

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089b.6
(647 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
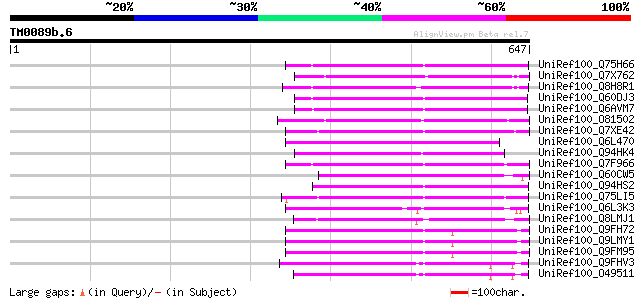
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q75H66 Hypothetical protein OSJNBb0007E22.22 [Oryza sa... 168 5e-40
UniRef100_Q7X762 OSJNBb0118P14.3 protein [Oryza sativa] 163 2e-38
UniRef100_Q8H8R1 Putative mutator-like transposase [Oryza sativa] 162 3e-38
UniRef100_Q60DJ3 MuDR family transposase protein [Oryza sativa] 155 3e-36
UniRef100_Q6AVM7 Putative polyprotein [Oryza sativa] 152 4e-35
UniRef100_O81502 F9D12.2 protein [Arabidopsis thaliana] 144 6e-33
UniRef100_Q7XE42 Putative mutator-like transposase [Oryza sativa] 140 1e-31
UniRef100_Q6L470 Putative mutator transposable element [Solanum ... 139 2e-31
UniRef100_Q94HK4 Mutator-like transposase [Oryza sativa] 138 6e-31
UniRef100_Q7F966 OSJNBa0091C07.2 protein [Oryza sativa] 134 8e-30
UniRef100_Q60CW5 Putative mutator transposable element-related p... 133 2e-29
UniRef100_Q94HS2 Putative transposon protein [Oryza sativa] 129 2e-28
UniRef100_Q75LI5 Putative transposon protein [Oryza sativa] 129 4e-28
UniRef100_Q6L3K3 Putative transposon MuDR mudrA-like protein [So... 127 1e-27
UniRef100_Q8LMJ1 Putative transposon protein [Oryza sativa] 125 3e-27
UniRef100_Q9FH72 Arabidopsis thaliana genomic DNA, chromosome 5,... 121 7e-26
UniRef100_Q9LMY1 F21F23.10 protein [Arabidopsis thaliana] 120 1e-25
UniRef100_Q9FM95 Similarity to mutator-like transposase [Arabido... 120 1e-25
UniRef100_Q9FHV3 Mutator-like transposase-like protein [Arabidop... 114 7e-24
UniRef100_O49511 MuDR transposable element - like protein [Arabi... 114 1e-23
>UniRef100_Q75H66 Hypothetical protein OSJNBb0007E22.22 [Oryza sativa]
Length = 981
Score = 168 bits (425), Expect = 5e-40
Identities = 97/303 (32%), Positives = 153/303 (50%), Gaps = 3/303 (0%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ L EH F +RHLY NF+ F G + ++N + A+++ Q W K E++A
Sbjct: 549 LIPAVQQLFPDSEHRFCVRHLYQNFQQSFKGEI-LKNQLWACARSSSVQEWNTKFEEMKA 607
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
L+ +AY WL Q+ + W + F+ + KCD+L+N E FN IL AR+ P+LTM++ I+
Sbjct: 608 LNEDAYNWLEQMAPNTWVRAFFSDFPKCDILLNNSCEVFNKYILEAREMPILTMLEKIKG 667
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
+M RF ++ K+ + PK ++L + + G +F+V+ ++I
Sbjct: 668 QLMTRFFNKQKEAQKWQGPICPKIRKKLLKIAEQANICYVLPAGKGVFQVEE--RGTKYI 725
Query: 524 VDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISG 583
VD+ C C W+L GIP H IA I + EDF+ Y A++A Y + P +
Sbjct: 726 VDVVTKHCDCRRWDLTGIPCCHAIACIREDHLSEEDFLPHCYSINAFKAVYAENIIPCND 785
Query: 584 QKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGH 643
+ W N IL P+Y + +GRPKK R + P E N +L S C + H
Sbjct: 786 KANWEKMNGPQILPPVYEKKVGRPKKSRRKQPQEVQGRNGPKLTKHGVTIHCSYCHEANH 845
Query: 644 NTR 646
N +
Sbjct: 846 NKK 848
>UniRef100_Q7X762 OSJNBb0118P14.3 protein [Oryza sativa]
Length = 939
Score = 163 bits (412), Expect = 2e-38
Identities = 98/293 (33%), Positives = 147/293 (49%), Gaps = 6/293 (2%)
Query: 356 EHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQI 415
E F +RHLY NF+ G + L A +T + W M +++AL EAY +L +I
Sbjct: 531 EQRFCVRHLYQNFQVLHKGETLKNQLWAIARSSTVPE-WNANMEKMKALSSEAYKYLEEI 589
Query: 416 PTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEK 475
