

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0089a.1
(119 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
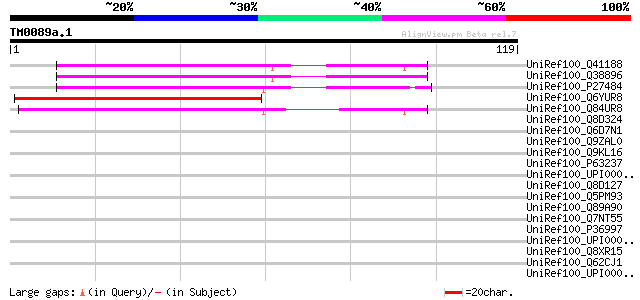
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q41188 Glycine-rich protein 2 [Arabidopsis thaliana] 56 2e-07
UniRef100_Q38896 Glycine-rich protein 2b [Arabidopsis thaliana] 54 6e-07
UniRef100_P27484 Glycine-rich protein 2 [Nicotiana sylvestris] 53 2e-06
UniRef100_Q6YUR8 Putative Glycine-rich protein 2 [Oryza sativa] 46 2e-04
UniRef100_Q84UR8 Putative cold shock protein-1 [Oryza sativa] 45 5e-04
UniRef100_Q8D324 CspE protein [Wigglesworthia glossinidia brevip... 44 0.001
UniRef100_Q6D7N1 Cold shock-like protein [Erwinia carotovora] 44 0.001
UniRef100_Q9ZAL0 Cold shock protein, CSPA [Vibrio cholerae] 44 0.001
UniRef100_Q9KL16 Cold shock protein cspV [Vibrio cholerae] 44 0.001
UniRef100_P63237 Cold shock-like protein cspE [Buchnera aphidicola] 44 0.001
UniRef100_UPI000025E3D1 UPI000025E3D1 UniRef100 entry 43 0.002
UniRef100_Q8D127 Cold shock protein [Yersinia pestis] 43 0.002
UniRef100_Q5PM93 Cold shock-like protein cspE [Salmonella enteri... 42 0.002
UniRef100_Q89A90 Cold shock-like protein cspE [Buchnera aphidicola] 42 0.002
UniRef100_Q7NT55 Cold shock transcription regulator protein [Chr... 42 0.003
UniRef100_P36997 Cold shock-like protein cspE [Escherichia coli] 42 0.003
UniRef100_UPI0000286E74 UPI0000286E74 UniRef100 entry 41 0.006
UniRef100_Q8XR15 PROBABLE COLD SHOCK-LIKE TRANSCRIPTION REGULATO... 41 0.006
UniRef100_Q62CJ1 Cold-shock domain family protein [Burkholderia ... 41 0.006
UniRef100_UPI0000334174 UPI0000334174 UniRef100 entry 41 0.007
>UniRef100_Q41188 Glycine-rich protein 2 [Arabidopsis thaliana]
Length = 203
Score = 56.2 bits (134), Expect = 2e-07
Identities = 40/98 (40%), Positives = 51/98 (51%), Gaps = 19/98 (19%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DDGG+DLF+ S I+S+GF LA E V+ + I I +
Sbjct: 15 VKWFDTQKGFGFITPDDGGDDLFVHQSSIRSEGFRSLAAEEAVEFEVEIDNNNRPKAIDV 74
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSS----GYGRGGG 98
+ PD AP+ G GG+ G GRGGG
Sbjct: 75 SG--------PDGAPVQGNSGGGSSGGRGGFGGGRGGG 104
>UniRef100_Q38896 Glycine-rich protein 2b [Arabidopsis thaliana]
Length = 201
Score = 54.3 bits (129), Expect = 6e-07
Identities = 37/94 (39%), Positives = 48/94 (50%), Gaps = 15/94 (15%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLTI-------LIPM 64
+K FD QKGF IT DGG+DLF+ S I+S+GF LA E V+ + + I +
Sbjct: 19 VKWFDTQKGFGFITPSDGGDDLFVHQSSIRSEGFRSLAAEESVEFDVEVDNSGRPKAIEV 78
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGG 98
+ PD AP+ G GG S G G GG
Sbjct: 79 SG--------PDGAPVQGNSGGGGSSGGRGGFGG 104
>UniRef100_P27484 Glycine-rich protein 2 [Nicotiana sylvestris]
Length = 214
Score = 52.8 bits (125), Expect = 2e-06
Identities = 39/95 (41%), Positives = 48/95 (50%), Gaps = 16/95 (16%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL-------TILIPM 64
+K F DQKGF IT DDGG DLF+ S I+S+GF LAE E V+ + T + +
Sbjct: 13 VKWFSDQKGFGFITPDDGGEDLFVHQSGIRSEGFRSLAEGETVEFEVESGGDGRTKAVDV 72
Query: 65 AALRR*RD*RPD*API*GTCCGGNDSSGYGRGGGY 99
PD A + G GG G G GGGY
Sbjct: 73 TG--------PDGAAVQGGRGGGGGGGGRG-GGGY 98
