

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.13
(195 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
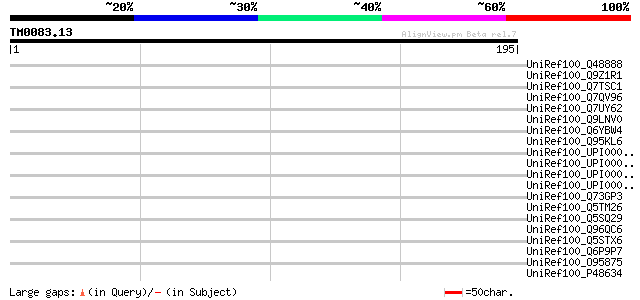
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q48888 Rep [Mycobacterium avium] 35 0.82
UniRef100_Q9Z1R1 BAT2 [Mus musculus] 34 2.4
UniRef100_Q7TSC1 HLA-B associated transcript 2 [Mus musculus] 34 2.4
UniRef100_Q7QV96 GLP_205_38349_33706 [Giardia lamblia ATCC 50803] 33 3.1
UniRef100_Q7UY62 Hypothetical protein [Rhodopirellula baltica] 33 4.1
UniRef100_Q9LNV0 F22G5.35 [Arabidopsis thaliana] 33 4.1
UniRef100_Q6YBW4 TACC3 [Oryctolagus cuniculus] 33 4.1
UniRef100_Q95KL6 Centrosome-associated AKAP350-binding protein T... 33 4.1
UniRef100_UPI000042F0A4 UPI000042F0A4 UniRef100 entry 33 5.3
UniRef100_UPI000023F00F UPI000023F00F UniRef100 entry 33 5.3
UniRef100_UPI0000457408 UPI0000457408 UniRef100 entry 32 6.9
UniRef100_UPI000036D337 UPI000036D337 UniRef100 entry 32 6.9
UniRef100_Q73GP3 Signal peptidase I [Wolbachia pipientis wMel] 32 6.9
UniRef100_Q5TM26 HLA-B associated transcript-2 [Macaca mulatta] 32 6.9
UniRef100_Q5SQ29 HLA-B associated transcript 2 [Homo sapiens] 32 6.9
UniRef100_Q96QC6 BAT2 protein [Homo sapiens] 32 6.9
UniRef100_Q5STX6 OTTHUMP00000062432 [Homo sapiens] 32 6.9
UniRef100_Q6P9P7 BAT2 protein [Homo sapiens] 32 6.9
UniRef100_O95875 BAT2 [Homo sapiens] 32 6.9
UniRef100_P48634 Large proline-rich protein BAT2 [Homo sapiens] 32 6.9
>UniRef100_Q48888 Rep [Mycobacterium avium]
Length = 360
Score = 35.4 bits (80), Expect = 0.82
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 3/70 (4%)
Query: 11 PIRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASH---PPQDKMTSPE 67
P+R HVR++ VT + A RG A++ P G K+ +H PP SP
Sbjct: 154 PVRDQHVRQDVVTTRNRRRTGAGNRGAARRARPDGYGLALAKTWRAHPQAPPWCHRHSPT 213
Query: 68 AYLKALRAEA 77
A+ L A A
Sbjct: 214 AWAAILAAPA 223
>UniRef100_Q9Z1R1 BAT2 [Mus musculus]
Length = 2157
Score = 33.9 bits (76), Expect = 2.4
Identities = 25/92 (27%), Positives = 35/92 (37%), Gaps = 14/92 (15%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 587 PVEPQLPSKEGPEPPEEVPPPTTPPAPKMEPKGDGVGSTRQPPSQGLGYPKYQKSLPPRF 646
Query: 62 KMTSPEAYLKALRAEALDCNLYYSYQTQVTAP 93
+ E LK + + + Q Q TAP
Sbjct: 647 QRQQQEQLLKQQQQQQ-----QWQQQQQGTAP 673
>UniRef100_Q7TSC1 HLA-B associated transcript 2 [Mus musculus]
Length = 2158
Score = 33.9 bits (76), Expect = 2.4
Identities = 25/92 (27%), Positives = 35/92 (37%), Gaps = 14/92 (15%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 587 PVEPQLPSKEGPEPPEEVPPPTTPPAPKMEPKGDGVGSTRQPPSQGLGYPKYQKSLPPRF 646
Query: 62 KMTSPEAYLKALRAEALDCNLYYSYQTQVTAP 93
+ E LK + + + Q Q TAP
Sbjct: 647 QRQQQEQLLKQQQQQQ-----QWQQQQQGTAP 673
>UniRef100_Q7QV96 GLP_205_38349_33706 [Giardia lamblia ATCC 50803]
Length = 1547
Score = 33.5 bits (75), Expect = 3.1
Identities = 32/120 (26%), Positives = 52/120 (42%), Gaps = 5/120 (4%)
Query: 61 DKMTSPEAYLKALR-AEALDCNL-YYSYQTQVTAPGRIRSLVSGLLTEDLDPQLKAGLLT 118
DK+ +++ K + E NL YY+Y Q GRI + G+L LD L +
Sbjct: 1369 DKLAYMDSFNKLYKHVELYAQNLAYYNYGVQELDRGRIAYSIGGML---LDAVLGDLVQY 1425
