

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
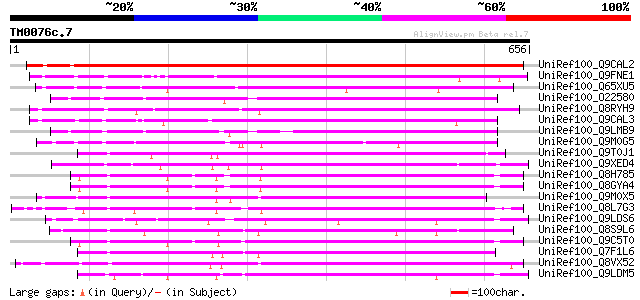
Query= TM0076c.7
(656 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9CAL2 Hypothetical protein F24J13.10 [Arabidopsis tha... 715 0.0
UniRef100_Q9FNE1 Receptor-like serine/threonine kinase [Arabidop... 482 e-134
UniRef100_Q65XU5 Hypothetical protein P0033D06.1 [Oryza sativa] 454 e-126
UniRef100_O22580 Receptor-like serine/threonine kinase [Arabidop... 434 e-120
UniRef100_Q8RYH9 Hypothetical protein OSJNBa0042P21.23 [Oryza sa... 423 e-117
UniRef100_Q9CAL3 Hypothetical protein F24J13.9 [Arabidopsis thal... 422 e-116
UniRef100_Q9LMB9 F14D16.24 [Arabidopsis thaliana] 412 e-113
UniRef100_Q9M0G5 Serine/threonine kinase-like protein [Arabidops... 375 e-102
UniRef100_Q9T0J1 Receptor-like protein kinase-like protein [Arab... 327 7e-88
UniRef100_Q9XED4 Receptor-like protein kinase homolog RK20-1 [Ph... 315 4e-84
UniRef100_Q8H785 Hypothetical protein [Arabidopsis thaliana] 312 2e-83
UniRef100_Q8GYA4 Putative receptor-like protein kinase 4 RLK4 [A... 312 2e-83
UniRef100_Q9M0X5 Receptor protein kinase-like protein [Arabidops... 309 1e-82
UniRef100_Q8L7G3 Putative serine/threonine kinase [Arabidopsis t... 308 3e-82
UniRef100_Q9LDS6 Serine/threonine kinase-like protein [Arabidops... 305 2e-81
UniRef100_Q8S9L6 AT4g21410/T6K22_140 [Arabidopsis thaliana] 304 5e-81
UniRef100_Q9C5T0 Receptor-like protein kinase 4 [Arabidopsis tha... 303 8e-81
UniRef100_Q7F1L6 Putative serine/threonine kinase receptor [Oryz... 299 2e-79
UniRef100_Q8VX52 Putative receptor-like serine-threonine protein... 299 2e-79
UniRef100_Q9LDM5 Serine/threonine kinase-like protein [Arabidops... 298 3e-79
>UniRef100_Q9CAL2 Hypothetical protein F24J13.10 [Arabidopsis thaliana]
Length = 646
Score = 715 bits (1846), Expect = 0.0
Identities = 358/629 (56%), Positives = 462/629 (72%), Gaps = 6/629 (0%)
Query: 22 PTLADPRATEAASLCTNRTSPMPQSQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTT 81
P+++D R A +C NRT+ PQ QR +F++NF A+++++ LV + +G VV G T
Sbjct: 17 PSVSDSRGDTVAQICNNRTTT-PQ-QRSLFVTNFLAAMDAVSPLVEAKGYGQVVNG---T 71
Query: 82 QNATVYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNF 141
N TVYA+GEC+KDL K DCD+CFAQ K +V RC P Q+G GG+ F DGCY+RYD YNF
Sbjct: 72 GNLTVYAYGECIKDLDKKDCDLCFAQIKAKVPRCLPFQKGTRGGQVFSDGCYIRYDDYNF 131
Query: 142 FNQSLSSQDLTLCGNIDFSG-NRSVHKANVVELVRNLSVEAPKNDGFFVGVLSKRNVSVY 200
+N++LS QD T+C + +G NR+V + N ELV+N+SVEA +N GF+ G + + NV+V+
Sbjct: 132 YNETLSLQDRTVCAPKEITGVNRTVFRDNAAELVKNMSVEAVRNGGFYAGFVDRHNVTVH 191
Query: 201 GLAQCWNFVNESVCQNCLVEAVTRIDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNIAPP 260
GLAQCW +N S C CL +A RI SC + EEGRVL+AGCY+RFST KFY+NS N
Sbjct: 192 GLAQCWETLNRSGCVECLSKASVRIGSCLVNEEGRVLSAGCYMRFSTQKFYNNSGNSTSD 251
Query: 261 GTKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNM 320
G G +G+I+A + + VA +L+V+ F ++K +++RE++Q G+ N S L
Sbjct: 252 GNGGHNHLGVILAVTSSVVAFVLLVSAAGFLLKKRHAKKQREKKQLGSLFMLANKSNLCF 311
Query: 321 PYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLI 380
YE LE+AT+YF D NKLG+GGSGSVYKG L +G TVA+KRL FNT QW DHFFNEVNLI
Sbjct: 312 SYENLERATDYFSDKNKLGQGGSGSVYKGVLTNGKTVAVKRLFFNTKQWVDHFFNEVNLI 371
Query: 381 KDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAE 440
+ HKNLVKLLGCSITGPESLLVYEY+ N SLHD+L VR+++Q L W R KIILGTAE
Sbjct: 372 SQVDHKNLVKLLGCSITGPESLLVYEYIANQSLHDYLFVRKDVQPLNWAKRFKIILGTAE 431
Query: 441 GLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYM 500
G+AYLHEES L+IIHRDIKLSNILL+D+FTP+IADFGLARLFPED++ ISTAI GTLGYM
Sbjct: 432 GMAYLHEESNLRIIHRDIKLSNILLEDDFTPRIADFGLARLFPEDKTHISTAIAGTLGYM 491
Query: 501 APEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVD 560
APEY+V GKLTEKAD+YSFGVL+IE+++GK +FVQ++ SIL VWSLY ++ + + VD
Sbjct: 492 APEYVVRGKLTEKADVYSFGVLMIEVITGKRNNAFVQDAGSILQSVWSLYRTSNVEEAVD 551
Query: 561 PILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPFLNFGSG 620
PIL N+ EA +LL+IGLLC QA+ + RP MSVVVKM+ +I PTQPPFLN GS
Sbjct: 552 PILGDNFNKIEASRLLQIGLLCVQAAFDQRPAMSVVVKMMKGSLEIHTPTQPPFLNPGSV 611
Query: 621 EFSRSSLPEESLQPGSNTQSSGDSMTESS 649
R + + +++ S D +TE S
Sbjct: 612 VEMRKMMMTPTTNQSNSSGSRSDYITEGS 640
>UniRef100_Q9FNE1 Receptor-like serine/threonine kinase [Arabidopsis thaliana]
Length = 651
Score = 482 bits (1240), Expect = e-134
Identities = 277/644 (43%), Positives = 370/644 (57%), Gaps = 38/644 (5%)
Query: 26 DPRATEAASLCTNRTSPMPQSQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNAT 85
D R T + C R+ S ++ F + + SL+ +TT+R T S+T+
Sbjct: 31 DQRTTVSGLFCGGRSK---SSADPNYIPTFVEDMHSLSLKLTTRRFATESLNSTTS---- 83
Query: 86 VYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQS 145
+YA +C DLS SDC +C+A +TR+ RC P R F DGC+LRY+ Y F+++S
Sbjct: 84 IYALIQCHDDLSPSDCQLCYAIARTRIPRCLPSS----SARIFLDGCFLRYETYEFYDES 139
Query: 146 LS-SQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVGVLSKRNVSVYGLAQ 204
+S + D C N R + V E ++V + GF GV + V + LAQ
Sbjct: 140 VSDASDSFSCSNDTVLDPRFGFQ--VSETAARVAV---RKGGF--GVAGENGV--HALAQ 190
Query: 205 CWNFVNESVCQNCLVEAVTRIDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNIAPPGTKG 264
CW + + C+ CL +AV + C + EGR +N GCYLR+S KFY+ +
Sbjct: 191 CWESLGKEDCRVCLEKAVKEVKRCVSRREGRAMNTGCYLRYSDHKFYNGDGHHK---FHV 247
Query: 265 STKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYEV 324
G+IVA A ++++ + + V + ++E+R G NNSK YE
Sbjct: 248 LFNKGVIVAIVLTTSAFVMLILLATYVIMTKVSKTKQEKRNLGLVSRKFNNSKTKFKYET 307
Query: 325 LEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQ 384
LEKAT+YF LG+GG+G+V+ G LP+G VA+KRL FNT W + FFNEVNLI +Q
Sbjct: 308 LEKATDYFSHKKMLGQGGNGTVFLGILPNGKNVAVKRLVFNTRDWVEEFFNEVNLISGIQ 367
Query: 385 HKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAY 444
HKNLVKLLGCSI GPESLLVYEYVPN SL L + L W R IILGTAEGLAY
Sbjct: 368 HKNLVKLLGCSIEGPESLLVYEYVPNKSLDQFLFDESQSKVLNWSQRLNIILGTAEGLAY 427
Query: 445 LHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEY 504
LH S ++IIHRDIK SN+LLDD PKIADFGLAR F D++ +ST I GTLGYMAPEY
Sbjct: 428 LHGGSPVRIIHRDIKTSNVLLDDQLNPKIADFGLARCFGLDKTHLSTGIAGTLGYMAPEY 487
Query: 505 IVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILE 564
+V G+LTEKAD+YSFGVLV+E+ G +FV + +L VW+LY NRL + +DP L+
Sbjct: 488 VVRGQLTEKADVYSFGVLVLEIACGTRINAFVPETGHLLQRVWNLYTLNRLVEALDPCLK 547
