK G G Y+C C+YDLH CA + P
Sbjct: 6 LKRHTVQHFTHI-HPLTEFNSIGDFICDGCKTYGSGKTYRCEPCNYDLHEYCATCPLTLP 64
Query: 58 TLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPM---- 113
T +HP + R C+ C++ V G Y CK C FD+HP C +LP
Sbjct: 65 TFIHPQHELSLVVRKQQSTRQNDRACDICDESVEGLFYRCKICEFDVHPLCTQLPQHVRH 124
Query: 114 VLDDGEVKLYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
VL L +S C C +SW YR + +++H+ C+
Sbjct: 125 VLHPAH-HLEFRPSGASTCMVCHGPCQSWRYRCELCRFDIHMECI 168
Score = 37.7 bits (86), Expect = 0.26
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 1/49 (2%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAI 54
+ H HP H L F + C C RY+C +C +D+HM+C +
Sbjct: 122 VRHVLHPAHHLEFRPSGAS-TCMVCHGPCQSWRYRCELCRFDIHMECIL 169
>UniRef100_Q6ZGE7 Hypothetical protein OJ1297_D02.2 [Oryza sativa]
Length = 576
Score = 100 bits (248), Expect = 4e-20
Identities = 82/293 (27%), Positives = 118/293 (39%), Gaps = 78/293 (26%)
Query: 6 ISHFSHPQHKLRFEHNA--FPFKCDGCKEIGIGSRYKCSIC-DYDLHMQCAIPSPTLVHP 62
I+H SHP+HKLR PFKCD CKE G G RY C C D ++H CA TL H
Sbjct: 271 ITHPSHPEHKLRMVTTTGEAPFKCDACKEPGDGPRYHCLTCEDLNMHKFCAHAPSTLYHH 330
Query: 63 FYTKCSFQFMSSPPGNIP-----------------RYCNACEKDVNGFVYHCKSCGFDLH 105
+ + +F+ ++ PP P R+C+ C V G VYHC DLH
Sbjct: 331 LFGR-TFELLAKPPQGRPEKPHPAATGGGRGESGGRWCDTCGDHVFGLVYHCSGANLDLH 389
Query: 106 PCCA---KLPM----VLDDGEVKLYLYRKV------------------------------ 128
PCCA +LP+ LD + L +K+
Sbjct: 390 PCCASLQELPVQNRETLDQPKESLATPQKLVKEGLAKITIDGVAFDIAAFSKCSLCSRQE 449
Query: 129 -SSPCHRCGRKGR--SWSYRSK------CKSYNLHVACVREMVVENWDHV--YVAGGKGR 177
P H C R R W Y + ++ +LHV+C++++ W GG+
Sbjct: 450 EEGPDHCCRRLRRQEQWCYYNSDVVDGGGEAVSLHVSCIKQIARRRWQAARDMKCGGQIM 509
Query: 178 NKVVETSVPGLKNTLYAAHNSRRSKGKVKKCCEIAGLAVQFVISAVLGDPTAL 230
+ E + G L+ + R I G V+ +I+ + GD TA+
Sbjct: 510 LAISEEMI-GEGGPLHGIPSERDR--------NIVGAVVRVIIAVIFGDRTAM 553
Score = 84.3 bits (207), Expect = 2e-15
Identities = 64/198 (32%), Positives = 93/198 (46%), Gaps = 42/198 (21%)
Query: 4 SEISHFSHPQHKLRFEHNA----FPFKCDGCKEIGIGSR-YKCSICDYDLHMQCAIPSPT 58
+EI+H HP HKLR N +PF+C GCKE+ G R Y+C CD L+ CA T
Sbjct: 13 AEITHPFHP-HKLRLADNVTDGHWPFRCGGCKELAAGRRRYRCEPCDLTLYTCCATAPLT 71
Query: 59 LVHPFYTKCSFQFMSSPP-------------GNIPRYCNACEKDV--NGFVYHCKS---- 99
L HP F+ + PP G C+AC + +GF YHC
Sbjct: 72 LEHPLLPGRHFRLLERPPPPPPWLAADDRGGGGWRPACDACGDLLRGDGFAYHCADGHGL 131
Query: 100 CGFDLHPCCA--KLPMVLDDGEVKLYLYRKVSSPCHRCG---------RKGRSWSYRSKC 148
G +LHP CA +LP+ G + L R+ ++P RCG R G W+ R +
Sbjct: 132 VGLNLHPRCARLRLPVAAARGAAAVKLCRR-AAPRRRCGVCMSGEDGYRHG-FWTCRFRR 189
Query: 149 KS----YNLHVACVREMV 162
++H++C++E++
Sbjct: 190 SGGDELVDVHLSCLKELM 207
