

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0074.9
(351 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
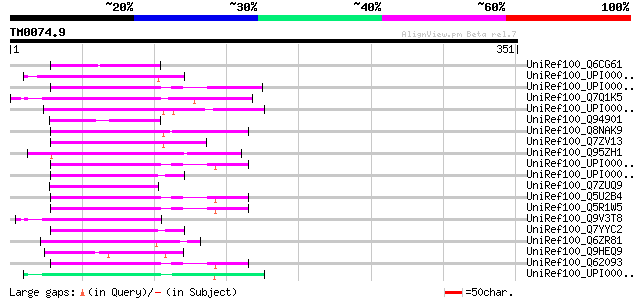
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6CG61 Similar to sp|P31209 Schizosaccharomyces pombe ... 54 8e-06
UniRef100_UPI000026EC47 UPI000026EC47 UniRef100 entry 53 1e-05
UniRef100_UPI00004364B1 UPI00004364B1 UniRef100 entry 52 3e-05
UniRef100_Q7Q1K5 ENSANGP00000010223 [Anopheles gambiae str. PEST] 51 5e-05
UniRef100_UPI0000363ED4 UPI0000363ED4 UniRef100 entry 50 7e-05
UniRef100_Q94901 CG8597-PA, isoform A [Drosophila melanogaster] 50 7e-05
UniRef100_Q8NAK9 Hypothetical protein FLJ35170 [Homo sapiens] 49 3e-04
UniRef100_Q7ZV13 Zgc:56283 [Brachydanio rerio] 48 4e-04
UniRef100_Q95ZH1 RBD protein [Chironomus tentans] 48 4e-04
UniRef100_UPI000024E83B UPI000024E83B UniRef100 entry 48 5e-04
UniRef100_UPI0000452ADA UPI0000452ADA UniRef100 entry 48 5e-04
UniRef100_Q7ZUQ9 Cold inducible RNA binding protein [Brachydanio... 48 5e-04
UniRef100_Q5U2B4 Hypothetical protein [Xenopus laevis] 48 5e-04
UniRef100_Q5R1W5 Arginine/serine-rich 2 splicing factor [Pan tro... 48 5e-04
UniRef100_Q9V3T8 SR family splicing factor SC35 [Drosophila mela... 48 5e-04
UniRef100_Q7YYC2 Splicing factor, possible [Cryptosporidium parvum] 48 5e-04
UniRef100_Q6ZR81 Hypothetical protein FLJ46566 [Homo sapiens] 48 5e-04
UniRef100_Q9HEQ9 Single-stranded TG1-3 binding protein [Schizosa... 48 5e-04
UniRef100_Q62093 Splicing factor, arginine/serine-rich 2 [Mus mu... 48 5e-04
UniRef100_UPI0000432ABB UPI0000432ABB UniRef100 entry 47 6e-04
>UniRef100_Q6CG61 Similar to sp|P31209 Schizosaccharomyces pombe
Polyadenylate-binding protein [Yarrowia lipolytica]
Length = 589
Score = 53.5 bits (127), Expect = 8e-06
Identities = 30/76 (39%), Positives = 44/76 (57%), Gaps = 1/76 (1%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
LFV GL R+T +R FERFG V S+ +P + R + FGFV F+ RD + A+ +L
Sbjct: 192 LFVRGLSRETTEEALRAEFERFGAVTSLLLPLD-ESHRAKGFGFVNFEEHRDAVTAVAAL 250
Query: 89 HGTRLNSFFLSINPAR 104
G L +S++ A+
Sbjct: 251 DGAELCGARISVSRAQ 266
>UniRef100_UPI000026EC47 UPI000026EC47 UniRef100 entry
Length = 171
Score = 53.1 bits (126), Expect = 1e-05
Identities = 37/122 (30%), Positives = 51/122 (41%), Gaps = 15/122 (12%)
Query: 10 PPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREK 69
PPPR V E RLFV L + Q+RK FE++G V F+P+ +
Sbjct: 2 PPPR-----PRRVVEPGSRLFVGNLSYEIDTDQLRKFFEKWGPVNDCFLPQDRTTGKPRG 56
Query: 70 FGFVLFQSKRDGIAAMRSLHGTRLNSFFLSIN----------PARFVDRKYKWNPPAKQH 119
FGFV F+ D A+ GT L+ L +N P + R KW +
Sbjct: 57 FGFVTFRDAADAKLAVAQADGTELDGRQLRVNVAEDTRGPPPPTQPTQRTSKWGTKEEDA 116
Query: 120 EE 121
E+
Sbjct: 117 ED 118
>UniRef100_UPI00004364B1 UPI00004364B1 UniRef100 entry
Length = 174
Score = 51.6 bits (122), Expect = 3e-05
Identities = 39/147 (26%), Positives = 63/147 (42%), Gaps = 22/147 (14%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 16 LKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 75