Query: 565 GNY-----PAEEACKLLKIGLLCTQASAELRPPMSVVVKMI-NNIHKIADPTQPPFLNF- 617
+ EACK+L++GLLCTQAS LRP M V++M+ + I PT PPFL
Sbjct: 548 DEFLQVQGSEAEACKVLRVGLLCTQASPSLRPSMEEVIRMLTERDYPIPSPTSPPFLRVS 607
Query: 618 -------GSGEFSRSSLPEESLQPGSNTQSSGDSMTESSEHRLV 654
GS S S+ + T + + +ESS R +
Sbjct: 608 SLTTDLEGSSTISHSTNSTTTFNTMVKTDQASYTSSESSTTRTI 651
>UniRef100_Q65XU5 Hypothetical protein P0033D06.1 [Oryza sativa]
Length = 702
Score = 454 bits (1167), Expect = e-126
Identities = 267/632 (42%), Positives = 365/632 (57%), Gaps = 41/632 (6%)
Query: 33 ASLCTNRTSPMPQSQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGEC 92
A++C N T+ + ++ F +F + LE + VT G G TVY G+C
Sbjct: 40 ATVCGNTTA----ADQEGFDVSFVNTLELIYQNVTRSGFGAAASGEGAD---TVYGLGQC 92
Query: 93 MKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQDLT 152
M LS +DC +C+AQ + ++ C P GGR + DGC+LRY A NF + + D
Sbjct: 93 MGYLSPTDCQLCYAQSRVKLPHCLP----ATGGRIYLDGCFLRYGADNFTAAATDASDTA 148
Query: 153 LCGNIDFSGNRSVHKANVVELVRNLSVEAP-KNDGFFVGVLSKRNVS--------VYGLA 203
+C N S S A L+RN++ AP D ++ S + S VY A
Sbjct: 149 VCSNATVSSPASF-AATSAALLRNVTAAAPGARDYYYYSASSSSSASALPSVSPRVYAAA 207
Query: 204 QCWNFVNESVCQNCLVEAVTRIDSCALKE--EGRVLNAGCYLRFSTDKFYDNSSNIAPPG 261
QCW +N + C C+ A R+ L EG LNAGC +R+ST FY ++ A G
Sbjct: 208 QCWRSLNATACAACVATARDRVVGRCLPRAAEGYGLNAGCVVRYSTQPFYLPANAAAAAG 267
Query: 262 TKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSE-NNSKLNM 320
+ V +++A F A+A + I +++ +N + + + S+L+
Sbjct: 268 SSTRHIVIVVIASVFCALAVIGIA--LVWAKMRNRRNDHHDDMDGSSEIIRTIAASQLSF 325
Query: 321 PYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLI 380
YE L KAT+ F+ NKLG+GG GSVYKG L DG +A+KRL FNT +WAD FFNEV L+
Sbjct: 326 KYEELCKATDDFNQINKLGQGGYGSVYKGVLLDGREIAVKRLFFNTREWADQFFNEVRLV 385
Query: 381 KDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVR-RNLQ----------QLTWE 429
+QHKNLVKLLGCSI GPESLLVYEY+ N SL +L R R+ L WE
Sbjct: 386 SQVQHKNLVKLLGCSIEGPESLLVYEYLCNTSLDHYLFDRFRSFMVDVADAFKKTALDWE 445
Query: 430 VRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQI 489
R +IILGTAEGL+YLH S+++IIHRDIK SN+LLD+ F PKIADFGLAR F EDQS +
Sbjct: 446 RRFEIILGTAEGLSYLHNASEIRIIHRDIKASNVLLDERFRPKIADFGLARNFMEDQSHL 505
Query: 490 STAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSC---SILHMV 546
ST + GT GYMAPEYIV G+LTEKADIYS+GVLV+E+++G+ + V +S S++ ++
Sbjct: 506 STGLAGTFGYMAPEYIVHGQLTEKADIYSYGVLVLEIITGRKSLNSVASSAEGHSLMSLI 565
Query: 547 WSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMI-NNIHK 605
W Y L +++DP L+ E A K+ +GLLC QAS LRPPM VV+M+ + ++
Sbjct: 566 WKHYNEGTLMELLDPNLQEQCTEEGALKVFHVGLLCAQASPNLRPPMWKVVEMLGSRNNE 625
Query: 606 IADPTQPPFLNFGSGEFSRSSLPEESLQPGSN 637
+ PTQPPF+N S SL+ S+
Sbjct: 626 LPRPTQPPFINIKGSNAKSDSSGSSSLKSSSD 657
>UniRef100_O22580 Receptor-like serine/threonine kinase [Arabidopsis thaliana]
Length = 617
Score = 434 bits (1117), Expect = e-120
Identities = 249/578 (43%), Positives = 347/578 (59%), Gaps = 33/578 (5%)
Query: 52 LSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKTR 111
L F A+ S+ +T + V S T + +Y F +C +DLS SDC CF + +
Sbjct: 48 LLGFLRAMSSVNDFITNDKLWVV--SSITDVSPPIYVFLQCREDLSVSDCRHCFNESRLE 105
Query: 112 VLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQ-DLTLCGNIDFSGNRSVHKANV 170
+ R V GR D C+LR+D +F + + D C +
Sbjct: 106 LER----NVRVPVGRIHSDRCFLRFDDRDFSEEFVDPTFDKANCEETGTGFGEFWRFLD- 160
Query: 171 VELVRNLSVEAPKNDGFFVGVLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRIDSCAL 230
E + N++++A KN GF + K +VY LAQCW ++E+ C+ CLV A + + +C
Sbjct: 161 -EALVNVTLKAVKNGGFGAASVIKTE-AVYALAQCWQTLDENTCRECLVNARSSLRACD- 217
Query: 231 KEEGRVLNAGCYLRFSTDKFYDNSSNIAPPGTKGSTKVGI----------IVAESFAAVA 280
E R GCYL++ST KF+D+ + P + + + AA++
Sbjct: 218 GHEARAFFTGCYLKYSTHKFFDDDAAEHKPDADQRNFIRSSFFPHLSDRDVTRLAIAAIS 277
Query: 281 TLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGE 340
++ + F + V R+R+ ++ S +N YE+LEKAT FHDS KLG+
Sbjct: 278 LSILTSLGAFISYRRVSRKRK----------AQVPSCVNFKYEMLEKATESFHDSMKLGQ 327
Query: 341 GGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPE 400
GG+GSVYKG LPDG VA+K+L FNT +WAD FFNEVNLI +QHKNLV+LLGCSI GP+
Sbjct: 328 GGAGSVYKGILPDGRIVAVKKLFFNTREWADQFFNEVNLISGVQHKNLVRLLGCSIEGPK 387
Query: 401 SLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKL 460
SLLVYEYV N SL L ++ + L+W+ R II+G +EGL YLH S++KIIHRDIK
Sbjct: 388 SLLVYEYVHNRSLDQILFMKNTVHILSWKQRFNIIIGISEGLEYLHRGSEVKIIHRDIKT 447
Query: 461 SNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFG 520
SNILLD N +PKIADFGL R D++Q +T I GTLGY+APEY++ G+LTEKAD+Y+FG
Sbjct: 448 SNILLDRNLSPKIADFGLIRSMGTDKTQTNTGIAGTLGYLAPEYLIKGQLTEKADVYAFG 507
Query: 521 VLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGL 580
VL+IE+++GK +F Q + S+L+ VW + +N L +DP L+G++ EEA K+L+IGL
Sbjct: 508 VLIIEIVTGKKNNAFTQGTSSVLYSVWEHFKANTLDRSIDPRLKGSFVEEEALKVLQIGL 567
Query: 581 LCTQASAELRP-PMSVVVKMI-NNIHKIADPTQPPFLN 616
LC Q+S ELRP MS +V M+ N K P QPPFL+
Sbjct: 568 LCVQSSVELRPSSMSEIVFMLQNKDSKFEYPKQPPFLS 605
>UniRef100_Q8RYH9 Hypothetical protein OSJNBa0042P21.23 [Oryza sativa]
Length = 663
Score = 423 bits (1087), Expect = e-117
Identities = 252/647 (38%), Positives = 353/647 (53%), Gaps = 40/647 (6%)
Query: 26 DPRATEAASLCTNRTSPMPQSQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNAT 85
DPR A C +P NF A++ L S V+ GT G++ N
Sbjct: 22 DPRTAVARQEC----APGGAVSGPALADNFVPAMDDLNSNVSANGFGTSAVGTTAGLNPN 77
Query: 86 -VYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQ 144
V+ G+C +DLS DC +CFA+ ++ + +C P GGR + DGC+ RY Y+FF++
Sbjct: 78 AVFGLGQCYRDLSPVDCKLCFAEVRSLLPKCYPSA----GGRLYLDGCFGRYANYSFFSE 133
Query: 145 SLSSQDLTLCGNID----------------FSGNRSVHKANVVELVRNLSVEAPKNDGFF 188
+L D CG + F+ ANV + + +V DG+
Sbjct: 134 TLGPDDAVTCGVVGGGGAGGGNYTGANPRGFADAVRAALANVTGVAASAAVPGG-GDGYA 192
Query: 189 VGVLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRIDSCA-LKEEGRVLNAGCYLRFST 247
VG S + + LAQCW +N + C CL A CA EGR L GCYLR+ST
Sbjct: 193 VGSASAGGATAFALAQCWGSLNATACGQCLRAAAAAAARCAPAAAEGRALYTGCYLRYST 252
Query: 248 DKFYDNSSNIAPPGTKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFG 307
F++ +S ++ + V I++ S A + IV + + +K + R ++ + F
Sbjct: 253 RLFWNLNSTAGSGSSRHNDVVWILLGSSLGAFVIVFIVVFLAW--KKKIFRNKKRSKSF- 309
Query: 308 AFLFSEN------NSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKR 361
++ + S LN YE L KATNYF +NKLG+G G+V+K L DG VA+KR