P + WC+ F+ + KCD+L+N E FN IL AR+ P+L+M++ IR+ IM R T ++
Sbjct: 590 PPNQWCRAFFSDFPKCDILLNNNSEVFNKYILDAREMPILSMLERIRNQIMNRLYTKQKE 649
Query: 476 MDK-Y*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCN 534
+++ + G+ PK +++ +M + G F+V I S Q+IV+L C C
Sbjct: 650 LERNWPCGICPKIKRKVEKNTEMANTCYVFPAGMGAFQVSDIGS--QYIVELNVKRCDCR 707
Query: 535 FWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEY 594
W+L GIP H I+ + H KPED V Y Y Y + + P+ W
Sbjct: 708 RWQLTGIPCNHAISCLRHERIKPEDVVSFCYSTRCYEQAYSYNIMPLRDSIHWEKMQGIE 767
Query: 595 ILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHNTRS 647
+ P+Y + +GRPKK R + P E H GV + S C GHN S
Sbjct: 768 VKPPVYEKKVGRPKKTRRKQPQELEGGTKISKH-GVEMH-CSYCKNGGHNKTS 818
>UniRef100_Q8H8R1 Putative mutator-like transposase [Oryza sativa]
Length = 746
Score = 162 bits (410), Expect = 3e-38
Identities = 96/309 (31%), Positives = 156/309 (50%), Gaps = 11/309 (3%)
Query: 341 LQDLLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMME 400
++ L+ D EH F +RHLY NF + G ++N + A++T W +
Sbjct: 392 VEGLIPAIKDEFPDSEHRFCVRHLYQNFAVLYKGEA-LKNQLWAIARSTTVPEWNVNTEK 450
Query: 401 LQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD* 460
++A++ +AY +L +IP + WC+ F ++KCD+L+N E IL AR+ +L+M++
Sbjct: 451 MKAVNKDAYGYLEEIPPNQWCRAFFRDFSKCDILLNNNLECHVRYILEARELTILSMLEK 510
Query: 461 IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQM--TSHLIPQCCG*DMFEVKHIVS 518
IRS +M R T E+ K+ + PK ++++ I+M T + +P G V +
Sbjct: 511 IRSKLMNRIYTKQEECKKWVFDICPKIKQKVEKNIEMSNTCYALPSRMG-----VFQVTD 565
Query: 519 HD-QFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHL 577
D QF+VD+K C C W+L+GIP H I+ + H KPED V Y +++ Y
Sbjct: 566 RDKQFVVDIKNKQCDCRRWQLIGIPCNHAISCLRHERIKPEDEVSFCYTIQSFKQAYMFN 625
Query: 578 VSPISGQKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSR 637
+ P+ + W N + P+Y + +GRPK R + P E H GV + S
Sbjct: 626 IMPVRDKTHWEKMNGVPVNPPVYEKKVGRPKTTRRKQPQELDAGTKISKH-GVQIH-CSY 683
Query: 638 CGQFGHNTR 646
C GHN +
Sbjct: 684 CKNVGHNKK 692
>UniRef100_Q60DJ3 MuDR family transposase protein [Oryza sativa]
Length = 1030
Score = 155 bits (393), Expect = 3e-36
Identities = 91/291 (31%), Positives = 144/291 (49%), Gaps = 4/291 (1%)
Query: 356 EHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQI 415
EH F +RHLY+NF+ F G V ++N + A+++ Q W K M ++ L+ AY WL ++
Sbjct: 596 EHRFCVRHLYSNFQLQFKGEV-LKNQLWACARSSSVQEWNKNMDVMRNLNKSAYEWLEKL 654
Query: 416 PTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGR-FATMNE 474
P + W + F+ + KCD+L+N E FN IL AR+ P+LTM++ I+ +M R F E
Sbjct: 655 PPNTWVRAFFSEFPKCDILLNNNCEVFNKYILEARELPILTMLEKIKGQLMTRHFNKQKE 714
Query: 475 KMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCN 534
D++ + PK +++ + G +F+V Q+IVD+ C C
Sbjct: 715 LADQFQGLICPKIRKKVLKNADAANTCYALPAGQGIFQVHE--REYQYIVDINAMYCDCR 772
Query: 535 FWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEY 594
W+L GIP H I+ + H E + Y A+ Y + P + + W N
Sbjct: 773 RWDLTGIPCNHAISCLRHERINAESILPNCYTTDAFSKAYGFNIWPCNDKSKWENVNGPE 832
Query: 595 ILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHNT 645
I P+Y + GRPKK R + P+E N +L + CG+ HN+
Sbjct: 833 IKPPVYEKKAGRPKKSRRKAPYEVIGKNGPKLTKHGVMMHYKYCGEENHNS 883
>UniRef100_Q6AVM7 Putative polyprotein [Oryza sativa]
Length = 1006
Score = 152 bits (383), Expect = 4e-35