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAPFSGTGGF 148
G L+ L + AR+ PP + R GAP GG+
Sbjct: 76 DGALLDGRELRVQMARY------GRPPDAHYS----------------RRGAPPRRYGGY 113
Query: 149 SQESQKVWRVKKSTRKSPEKKVEEQER 175
+ S+ + + RK +K ++++R
Sbjct: 114 GRRSRSREKRRPEKRKHRRRKKQKKKR 140
>UniRef100_Q7Q1K5 ENSANGP00000010223 [Anopheles gambiae str. PEST]
Length = 172
Score = 50.8 bits (120), Expect = 5e-05
Identities = 44/170 (25%), Positives = 75/170 (43%), Gaps = 16/170 (9%)
Query: 1 MRPTGSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPK 60
++ +G AR PPPR + I L VD L T +R++FER G V +++P+
Sbjct: 6 IKMSGYAR-PPPR---------IDGMISLKVDNLTYRTTPDDLRRVFERCGEVGDIYIPR 55
Query: 61 TIKRVRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHE 120
F FV F KRD A+ ++ G L+ L + AR+ + P ++
Sbjct: 56 DRHTRESRGFAFVRFYDKRDAEDALDAMDGRMLDGRELRVQMARY----GRPTSPQRRGN 111
Query: 121 ELPGRK--KIRQEWRIKHRVGAPFSGTGGFSQESQKVWRVKKSTRKSPEK 168
GR+ + R+ R + R +P +S+ + R + ++ S K
Sbjct: 112 RYNGRERGRSRERHRRRSRSRSPEHRRRSYSRSKSRTPRSRSGSKSSRGK 161
>UniRef100_UPI0000363ED4 UPI0000363ED4 UniRef100 entry
Length = 273
Score = 50.4 bits (119), Expect = 7e-05
Identities = 43/158 (27%), Positives = 67/158 (42%), Gaps = 10/158 (6%)
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
E + L VD L T +R++FE++G V V++P+ F FV F KRD
Sbjct: 11 EGMVSLKVDNLTYRTAPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFFDKRDAED 70
Query: 84 AMRSLHGTRLNSFFLSINPARF---VDRKYKW--NPPAKQHEELPGRKKIRQEWRIKHRV 138
AM ++ G L+ L + AR+ D Y P K+H R K R R +
Sbjct: 71 AMDAMDGALLDGRELRVQMARYGRPPDSHYGGGRRGPPKKHSGRRSRSKSRSRSRSR--- 127
Query: 139 GAPFSGTGGFSQESQKVWRVKKSTRKSPEKKVEEQERS 176
+S S+ R K ++ K + + + + RS
Sbjct: 128 --SYSPARRSKSRSRSYSRSKSTSPKGRKMEAKSKSRS 163
>UniRef100_Q94901 CG8597-PA, isoform A [Drosophila melanogaster]
Length = 352
Score = 50.4 bits (119), Expect = 7e-05
Identities = 26/77 (33%), Positives = 46/77 (58%), Gaps = 8/77 (10%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LF+ LD T+ +++R LFE++GTVV V K +GFV ++++ G A+++
Sbjct: 8 KLFIGNLDEKTQATELRALFEKYGTVVECDVVK--------NYGFVHMETEQQGRDAIQN 59
Query: 88 LHGTRLNSFFLSINPAR 104
L+G LN F + + A+
Sbjct: 60 LNGYTLNEFAIKVEAAK 76
>UniRef100_Q8NAK9 Hypothetical protein FLJ35170 [Homo sapiens]
Length = 201
Score = 48.5 bits (114), Expect = 3e-04
Identities = 38/139 (27%), Positives = 62/139 (44%), Gaps = 3/139 (2%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 16 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 75
Query: 89 HGTRLNSFFLSINPARF--VDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAPFSGTG 146
G L+ L + AR R+ + +P ++ R + R R ++ S T
Sbjct: 76 DGAVLDGRELRVQMARCGGYGRRSR-SPRRRRRSRSRSRSRSRSRSRSRYSRSKSRSRTR 134
Query: 147 GFSQESQKVWRVKKSTRKS 165
S+ + K ++S KS
Sbjct: 135 SRSRSTSKSRSARRSKSKS 153
>UniRef100_Q7ZV13 Zgc:56283 [Brachydanio rerio]
Length = 225