Sbjct: 310 IDIYGDGVPVRIAQSSLNFKYEELRKATNYFDPANKLGQGSYGAVFKAILLDGKQVAVKR 369
Query: 362 LSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRR 421
L NT +W D FFNEV LI ++HKNLVKLLGCS+ GPESLLVYEY N SL L
Sbjct: 370 LFLNTREWVDQFFNEVELISQVRHKNLVKLLGCSVNGPESLLVYEYYFNKSLDLFLFDAS 429
Query: 422 NLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARL 481
+ LTW +R II G AEGL+YLHEES+ +IIHRDIK SNILLDD F PKI DFGLAR
Sbjct: 430 RSRNLTWNLRVDIIQGIAEGLSYLHEESETRIIHRDIKASNILLDDKFKPKITDFGLARA 489
Query: 482 FPEDQSQISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFV--QNS 539
F ED++ ++T + GTLGYMAPEY+ G LTEKAD+YS+G+LV+EL++G+ + +
Sbjct: 490 FGEDRTHLTTGVAGTLGYMAPEYLAHGHLTEKADVYSYGILVLELVTGQRCSGSIGSHGG 549
Query: 540 CSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKM 599
+L VW+ Y + + I D + + +E +++IGL CTQA+A RP M+ VV++
Sbjct: 550 HFLLTKVWNHYKNKAVEMIADRSIYEDTIRDEVMHVVQIGLSCTQANAGDRPTMTKVVEL 609
Query: 600 INNIHKIAD--PTQPPFLNFGSGEFSRSSLPEESLQPGSNTQSSGDS 644
+ + + + PPFL+ + E + L S SG S
Sbjct: 610 LRSHRHDVEIILSDPPFLDVEAFEDIKQGEQSRLLSARSAHSVSGSS 656
>UniRef100_Q9CAL3 Hypothetical protein F24J13.9 [Arabidopsis thaliana]
Length = 649
Score = 422 bits (1085), Expect = e-116
Identities = 242/606 (39%), Positives = 345/606 (55%), Gaps = 32/606 (5%)
Query: 26 DPRATEAASLCTNRTSPMPQSQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNAT 85
D RA C SP+ + ++ NF +E +++ V T G + G+ N
Sbjct: 30 DARARAVKVTC----SPLLEHNETAYVPNFVATMEKISTQVQTSGFGVALTGTGPDAN-- 83
Query: 86 VYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQS 145
Y +C DL +DC +C+A+ +T + +C P GGR F DGC++R + Y+F+N+
Sbjct: 84 -YGLAQCYGDLPLNDCVLCYAEARTMLPQCYP----QNGGRIFLDGCFMRAENYSFYNEY 138
Query: 146 LSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVGVLS---KRNVSVYGL 202
+D +CGN N++ A V + +RN EA G+ + S + L
Sbjct: 139 KGPEDSIVCGNTTRK-NKTFGDA-VRQGLRNAVTEASGTGGYARASAKAGESESESAFVL 196
Query: 203 AQCWNFVNESVCQNCLVEA-VTRIDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNIAPPG 261
A CW ++ C+ CL A + + C EGR L+ GC+LR+S F N P
Sbjct: 197 ANCWRTLSPDSCKQCLENASASVVKGCLPWSEGRALHTGCFLRYSDQDFL----NKIPRN 252
Query: 262 TKGSTKVGIIVAESFAAVATLLIVATIIFFV--RKNVLRQRRERRQFGAFLFSENNSKLN 319
+ V +IV ++V +I + ++ R+ + R+RR + + +S LN
Sbjct: 253 GRSRGSVVVIVVSVLSSVVVFMIGVAVSVYICKRRTIKRKRRGSKDVEKMAKTLKDSSLN 312
Query: 320 MPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNL 379
Y LEKAT F ++NKLG+GG G+VYKG LPDG +A+KRL FN A F+NEVN+
Sbjct: 313 FKYSTLEKATGSFDNANKLGQGGFGTVYKGVLPDGRDIAVKRLFFNNRHRATDFYNEVNM 372
Query: 380 IKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTA 439
I ++HKNLV+LLGCS +GPESLLVYEY+ N SL + + L W+ R+ II+GTA
Sbjct: 373 ISTVEHKNLVRLLGCSCSGPESLLVYEYLQNKSLDRFIFDVNRGKTLDWQRRYTIIVGTA 432
Query: 440 EGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGY 499
EGL YLHE+S +KIIHRDIK SNILLD KIADFGLAR F +D+S ISTAI GTLGY
Sbjct: 433 EGLVYLHEQSSVKIIHRDIKASNILLDSKLQAKIADFGLARSFQDDKSHISTAIAGTLGY 492
Query: 500 MAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQN--SCSILHMVWSLYGSNRLCD 557
MAPEY+ G+LTE D+YSFGVLV+E+++GK T + S S++ W + S L
Sbjct: 493 MAPEYLAHGQLTEMVDVYSFGVLVLEIVTGKQNTKSKMSDYSDSLITEAWKHFQSGELEK 552
Query: 558 IVDPIL------EGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIAD-PT 610
I DP L + + +E ++++IGLLCTQ LRPPMS ++ M+ N ++ P+
Sbjct: 553 IYDPNLDWKSQYDSHIIKKEIARVVQIGLLCTQEIPSLRPPMSKLLHMLKNKEEVLPLPS 612
Query: 611 QPPFLN 616
PPF++
Sbjct: 613 NPPFMD 618
>UniRef100_Q9LMB9 F14D16.24 [Arabidopsis thaliana]
Length = 668
Score = 412 bits (1059), Expect = e-113
Identities = 242/582 (41%), Positives = 340/582 (57%), Gaps = 58/582 (9%)
Query: 52 LSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKTR 111
L F A+ S+ +T + V S T + +Y F +C +DLS SDC CF + +
Sbjct: 116 LLGFLRAMSSVNDFITNDKLWVV--SSITDVSPPIYVFLQCREDLSVSDCRHCFNESRLE 173
Query: 112 VLR-CSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQ-DLTLCGNIDFSGNRSVHKAN 169
+ R CS GGR D C+LR+D +F + + D C +
Sbjct: 174 LERKCSG-----SGGRIHSDRCFLRFDDRDFSEEFVDPTFDKANCEETGTGFGEFWRFLD 228
Query: 170 VVELVRNLSVEAPKNDGFFVGVLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRIDSCA 229
E + N++++A KN GF + K +VY LAQCW ++E+ C+ CLV A + + +C
Sbjct: 229 --EALVNVTLKAVKNGGFGAASVIKTE-AVYALAQCWQTLDENTCRECLVNARSSLRACD 285
Query: 230 LKEEGRVLNAGCYLRFSTDKFYDNSSNIAPPGTKGSTKVGIIVAESF------------- 276
E R GCYL++ST KF+D+++ P + + + SF
Sbjct: 286 -GHEARAFFTGCYLKYSTHKFFDDAAEHKPDADQRN-----FIRSSFFPHLSDRDVTRLA 339
Query: 277 -AAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDS 335
AA++ ++ + F + V R+R+ ++ S +N YE+LEKAT FHDS
Sbjct: 340 IAAISLSILTSLGAFISYRRVSRKRK----------AQVPSCVNFKYEMLEKATESFHDS 389
Query: 336 NKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCS 395
KLG+GG A+K+L FNT +WAD FFNEVNLI +QHKNLV+LLGCS
Sbjct: 390 MKLGQGG---------------AVKKLFFNTREWADQFFNEVNLISGVQHKNLVRLLGCS 434
Query: 396 ITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIH 455
I GP+SLLVYEYV N SL L ++ + L+W+ R II+G +EGL YLH S++KIIH
Sbjct: 435 IEGPKSLLVYEYVHNRSLDQILFMKNTVHILSWKQRFNIIIGISEGLEYLHRGSEVKIIH 494
Query: 456 RDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTAICGTLGYMAPEYIVLGKLTEKAD 515
RDIK SNILLD N +PKIADFGL R D++Q +T I GTLGY+APEY++ G+LTEKAD
Sbjct: 495 RDIKTSNILLDRNLSPKIADFGLIRSMGTDKTQTNTGIAGTLGYLAPEYLIKGQLTEKAD 554
Query: 516 IYSFGVLVIELLSGKSRTSFVQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKL 575
+Y+FGVL+IE+++GK +F Q + S+L+ VW + +N L +DP L+G++ EEA K+
Sbjct: 555 VYAFGVLIIEIVTGKKNNAFTQGTSSVLYSVWEHFKANTLDRSIDPRLKGSFVEEEALKV 614
Query: 576 LKIGLLCTQASAELRPPMSVVVKMI-NNIHKIADPTQPPFLN 616
L+IGLLC Q+S ELRP MS +V M+ N K P QPPFL+
Sbjct: 615 LQIGLLCVQSSVELRPSMSEIVFMLQNKDSKFEYPKQPPFLS 656
>UniRef100_Q9M0G5 Serine/threonine kinase-like protein [Arabidopsis thaliana]
Length = 625
Score = 375 bits (963), Expect = e-102
Identities = 222/607 (36%), Positives = 338/607 (55%), Gaps = 44/607 (7%)
Query: 35 LCTNRTSPMPQSQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGECMK 94
+C N T ++ R+ + N L+++ + + GT G + +Y +C+
Sbjct: 35 ICNNGTVSNEEAYRRSYQIN----LDAIRGDMRHVKFGTHEHGDPPER---MYVLSQCVS 87
Query: 95 DLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQDLTLC 154
DLS +C +C+++ + +C P GG F DGC++R D Y+F+ + +S QD +C
Sbjct: 88 DLSSDECSLCWSRATDLLSQCFP----ATGGWFHLDGCFVRADNYSFYQEPVSHQDTKIC 143