Identities = 91/291 (31%), Positives = 142/291 (48%), Gaps = 4/291 (1%)
Query: 356 EHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQI 415
EH F +RHLY+NF+ F G V +N + A+++ Q W K M ++ L+ AY WL ++
Sbjct: 572 EHRFCVRHLYSNFQLQFKGEVP-KNQLWACARSSSVQEWNKNMDVMRNLNKSAYEWLEKL 630
Query: 416 PTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGR-FATMNE 474
P + W + F+ + KCD+L+N E FN IL AR+ P+LTM++ I+ +M R F E
Sbjct: 631 PPNTWVRAFFSEFPKCDILLNNNCEVFNKYILEARELPILTMLEKIKGQLMTRHFNKQKE 690
Query: 475 KMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCN 534
D++ + PK +++ + G +F+V Q+IVD+ C C
Sbjct: 691 LADQFQGLICPKIRKKVLKNADAANTCYALPAGQGIFQVHE--REYQYIVDINAMHCDCR 748
Query: 535 FWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEY 594
W+L GIP H I+ + H E + Y A+ Y + P + + W N
Sbjct: 749 RWDLTGIPCNHAISCLRHERINAESILPNCYTTDAFSKAYGFNIWPCNDKSKWENINGPE 808
Query: 595 ILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGHNT 645
I P+Y + GRPKK R + P+E N +L CG+ HN+
Sbjct: 809 IKPPVYEKKAGRPKKSRRKAPYEVIGKNGPKLTKHGVMMHCKYCGEENHNS 859
>UniRef100_O81502 F9D12.2 protein [Arabidopsis thaliana]
Length = 940
Score = 144 bits (364), Expect = 6e-33
Identities = 98/314 (31%), Positives = 153/314 (48%), Gaps = 5/314 (1%)
Query: 335 TEVSLFLQDLLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLW 394
T +S + L+ DL+ EH RH+YAN+K +G A AT +
Sbjct: 500 TVISDKQKGLINAVADLLPQAEHRHCARHVYANWKKVYGDYCHESYFWAIAYSATEGD-Y 558
Query: 395 TKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPV 454
+ M L++ P+A L++ + WC+ F+ ++ C+ + N L ESFN TI AR PV
Sbjct: 559 SYNMDALRSYDPDACDDLLKTDPTTWCRAFFSTHSSCEDVSNNLSESFNRTIREARKLPV 618
Query: 455 LTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVK 514
+ M++ +R M R + + +K K PK ++ L+ + + + G + FE+
Sbjct: 619 VNMLEEVRRISMKRISKLCDKTAKCETRFPPKIMQILEGNRKSSKYCQVLKSGENKFEI- 677
Query: 515 HIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATY 574
+ + VDL TC C WEL GIP H I I+ + + PED+V ++ATY
Sbjct: 678 -LEGSGYYSVDLLTRTCGCRQWELTGIPCPHAIYVITEHNRDPEDYVDRLLETQVWKATY 736
Query: 575 EHLVSPISGQKLWPTTNDEYILSPLYRRALGRPKKL-RMRDPHEDTTNNHTRLHIGVSRN 633
+ + P++G++LW I P R GRPKK R+++P E +T N T+L
Sbjct: 737 KDNIEPMNGERLWKRRGKGRIEVPDKRGKRGRPKKFARIKEPGESST-NPTKLTREGKTV 795
Query: 634 KFSRCGQFGHNTRS 647
S C Q GHN S
Sbjct: 796 TCSNCKQIGHNKGS 809
>UniRef100_Q7XE42 Putative mutator-like transposase [Oryza sativa]
Length = 1005
Score = 140 bits (353), Expect = 1e-31
Identities = 85/304 (27%), Positives = 145/304 (46%), Gaps = 4/304 (1%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
LL++ L EH RH+YAN++ + + AKA QL+ +L A
Sbjct: 554 LLSIVSTLFPFAEHRMCARHIYANWRKKHRLQEYQKRFWK-IAKAPNEQLFNHYKRKLAA 612
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
P + L + W + F + C+ + N + ESFN+ I+ +R KP++TM++ IR
Sbjct: 613 KTPRGWQDLEKTNPIHWSRAWFRLGSNCESVDNNMSESFNSWIIESRFKPIITMLEDIRI 672
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
+ R +++ V P + + T G FEV+ +F
Sbjct: 673 QVTRRIQENRSNSERWTMTVCPNIIRKFNKIRHRTQFCHVLWNGDAGFEVRD--KKWRFT 730
Query: 524 VDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISG 583
VDL TC C +W++ GIP +H AA+ Q+P + +H + Y+ TY+H++ P+
Sbjct: 731 VDLTSKTCSCRYWQVSGIPCQHACAALFKMAQEPNNCIHECFSLERYKKTYQHVLQPVEH 790
Query: 584 QKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGH 643
