

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0079c.14
(171 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
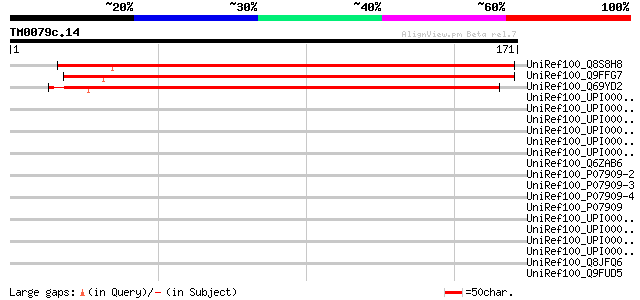
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S8H8 Hypothetical protein At2g15695 [Arabidopsis tha... 166 2e-40
UniRef100_Q9FFG7 Similarity to unknown protein [Arabidopsis thal... 155 5e-37
UniRef100_Q69YD2 Hypothetical protein P0701E03.39 [Oryza sativa] 148 5e-35
UniRef100_UPI0000432747 UPI0000432747 UniRef100 entry 35 1.0
UniRef100_UPI0000432745 UPI0000432745 UniRef100 entry 35 1.0
UniRef100_UPI0000432744 UPI0000432744 UniRef100 entry 35 1.0
UniRef100_UPI0000432743 UPI0000432743 UniRef100 entry 35 1.0
UniRef100_UPI0000432742 UPI0000432742 UniRef100 entry 35 1.0
UniRef100_UPI0000432741 UPI0000432741 UniRef100 entry 35 1.0
UniRef100_Q6ZAB6 Hypothetical protein P0429B05.11 [Oryza sativa] 35 1.0
UniRef100_P07909-2 Splice isoform A of P07909 [Drosophila melano... 33 2.3
UniRef100_P07909-3 Splice isoform E of P07909 [Drosophila melano... 33 2.3
UniRef100_P07909-4 Splice isoform D of P07909 [Drosophila melano... 33 2.3
UniRef100_P07909 Heterogeneous nuclear ribonucleoprotein A1 [Dro... 33 2.3
UniRef100_UPI0000361EAE UPI0000361EAE UniRef100 entry 33 3.0
UniRef100_UPI0000361EAD UPI0000361EAD UniRef100 entry 33 3.0
UniRef100_UPI000023F27A UPI000023F27A UniRef100 entry 33 3.9
UniRef100_UPI000028601A UPI000028601A UniRef100 entry 32 5.0
UniRef100_Q8JFQ6 Keratin 13 [Oncorhynchus mykiss] 32 5.0
UniRef100_Q9FUD5 Glycine-rich RNA-binding protein [Sorghum bicolor] 32 5.0
>UniRef100_Q8S8H8 Hypothetical protein At2g15695 [Arabidopsis thaliana]
Length = 420
Score = 166 bits (420), Expect = 2e-40
Identities = 80/165 (48%), Positives = 109/165 (65%), Gaps = 11/165 (6%)
Query: 17 VRGGNIYWGRKHATDFR-----------GIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLV 65
+ GG +YWG+K + G+VVIF W S+ + L +FV+LYSSLGWNSLV
Sbjct: 2 IGGGRVYWGKKLDKEMEDAAVVDGGGSNGVVVIFVWSSINENQLMNFVDLYSSLGWNSLV 61
Query: 66 CYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRC 125
C A +L+A E + LAF ++ EL+EELK++ CPV+F AFS KAC+YKVLQ+I C
Sbjct: 62 CRADFLTAVYPEMALSLAFHLLSELVEELKSRPCPVIFLAFSGAPKACMYKVLQVIMDDC 121
Query: 126 ETPHCLHNYHLLRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
E + L+R C+SGH+YDSGPLD TSD +FALHP++ ++
Sbjct: 122 EAQIHPDDSQLVRTCLSGHVYDSGPLDFTSDLNVKFALHPTIRRM 166
>UniRef100_Q9FFG7 Similarity to unknown protein [Arabidopsis thaliana]
Length = 403
Score = 155 bits (391), Expect = 5e-37
Identities = 76/154 (49%), Positives = 106/154 (68%), Gaps = 2/154 (1%)
Query: 19 GGNIYWGRKHAT--DFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAHYLSAFRD 76
GGN YW +K+ + IVV+FAW+S + L++ V+LYSSL W+SLVC++ +L+ F
Sbjct: 5 GGNYYWRKKNNNGGESEAIVVVFAWMSSEERNLKNHVDLYSSLLWDSLVCHSQFLNMFLP 64
Query: 77 ESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPHCLHNYHL 136
+ LA VV EL++ELK K P+VFA+FS G AC+YKVLQ+++G CET + L
Sbjct: 65 DKAADLASNVVSELVKELKAKPVPLVFASFSGGPNACMYKVLQILEGTCETGLNPDDCRL 124
Query: 137 LRNCVSGHIYDSGPLDVTSDFGFRFALHPSMAKV 170
+RNC+SG IYDS P+D TSD G R A+HP+ K+
Sbjct: 125 VRNCISGFIYDSCPVDFTSDLGARLAVHPTTLKM 158