Query: 155 GNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVGVLSKRNVSVYGLAQCWNFVNESVC 214
+ + K V E+ +++ AP + GF V + R+++VYGL CW +N+ +C
Sbjct: 144 ASD--KEKSAEFKGLVKEVTKSIVEAAPYSRGFSVAKMGIRDLTVYGLGVCWRTLNDELC 201
Query: 215 QNCLVEAVTRIDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNIAPPGTKGSTKVGIIVAE 274
+ CL + + SC +EG LNAGCYLR+S FY+ +A TK + +++
Sbjct: 202 KLCLADGALSVTSCLPSKEGFALNAGCYLRYSNYTFYNERGLLAMSFTKENLTYIFVIS- 260
Query: 275 SFAAVATLLIVATI----IFFVR----KNVLRQRRERRQFGAFLFSENNSK--------L 318
V L I A F++R K + + ++ L E S+ +
Sbjct: 261 ---MVGVLAIAAGFWCGKCFYMRTSPKKKIKGTKTKKFHLFGHLRIEKESESICTESHLM 317
Query: 319 NMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVN 378
+ Y L+KATN F++S KLG GG G V+KG L DG +AIKRL + + D NE++
Sbjct: 318 SFEYSTLKKATNNFNESCKLGVGGYGEVFKGTLSDGREIAIKRLHVSGKKPRDEIHNEID 377
Query: 379 LIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGT 438
+I QHKNLV+LLGC T S +VYE++ N SL L ++L W+ R IILGT
Sbjct: 378 VISRCQHKNLVRLLGCCFTNMNSFIVYEFLANTSLDHILFNPEKKKELDWKKRRTIILGT 437
Query: 439 AEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQI------STA 492
AEGL YLHE KIIHRDIK SNILLD + PKI+DFGLA+ +PE I ++
Sbjct: 438 AEGLEYLHE--TCKIIHRDIKASNILLDLKYKPKISDFGLAKFYPEGGKDIPASSLSPSS 495
Query: 493 ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNSC--SILHMVWSLY 550
I GTLGYMAPEYI G+L+ K D YSFGVLV+E+ SG F ++ +++ VW +
Sbjct: 496 IAGTLGYMAPEYISKGRLSNKIDAYSFGVLVLEITSGFRNNKFRSDNSLETLVTQVWKCF 555
Query: 551 GSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIH-KIADP 609
SN++ +++D + + +E ++++IGLLCTQ S +LRP MS V++M+++ + P
Sbjct: 556 ASNKMEEMIDKDMGEDTDKQEMKRVMQIGLLCTQESPQLRPTMSKVIQMVSSTDIVLPTP 615
Query: 610 TQPPFLN 616
T+PPFL+
Sbjct: 616 TKPPFLH 622
>UniRef100_Q9T0J1 Receptor-like protein kinase-like protein [Arabidopsis thaliana]
Length = 665
Score = 327 bits (838), Expect = 7e-88
Identities = 211/573 (36%), Positives = 309/573 (53%), Gaps = 36/573 (6%)
Query: 86 VYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQS 145
V + +C D+ C C A R++ P+Q+ ++D C RY FN+
Sbjct: 77 VNSISQCRGDVKLEVCINCIAMAGKRLVTLCPVQKEAI---IWYDKCTFRYSNRTIFNRL 133
Query: 146 LSSQDLTLCGNIDFSGNRSVHKANVVELVRNL----SVEAPKNDGFFVGVLSKRNV-SVY 200
S ++ G +F+G+R + ++ L+ L SV F VG S + +++
Sbjct: 134 EISPHTSITGTRNFTGDRDSWEKSLRGLLEGLKNRASVIGRSKKNFVVGETSGPSFQTLF 193
Query: 201 GLAQCWNFVNESVCQNCLVEAVTRIDSCA-LKEEGRVLNAGCYLRFSTDKFYDN------ 253
GL QC ++E C CL + + +I SC +K V++ C L ++ +FYD
Sbjct: 194 GLVQCTPDISEEDCSYCLSQGIAKIPSCCDMKMGSYVMSPSCMLAYAPWRFYDPVDTDDP 253
Query: 254 SSNIAPPG----------TKGSTKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRER 303
SS A P T+G G+ A FA+ + ++V I+ V LR++
Sbjct: 254 SSVPATPSRPPKNETRSVTQGDKNRGVPKALIFASASVAIVVLFIVLLVVFLKLRRKENI 313
Query: 304 RQFGAFLFSENNSKLNMPYE--VLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKR 361
R +EN S +M ++ VL+ AT++F NKLGEGG G+VYKG L DG +A+KR
Sbjct: 314 RNSENKHENENISTDSMKFDFSVLQDATSHFSLENKLGEGGFGAVYKGVLSDGQKIAVKR 373
Query: 362 LSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRR 421
LS N Q F NE L+ LQH+NLVKLLG SI G E LLVYE++P+ SL +
Sbjct: 374 LSKNAQQGETEFKNEFLLVAKLQHRNLVKLLGYSIEGTERLLVYEFLPHTSLDKFIFDPI 433
Query: 422 NLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARL 481
+L WE+R+KII G A GL YLH++S+L+IIHRD+K SNILLD+ TPKIADFG+ARL
Sbjct: 434 QGNELEWEIRYKIIGGVARGLLYLHQDSRLRIIHRDLKASNILLDEEMTPKIADFGMARL 493
Query: 482 FPEDQS--QISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNS 539
F D + + + I GT GYMAPEY++ G+ + K D+YSFGVLV+E++SGK + F
Sbjct: 494 FDIDHTTQRYTNRIVGTFGYMAPEYVMHGQFSFKTDVYSFGVLVLEIISGKKNSGFSSED 553
Query: 540 C--SILHMVWSLYGSNRLCDIVDPIL--EGNYPAEEACKLLKIGLLCTQASAELRPPMSV 595
++ W + ++VD IL +Y + + + IGLLC Q RP M+
Sbjct: 554 SMGDLISFAWRNWKEGVALNLVDKILMTMSSYSSNMIMRCINIGLLCVQEKVAERPSMAS 613
Query: 596 VVKMINNIHKIA--DPTQPPFLNFGSGEFSRSS 626
VV M++ H IA +P++P F + + SS
Sbjct: 614 VVLMLDG-HTIALSEPSKPAFFSHSNAVSDSSS 645
Score = 38.1 bits (87), Expect = 0.83
Identities = 20/92 (21%), Positives = 40/92 (42%), Gaps = 3/92 (3%)
Query: 52 LSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKTR 111
L + L++ S++ + VV +S T++ +C D+S+ DC C +Q +
Sbjct: 158 LRGLLEGLKNRASVIGRSKKNFVVGETSGPSFQTLFGLVQCTPDISEEDCSYCLSQGIAK 217
Query: 112 VLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFN 143
+ C ++ G C L Y + F++
Sbjct: 218 IPSCCDMKM---GSYVMSPSCMLAYAPWRFYD 246
Score = 37.0 bits (84), Expect = 1.8
Identities = 29/111 (26%), Positives = 52/111 (46%), Gaps = 7/111 (6%)
Query: 153 LCGNIDFSGNRSV---HKANVVELVRNLSVEAPKNDGFFVGVLSKRNVSVYGLAQCWNFV 209
+C N+ +GN +V + N+ L+ +LS +GF+ + + V ++QC V
Sbjct: 30 ICSNV--TGNFTVNTPYAVNLDRLISSLSSLRRNVNGFYNISVGDSDEKVNSISQCRGDV 87
Query: 210 NESVCQNCLVEAVTR-IDSCALKEEGRVLNAGCYLRFSTDKFYDNSSNIAP 259
VC NC+ A R + C +++E + C R+S + N I+P
Sbjct: 88 KLEVCINCIAMAGKRLVTLCPVQKEAIIWYDKCTFRYSNRTIF-NRLEISP 137
>UniRef100_Q9XED4 Receptor-like protein kinase homolog RK20-1 [Phaseolus vulgaris]
Length = 666
Score = 315 bits (806), Expect = 4e-84
Identities = 217/631 (34%), Positives = 325/631 (51%), Gaps = 36/631 (5%)
Query: 53 SNFYDALESLTSLVT--TQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKT 110
S +++ L +L S ++ TQ + S N V A G C D+ +C C
Sbjct: 39 STYHNNLNTLLSTLSSHTQINYGFYNFSHGQNNDKVNAIGLCRGDVKPDECRRCLNDSAL 98
Query: 111 RVLRCSPIQRGVEGGRFFFDG-CYLRYDAYNFFNQSLSSQDLTLCGNIDFSGNRSVHKAN 169
+ + P Q+ E + C LRY F SS L N++
Sbjct: 99 TITQLCPNQK--EALLWLNTSKCLLRYSHRTIFGVMESSPGFYLT-NVNNVTEADKFNQA 155
Query: 170 VVELVRNLSVEAPKNDGFFV----GVLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRI 225
+ L+RN +V A D ++ +VYGL QC ++E+ C CL A++ I
Sbjct: 156 LSNLMRNPTVVAASGDSRLKYAADSAIAANFQTVYGLVQCTPDLSETDCNRCLDGAISEI 215
Query: 226 DSCA-LKEEGRVLNAGCYLRFSTDKFYDNSS----NIAPPGTKGSTKVGIIVAES----- 275
SC K GRVL C +RF + FYD+++ ++ PP S+ ES
Sbjct: 216 PSCCGNKMGGRVLRPSCNIRFESAIFYDSNAKLDPDVTPPSPPPSSFTNTSPKESNNTIT 275
Query: 276 --FAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNS-----KLNMPYEVLEKA 328
A V ++++VA + LR+++ R+ +++ L ++ + A
Sbjct: 276 IVIAVVVSIVVVAVVSLLGLCIYLRRKKARKSPTVNQDDDDDDIEISQSLQFDFDTIRVA 335
Query: 329 TNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNL 388
T F +SNKLG+GG G+VY+G LP+G +A+KRLS ++Q F NEV L+ LQH+NL
Sbjct: 336 TEDFSNSNKLGQGGFGAVYRGRLPNGQMIAVKRLSSGSSQGDTEFKNEVLLMAKLQHRNL 395