>UniRef100_Q6Z584 Hypothetical protein OSJNBa0005L24.29 [Oryza sativa]
Length = 232
Score = 96.7 bits (239), Expect = 5e-19
Identities = 73/247 (29%), Positives = 112/247 (44%), Gaps = 29/247 (11%)
Query: 5 EISHFSHPQHKLRF--EHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHP 62
E+ H + P L + +A F+C GC E G G RY D+ LH CA+ +PTL HP
Sbjct: 4 EVRHPARPGCMLTLHGDADAMAFQCTGCMETGKGPRYTSG--DHVLHTYCALATPTLQHP 61
Query: 63 FYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLD-DGEVK 121
+ +P G C+AC V GF YH + G DLHP CAK+P + G
Sbjct: 62 LVEGIMELRLVAPTGGDAVRCDACYDAVRGFHYHSSTSGVDLHPGCAKMPRSITLRGGTI 121
Query: 122 LYLYRKVSSPCHRC-GRKG--RSWSYRSK---CKSYNLHVACVREMVVENWDHVYVAGGK 175
L +VS C C +G R W YRS+ + LHV C++E + AG
Sbjct: 122 FDLRTEVSHRCTSCKAMEGFYRPWFYRSENNPDQRMYLHVKCIKE--------IQDAGDD 173
Query: 176 GRNKVVETSVPGLKNTLYAAHNSRRSKGKVKKCCEIAGLAVQFVISAVLGDPTALIAGVV 235
+++ + + R+ ++ C+ + V+ V ++GDPT ++ V
Sbjct: 174 DEVRMM----------VRLQERAGRNVRLERRVCKTLVIMVRIVFRLLIGDPTPILTEGV 223
Query: 236 GSLMSRA 242
+++S A
Sbjct: 224 NAIVSMA 230
>UniRef100_Q7XJP4 At2g37800 protein [Arabidopsis thaliana]
Length = 396
Score = 94.7 bits (234), Expect = 2e-18
Identities = 51/160 (31%), Positives = 74/160 (45%), Gaps = 8/160 (5%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYT 65
+ HF+H H L F CDGCK G G Y+C+ CDY+LH CA TL +
Sbjct: 145 VQHFTHI-HPLTKVDGYGEFTCDGCKTYGFGKTYRCTRCDYNLHDHCATCPSTLATFMHP 203
Query: 66 KCSFQFMSSPPGNI---PRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPMVLDDGEVKL 122
+ + + P + R C+ C++ G Y C+ CGFD+HP C +LP +
Sbjct: 204 QHELRLVFRGPEHTHQNKRMCDICDESAEGLYYQCEPCGFDVHPLCTQLPQHVRHVPHPA 263
Query: 123 YLYR----KVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
+L SS C C RSW Y+ ++H+ C+
Sbjct: 264 HLLELSQWGASSICMVCRGAIRSWRYKCGPCGLDVHMECI 303
Score = 51.6 bits (122), Expect = 2e-05
Identities = 35/114 (30%), Positives = 50/114 (43%), Gaps = 15/114 (13%)
Query: 4 SEISHFSHPQHKLRF-----EH-NAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCA-IPS 56
S ++ F HPQH+LR EH + CD C E G Y+C C +D+H C +P
Sbjct: 195 STLATFMHPQHELRLVFRGPEHTHQNKRMCDICDESAEGLYYQCEPCGFDVHPLCTQLPQ 254
Query: 57 PT--LVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCC 108
+ HP + Q+ +S C C + + Y C CG D+H C
Sbjct: 255 HVRHVPHPAHLLELSQWGAS------SICMVCRGAIRSWRYKCGPCGLDVHMEC 302
Score = 48.1 bits (113), Expect = 2e-04
Identities = 16/32 (50%), Positives = 21/32 (65%)
Query: 81 RYCNACEKDVNGFVYHCKSCGFDLHPCCAKLP 112
R C+ C++ G Y CK CGFD+HP C +LP
Sbjct: 34 RMCDICDESAEGLYYQCKPCGFDVHPLCTQLP 65
Score = 36.6 bits (83), Expect = 0.58
Identities = 17/54 (31%), Positives = 23/54 (42%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTL 59