+ WP + + L P ++ GRPKK R +DP + + +G ++ + RCG +GH
Sbjct: 791 ESAWPVSPNPKPLPPRVKKMPGRPKKNRRKDPSKPVKSGTKSSKVG-TKIRCRRCGNYGH 849
Query: 644 NTRS 647
N+R+
Sbjct: 850 NSRT 853
>UniRef100_Q6L470 Putative mutator transposable element [Solanum demissum]
Length = 707
Score = 139 bits (351), Expect = 2e-31
Identities = 85/268 (31%), Positives = 137/268 (50%), Gaps = 1/268 (0%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L + +DL +++F +RHL+ NFK + ++N + AA AT + M +L
Sbjct: 433 LTYLMNDLEIEEQYLFCVRHLHNNFKRAGYSGMALKNALWKAASATTIDRFDACMTDLFE 492
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
L +AY WL S W + F+ KCD+L+N E FN IL ARDKP++ +++ IR
Sbjct: 493 LDKDAYAWLSAKVPSEWSRSHFSPLPKCDILLNNQCEVFNKFILDARDKPIVKLLETIRH 552
Query: 464 YIMGRFATMNEKMDKY-*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQF 522
+M R ++ EK +K+ + P ++L ++ ++ IPQ +EV V D +
Sbjct: 553 LLMTRINSIREKAEKWNLNDICPTIKKKLAKTMKKAANYIPQRSNMWNYEVIGPVEGDNW 612
Query: 523 IVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPIS 582
VDL TC C WEL G+P +H I++I + ++V Y+ YR YE + P++
Sbjct: 613 AVDLYNRTCSCRQWELSGVPCKHAISSIWLKNDEVLNYVDDCYKVDTYRKIYEASILPMN 672
Query: 583 GQKLWPTTNDEYILSPLYRRALGRPKKL 610
G LWP + + P Y + KK+
Sbjct: 673 GLDLWPKSLNPPPFPPSYLNNKKKGKKM 700
>UniRef100_Q94HK4 Mutator-like transposase [Oryza sativa]
Length = 725
Score = 138 bits (347), Expect = 6e-31
Identities = 86/262 (32%), Positives = 132/262 (49%), Gaps = 2/262 (0%)
Query: 356 EHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQI 415
EH + HL N N + + A AT + K M +L+ L+ +A+ WL+ I
Sbjct: 424 EHRYCKMHLLQNMGNKGWRGEKYKGFVDAAIYATTVWDYDKAMEDLKKLNLKAWEWLIAI 483
Query: 416 PTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNEK 475
+ + AF+ K D+++N L E FN IL ARDKP++TMV+ IR +M R + +
Sbjct: 484 GKEHFSRHAFSPKAKSDLVVNNLSEVFNKYILDARDKPIVTMVEHIRRKVMVRLSLKRQG 543
Query: 476 MDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNF 535
D + P +L+ E + +EV H + F VD+ TC C+
Sbjct: 544 GDAAQWEITPIVAGKLEMEKNHARYCWCYQSNLTTWEV-HCLDRS-FAVDISARTCACHK 601
Query: 536 WELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEYI 595
W+L GIP +H++ A+ G PED+V Y+R+ AY TY ++ P+ + W T+ YI
Sbjct: 602 WQLTGIPCKHDVCALYKAGHTPEDYVADYFRKDAYMRTYTAVIYPVPDEHRWTKTDSPYI 661
Query: 596 LSPLYRRALGRPKKLRMRDPHE 617
P + + +GRPKK R R P E
Sbjct: 662 DPPKFDKHVGRPKKSRRRGPDE 683
>UniRef100_Q7F966 OSJNBa0091C07.2 protein [Oryza sativa]
Length = 879
Score = 134 bits (337), Expect = 8e-30
Identities = 85/304 (27%), Positives = 140/304 (45%), Gaps = 5/304 (1%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L++ ++ + +EH RH+YAN++ + + + AKA+ + +L
Sbjct: 515 LVSAVEEFLPQIEHRMCTRHIYANWRKKYRDQA-FQKPFWKCAKASCRPFFNFCRAKLAQ 573
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
L P W + F + CD + N + ESFN I+ R P+++M + IR+
Sbjct: 574 LTPAGAKXXXSTEPMHWSRAWFRIGSNCDSVDNNMCESFNNWIIDIRAHPIISMFEGIRT 633
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
+ R K + + P L++L I ++ + G D FEV ++
Sbjct: 634 KVYVRIQQNRSKAKGWLGRICPNILKKLNKYIDLSGNCEAIWNGKDGFEVTD--KDKRYT 691
Query: 524 VDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISG 583
VDL+K TC C +W+L GIP H I A+ + ++PED++ Y Y Y+H + P+ G
Sbjct: 692 VDLEKRTCSCRYWQLAGIPCAHAITALFVSSKQPEDYIADCYSVEVYNKIYDHCMMPMEG 751
Query: 584 QKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKFSRCGQFGH 643