Query: 389 VKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEE 448
V+LLG + G E LL+YE+VPN SL + QL WE+R+KII G A GL YLHE+
Sbjct: 396 VRLLGFCLEGRERLLIYEFVPNKSLDYFIFDPVKKAQLDWEMRYKIIRGIARGLLYLHED 455
Query: 449 SQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTLGYMAPEYIVL 507
S L+IIHRD+K SNILLD+ PKIADFG+ARL D++ +T + GT GYMAPEYI+
Sbjct: 456 SLLRIIHRDLKASNILLDEEMNPKIADFGMARLVLLDETHANTNRVVGTYGYMAPEYIMQ 515
Query: 508 GKLTEKADIYSFGVLVIELLSGKSRTSF--VQNSCSILHMVWSLYGSNRLCDIVDPILEG 565
G+ + K+DI+SFGVL++E++SG+ + F +N +L W + +IVDP LE
Sbjct: 516 GQFSVKSDIFSFGVLLLEIVSGQKNSGFRHGENVEDLLSFTWRNWRDGTAVNIVDPSLEN 575
Query: 566 NYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIH-KIADPTQPPFLNFGSGEFSR 624
N E + + IGLLC Q + RP M+ ++ M+++ + P++P F +
Sbjct: 576 N-SRNEVMRCIHIGLLCVQENLTDRPTMATIMLMLSSYSLGLPIPSEPAFY----ANSTA 630
Query: 625 SSLPEESLQPGSNTQSSGDSMTESSEHRLVT 655
SLP S S+ ++ S ES +T
Sbjct: 631 RSLPATSSWGHSSRATANQSAQESENENSIT 661
>UniRef100_Q8H785 Hypothetical protein [Arabidopsis thaliana]
Length = 658
Score = 312 bits (800), Expect = 2e-83
Identities = 210/607 (34%), Positives = 313/607 (50%), Gaps = 54/607 (8%)
Query: 78 SSTTQNATV-------YAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFD 130
S+ QNATV C D+S C C + L P Q+ ++D
Sbjct: 64 STGFQNATVGQAPDRVTGLFNCRGDVSTEVCRRCVSFAVNDTLTRCPNQKEAT---LYYD 120
Query: 131 GCYLRYDAYNFFNQSLSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVG 190
C LRY N + +++ + L + + N+ +++V N + N G
Sbjct: 121 ECVLRYSNQNILSTLITTGGVILVNTRNVTSNQLDLLSDLVLPTLNQAATVALNSSKKFG 180
Query: 191 VLSKRNV----SVYGLAQCWNFVNESVCQNCLVEAVTRIDSCALKEEGRVLNAGCYLRFS 246
K N S YGL QC + C CL + +I + + R++N C R+
Sbjct: 181 T-RKNNFTALQSFYGLVQCTPDLTRQDCSRCLQLVINQIPTDRIG--ARIINPSCTSRYE 237
Query: 247 TDKFYDNSSNIAPP----------------GTKGSTKVGIIVAESFAAVATLLIVATIIF 290
FY S+ PP G G++KV +I A+ +IVA ++F
Sbjct: 238 IYAFYTESAVPPPPPPPSICTPPVSAPPRSGKDGNSKVLVI------AIVVPIIVAVLLF 291
Query: 291 FVRKNVLRQRRERRQFGAFLFSENN----SKLNMPYEVLEKATNYFHDSNKLGEGGSGSV 346
L +R + + F+ ++ L + Y ++ AT+ F +SNK+G+GG G V
Sbjct: 292 IAGYCFLTRRARKSYYTPSAFAGDDITTADSLQLDYRTIQTATDDFVESNKIGQGGFGEV 351
Query: 347 YKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYE 406
YKG L DGT VA+KRLS ++ Q F NEV L+ LQH+NLV+LLG + G E +LVYE
Sbjct: 352 YKGTLSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYE 411
Query: 407 YVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLD 466
YVPN SL L QL W R+KII G A G+ YLH++S+L IIHRD+K SNILLD
Sbjct: 412 YVPNKSLDYFLFDPAKKGQLDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASNILLD 471
Query: 467 DNFTPKIADFGLARLFPEDQSQISTA-ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIE 525
+ PKIADFG+AR+F DQ++ +T+ I GT GYM+PEY + G+ + K+D+YSFGVLV+E
Sbjct: 472 ADMNPKIADFGMARIFGLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLE 531
Query: 526 LLSGKSRTSFVQ--NSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCT 583
++SGK +SF Q + ++ W L+ + R ++VDP + N E + + IGLLC
Sbjct: 532 IISGKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCV 591
Query: 584 QASAELRPPMSVVVKMI-NNIHKIADPTQPPFLNFGSGEFSRSSLPEESLQPGSNTQSSG 642
Q RP +S +V M+ +N + P QP G F +S + ++ L + ++S
Sbjct: 592 QEDPAERPTLSTIVLMLTSNTVTLPVPRQP-------GLFFQSRIGKDPLDTDTTSKSLL 644
Query: 643 DSMTESS 649
S+ ++S
Sbjct: 645 GSVDDAS 651
>UniRef100_Q8GYA4 Putative receptor-like protein kinase 4 RLK4 [Arabidopsis thaliana]
Length = 669
Score = 312 bits (800), Expect = 2e-83
Identities = 210/607 (34%), Positives = 313/607 (50%), Gaps = 54/607 (8%)
Query: 78 SSTTQNATV-------YAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFD 130
S+ QNATV C D+S C C + L P Q+ ++D
Sbjct: 75 STGFQNATVGQAPDRVTGLFNCRGDVSTEVCRRCVSFAVNDTLTRCPNQKEAT---LYYD 131
Query: 131 GCYLRYDAYNFFNQSLSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVG 190
C LRY N + +++ + L + + N+ +++V N + N G
Sbjct: 132 ECVLRYSNQNILSTLITTGGVILVNTRNVTSNQLDLLSDLVLPTLNQAATVALNSSKKFG 191
Query: 191 VLSKRNV----SVYGLAQCWNFVNESVCQNCLVEAVTRIDSCALKEEGRVLNAGCYLRFS 246
K N S YGL QC + C CL + +I + + R++N C R+
Sbjct: 192 T-RKNNFTALQSFYGLVQCTPDLTRQDCSRCLQLVINQIPTDRIG--ARIINPSCTSRYE 248
Query: 247 TDKFYDNSSNIAPP----------------GTKGSTKVGIIVAESFAAVATLLIVATIIF 290
FY S+ PP G G++KV +I A+ +IVA ++F
Sbjct: 249 IYAFYTESAVPPPPPPPSISTPPVSAPPRSGKDGNSKVLVI------AIVVPIIVAVLLF 302
Query: 291 FVRKNVLRQRRERRQFGAFLFSENN----SKLNMPYEVLEKATNYFHDSNKLGEGGSGSV 346
L +R + + F+ ++ L + Y ++ AT+ F +SNK+G+GG G V
Sbjct: 303 IAGYCFLTRRARKSYYTPSAFAGDDITTADSLQLDYRTIQTATDDFVESNKIGQGGFGEV 362
Query: 347 YKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYE 406
YKG L DGT VA+KRLS ++ Q F NEV L+ LQH+NLV+LLG + G E +LVYE
Sbjct: 363 YKGTLSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYE 422
Query: 407 YVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLD 466
YVPN SL L QL W R+KII G A G+ YLH++S+L IIHRD+K SNILLD
Sbjct: 423 YVPNKSLDYFLFDPAKKGQLDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASNILLD 482
Query: 467 DNFTPKIADFGLARLFPEDQSQISTA-ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIE 525
+ PKIADFG+AR+F DQ++ +T+ I GT GYM+PEY + G+ + K+D+YSFGVLV+E
Sbjct: 483 ADMNPKIADFGMARIFGLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLE 542
Query: 526 LLSGKSRTSFVQ--NSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCT 583
++SGK +SF Q + ++ W L+ + R ++VDP + N E + + IGLLC
Sbjct: 543 IISGKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCV 602
Query: 584 QASAELRPPMSVVVKMI-NNIHKIADPTQPPFLNFGSGEFSRSSLPEESLQPGSNTQSSG 642
Q RP +S +V M+ +N + P QP G F +S + ++ L + ++S
Sbjct: 603 QEDPAERPTLSTIVLMLTSNTVTLPVPRQP-------GLFFQSRIGKDPLDTDTTSKSLL 655
Query: 643 DSMTESS 649
S+ ++S
Sbjct: 656 GSVDDAS 662
>UniRef100_Q9M0X5 Receptor protein kinase-like protein [Arabidopsis thaliana]
Length = 675
Score = 309 bits (792), Expect = 1e-82
Identities = 206/598 (34%), Positives = 302/598 (50%), Gaps = 40/598 (6%)
Query: 35 LCTNRTSPMPQSQRQVFLSNFYDALESLTS--LVTTQRHGTVVKGSSTTQNATVYAFGEC 92
+C N T+ S+ +L+N L SL+S G N VY C
Sbjct: 33 ICPNTTT---YSRNSSYLTNLRTVLSSLSSPNAAYASLFDNAAAGEENDSNR-VYGVFLC 88
Query: 93 MKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQDLT 152
D+S C C A L+ P ++ ++D C +RY + Q +
Sbjct: 89 RGDVSAEICRDCVAFAANETLQRCPREKVAV---IWYDECMVRYSNQSIVGQMRIRPGVF 145
Query: 153 LCGNIDFSGNR-SVHKANVVELVRNLSVEAPKNDGFFVGVLSKRNV--SVYGLAQCWNFV 209