>UniRef100_Q69YD2 Hypothetical protein P0701E03.39 [Oryza sativa]
Length = 405
Score = 148 bits (374), Expect = 5e-35
Identities = 72/156 (46%), Positives = 95/156 (60%), Gaps = 7/156 (4%)
Query: 14 GCAVRGGNIYWG----RKHATDFRGIVVIFAWVSVPQTLLQDFVNLYSSLGWNSLVCYAH 69
GC GG YW A RG+VV+F WV + L+ FV LY+SLGW LVC+
Sbjct: 4 GC---GGRFYWAPAPPSPSAAGARGVVVVFGWVWSDEAQLRPFVELYASLGWRCLVCHPD 60
Query: 70 YLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACLYKVLQLIDGRCETPH 129
++ + E LA V+ EL++E K K P VFA+FS GSK C+YKV+QL+DG CE
Sbjct: 61 LVALYLSEKAASLASGVISELVKEFKVKPLPTVFASFSGGSKGCMYKVIQLLDGNCEGDA 120
Query: 130 CLHNYHLLRNCVSGHIYDSGPLDVTSDFGFRFALHP 165
+ +Y L+RNC+ G IYDSGP+D SD G +F +P
Sbjct: 121 TMKDYRLVRNCICGQIYDSGPVDFFSDVGTQFLQNP 156
>UniRef100_UPI0000432747 UPI0000432747 UniRef100 entry
Length = 1302
Score = 34.7 bits (78), Expect = 1.0
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1039 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1092
>UniRef100_UPI0000432745 UPI0000432745 UniRef100 entry
Length = 1559
Score = 34.7 bits (78), Expect = 1.0
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1050 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1103
>UniRef100_UPI0000432744 UPI0000432744 UniRef100 entry
Length = 1520
Score = 34.7 bits (78), Expect = 1.0
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1034 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1087
>UniRef100_UPI0000432743 UPI0000432743 UniRef100 entry
Length = 1674
Score = 34.7 bits (78), Expect = 1.0
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1127 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1180
>UniRef100_UPI0000432742 UPI0000432742 UniRef100 entry
Length = 1658
Score = 34.7 bits (78), Expect = 1.0
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1113 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1166
>UniRef100_UPI0000432741 UPI0000432741 UniRef100 entry
Length = 1624
Score = 34.7 bits (78), Expect = 1.0
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Query: 60 GWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAAFSAGSKACL 114
GW ++ H+ ++ RDE+ V LAF + ++I EL + ++ +F K CL
Sbjct: 1113 GWKNIFSVFHHAASDRDEAVVELAFSMTGKIINELYAEDFSIMVDSFQDAVK-CL 1166
>UniRef100_Q6ZAB6 Hypothetical protein P0429B05.11 [Oryza sativa]
Length = 330
Score = 34.7 bits (78), Expect = 1.0
Identities = 13/21 (61%), Positives = 15/21 (70%)
Query: 4 AESEGGGGGGGCAVRGGNIYW 24
A S+GGGGGGGC GG Y+
Sbjct: 47 ASSDGGGGGGGCRSNGGRFYF 67
>UniRef100_P07909-2 Splice isoform A of P07909 [Drosophila melanogaster]
Length = 364
Score = 33.5 bits (75), Expect = 2.3
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 256 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 284
>UniRef100_P07909-3 Splice isoform E of P07909 [Drosophila melanogaster]
Length = 360
Score = 33.5 bits (75), Expect = 2.3
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 252 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 280
>UniRef100_P07909-4 Splice isoform D of P07909 [Drosophila melanogaster]
Length = 361
Score = 33.5 bits (75), Expect = 2.3