Score = 48.1 bits (113), Expect = 4e-04
Identities = 34/112 (30%), Positives = 52/112 (46%), Gaps = 4/112 (3%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 16 LKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 75
Query: 89 HGTRLNSFFLSINPARF---VDRKY-KWNPPAKQHEELPGRKKIRQEWRIKH 136
G L+ L + AR+ D Y + P +++ R + R R KH
Sbjct: 76 DGALLDGRELRVQMARYGRPPDAHYSRRGAPPRRYGGYGRRSRSRSPRRRKH 127
>UniRef100_Q95ZH1 RBD protein [Chironomus tentans]
Length = 849
Score = 48.1 bits (113), Expect = 4e-04
Identities = 38/152 (25%), Positives = 62/152 (40%), Gaps = 5/152 (3%)
Query: 13 RWSCHERSEVAEENI----RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRRE 68
RW E EE+I +LF L V+ +FE++G VV V VP + +
Sbjct: 299 RWQEQEEKLKGEEDICESGKLFFRNLPYTVTEDDVQTVFEKYGNVVEVNVPIDPTTRKIK 358
Query: 69 KFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKI 128
FG V F + + A L+GT + + P + D+ K ++ RK++
Sbjct: 359 GFGTVTFLMPENAVQAYNELNGTMFHGRMFHLLPGKSNDKTEADESDPKNFKD-KRRKEL 417
Query: 129 RQEWRIKHRVGAPFSGTGGFSQESQKVWRVKK 160
++ H F GT ++ KV+ K
Sbjct: 418 KKTASSAHNWNTLFMGTNAVAEIISKVYGKSK 449
Score = 39.3 bits (90), Expect = 0.16
Identities = 38/158 (24%), Positives = 69/158 (43%), Gaps = 14/158 (8%)
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREK------FGFVLFQS 77
E N LF+ L++DT +R++F+ GT+ S+ + K K EK +GF+ F+
Sbjct: 613 EPNTTLFIKNLNKDTVEETIREIFKNIGTIRSIQIAKK-KSTDDEKKLIPLGYGFIQFKQ 671
Query: 78 KRDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHR 137
A++++ ++ + + + DR N PA + KK +I R
Sbjct: 672 ASAADKALKTMQHKEIDGIKIELKRS---DRTL--NTPAHVSRKKTDNKKQEGSTKIMVR 726
Query: 138 VGAPFSGTGGFSQESQKVWRVKKSTRKSPEKKVEEQER 175
PF ++ +V+ K+ R P+K +Q R
Sbjct: 727 -NIPFQANANEIRQLFQVFGELKAVR-LPKKPGIDQHR 762
Score = 36.6 bits (83), Expect = 1.1
Identities = 19/66 (28%), Positives = 36/66 (53%)
Query: 24 EENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIA 83
E + ++ V + +++R+LF+ FG + +V +PK + FGF+ F +K D +
Sbjct: 718 EGSTKIMVRNIPFQANANEIRQLFQVFGELKAVRLPKKPGIDQHRGFGFIDFVTKSDAKS 777
Query: 84 AMRSLH 89
A +LH
Sbjct: 778 AFDALH 783
>UniRef100_UPI000024E83B UPI000024E83B UniRef100 entry
Length = 252
Score = 47.8 bits (112), Expect = 5e-04
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 16 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 75
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 76 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 113
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 114 GYGRRSRSPRRRRRSRSRS 132
>UniRef100_UPI0000452ADA UPI0000452ADA UniRef100 entry
Length = 330
Score = 47.8 bits (112), Expect = 5e-04
Identities = 28/93 (30%), Positives = 48/93 (51%), Gaps = 4/93 (4%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L + L DT S VR+ FERFG + V++P + R FGFV + + D AA+ +
Sbjct: 15 LLIRSLRFDTPTSLVRREFERFGAIRDVYLPLDYRSRRPRGFGFVEYVEEEDARAALEKM 74
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEE 121
G L+ +++ A ++ + +P + +H E
Sbjct: 75 DGATLDGVTINVTFA----QEGRKSPESMRHRE 103