>UniRef100_Q6YUR8 Putative Glycine-rich protein 2 [Oryza sativa]
Length = 241
Score = 45.8 bits (107), Expect = 2e-04
Identities = 25/58 (43%), Positives = 35/58 (60%)
Query: 2 MMSVSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
M + + R +K F+D KGF I+ DDG DLF+ S IK+ GF LAE E V+ ++
Sbjct: 1 MAAAARHRGTVKWFNDTKGFGFISPDDGSEDLFVHQSSIKADGFRSLAEGEQVEFAIS 58
>UniRef100_Q84UR8 Putative cold shock protein-1 [Oryza sativa]
Length = 197
Score = 44.7 bits (104), Expect = 5e-04
Identities = 36/108 (33%), Positives = 49/108 (45%), Gaps = 24/108 (22%)
Query: 3 MSVSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL---- 58
M+ V+ +K FD KGF IT DDGG DLF+ S +KS G+ L + + V+ +
Sbjct: 1 MASERVKGTVKWFDATKGFGFITPDDGGEDLFVHQSSLKSDGYRSLNDGDVVEFSVGSGN 60
Query: 59 ---TILIPMAALRR*RD*RPD*API*GTCCGGNDSS-----GYGRGGG 98
T + + AP G GG+ S GYG GGG
Sbjct: 61 DGRTKAVDVT------------APGGGALTGGSRPSGGGDRGYGGGGG 96
>UniRef100_Q8D324 CspE protein [Wigglesworthia glossinidia brevipalpis]
Length = 69
Score = 43.5 bits (101), Expect = 0.001
Identities = 22/55 (40%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQRVEFEIT 55
>UniRef100_Q6D7N1 Cold shock-like protein [Erwinia carotovora]
Length = 69
Score = 43.5 bits (101), Expect = 0.001
Identities = 22/55 (40%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 MSKIKGSVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQRVEFEIT 55
>UniRef100_Q9ZAL0 Cold shock protein, CSPA [Vibrio cholerae]
Length = 70
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/47 (42%), Positives = 33/47 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL 58
+K F++ KGF +T D+GGND+F+ + I+S+GF LAE + V ++
Sbjct: 9 VKWFNETKGFGFLTQDNGGNDVFVHLNSIQSEGFKTLAEGQRVSFIV 55
>UniRef100_Q9KL16 Cold shock protein cspV [Vibrio cholerae]
Length = 70
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/47 (42%), Positives = 33/47 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLL 58
+K F++ KGF +T D+GGND+F+ + I+S+GF LAE + V ++
Sbjct: 9 VKWFNETKGFGFLTQDNGGNDVFVHFNSIQSEGFKTLAEGQRVSFIV 55
>UniRef100_P63237 Cold shock-like protein cspE [Buchnera aphidicola]
Length = 68
Score = 43.5 bits (101), Expect = 0.001
Identities = 22/54 (40%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I+S GF LAE + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLAEGQSVEFEIT 54
>UniRef100_UPI000025E3D1 UPI000025E3D1 UniRef100 entry
Length = 67
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/43 (51%), Positives = 30/43 (69%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F+D KGF IT D+GG+DLF S+I++ GF LAE + V
Sbjct: 6 VKWFNDSKGFGFITPDNGGDDLFAHFSEIRADGFKTLAENQKV 48
>UniRef100_Q8D127 Cold shock protein [Yersinia pestis]
Length = 84
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/55 (40%), Positives = 34/55 (61%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I S GF LAE + V+ +T
Sbjct: 16 MSKIKGSVKWFNESKGFGFITPEDGSKDVFVHFSAIASNGFKTLAEGQRVEFEIT 70
>UniRef100_Q5PM93 Cold shock-like protein cspE [Salmonella enterica subsp. enterica
serovar Paratypi A str. ATCC 9150]
Length = 69
Score = 42.4 bits (98), Expect = 0.002
Identities = 21/55 (38%), Positives = 35/55 (63%)
Query: 5 VSMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