L + + N+ S ++ L+ +++V+A + F + V ++Y L QC +
Sbjct: 146 LTNKQNITENQVSRFNESLPALLIDVAVKAALSSRKFATEKANFTVFQTIYSLVQCTPDL 205
Query: 210 NESVCQNCLVEAVTRIDSCALKEEG-RVLNAGCYLRFSTDKFYDNSSNIAP--------- 259
C++CL + + + C + G RV+ C R+ FY+ + AP
Sbjct: 206 TNQDCESCLRQVINYLPRCCDRSVGGRVIAPSCSFRYELYPFYNETIAAAPMAPPPSSTV 265
Query: 260 -------PGTKGSTKVGIIVAESFA---AVATLLIVATIIFFVRK--NVLRQRRERRQFG 307
P KG K ++ + A +V LL+ A R+ N L E
Sbjct: 266 TAPPLNIPSEKGKGKNLTVIVTAIAVPVSVCVLLLGAMCWLLARRRNNKLSAETEDLDED 325
Query: 308 AFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTT 367
+E L + +E ATN F +SNKLG GG G VYKG L G TVAIKRLS +T
Sbjct: 326 GITSTET---LQFQFSAIEAATNKFSESNKLGHGGFGEVYKGQLITGETVAIKRLSQGST 382
Query: 368 QWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLT 427
Q A+ F NEV+++ LQH+NL KLLG + G E +LVYE+VPN SL L + L
Sbjct: 383 QGAEEFKNEVDVVAKLQHRNLAKLLGYCLDGEEKILVYEFVPNKSLDYFLFDNEKRRVLD 442
Query: 428 WEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQS 487
W+ R+KII G A G+ YLH +S+L IIHRD+K SNILLD + PKI+DFG+AR+F DQ+
Sbjct: 443 WQRRYKIIEGIARGILYLHRDSRLTIIHRDLKASNILLDADMHPKISDFGMARIFGVDQT 502
Query: 488 QIST-AICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQNS--CSILH 544
Q +T I GT GYM+PEY + GK + K+D+YSFGVLV+EL++GK +SF + ++
Sbjct: 503 QANTKRIVGTYGYMSPEYAIHGKYSVKSDVYSFGVLVLELITGKKNSSFYEEDGLGDLVT 562
Query: 545 MVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINN 602
VW L+ N ++VD + GN+ E + + I LLC Q + RP M ++ M+N+
Sbjct: 563 YVWKLWVENSPLELVDEAMRGNFQTNEVIRCIHIALLCVQEDSSERPSMDDILVMMNS 620
>UniRef100_Q8L7G3 Putative serine/threonine kinase [Arabidopsis thaliana]
Length = 659
Score = 308 bits (790), Expect = 3e-82
Identities = 224/681 (32%), Positives = 332/681 (47%), Gaps = 76/681 (11%)
Query: 3 FPFPWFLLLIFLCHHHHPLPTLADPRATEAASLCTNRTSPMPQSQRQVFLSNFYDALESL 62
FPF + L FL + DPR A C N T+ S +L+N L SL
Sbjct: 5 FPFIFLFLFSFLTSFR---ASAQDPRFL--AYYCPNATT---YSSNSTYLTNLKTLLSSL 56
Query: 63 TSLVTTQRHGTVVKGSSTTQNATVYAFGE-------CMKDLSKSDCDVCFAQCKTRVLRC 115
+S + G QNATV + C D+S C C
Sbjct: 57 SSRNASYSTGF--------QNATVGQALDRVTGLFLCRGDVSPEVCRNCVTFAVNNTFSR 108
Query: 116 SPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQDLTLCGN---IDFSGNRSVHKANVV- 171
P QR F+++ C LRY N + +++++ + N I N+ N+V
Sbjct: 109 CPNQREAV---FYYEECILRYSHKNILSTAITNEGEFILRNPNHISPIQNQINQFTNLVL 165
Query: 172 ELVRNLSVEAPKNDGFFVGVLSKRNV--SVYGLAQCWNFVNESVCQNCLVEAVTRIDSCA 229
+ +++EA N F + ++ + YGL QC ++ C NCL ++ R+
Sbjct: 166 SNMNQIAIEAADNPRKFSTIKTELTALQTFYGLVQCTPDLSRQNCMNCLTSSINRMPFSR 225
Query: 230 LKEEGRVLNAGCYLRFSTDKFYDNSSNIAPP------------GTKGSTKVGIIVAESFA 277
+ R C R+ FY+ ++ PP G++ V ++
Sbjct: 226 IG--ARQFWPSCNSRYELYDFYNETAIGTPPPPLPPLASPSLSDKSGNSNVVVVAVVVPI 283
Query: 278 AVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSK-----LNMPYEVLEKATNYF 332
VA L+ +A FF + R ++ +G + + K L + Y ++ ATN F
Sbjct: 284 IVAVLIFIAGYCFFAK-------RAKKTYGTTPALDEDDKTTIESLQLDYRAIQAATNDF 336
Query: 333 HDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLL 392
++NK+G GG G VYKG +GT VA+KRLS + Q F NEV ++ +L+HKNLV++L
Sbjct: 337 SENNKIGRGGFGDVYKGTFSNGTEVAVKRLSKTSEQGDTEFKNEVVVVANLRHKNLVRIL 396
Query: 393 GCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLK 452
G SI E +LVYEYV N SL + L QL W R+ II G A G+ YLH++S+L
Sbjct: 397 GFSIEREERILVYEYVENKSLDNFLFDPAKKGQLYWTQRYHIIGGIARGILYLHQDSRLT 456
Query: 453 IIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQISTA-ICGTLGYMAPEYIVLGKLT 511
IIHRD+K SNILLD + PKIADFG+AR+F DQ+Q +T+ I GT GYM+PEY + G+ +
Sbjct: 457 IIHRDLKASNILLDADMNPKIADFGMARIFGMDQTQQNTSRIVGTYGYMSPEYAMRGQFS 516
Query: 512 EKADIYSFGVLVIELLSGKSRTSFVQ--NSCSILHMVWSLYGSNRLCDIVDPILEGNYPA 569
K+D+YSFGVLV+E++SG+ SF++ ++ ++ W L+ + D+VDP + +
Sbjct: 517 MKSDVYSFGVLVLEIISGRKNNSFIETDDAQDLVTHAWRLWRNGTALDLVDPFIADSCRK 576
Query: 570 EEACKLLKIGLLCTQASAELRPPMSVV-VKMINNIHKIADPTQPPFLNFGSGEFSRSSLP 628
E + IGLLC Q RP MS + V + +N + P QP F F RS
Sbjct: 577 SEVVRCTHIGLLCVQEDPVKRPAMSTISVMLTSNTMALPAPQQPGF-------FVRS--- 626
Query: 629 EESLQPGSNTQSSGDSMTESS 649
+PG+N S S T S
Sbjct: 627 ----RPGTNRLDSDQSTTNKS 643
>UniRef100_Q9LDS6 Serine/threonine kinase-like protein [Arabidopsis thaliana]
Length = 656
Score = 305 bits (782), Expect = 2e-81
Identities = 211/639 (33%), Positives = 329/639 (51%), Gaps = 56/639 (8%)
Query: 46 SQRQVFLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCF 105
+ R + LSN L S S +G+V +G +YA G C+ CD C
Sbjct: 40 TNRHLILSN----LASNVSSRDGYYNGSVGEGPDR-----IYALGLCIPGTDPKVCDDCM 90
Query: 106 AQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQDLTLCGNID---FSGN 162
T +L+ P Q R C++RY +FFN+ + + + G+++ F G+
Sbjct: 91 QIASTGILQNCPNQTDSYDWRSQKTLCFVRYSNSSFFNK-MDLEPTMVIGDLNSGLFQGD 149
Query: 163 RSVHKANVVELVRNLSVEAPKNDGFFVGVLSKR--NVSVYGLAQCWNFVNESVCQNCLVE 220
+ + E + ++ + ++ +S R + +Y L QC ++ C+ C+ +
Sbjct: 150 LAAYTRTWEEFMNSMITRVGRTR--YLADISPRIGSARIYALMQCIRGISSMECETCIRD 207
Query: 221 AVTRIDSCALKEEGRVLNAG-CYLRFSTDKF---YDNSSNIAPPGTKGST-KVGIIVAES 275
V SC G + C+ R+ ++ + ++ ++ PP G T G IVA
Sbjct: 208 NVRMYQSCCNGFIGGTIRKPVCFFRWDGSEYLGAFGDTPSLPPPSPDGKTISTGAIVA-- 265
Query: 276 FAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMP------YEVLEKAT 329
++V+ +IF V ++ R+RRQ L + + + P + LE AT
Sbjct: 266 -------VVVSVVIFVVLLALVLVIRKRRQSYKTLKPKTDDDMTSPQSLQFDFMTLEAAT 318
Query: 330 NYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLV 389
+ F +NKLG+GG G VYKG LP+ T VA+KRLS N+ Q F NEV ++ LQHKNLV
Sbjct: 319 DKFSRNNKLGKGGFGEVYKGMLPNETEVAVKRLSSNSGQGTQEFKNEVVIVAKLQHKNLV 378
Query: 390 KLLGCSITGPESLLVYEYVPNLSLH--------DHLSVRRNLQQLTWEVRHKIILGTAEG 441
+LLG + E +LVYE+VPN SL+ HL QL W+ R+ II G G
Sbjct: 379 RLLGFCLERDEQILVYEFVPNKSLNYFLFGNKQKHLLDPTKKSQLDWKRRYNIIGGITRG 438
Query: 442 LAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTLGYM 500
L YLH++S+L IIHRDIK SNILLD + PKIADFG+AR F DQ++ +T + GT GYM
Sbjct: 439 LLYLHQDSRLTIIHRDIKASNILLDADMNPKIADFGMARNFRVDQTEDNTRRVVGTFGYM 498
Query: 501 APEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQ---NSCSILHMVWSLYGSNRLCD 557
PEY+ G+ + K+D+YSFGVL++E++ GK +SF + + +++ VW L+ ++ D
Sbjct: 499 PPEYVTHGQFSTKSDVYSFGVLILEIVCGKKNSSFYKIDDSGGNLVTHVWRLWNNDSPLD 558
Query: 558 IVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNIHKIADPTQPPFLNF 617
++DP +E + ++ + + IGLLC Q + RP MS + +M+ N +PP