Query: 119 FLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGEETGTMGEIEAELTSLLATREGLDRD 178
L LT++ HRR L G+ S+ + K + T A++ + + G +D
Sbjct: 1426 TLRLTKKQSAQHRRMLLTESSTGGIAESSLDCKAQHAVTSISSTAQMCASAGDQHGTPKD 1485
>UniRef100_Q7UY62 Hypothetical protein [Rhodopirellula baltica]
Length = 356
Score = 33.1 bits (74), Expect = 4.1
Identities = 38/138 (27%), Positives = 57/138 (40%), Gaps = 34/138 (24%)
Query: 43 PPGNDTYHPK----SQASHPPQDKMTSPEAYLKALRAEALDCNLYYSYQTQVTAPGRIRS 98
P G T PK SQ S Q M + A +++R E N+ Y ++ R R
Sbjct: 61 PEGLPTPEPKLATDSQLSELAQTAMRAAAAQRESMRLE----NMVKRYDMELEQAKRERG 116
Query: 99 LVSGLLTEDLDPQLKAGLLTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGEETGTM 158
+ LLT L EE WE +++L+ + +V A + E M
Sbjct: 117 ALLDLLT----------------LAEEAWEQKQKQLDQD------KVRAAKLGSE----M 150
Query: 159 GEIEAELTSLLATREGLD 176
E+ AEL +L T+E L+
Sbjct: 151 REVSAELATLAGTKERLE 168
>UniRef100_Q9LNV0 F22G5.35 [Arabidopsis thaliana]
Length = 352
Score = 33.1 bits (74), Expect = 4.1
Identities = 14/50 (28%), Positives = 23/50 (46%)
Query: 17 VRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSP 66
++ + +H PP+ + + L P GN Y P PPQ +T+P
Sbjct: 135 IKDRPIPPPQHPPPRPQSQPLDYYSAPQGNHYYSPSPPPPPPPQAPITAP 184
>UniRef100_Q6YBW4 TACC3 [Oryctolagus cuniculus]
Length = 736
Score = 33.1 bits (74), Expect = 4.1
Identities = 23/81 (28%), Positives = 35/81 (42%)
Query: 107 DLDPQLKAGLLTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGEETGTMGEIEAELT 166
DLD ++A LEL ELH + LE+ ++ G E + E AE+
Sbjct: 541 DLDAAVQAAQQENLELRSRCEELHTKNLEMGKIMDGFEGIVYQAMEEVQRQKEAARAEVQ 600
Query: 167 SLLATREGLDRDLGAVHLRYN 187
+L RE L DL + ++
Sbjct: 601 KVLKDREQLAADLSSTEKSFS 621
>UniRef100_Q95KL6 Centrosome-associated AKAP350-binding protein TACC4 [Oryctolagus
cuniculus]
Length = 454
Score = 33.1 bits (74), Expect = 4.1
Identities = 23/81 (28%), Positives = 35/81 (42%)
Query: 107 DLDPQLKAGLLTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGEETGTMGEIEAELT 166
DLD ++A LEL ELH + LE+ ++ G E + E AE+
Sbjct: 259 DLDAAVQAAQQENLELRSRCEELHTKNLEMGKIMDGFEGIVYQAMEEVQRQKEAARAEVQ 318
Query: 167 SLLATREGLDRDLGAVHLRYN 187
+L RE L DL + ++
Sbjct: 319 KVLKDREQLAADLSSTEKSFS 339
>UniRef100_UPI000042F0A4 UPI000042F0A4 UniRef100 entry
Length = 534
Score = 32.7 bits (73), Expect = 5.3
Identities = 16/38 (42%), Positives = 18/38 (47%)
Query: 29 PPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDKMTSP 66
P + RGRG A R G P S PPQD +SP
Sbjct: 469 PARGRGRGKAYSRGRSGGGGPRPTSDTRTPPQDGQSSP 506
>UniRef100_UPI000023F00F UPI000023F00F UniRef100 entry
Length = 761
Score = 32.7 bits (73), Expect = 5.3
Identities = 21/78 (26%), Positives = 40/78 (50%)
Query: 94 GRIRSLVSGLLTEDLDPQLKAGLLTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHKGE 153
G +++ G L P + A LT ++ E++ EL R+ + WG ++ ++++
Sbjct: 648 GFFSNIIGGGLWSRGTPSVTAKTLTVEDMNEQLDELLRQSIYNFTHRWGSLMNDSQNRKP 707
Query: 154 ETGTMGEIEAELTSLLAT 171
+ ++EAEL SLL T
Sbjct: 708 GVKPIAKVEAELESLLMT 725
>UniRef100_UPI0000457408 UPI0000457408 UniRef100 entry
Length = 2149
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_UPI000036D337 UPI000036D337 UniRef100 entry