WP T P Y + GRP+K R RDP E +L ++ + S+C Q H
Sbjct: 752 MMQWPITGHPKPGPPGYVKMPGRPRKERRRDPLE--AKKGKKLSKTGTKGRCSQCKQTTH 809
Query: 644 NTRS 647
N R+
Sbjct: 810 NIRT 813
>UniRef100_Q60CW5 Putative mutator transposable element-related protein [Solanum
tuberosum]
Length = 493
Score = 133 bits (334), Expect = 2e-29
Identities = 85/275 (30%), Positives = 129/275 (46%), Gaps = 22/275 (8%)
Query: 385 AAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNA 444
AAKAT + + M+ ++ L P A WL S W + + KCD+L+N + E FN+
Sbjct: 169 AAKATTVKEFDACMVRIRELDPNAVDWLNDKEPSQWSRSHLSSDAKCDILLNNICEVFNS 228
Query: 445 TILLARDKPVLTMVD*IRSYIMGRFATMNEKMDKY*RG-VMPKPLERLKWEIQMTSHLIP 503
I ARDKP++T+++ +R +M R EK K+ V PK + L IP
Sbjct: 229 MIFDARDKPIVTLLEKLRYLLMARMLANREKAHKWSSNDVCPKIKDILHKNQTAAGEYIP 288
Query: 504 QCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHA 563
+ +E+ HD + VDL+ C C W ++GIP +H IAAI D+V
Sbjct: 289 RKSNQRKYEIIGATIHDSWAVDLENRICSCTKWSIMGIPCKHAIAAIRAKKDNILDYVDD 348
Query: 564 YYRRAAYRATYEHLVSPISGQKLWP--TTNDEYILSPLYRRALGRPKKLRMRDPHEDTTN 621
Y+ YR YEH + I+G ++WP T L+ + + GR +K R ++ E
Sbjct: 349 CYKVETYRRIYEHAILSINGPQMWPKSTKVPPLPLTIVGNKKTGRKQKARRKEADE---- 404
Query: 622 NHTRLHIGVSRNKFSR---------CGQFGHNTRS 647
+G SR K R C + GHN ++
Sbjct: 405 ------VGASRTKIKRKQQSLDCSTCNKPGHNKKT 433
>UniRef100_Q94HS2 Putative transposon protein [Oryza sativa]
Length = 841
Score = 129 bits (325), Expect = 2e-28
Identities = 74/270 (27%), Positives = 130/270 (47%), Gaps = 3/270 (1%)
Query: 378 IRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNK 437
++N + A+++ W M E+++L+ +AY WL ++P W K F+ + KCD+L+N
Sbjct: 436 LKNQLWACARSSSEVEWNANMEEMKSLNQDAYEWLQKMPPKTWVKAYFSEFPKCDILLNN 495
Query: 438 LYESFNATILLARDKPVLTMVD*IRSYIMGR-FATMNEKMDKY*RGVMPKPLERLKWEIQ 496
E FN IL AR+ P+L+M + I+S ++ R ++ E +++ + PK +++
Sbjct: 496 NCEVFNKYILEARELPILSMFEKIKSQLISRHYSKQKEVAEQWHGPICPKIRKKVLKNAD 555
Query: 497 MTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQK 556
M + G +F+V+ + ++IVDL C C W+L GIP H I+ +
Sbjct: 556 MANTCYVLPAGKGIFQVED--RNFKYIVDLSAKHCDCRRWDLTGIPCNHAISCLRSERIS 613
Query: 557 PEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTNDEYILSPLYRRALGRPKKLRMRDPH 616
E + Y A+ Y + P + W N + P+Y + +GRP K R + PH
Sbjct: 614 AESILPPCYSLEAFSRAYAFNIWPYNDMTKWVQVNGPEVKPPIYEKKVGRPPKSRRKAPH 673
Query: 617 EDTTNNHTRLHIGVSRNKFSRCGQFGHNTR 646
E N ++ S C + HN +
Sbjct: 674 EVQGKNGPKMSKHGVEMHCSFCKEPRHNKK 703
>UniRef100_Q75LI5 Putative transposon protein [Oryza sativa]
Length = 839
Score = 129 bits (323), Expect = 4e-28
Identities = 86/312 (27%), Positives = 146/312 (46%), Gaps = 8/312 (2%)
Query: 339 LFLQD----LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLW 394
+FL D LL+V + L EH RH+YAN++ + + A+++ L+
Sbjct: 430 VFLTDQQKGLLSVIEHLFPKAEHRMCARHIYANWRKRHRLQEYQKRFWK-IARSSNAVLF 488
Query: 395 TKKMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPV 454
+L P + L + WC+ F + CD + N + ESFN I+ AR KP+
Sbjct: 489 NHYKSKLANKTPMGWEDLEKTNPIHWCRAWFKLGSNCDSVENNICESFNNWIIEARFKPI 548
Query: 455 LTMVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVK 514
+TM++ IR + R +++ G+ P L+++ T G FEV+
Sbjct: 549 ITMLEDIRMKVTRRIQENKTNSERWTMGICPNILKKINKIRHATQFCHVLWNGSSGFEVR 608
Query: 515 HIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATY 574
+F VDL +TC C +W++ GIP +H AA ++P + V+ + YR TY