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 253 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 281
>UniRef100_P07909 Heterogeneous nuclear ribonucleoprotein A1 [Drosophila
melanogaster]
Length = 365
Score = 33.5 bits (75), Expect = 2.3
Identities = 15/29 (51%), Positives = 16/29 (54%)
Query: 3 GAESEGGGGGGGCAVRGGNIYWGRKHATD 31
G S GGGGGGG GGN WG + D
Sbjct: 257 GNNSFGGGGGGGGGYGGGNNSWGNNNPWD 285
>UniRef100_UPI0000361EAE UPI0000361EAE UniRef100 entry
Length = 1109
Score = 33.1 bits (74), Expect = 3.0
Identities = 21/67 (31%), Positives = 32/67 (47%), Gaps = 5/67 (7%)
Query: 46 QTLLQDFVNLYSSLGWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAA 105
Q LL+D + WN+L+ + DES++ +FCV+ I+ K +C V
Sbjct: 1006 QQLLKDINRV-----WNNLMGFMSLAKLTPDESSLDFSFCVLRHGIKNAKELACGVCLLN 1060
Query: 106 FSAGSKA 112
A SKA
Sbjct: 1061 VDARSKA 1067
>UniRef100_UPI0000361EAD UPI0000361EAD UniRef100 entry
Length = 1266
Score = 33.1 bits (74), Expect = 3.0
Identities = 21/67 (31%), Positives = 32/67 (47%), Gaps = 5/67 (7%)
Query: 46 QTLLQDFVNLYSSLGWNSLVCYAHYLSAFRDESTVPLAFCVVDELIEELKTKSCPVVFAA 105
Q LL+D + WN+L+ + DES++ +FCV+ I+ K +C V
Sbjct: 1163 QQLLKDINRV-----WNNLMGFMSLAKLTPDESSLDFSFCVLRHGIKNAKELACGVCLLN 1217
Query: 106 FSAGSKA 112
A SKA
Sbjct: 1218 VDARSKA 1224
>UniRef100_UPI000023F27A UPI000023F27A UniRef100 entry
Length = 657
Score = 32.7 bits (73), Expect = 3.9
Identities = 16/35 (45%), Positives = 22/35 (62%), Gaps = 4/35 (11%)
Query: 2 RGAESEGGGGGGGCA----VRGGNIYWGRKHATDF 32
+G + EGGGGGGG + + GG IY+G A +F
Sbjct: 11 QGEQREGGGGGGGFSFNKLILGGAIYFGLNAAMNF 45
>UniRef100_UPI000028601A UPI000028601A UniRef100 entry
Length = 255
Score = 32.3 bits (72), Expect = 5.0
Identities = 25/64 (39%), Positives = 33/64 (51%), Gaps = 15/64 (23%)
Query: 87 VDELIEELKTKSCPVVFAAF----SAGSKACLYKVLQLIDGRCETPHCLH---------N 133
VDEL EE+K K+ PV+F+A S KA YK ++ ID C +H N
Sbjct: 63 VDEL-EEIKDKTRPVIFSAHGVPKSVPKKAAEYK-MEYIDATCPLVSKVHKEAENLHKNN 120
Query: 134 YHLL 137
YH+L
Sbjct: 121 YHIL 124
>UniRef100_Q8JFQ6 Keratin 13 [Oncorhynchus mykiss]
Length = 492
Score = 32.3 bits (72), Expect = 5.0
Identities = 13/20 (65%), Positives = 16/20 (80%)
Query: 6 SEGGGGGGGCAVRGGNIYWG 25
S GGGGGGG A+R G++Y G
Sbjct: 41 SMGGGGGGGGAMRSGSVYGG 60
>UniRef100_Q9FUD5 Glycine-rich RNA-binding protein [Sorghum bicolor]
Length = 170
Score = 32.3 bits (72), Expect = 5.0
Identities = 15/31 (48%), Positives = 18/31 (57%)
Query: 4 AESEGGGGGGGCAVRGGNIYWGRKHATDFRG 34
A+S GGGGGGG GG Y GR+ + G
Sbjct: 84 AQSRGGGGGGGGYGGGGGGYGGRREGGGYGG 114
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.141 0.461
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 311,385,112
Number of Sequences: 2790947
Number of extensions: 12931700
Number of successful extensions: 83339
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 83276
Number of HSP's gapped (non-prelim): 40
length of query: 171
length of database: 848,049,833
effective HSP length: 118
effective length of query: 53
effective length of database: 518,718,087
effective search space: 27492058611
effective search space used: 27492058611
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0079c.14