Length = 1405
Score = 32.3 bits (72), Expect = 6.9
Identities = 39/177 (22%), Positives = 66/177 (37%), Gaps = 25/177 (14%)
Query: 12 IRPCHVRKEAVTAKRHTPPKARGRGLAKQRDPPGNDTYHPKS------------------ 53
+ P + + T K + P LA + DP G P+
Sbjct: 1136 LTPANSQASKATPKLDSSPSVSST-LAAKDDPDGKQEAKPQQAAGMLSPKTGGKEAASGT 1194
Query: 54 --QASHPPQDKMTSPEAYLKALRAEALDCNLYYSYQTQVTAPGRIRSLVSGLLTEDLDPQ 111
Q S P+ +P+A AL++ C L + ++++ V +LTE L+ +
Sbjct: 1195 TPQKSRKPKKGAGNPQASTLALQSNIAQCLLGQPWPLN---EAQVQASVVKVLTELLEQE 1251
Query: 112 LKAGLLTFLELTEEMWELHRRRLEVNMLIWGLEVSATEHK-GEETGTMGEIEAELTS 167
K + T E + + WE +R+L + L S + K G G + E TS
Sbjct: 1252 RKKVVDTTKESSRKGWESRKRKLSGDQLAARTPRSKKKKKLGAREGGEASVSPEKTS 1308
>UniRef100_Q73GP3 Signal peptidase I [Wolbachia pipientis wMel]
Length = 248
Score = 32.3 bits (72), Expect = 6.9
Identities = 26/92 (28%), Positives = 40/92 (43%), Gaps = 9/92 (9%)
Query: 27 HTPPKARGRGLAKQRDPPGNDTYHPKSQASHPPQDK--MTSPEAYLKALRAEALDCNLYY 84
+TPPK RG P ND+ + P DK M E YL + E ++
Sbjct: 81 YTPPK---RGDIVVFKPTRNDSIRFVKRVIGTPGDKVQMIEGELYLNDQKVERRQIESFF 137
Query: 85 SYQTQVTAPGRIRSLVSG----LLTEDLDPQL 112
Y++ P I +L+SG +L +D+ +L
Sbjct: 138 DYESNRNIPRYIETLLSGKEHEILVDDISNKL 169
>UniRef100_Q5TM26 HLA-B associated transcript-2 [Macaca mulatta]
Length = 2160
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPAPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_Q5SQ29 HLA-B associated transcript 2 [Homo sapiens]
Length = 2157
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_Q96QC6 BAT2 protein [Homo sapiens]
Length = 2157
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_Q5STX6 OTTHUMP00000062432 [Homo sapiens]
Length = 2157
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_Q6P9P7 BAT2 protein [Homo sapiens]
Length = 2157
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_O95875 BAT2 [Homo sapiens]
Length = 2157
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 588 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 647
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 648 QRQQQEQLLK 657
>UniRef100_P48634 Large proline-rich protein BAT2 [Homo sapiens]
Length = 2142
Score = 32.3 bits (72), Expect = 6.9
Identities = 20/70 (28%), Positives = 27/70 (38%), Gaps = 9/70 (12%)
Query: 11 PIRPCHVRKEAVTAKRHTPP---------KARGRGLAKQRDPPGNDTYHPKSQASHPPQD 61
P+ P KE PP + +G G+ R PP +PK Q S PP+
Sbjct: 576 PVEPQLPSKEGPEPPEEVPPPTTPPVPKVEPKGDGIGPTRQPPSQGLGYPKYQKSLPPRF 635
Query: 62 KMTSPEAYLK 71
+ E LK
Sbjct: 636 QRQQQEQLLK 645
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 330,451,977
Number of Sequences: 2790947
Number of extensions: 12953390
Number of successful extensions: 32485
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 32464
Number of HSP's gapped (non-prelim): 42
length of query: 195
length of database: 848,049,833
effective HSP length: 121
effective length of query: 74
effective length of database: 510,345,246
effective search space: 37765548204
effective search space used: 37765548204
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0083.13