+ H HP H L C C+ RYKC C D+HM+C S ++
Sbjct: 256 VRHVPHPAHLLELSQWGASSICMVCRGAIRSWRYKCGPCGLDVHMECISSSASV 309
>UniRef100_Q6Z586 Hypothetical protein OSJNBa0005L24.22 [Oryza sativa]
Length = 259
Score = 92.8 bits (229), Expect = 7e-18
Identities = 75/238 (31%), Positives = 112/238 (46%), Gaps = 33/238 (13%)
Query: 22 AFPFKCDGCKEIGIGSRYKCSICDYD----LHMQCAIPSPTLVHPFYTKCSFQFMSSPPG 77
A F+CDGC G G+RY + ++ LH CA+ +PTL H K + + P
Sbjct: 34 AVAFQCDGCMLPGEGTRYTSVVDNHPTHLALHTSCALATPTLQHAL-VKGTMELRHEAPA 92
Query: 78 NIPRYCNACEKDVNGFVYHC-----KSCGFDLHPCCAKLPM-VLDDGEVKLYLYRKVSSP 131
C+AC + V GF Y+ K LHPCCA+LP+ + G + L +VS
Sbjct: 93 GGAGVCSACFETVRGFHYYGSRKTGKGEHPKLHPCCARLPVSIAVRGGLTFELRAEVS-- 150
Query: 132 CHRC-GRKGRSWSYRSKC-KSYN-------LHVACVREMVVENWDHVYVAGGKGRNKVVE 182
HRC G + W YR C +S N LHV C+RE++ GG G +
Sbjct: 151 -HRCTGCRAMEWYYRPWCYRSTNSPDHRVYLHVKCIREIMES-------PGGGGGGGAGD 202
Query: 183 TSVPGLKNTLYAAHNSRRSKGKVKKCCEIAGLAVQFVISAVLGDPTALIAGVVGSLMS 240
+ L A S + + +V C+I + V+ V+ ++GDPTAL+ V +++S
Sbjct: 203 EDDRVVARLLERADQSSKLERRV---CKILVILVRVVVRMLIGDPTALLTEGVSAIVS 257
>UniRef100_Q7XJP5 At2g37810 protein [Arabidopsis thaliana]
Length = 233
Score = 89.0 bits (219), Expect = 1e-16
Identities = 51/157 (32%), Positives = 71/157 (44%), Gaps = 8/157 (5%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYT 65
+ HFSH H L F C GCK G G Y+C+ CDY LH CA TL+ +
Sbjct: 11 VQHFSHT-HPLTKVDEYGEFTCGGCKTYGFGKTYRCAPCDYVLHDHCATCPFTLISFMHP 69
Query: 66 KCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAK----LPMVLDDGEVK 121
+ + + N+ CN C + V G YHC++CGFD+HP C + + V +
Sbjct: 70 QHELRLFVNGSENM---CNICHRLVEGVYYHCETCGFDVHPLCTQHHQHVSYVPHPAHLL 126
Query: 122 LYLYRKVSSPCHRCGRKGRSWSYRSKCKSYNLHVACV 158
S+ C C SW Y+ ++HV CV
Sbjct: 127 ELSQCGASNTCMVCRGAILSWRYKCGPCMLDVHVECV 163
Score = 37.4 bits (85), Expect = 0.34
Identities = 18/61 (29%), Positives = 26/61 (42%), Gaps = 1/61 (1%)
Query: 3 YSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPS-PTLVH 61
+ +S+ HP H L C C+ + RYKC C D+H++C S P
Sbjct: 113 HQHVSYVPHPAHLLELSQCGASNTCMVCRGAILSWRYKCGPCMLDVHVECVNSSAPAATE 172
Query: 62 P 62
P
Sbjct: 173 P 173
>UniRef100_Q8H5Y9 CHP-rich zinc finger protein-like [Oryza sativa]
Length = 202
Score = 78.6 bits (192), Expect = 1e-13
Identities = 51/165 (30%), Positives = 75/165 (44%), Gaps = 16/165 (9%)
Query: 6 ISHFSHPQH---KLRFEHNAFPFKCDGCKEIGIGS-RYKCSICDYDLHMQCAIPSPTLV- 60
I+HF+HPQH K R++ + CD C+ G Y+C+ CD+D+H CA +
Sbjct: 7 ITHFAHPQHLLLKTRYDSTS-RHVCDICRAKLSGLVGYRCNACDFDIHQACADYFKKTIS 65
Query: 61 ---HPFYTKCSFQFMSSPPGNIPRYCNACEKDV--NGFVYHCKSCGFDLHPCCAKLPMVL 115