Sbjct: 559 LIDPAIEESCDNDKVIRCIHIGLLCVQETPVDRPEMSTIFQMLTNSSITLPVPRPP---- 614
Query: 618 GSGEFSRSSLPEESLQPGSNT-QSSGDSMTESSEHRLVT 655
G F R+ + L GS QSS S+ + + +T
Sbjct: 615 --GFFFRNRSNLDPLTYGSELGQSSSKSIPYTIDSASIT 651
>UniRef100_Q8S9L6 AT4g21410/T6K22_140 [Arabidopsis thaliana]
Length = 679
Score = 304 bits (779), Expect = 5e-81
Identities = 207/629 (32%), Positives = 304/629 (47%), Gaps = 49/629 (7%)
Query: 51 FLSNFYDALESLTSLVTTQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKT 110
F N + SL+SL +Q +G S + YA G C +++ + DC C
Sbjct: 48 FAGNLNRLVSSLSSL-KSQAYGFYNLSSGDSSGERAYAIGLCRREVKRDDCVSCIQTAAR 106
Query: 111 RVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFFNQSLSSQDLTLCGNIDFSGNRSVHKA-- 168
+ + P+ + ++ C RY + + ++ + S NR +
Sbjct: 107 NLTKQCPLTKQAV---VWYTHCMFRYSNRTIYGRKETNPTKAFIAGEEISANRDDFERLQ 163
Query: 169 -NVVELVRNLSVEAPKNDGFFVG--VLSKRNVSVYGLAQCWNFVNESVCQNCLVEAVTRI 225
+++ ++ ++ N + G S YG QC ++E C +CLV I
Sbjct: 164 RGLLDRLKGIAAAGGPNRKYAQGNGSASAGYRRFYGTVQCTPDLSEQDCNDCLVFGFENI 223
Query: 226 DSCALKEEG-RVLNAGCYLRFSTDKFYDNSSNIAPPGT--------------------KG 264
SC E G R + C RF T +FY+ +++ P KG
Sbjct: 224 PSCCDAEIGLRWFSPSCNFRFETWRFYEFDADLEPDPPAIQPADSPQSAARTERTGKGKG 283
Query: 265 STKVGIIVAESFAAVATLLIVATIIFFVRKNVLRQRRERRQFGAFLFS--------ENNS 316
+KV I + VA L I ++ RKN + + + G S N
Sbjct: 284 GSKVIIAIVIPILLVALLAICLCLVLKWRKN--KSGYKNKVLGKSPLSGSIAEDEFSNTE 341
Query: 317 KLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNE 376
L + +E L+ AT+ F N+LG GG GSVYKG P G +A+KRLS N+ Q + F NE
Sbjct: 342 SLLVHFETLKTATDNFSSENELGRGGFGSVYKGVFPQGQEIAVKRLSGNSGQGDNEFKNE 401
Query: 377 VNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIIL 436
+ L+ LQH+NLV+L+G I G E LLVYE++ N SL + Q L W VR+K+I
Sbjct: 402 ILLLAKLQHRNLVRLIGFCIQGEERLLVYEFIKNASLDQFIFDTEKRQLLDWVVRYKMIG 461
Query: 437 GTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIADFGLARLFPEDQS---QISTAI 493
G A GL YLHE+S+ +IIHRD+K SNILLD PKIADFGLA+LF Q+ + ++ I
Sbjct: 462 GIARGLLYLHEDSRFRIIHRDLKASNILLDQEMNPKIADFGLAKLFDSGQTMTHRFTSRI 521
Query: 494 CGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRTSFVQN----SCSILHMVWSL 549
GT GYMAPEY + G+ + K D++SFGVLVIE+++GK + N + +L VW
Sbjct: 522 AGTYGYMAPEYAMHGQFSVKTDVFSFGVLVIEIITGKRNNNGGSNGDEDAEDLLSWVWRS 581
Query: 550 YGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELRPPMSVVVKMINNI-HKIAD 608
+ + + ++DP L E + + IGLLC Q SA RP M+ V M+N+ +
Sbjct: 582 WREDTILSVIDPSLTAG-SRNEILRCIHIGLLCVQESAATRPTMATVSLMLNSYSFTLPT 640
Query: 609 PTQPPFLNFGSGEFSRSSLPEESLQPGSN 637
P +P F+ S S E LQ SN
Sbjct: 641 PLRPAFVLESVVIPSNVSSSTEGLQMSSN 669
>UniRef100_Q9C5T0 Receptor-like protein kinase 4 [Arabidopsis thaliana]
Length = 658
Score = 303 bits (777), Expect = 8e-81
Identities = 209/605 (34%), Positives = 307/605 (50%), Gaps = 50/605 (8%)
Query: 78 SSTTQNATV-------YAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFD 130
S+ QNATV C D+S C C + L P Q+ ++D
Sbjct: 64 STGFQNATVGQAPDRVTGLFNCRGDVSTEVCRRCVSFAVNDTLTRCPNQKEAT---LYYD 120
Query: 131 GCYLRYDAYNFFNQSLSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVG 190
C LRY N + +++ + L + + N+ +++V N + N G
Sbjct: 121 ECVLRYSNQNILSTLITTGGVILVNTRNVTSNQLDLLSDLVLPTLNQAATVALNSSKKFG 180
Query: 191 VLSKRNV----SVYGLAQCWNFVNESVCQNCLVEAVTRIDSCALKEEGRVLNAGCYLRFS 246
K N S YGL QC + C CL + +I + + R++N C R+
Sbjct: 181 T-RKNNFTALQSFYGLVQCTPDLTRQDCSRCLQLVINQIPTDRIG--ARIINPSCTSRYE 237
Query: 247 TDKFYDNSSNIAPPGT----------------KGSTKVGIIVAESFAAVATLLIVATIIF 290
FY S+ PP +G++KV +I VA L +A F
Sbjct: 238 IYAFYTESAVPPPPPPPSISTPPVSAPPRSEKEGNSKVLVIAIVVPIIVAVRLFIAGYCF 297
Query: 291 FVRKNVLRQRRERRQFGAFLFSENNS--KLNMPYEVLEKATNYFHDSNKLGEGGSGSVYK 348
R R R+ AF + + L + Y ++ AT+ F +SNK+G+GG G VYK
Sbjct: 298 LTR----RARKSYSTPSAFAGDDITTADSLQLDYRTIQTATDDFVESNKIGQGGFGEVYK 353
Query: 349 GALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYV 408
G L DGT VA+KRLS ++ Q F NEV L+ LQH+NLV+LLG + G E +LVYEYV
Sbjct: 354 GTLSDGTEVAVKRLSKSSGQGEVEFKNEVVLVAKLQHRNLVRLLGFCLDGEERVLVYEYV 413
Query: 409 PNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDN 468
PN SL L Q W R+KII G A G+ YLH++S+L IIHRD+K S ILLD +
Sbjct: 414 PNKSLDYFLFDPAKKGQXDWTRRYKIIGGVARGILYLHQDSRLTIIHRDLKASTILLDAD 473
Query: 469 FTPKIADFGLARLFPEDQSQISTA-ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELL 527
PKIADFG+AR+F DQ++ +T+ I GT GYM+PEY + G+ + K+D+YSFGVLV+E++
Sbjct: 474 MNPKIADFGMARIFGLDQTEENTSRIVGTYGYMSPEYAMHGQYSMKSDVYSFGVLVLEII 533
Query: 528 SGKSRTSFVQ--NSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQA 585
SGK +SF Q + ++ W L+ + R ++VDP + N E + + IGLLC Q
Sbjct: 534 SGKKNSSFYQTDGAHDLVSYAWGLWSNGRPLELVDPAIVENCQRNEVVRCVHIGLLCVQE 593
Query: 586 SAELRPPMSVVVKMI-NNIHKIADPTQPPFLNFGSGEFSRSSLPEESLQPGSNTQSSGDS 644
RP +S +V M+ +N + P QP G F +S + ++ L + ++S S
Sbjct: 594 DPAERPTLSTIVLMLTSNTVTLPVPRQP-------GLFFQSRIGKDPLDTDTTSKSLLGS 646
Query: 645 MTESS 649
+ ++S
Sbjct: 647 VDDAS 651
>UniRef100_Q7F1L6 Putative serine/threonine kinase receptor [Oryza sativa]
Length = 670
Score = 299 bits (766), Expect = 2e-79
Identities = 204/567 (35%), Positives = 291/567 (50%), Gaps = 43/567 (7%)
Query: 86 VYAFGECMKDLSKSD-CDVCFAQCKTRVLRCSPIQRGVEGGRFFFDGCYLRYDAYNFF-N 143
VYA C D + + C C A + P + F+D C LR+ NF +
Sbjct: 85 VYALALCRGDTANATACAGCVAAAFQDAQQLCPYNKDAT---VFYDACALRFSNQNFLAS 141
Query: 144 QSLSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDG--FFVGVLSKRNVSVYG 201
+ ++ L L + S V A V L+ + A N F G +YG
Sbjct: 142 TNGDNKFLILMNTQNVSAPAKVFDAAVGVLINATADYAAANSSRRFGTGEEGFNGSKIYG 201
Query: 202 LAQCWNFVNESVCQNCLVEAVTRIDSC-ALKEEGRVLNAGCYLRFSTDKFYDNSS----- 255
LAQC + + C++CL V + + K+ GRVL C R+ F+D S
Sbjct: 202 LAQCTPDMATATCRSCLGGIVGMMPKYFSGKQGGRVLGLRCNYRYEIYPFFDGVSLLQLP 261
Query: 256 ---------------NIAPPGTKGSTK-----VGIIVAESFAAVATLLIVATIIFFVRKN 295
N+ PP T T+ G ++A + VA +L I F++ K
Sbjct: 262 AASLGAPPAPSPAAVNVTPPATTTGTRRRGNTTGRVLAIALPIVAAILAAVVICFYIWKR 321
Query: 296 VLRQRRERRQFGAFLFS----ENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGAL 351
+ R R+ A E+ L + L ATN F DSNKLGEGG G+VYKG L
Sbjct: 322 --KTERARKPSIADPTDPADIESIDSLILSISTLRVATNNFDDSNKLGEGGFGAVYKGVL 379
Query: 352 PDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNL 411
P +A+KRLS ++ Q + NE+ L+ LQHKNLV+LLG + E LLVYEY+PN
Sbjct: 380 PSDQEIAVKRLSQSSRQGIEELKNELVLVAKLQHKNLVRLLGVCLEEHEKLLVYEYMPNK 439