Sbjct: 609 E--KKWRFTVDLSANTCSCRYWQISGIPCQHACAAYFKMAEEPNNHVNMCFSIDQYRNTY 666
Query: 575 EHLVSPISGQKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNK 634
+ ++ P+ + +WP + + L P ++ G PK+ R +DP E + T+ K
Sbjct: 667 QDVLQPVEHESVWPLSTNPRPLPPRVKKMPGSPKRARRKDPTE-AAGSSTKSSKRGGSVK 725
Query: 635 FSRCGQFGHNTR 646
C + GHN+R
Sbjct: 726 CGFCHEKGHNSR 737
>UniRef100_Q6L3K3 Putative transposon MuDR mudrA-like protein [Solanum demissum]
Length = 873
Score = 127 bits (319), Expect = 1e-27
Identities = 93/315 (29%), Positives = 140/315 (43%), Gaps = 26/315 (8%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
LL + H + RH+ AN+ + G V +R L+ +A +TY + + ++ + A
Sbjct: 411 LLDAVSQVFPKAHHRWCARHIEANWSKAWKG-VQMRKLLWWSAWSTYEEEFHDQLKVMGA 469
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+ +A L+ P WC+ F K N ESFN IL AR KP++ M++ IR
Sbjct: 470 VSKQAAKDLVWYPAQNWCRAYFDTVCKNHSCENNFTESFNKWILEARAKPIIKMLENIRI 529
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCC-----G*DMFEVKHIVS 518
IM R + E+ + P +E + ++I QCC G +EV +
Sbjct: 530 KIMNRLQKLEEEGKNWKGDFSPYAME-----LYNDFNIIAQCCQVQSNGDQGYEV--VEG 582
Query: 519 HDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLV 578
D+ +V+L + C C W+L GI H I A H+ Q+P D + +Y R AY Y H +
Sbjct: 583 EDRHVVNLNRKKCTCRTWDLTGILCPHAIKAYLHDKQEPLDQLSWWYSREAYMLVYMHKI 642
Query: 579 SPISGQKLWPTTNDEYILSPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGV---SRNKF 635
P+ G+K W + P + +GRPK R R+ E R GV SR
Sbjct: 643 QPVRGEKFWKVDPSHAMEPPEIHKLVGRPKLKRKREKDE------ARKREGVWSASRKGL 696
Query: 636 ----SRCGQFGHNTR 646
C GHN R
Sbjct: 697 KMTCGHCSATGHNQR 711
>UniRef100_Q8LMJ1 Putative transposon protein [Oryza sativa]
Length = 835
Score = 125 bits (315), Expect = 3e-27
Identities = 90/307 (29%), Positives = 138/307 (44%), Gaps = 31/307 (10%)
Query: 354 GVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLM 413
GVEH +RHL NF+ F G V RNL A++A ++ E++ P A W+
Sbjct: 505 GVEHRECMRHLVKNFQKRFSGEVFERNLW-SASRAYRQDIYEGHYNEMKEACPRATEWID 563
Query: 414 QIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMN 473
W + F+ +KCD + N + E+FN+ I + PV+ ++D IR IM R +
Sbjct: 564 DFHKHIWTRSKFSPVSKCDYVTNNIAETFNSWIRHEKSLPVVDLMDKIRHMIMERISIRK 623
Query: 474 EKMDKY*RGVMPKPLERLKWEIQMTSHLIPQC---------CG*DMFEVKHIVSHDQFIV 524
+K ++P ++ L + + + G D+ +H V
Sbjct: 624 RLAEKLTGQILPSVMKTLYARSRDLGYKLYSAHKHLGEIGGTGRDLKTWRH-------TV 676
Query: 525 DLKKHTCMCNFWELVGIPFRHEI-AAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISG 583
DL C C W++ GIP H I IS G + E FV YY AA++ Y V P++
Sbjct: 677 DLNTRECSCRQWQITGIPCTHVIFLIISRRGLELEQFVDEYYYVAAFKRAYAGHVVPMTD 736
Query: 584 QKLWPTTNDEYIL-SPLYRRALGRPKKLRMRDPHEDTTNNHTRLHIGVSRNKF--SRCGQ 640
+ W N L PL +R+ GRP+ R++ E GV + K+ RCGQ
Sbjct: 737 KAQWAKGNIGLKLHPPLLKRSAGRPRSRRIKGVEEG----------GVGKRKYRCKRCGQ 786
Query: 641 FGHNTRS 647
FGH ++
Sbjct: 787 FGHTKKT 793
>UniRef100_Q9FH72 Arabidopsis thaliana genomic DNA, chromosome 5, P1 clone:MRD20
[Arabidopsis thaliana]
Length = 733
Score = 121 bits (303), Expect = 7e-26
Identities = 79/309 (25%), Positives = 156/309 (49%), Gaps = 11/309 (3%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ +++ EH RH++AN + + + + A+A ++ K++ +++
Sbjct: 367 LIYAIKNVLPYAEHRMCARHIFANLQKRYKQMGPLHKVFWKCARAYNETVFWKQLEKMKT 426