HP++T S P G+ C+ C ++ FVY C C FD+HP C LP +
Sbjct: 66 FFAHPWHT---LTLSSIPDGSTTWSCDLCRENCPRGNFVYRCIQCAFDVHPLCILLPQTI 122
Query: 116 DDGEVKLYLYRKVSS--PCHRCGRKGRSWSYRSKCKSYNLHVACV 158
+ + V S C C W Y+ + LH+ CV
Sbjct: 123 RSPLHQQHDIHMVPSWGRCSACREDLDLWYYQCGFCLFKLHIRCV 167
>UniRef100_Q8H5Y4 CHP-rich zinc finger protein-like [Oryza sativa]
Length = 259
Score = 76.6 bits (187), Expect = 5e-13
Identities = 53/169 (31%), Positives = 74/169 (43%), Gaps = 17/169 (10%)
Query: 4 SEISHFSHPQHKLRFE--HNAFPFKCDGCKEIGIGS-RYKCSICDYDLHMQCAIPSPTLV 60
+ I+HF+HPQH+L +A CD C+ G Y+C+ CD+D+H CA +
Sbjct: 5 AHITHFAHPQHRLLKTPYSSASRHVCDICEAKLSGLVGYRCNACDFDIHEACADYFKETI 64
Query: 61 ----HPFYTKCSFQFMSSPPGNIPRYCNACEKDV--NGFVYHCKSCGFDLHPCCAKLPMV 114
HP++T PP N C+ C + FVY C C FD+HP C LP
Sbjct: 65 SFFAHPWHT---LTLCRMPPENKGWVCDLCMEHCPPGNFVYRCIQCKFDVHPLCTLLPQT 121
Query: 115 LDDGEVKLYLYRKV--SSPCHRCGRKGRSWSY---RSKCKSYNLHVACV 158
+ + V S C+ C + W Y SY LH+ CV
Sbjct: 122 IRSPLHPHHDLNMVPSSGHCNACPERLPVWHYICGPCTSPSYRLHIGCV 170
>UniRef100_Q6YTW3 CHP-rich zinc finger protein-like [Oryza sativa]
Length = 237
Score = 75.9 bits (185), Expect = 9e-13
Identities = 52/162 (32%), Positives = 68/162 (41%), Gaps = 15/162 (9%)
Query: 8 HFSHPQHKLRFEHNAFPFK--CDGCKEIGIGS-RYKCSICDYDLHMQCAIPSPTLV---- 60
HF+HPQH L H + CD C+ G Y+C+ CD D+H CA V
Sbjct: 8 HFAHPQHLLLRTHYSTTSSHFCDICRTNVAGMVGYRCNTCDIDIHDACADYFEETVSFFA 67
Query: 61 HPFYTKCSFQFMSSPPGNIPRY-CNACEKDV--NGFVYHCKSCGFDLHPCCAKLPMVLDD 117
HP++T + P + R+ C+ C + FVY C CGFD HP C LP +
Sbjct: 68 HPWHT---LTLSTIPDDDTNRWSCDLCTESCPRGSFVYRCAQCGFDAHPLCTLLPQTIRS 124
Query: 118 GEVKLYLYRKVSS--PCHRCGRKGRSWSYRSKCKSYNLHVAC 157
+ S C C + W YR Y LHV C
Sbjct: 125 PLHPQHDLNMAPSWGTCSACHQGLNMWHYRCGFCLYKLHVVC 166
>UniRef100_Q9ZUW8 Hypothetical protein At2g27660 [Arabidopsis thaliana]
Length = 718
Score = 75.5 bits (184), Expect = 1e-12
Identities = 48/167 (28%), Positives = 73/167 (42%), Gaps = 13/167 (7%)
Query: 4 SEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSR-YKCSICDYDLHMQCAIPSPTLVHP 62
S I+HFSHP H+L+ C CK G R Y C C++ LH C+ + HP
Sbjct: 10 SFINHFSHP-HRLQLTPATSSPPCSACKLTGGNGRVYSCRPCNFSLHESCSKMKQVITHP 68
Query: 63 FYTKCSFQFMSSPPGNIPRY-CNACEKDVNGFVYHCKSCGFDLHPCCAKLPM-VLDDGEV 120
+ + + +P + + C+ C GF Y C C FD+H CA P+ ++
Sbjct: 69 SHPSHTLSLLVAPVYDGGYFNCDGCGIHGTGFSYQCSVCDFDIHALCAYKPLSIIHKSHP 128
Query: 121 KLYLYRKVSSP--------CHRCGRKGRS-WSYRSKCKSYNLHVACV 158
+ L SP C C + G++ W YR ++ HV C+
Sbjct: 129 QHNLKLAFQSPYGANKGFSCDICRKIGKNQWLYRCIPCEFDAHVGCI 175
Score = 70.9 bits (172), Expect = 3e-11
Identities = 39/114 (34%), Positives = 56/114 (48%), Gaps = 7/114 (6%)