Query: 412 SLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTP 471
SL L L W R KI+ A GL YLHE+SQLKIIHRD+K SN+LLD +F P
Sbjct: 440 SLDTILFDPDRSNVLDWWKRLKIVNAIARGLQYLHEDSQLKIIHRDLKASNVLLDSDFNP 499
Query: 472 KIADFGLARLFPEDQSQ-ISTAICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSG- 529
KI+DFGLARLF DQSQ ++ + GT GYMAPEY + G + K+D++SFGVL++E+++G
Sbjct: 500 KISDFGLARLFGNDQSQDVTNRVVGTYGYMAPEYAMRGHYSIKSDVFSFGVLILEIVTGR 559
Query: 530 KSRTSF-VQNSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAE 588
K+ S+ + S +L +VW + + + ++ D + G+ P ++ K + IGLLC Q
Sbjct: 560 KNNVSYDSEQSVDLLTLVWEHWLAGTVVELADSSMAGHCPGDQILKCVHIGLLCVQEDPT 619
Query: 589 LRPPMSVV-VKMINNIHKIADPTQPPF 614
RP MS+V V + ++ + P++P F
Sbjct: 620 ERPMMSMVNVMLSSSTVSLQAPSRPAF 646
>UniRef100_Q8VX52 Putative receptor-like serine-threonine protein kinase [Solanum
tuberosum]
Length = 676
Score = 299 bits (765), Expect = 2e-79
Identities = 217/679 (31%), Positives = 332/679 (47%), Gaps = 52/679 (7%)
Query: 8 FLLLIFLCHHHHPLPTLADPRATEAASLCTNRTSPMPQSQRQVFLSNFYDALESLTSLVT 67
+LL++FL H H L +A S N ++ + +N L SL+S +
Sbjct: 6 WLLILFL--HIHVLNIVAQLPDLRFGSCGGNGN----YTENSTYKNNLNTLLTSLSSKID 59
Query: 68 TQRHGTVVKGSSTTQNATVYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRF 127
G ++ + + + C D+ +DC C ++ + P Q+ V GG
Sbjct: 60 NYGFYNASIGQNSDRASVIVL---CRGDVELADCRGCVDNVVQKIAQLCPNQKEVFGG-- 114
Query: 128 FFDGCYLRYDAYNFFNQSLSSQDLTLCG--NIDFSGNRSVHKANVVELVRNLSVEAPKND 185
+DGC L+Y + + S N G + ++E +R+ +V+
Sbjct: 115 -YDGCMLQYSNQSILETTSFSLKYYFWNPANATKPGEFNQELGRLLENLRDRAVDDGPLQ 173
Query: 186 GFFVGVLSKRNV-SVYGLAQCWNFVNESVCQNCLVEAVTRIDSCAL--KEEGRVLNAGCY 242
+ G + + ++Y L QC ++ C +CL +A + C K GR++ C
Sbjct: 174 KYASGNATGPDFQAIYALVQCTPDLSRQSCFSCLSDAYGNMPRCPCLGKRGGRIIGVRCN 233
Query: 243 LRFSTDKFYD-----------NSSNIAPPGTKGST-----------KVGIIVAESFAAVA 280
R+ + +F++ N + P GT T + IIV + V
Sbjct: 234 FRYESSRFFEDVPLEAPPPAGNDNTTVPTGTDDKTVPTGEDDKTTRTIIIIVVSTVTVVI 293
Query: 281 TLLIVATIIFFVRKNVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGE 340
++ +A I+ RK L + +E+ + + AT+ F D+NKLGE
Sbjct: 294 LIICIAVILIRRRKRKLVNEIQSTSVDDTSIAES---FQYDFSAIRAATDDFSDANKLGE 350
Query: 341 GGSGSVYKGALPDGTTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPE 400
GG G VYKG L +G VA+KRLS ++ Q F NEV L+ LQH+NLV+LLG + G E
Sbjct: 351 GGFGPVYKGKLQNGQEVAVKRLSADSGQGDLEFKNEVLLVARLQHRNLVRLLGFCLDGTE 410
Query: 401 SLLVYEYVPNLSLHDHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKL 460
LLVYE+VPN SL L +QL WE R KII G A+G+ YLHE+S+L+IIHRD+K
Sbjct: 411 RLLVYEFVPNASLDHFLFDSVKRRQLDWERRSKIIGGIAKGILYLHEDSRLRIIHRDLKA 470
Query: 461 SNILLDDNFTPKIADFGLARLFPEDQSQIST-AICGTLGYMAPEYIVLGKLTEKADIYSF 519
SN+LLD PKI+DFG+ARLF D++Q ST I GT GYMAPEY + G+ + K+D++SF
Sbjct: 471 SNVLLDAEMNPKISDFGMARLFELDETQGSTNRIVGTYGYMAPEYAMHGQFSVKSDVFSF 530
Query: 520 GVLVIELLSGKSRTSFVQNSC--SILHMVWSLYGSNRLCDIVDPIL-EGNYPAEEACKLL 576
GVLV+E+LSG+ T F +L WS + + + VDP+L E + + +
Sbjct: 531 GVLVLEILSGQKNTCFRNGESVEDLLSFAWSSWRNGTTINFVDPMLKESTGLIRDIMRNI 590
Query: 577 KIGLLCTQASAELRPPMSVVVKMINNIH-KIADPTQPPFLNFGSGEFSRSSLPEESLQ-- 633
I LLC Q S RP M+ VV M+++ + P+ P F + S + E + +
Sbjct: 591 HIALLCVQESVADRPTMAAVVLMLSSFSLSLPMPSGPAFYMHSNITAGTSLIQEYNTRVT 650
Query: 634 ---PGSNTQSSGDSMTESS 649
+ ++S G S E+S
Sbjct: 651 DSSERAKSKSIGSSRNEAS 669
>UniRef100_Q9LDM5 Serine/threonine kinase-like protein [Arabidopsis thaliana]
Length = 666
Score = 298 bits (763), Expect = 3e-79
Identities = 203/606 (33%), Positives = 303/606 (49%), Gaps = 52/606 (8%)
Query: 86 VYAFGECMKDLSKSDCDVCFAQCKTRVLRCSPIQRGVEGGRFFFDG-------CYLRYDA 138
+YA G C+ C C +L+ P Q F++ G C++RY
Sbjct: 72 IYALGLCIPGSDPRVCSDCIQLASQGLLQTCPNQTD----SFYWTGDNADKTLCFVRYSN 127
Query: 139 YNFFNQSLSSQDLTLCGNIDFSGNRSVHKANVVELVRNLSVEAPKNDGFFVGVLSKRNVS 198
+FFN+ + + F GN + + + N + + + N
Sbjct: 128 NSFFNKMALEPTHAVYNTMRFQGNLTAY-TRTWDAFMNFMFTRVGQTRYLADISPRINQE 186
Query: 199 ------VYGLAQCWNFVNESVCQNCLVEAVTRIDSCALKEEGRVLNAG-CYLRFSTDKFY 251
+Y L QC ++ C+ CL + V SC G V+N CY R+ K+Y
Sbjct: 187 PLSPDLIYALMQCIPGISSEDCETCLGKCVDDYQSCCNGFIGGVVNKPVCYFRWDGYKYY 246
Query: 252 DNSSNIAPP-----------------GTKGSTKVGIIVAESFAAVATLLIVATIIFFVRK 294
+ AP G K ST G+IVA +AV +++VA + ++
Sbjct: 247 GAFGDEAPSQPPTPLPLPPPPPRDPDGKKIST--GVIVAIVVSAVIFVVLVALGLVIWKR 304
Query: 295 NVLRQRRERRQFGAFLFSENNSKLNMPYEVLEKATNYFHDSNKLGEGGSGSVYKGALPDG 354
RQ + ++ + L + +E AT+ F +NKLG+GG G VYKG LP+
Sbjct: 305 ---RQSYKTLKYHTDDDMTSPQSLQFDFTTIEVATDNFSRNNKLGQGGFGEVYKGMLPNE 361
Query: 355 TTVAIKRLSFNTTQWADHFFNEVNLIKDLQHKNLVKLLGCSITGPESLLVYEYVPNLSLH 414
T +A+KRLS N+ Q F NEV ++ LQHKNLV+LLG I E +LVYE+V N SL
Sbjct: 362 TEIAVKRLSSNSGQGTQEFKNEVVIVAKLQHKNLVRLLGFCIERDEQILVYEFVSNKSLD 421
Query: 415 DHLSVRRNLQQLTWEVRHKIILGTAEGLAYLHEESQLKIIHRDIKLSNILLDDNFTPKIA 474
L + QL W+ R+ II G GL YLH++S+L IIHRDIK SNILLD + PKIA
Sbjct: 422 YFLFDPKMKSQLDWKRRYNIIGGVTRGLLYLHQDSRLTIIHRDIKASNILLDADMNPKIA 481
Query: 475 DFGLARLFPEDQSQISTA-ICGTLGYMAPEYIVLGKLTEKADIYSFGVLVIELLSGKSRT 533
DFG+AR F DQ++ T + GT GYM PEY+ G+ + K+D+YSFGVL++E++ GK +
Sbjct: 482 DFGMARNFRVDQTEDQTGRVVGTFGYMPPEYVTHGQFSTKSDVYSFGVLILEIVCGKKNS 541
Query: 534 SFVQ---NSCSILHMVWSLYGSNRLCDIVDPILEGNYPAEEACKLLKIGLLCTQASAELR 590
SF Q + +++ VW L+ ++ D++DP ++ +Y +E + + IG+LC Q + R
Sbjct: 542 SFFQMDDSGGNLVTHVWRLWNNDSPLDLIDPAIKESYDNDEVIRCIHIGILCVQETPADR 601
Query: 591 PPMSVVVKMINNIHKIADPTQPPFLNFGSGEFSRSSLPEESLQPGSNT-QSSGDSMTESS 649
P MS + +M+ N +PP G F R+ + L GS QSS S+ S
Sbjct: 602 PEMSTIFQMLTNSSITLPVPRPP------GFFFRNRPNLDPLTYGSEQGQSSSMSVPFSI 655
Query: 650 EHRLVT 655
+ +T
Sbjct: 656 DSASIT 661
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,081,610,093
Number of Sequences: 2790947
Number of extensions: 45152706
Number of successful extensions: 154634
Number of sequences better than 10.0: 18182
Number of HSP's better than 10.0 without gapping: 9430
Number of HSP's successfully gapped in prelim test: 8756
Number of HSP's that attempted gapping in prelim test: 115869
Number of HSP's gapped (non-prelim): 21167
length of query: 656
length of database: 848,049,833
effective HSP length: 134
effective length of query: 522
effective length of database: 474,062,935
effective search space: 247460852070
effective search space used: 247460852070
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0076c.7