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+ EAY + + S W + F+ TK + N + ES+NA + AR+KPV+ +++ IR
Sbjct: 427 IKFEAYDEVKRSVGSNWSRAFFSDITKSAAVENNISESYNAVLKDAREKPVVALLEDIRR 486
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
+IM ++M + PK + ++ + P G ++EV H ++++
Sbjct: 487 HIMASNLVKIKEMQNVTGLITPKAIAIMEKRKKSLKWCYPFSNGRGIYEVDH--GKNKYV 544
Query: 524 VDLK-KHTCMCNFWELVGIPFRHEIAAI---SHNGQKPEDFVHAYYRRAAYRATYEHLVS 579
V ++ K +C C +++ GIP H ++A+ + PE + +Y ++ Y L+
Sbjct: 545 VHVRDKTSCTCREYDVSGIPCCHIMSAMWAEYKETKLPETAILDWYSVEKWKLCYNSLLF 604
Query: 580 PISGQKLWPTTNDEYILSPLYRRALGRPKKL-RMRDPHEDTTNNHTRLHIGVSRNKFSRC 638
P++G +LW T +D ++ P R GRPKK R++DP E+ ++ +++ + N C
Sbjct: 605 PVNGMELWETHSDVVVMPPPDRIMPGRPKKNDRIKDPSEEASSENSQKALVTCSN----C 660
Query: 639 GQFGHNTRS 647
GQ GHN R+
Sbjct: 661 GQIGHNKRT 669
>UniRef100_Q9LMY1 F21F23.10 protein [Arabidopsis thaliana]
Length = 753
Score = 120 bits (301), Expect = 1e-25
Identities = 79/309 (25%), Positives = 156/309 (49%), Gaps = 11/309 (3%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ +++ EH RH++AN + + + + A+A ++ K++ +++
Sbjct: 387 LIYAIKNVLPYAEHRMCARHIFANLQKRYKQMGPLHKVFWKCARAYNETVFWKQLEKMKT 446
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+ EAY + + S W + F+ TK + N + ES+NA + AR+KPV+ +++ IR
Sbjct: 447 IKFEAYDEVKRSVGSNWSRAFFSDITKSAAVENNISESYNAVLKDAREKPVVALLEDIRR 506
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
+IM ++M + PK + ++ + P G ++EV H ++++
Sbjct: 507 HIMASNLVKIKEMQNVTGLITPKAIAIMEKRKKSLKWCYPFSNGRGIYEVDH--GKNKYV 564
Query: 524 VDLK-KHTCMCNFWELVGIPFRHEIAAI---SHNGQKPEDFVHAYYRRAAYRATYEHLVS 579
V ++ K +C C +++ GIP H ++A+ + PE + +Y ++ Y L+
Sbjct: 565 VHVRDKTSCTCREYDVSGIPCCHIMSAMWAEYKETKLPETAILDWYSVEKWKLCYNSLLF 624
Query: 580 PISGQKLWPTTNDEYILSPLYRRALGRPKKL-RMRDPHEDTTNNHTRLHIGVSRNKFSRC 638
P++G +LW T +D ++ P R GRPKK R++DP E+ ++ +++ + N C
Sbjct: 625 PVNGMELWETHSDVVVMPPPDRIMPGRPKKNDRIKDPSEEASSENSQKALVTCSN----C 680
Query: 639 GQFGHNTRS 647
GQ GHN R+
Sbjct: 681 GQTGHNKRT 689
>UniRef100_Q9FM95 Similarity to mutator-like transposase [Arabidopsis thaliana]
Length = 733
Score = 120 bits (301), Expect = 1e-25
Identities = 79/309 (25%), Positives = 156/309 (49%), Gaps = 11/309 (3%)
Query: 344 LLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQA 403
L+ +++ EH RH++AN + + + + A+A ++ K++ +++
Sbjct: 367 LIYAIKNVLPYAEHRMCARHIFANLQKRYKQMGPLHKVFWKCARAYNETVFWKQLEKMKT 426
Query: 404 LHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRS 463
+ EAY + + S W + F+ TK + N + ES+NA + AR+KPV+ +++ IR
Sbjct: 427 IKFEAYDEVKRSVGSNWSRAFFSDITKSAAVENNISESYNAVLKDAREKPVVALLEDIRR 486
Query: 464 YIMGRFATMNEKMDKY*RGVMPKPLERLKWEIQMTSHLIPQCCG*DMFEVKHIVSHDQFI 523
+IM ++M + PK + ++ + P G ++EV H ++++
Sbjct: 487 HIMASNLVKLKEMQNVTGLITPKAIAIMEKRKKSLKWCYPFSNGRGIYEVDH--GKNKYV 544
Query: 524 VDLK-KHTCMCNFWELVGIPFRHEIAAI---SHNGQKPEDFVHAYYRRAAYRATYEHLVS 579
V ++ K +C C +++ GIP H ++A+ + PE + +Y ++ Y L+
Sbjct: 545 VHVRDKTSCTCREYDVSGIPCCHIMSAMWAEYKETKLPETAILDWYSVEKWKLCYNSLLF 604
Query: 580 PISGQKLWPTTNDEYILSPLYRRALGRPKKL-RMRDPHEDTTNNHTRLHIGVSRNKFSRC 638
P++G +LW T +D ++ P R GRPKK R++DP E+ ++ +++ + N C
Sbjct: 605 PVNGMELWETHSDVVVMPPPDRIMPGRPKKNDRIKDPSEEASSENSQKALVTCSN----C 660
Query: 639 GQFGHNTRS 647
GQ GHN R+
Sbjct: 661 GQTGHNKRT 669
>UniRef100_Q9FHV3 Mutator-like transposase-like protein [Arabidopsis thaliana]