+S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 1 MSKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 55
>UniRef100_Q89A90 Cold shock-like protein cspE [Buchnera aphidicola]
Length = 68
Score = 42.4 bits (98), Expect = 0.002
Identities = 21/54 (38%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I+S GF L+E + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQSNGFKTLSEGQSVEFEIT 54
>UniRef100_Q7NT55 Cold shock transcription regulator protein [Chromobacterium
violaceum]
Length = 110
Score = 42.0 bits (97), Expect = 0.003
Identities = 22/48 (45%), Positives = 31/48 (63%)
Query: 7 MVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
M +K F+D KGF IT D+GG+D+F S+I ++GF LAE + V
Sbjct: 44 MATGTVKWFNDSKGFGFITPDEGGDDVFAHFSQINAKGFRSLAENQRV 91
>UniRef100_P36997 Cold shock-like protein cspE [Escherichia coli]
Length = 68
Score = 42.0 bits (97), Expect = 0.003
Identities = 21/54 (38%), Positives = 34/54 (62%)
Query: 6 SMVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
S ++ +K F++ KGF IT +DG D+F+ S I++ GF LAE + V+ +T
Sbjct: 1 SKIKGNVKWFNESKGFGFITPEDGSKDVFVHFSAIQTNGFKTLAEGQRVEFEIT 54
>UniRef100_UPI0000286E74 UPI0000286E74 UniRef100 entry
Length = 77
Score = 41.2 bits (95), Expect = 0.006
Identities = 22/48 (45%), Positives = 28/48 (57%)
Query: 7 MVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
M +K F+D KGF IT DDGG DLF S+++ GF L E + V
Sbjct: 11 MATGTVKWFNDAKGFGFITPDDGGEDLFAHFSEVQGSGFKSLQENQKV 58
>UniRef100_Q8XR15 PROBABLE COLD SHOCK-LIKE TRANSCRIPTION REGULATOR PROTEIN
[Ralstonia solanacearum]
Length = 67
Score = 41.2 bits (95), Expect = 0.006
Identities = 24/53 (45%), Positives = 29/53 (54%)
Query: 7 MVREKLK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFVDLLLT 59
M +K F+D KGF IT D+GG DLF S I QGF L E + V +T
Sbjct: 1 MATGTVKWFNDAKGFGFITPDEGGEDLFAHFSAINMQGFKTLKEGQRVSFEIT 53
>UniRef100_Q62CJ1 Cold-shock domain family protein [Burkholderia mallei]
Length = 67
Score = 41.2 bits (95), Expect = 0.006
Identities = 21/43 (48%), Positives = 30/43 (68%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F++ KGF IT D GG+DLF S+I+S+G+ LAE + V
Sbjct: 6 VKWFNETKGFGFITPDSGGDDLFAHFSEIRSEGYKTLAENQKV 48
>UniRef100_UPI0000334174 UPI0000334174 UniRef100 entry
Length = 67
Score = 40.8 bits (94), Expect = 0.007
Identities = 22/43 (51%), Positives = 27/43 (62%)
Query: 12 LK*FDDQKGFRLITHDDGGNDLFI**SKIKSQGF*CLAEREFV 54
+K F+D KGF IT D+GG DLF S+IK GF L E + V
Sbjct: 6 VKWFNDAKGFGFITSDNGGEDLFAHFSEIKMDGFKTLKENQRV 48
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.359 0.166 0.627
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 181,475,224
Number of Sequences: 2790947
Number of extensions: 6045539
Number of successful extensions: 36256
Number of sequences better than 10.0: 273
Number of HSP's better than 10.0 without gapping: 229
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 35937
Number of HSP's gapped (non-prelim): 319
length of query: 119
length of database: 848,049,833
effective HSP length: 95
effective length of query: 24
effective length of database: 582,909,868
effective search space: 13989836832
effective search space used: 13989836832
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0089a.1