Query: 6 ISHFSHPQHKLRF----EHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVH 61
I+H SHP H L ++ F CDGC G G Y+CS+CD+D+H CA +++H
Sbjct: 65 ITHPSHPSHTLSLLVAPVYDGGYFNCDGCGIHGTGFSYQCSVCDFDIHALCAYKPLSIIH 124
Query: 62 PFYTKCSFQ--FMSSPPGNIPRYCNACEK-DVNGFVYHCKSCGFDLHPCCAKLP 112
+ + + + F S N C+ C K N ++Y C C FD H C P
Sbjct: 125 KSHPQHNLKLAFQSPYGANKGFSCDICRKIGKNQWLYRCIPCEFDAHVGCITGP 178
>UniRef100_Q7XJP2 At2g37780 protein [Arabidopsis thaliana]
Length = 286
Score = 75.5 bits (184), Expect = 1e-12
Identities = 44/138 (31%), Positives = 61/138 (43%), Gaps = 8/138 (5%)
Query: 6 ISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSICDYDLHMQCAIPSPTLVHPFYT 65
I HF+H H L + + CDGCK G G Y+CS CDYDLH CA L++ +
Sbjct: 7 IQHFTH-NHLLTQVNGIGTYTCDGCKLYGEGRTYRCSDCDYDLHEYCATCPSILLNSCHG 65
Query: 66 KCSFQFMSSPPGNIPRYCNACEKDVNGFVYHCKSCGFDLHPCCAKLPM------VLDDGE 119
+ R C C + G Y C+ C F+ HP C PM +L +
Sbjct: 66 P-DHELSLFNGHMTERSCYVCRVSIQGMFYKCRQCSFEAHPLCTYAPMHASSPDLLVTKQ 124
Query: 120 VKLYLYRKVSSPCHRCGR 137
L+ + SP H+ G+
Sbjct: 125 RSLHGHAGQPSPPHQYGQ 142
>UniRef100_Q8S092 P0470A12.27 protein [Oryza sativa]
Length = 280
Score = 72.4 bits (176), Expect = 9e-12
Identities = 59/190 (31%), Positives = 82/190 (43%), Gaps = 33/190 (17%)
Query: 3 YSEISHFSHPQHKLRFEHNAFPFKCDGCKEIGIGSRYKCSI----CDYDLHMQCAIP--S 56
+S++ H H+L F+CDGCKE G RY C + + LH CA
Sbjct: 4 HSKLRAPEHHDHELTLTAGKESFRCDGCKEHGYHMRYVCKLGGCRAGFHLHEACAQHRFG 63
Query: 57 PTLVHPFYTKCSFQFMSSPPGNIPRYCNACEKDVNGFVY--------HCKSCGFDLHPCC 108
+ PF + S F S P + VNG+ Y G LHPCC
Sbjct: 64 DSYQDPF-KRYSLVFHKSLP-------SCARLQVNGYAYVRDIGRLRTLLKRGRVLHPCC 115
Query: 109 AKLPMVLD-DGEV-KLYLYRKVSSPCHRC-----GRKGRSWSYRSK----CKSYNLHVAC 157
A LP V++ +G V KL L RK+ SPC +C G + +W Y S +HVAC
Sbjct: 116 AALPKVIEAEGSVTKLRLTRKLRSPCCKCRHVKLGDRRHTWGYVSDGGGGAGVVQIHVAC 175
Query: 158 VREMVVENWD 167
++ E ++
Sbjct: 176 ANDLFREEYE 185
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.138 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 439,463,568
Number of Sequences: 2790947
Number of extensions: 18909275
Number of successful extensions: 55274
Number of sequences better than 10.0: 369
Number of HSP's better than 10.0 without gapping: 123
Number of HSP's successfully gapped in prelim test: 246
Number of HSP's that attempted gapping in prelim test: 53314
Number of HSP's gapped (non-prelim): 1211
length of query: 242
length of database: 848,049,833
effective HSP length: 124
effective length of query: 118
effective length of database: 501,972,405
effective search space: 59232743790
effective search space used: 59232743790
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0091a.6