Length = 599
Score = 114 bits (286), Expect = 7e-24
Identities = 81/322 (25%), Positives = 146/322 (45%), Gaps = 23/322 (7%)
Query: 337 VSLFLQDLLTVFDDLMGGVEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTK 396
V L + L+ + +EH ++H+Y N K +G + +I+ L+ + + + + +
Sbjct: 270 VCLIVSGLIAAVQLELPKIEHRMCVQHIYGNSKKIYGSKTMIKPLLWNLDWSYNEKEYKQ 329
Query: 397 KMMELQALHPEAYTWLMQIPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLT 456
+ +++ + Y +M+ +W + + C+ + N ESFN ++ AR+KP +
Sbjct: 330 HLEKIRCYDTKVYESVMKTKPRSWVRAFQKIGSFCEDVDNNSVESFNGSLNKAREKPFVA 389
Query: 457 MVD*IRSYIMGRFATMNEKMDKY*RGVMPKPLERLKWE--IQMTSHLIPQCCG*DMFEVK 514
M++ IR M R A + + + P + L E + + + P G M+EV+
Sbjct: 390 MLETIRRMAMVRIAKRSVESHTHTGVCTPYVTKFLAGEHKVAAAAKVSPSTNG--MYEVR 447
Query: 515 HIVSHDQFIVDLKKHTCMCNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATY 574
H D VDL+ +TC C W++ GIP H + I H +PE+FV ++R A +R Y
Sbjct: 448 H--GGDLHRVDLQAYTCTCIKWQICGIPCEHAYSVIIHKKLEPENFVCCWFRTAMWRKNY 505
Query: 575 EHLVSPISGQKLWPTTNDEYILSP-----LYRRALGRPKKLRMRDPHEDTTNNHTR---- 625
+ P G K WP +N + P + + + +K R +E T +
Sbjct: 506 TDGLFPQRGPKFWPESNGLRVYVPPPTEGEEDKKMTKAEKKRKNGVNESPTKKQPKGKKR 565
Query: 626 -LHIGVSRNKFSRCGQFGHNTR 646
+H GV CG HN+R
Sbjct: 566 IMHCGV-------CGAADHNSR 580
>UniRef100_O49511 MuDR transposable element - like protein [Arabidopsis thaliana]
Length = 633
Score = 114 bits (284), Expect = 1e-23
Identities = 78/304 (25%), Positives = 140/304 (45%), Gaps = 23/304 (7%)
Query: 355 VEHMFYLRHLYANFKNFFGGRVVIRNLMMDAAKATYHQLWTKKMMELQALHPEAYTWLMQ 414
+EH ++H+Y N K +G + +I+ L+ + A + + + + +++ + Y +M+
Sbjct: 316 IEHRMCVQHIYGNLKKTYGSKTMIKPLLWNLAWSYNETEYKQHLEKIRCYDTKVYESVMK 375
Query: 415 IPTSAWCKQAFTFYTKCDVLMNKLYESFNATILLARDKPVLTMVD*IRSYIMGRFATMNE 474
+W + + C+ + N ESFN ++ AR+K + M++ IR M R A +
Sbjct: 376 TNPRSWVRAFQKIGSFCEDVDNNSVESFNGSLNKAREKQFVAMLETIRRMAMVRIAKRSV 435
Query: 475 KMDKY*RGVMPKPLERLKWE--IQMTSHLIPQCCG*DMFEVKHIVSHDQFIVDLKKHTCM 532
+ + P ++ L E + T+ + P G M+EV+H D VDL +TC
Sbjct: 436 ESHTHTGVCTPYVMKFLAGEHKVASTAKVAPGTNG--MYEVRH--GGDTHRVDLAAYTCT 491
Query: 533 CNFWELVGIPFRHEIAAISHNGQKPEDFVHAYYRRAAYRATYEHLVSPISGQKLWPTTND 592
C W++ GIP H I H +PEDFV ++R A ++ Y + P G K WP +N
Sbjct: 492 CIKWQICGIPCEHAYGVILHKKLQPEDFVCQWFRTAMWKKNYTDGLFPQRGPKFWPESNG 551
Query: 593 EYILSP-----LYRRALGRPKKLRMRDPHEDTTNNHTR-----LHIGVSRNKFSRCGQFG 642
+ + + + + +K R + +E T + +H GV CG
Sbjct: 552 PRVFAAEPPEGEEDKKMTKAEKKRKKGVNESPTKKQPKAKKRIMHCGV-------CGAAD 604
Query: 643 HNTR 646
HN+R
Sbjct: 605 HNSR 608
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.341 0.148 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,002,652,393
Number of Sequences: 2790947
Number of extensions: 39486932
Number of successful extensions: 110069
Number of sequences better than 10.0: 145
Number of HSP's better than 10.0 without gapping: 79
Number of HSP's successfully gapped in prelim test: 66
Number of HSP's that attempted gapping in prelim test: 109778
Number of HSP's gapped (non-prelim): 181
length of query: 647
length of database: 848,049,833
effective HSP length: 134
effective length of query: 513
effective length of database: 474,062,935
effective search space: 243194285655
effective search space used: 243194285655
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0089b.6