>UniRef100_Q7ZUQ9 Cold inducible RNA binding protein [Brachydanio rerio]
Length = 184
Score = 47.8 bits (112), Expect = 5e-04
Identities = 22/76 (28%), Positives = 42/76 (54%)
Query: 28 RLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRS 87
+LF+ GL DT + + F ++GT+ V V + + R FGFV F++ D AM +
Sbjct: 6 KLFIGGLSYDTTEQSLEEAFSKYGTIAKVDVIRDRETDRSRGFGFVTFENPEDAKDAMAA 65
Query: 88 LHGTRLNSFFLSINPA 103
++G +++ + ++ A
Sbjct: 66 MNGKQVDGRMIRVDEA 81
>UniRef100_Q5U2B4 Hypothetical protein [Xenopus laevis]
Length = 271
Score = 47.8 bits (112), Expect = 5e-04
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 66 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 125
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 126 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 163
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 164 GYGRRSRSPRRRRRSRSRS 182
>UniRef100_Q5R1W5 Arginine/serine-rich 2 splicing factor [Pan troglodytes]
Length = 221
Score = 47.8 bits (112), Expect = 5e-04
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 16 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 75
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 76 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 113
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 114 GYGRRSRSPRRRRRSRSRS 132
>UniRef100_Q9V3T8 SR family splicing factor SC35 [Drosophila melanogaster]
Length = 195
Score = 47.8 bits (112), Expect = 5e-04
Identities = 32/101 (31%), Positives = 48/101 (46%), Gaps = 10/101 (9%)
Query: 5 GSARAPPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKR 64
G+AR PPPR + + L VD L T +R++FER G V +++P+
Sbjct: 11 GAAR-PPPR---------IDGMVSLKVDNLTYRTTPEDLRRVFERCGEVGDIYIPRDRYT 60
Query: 65 VRREKFGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPARF 105
F FV F KRD A+ ++ G L+ L + AR+
Sbjct: 61 RESRGFAFVRFYDKRDAEDALEAMDGRMLDGRELRVQMARY 101
>UniRef100_Q7YYC2 Splicing factor, possible [Cryptosporidium parvum]
Length = 330
Score = 47.8 bits (112), Expect = 5e-04
Identities = 28/93 (30%), Positives = 48/93 (51%), Gaps = 4/93 (4%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L + L DT S VR+ FERFG + V++P + R FGFV + + D AA+ +
Sbjct: 15 LLIRSLRFDTPTSLVRREFERFGAIRDVYLPLDYRSRRPRGFGFVEYVEEEDARAALEKM 74
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEE 121
G L+ +++ A ++ + +P + +H E
Sbjct: 75 DGATLDGVTINVTFA----QEGRKSPESMRHRE 103
>UniRef100_Q6ZR81 Hypothetical protein FLJ46566 [Homo sapiens]
Length = 135
Score = 47.8 bits (112), Expect = 5e-04
Identities = 33/121 (27%), Positives = 57/121 (46%), Gaps = 14/121 (11%)
Query: 22 VAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDG 81
+A + +LFV GL DT + ++F ++G + V V K + R FGFV F++ D
Sbjct: 1 MASDEGKLFVGGLSFDTNEQSLEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDA 60
Query: 82 IAAMRSLHG-TRLNSFFLSI---------NPARFVDRKYKWNPPAKQHEELPGRKKIRQE 131
AM +++G R+ S+ NP+R + +W + PGR++ +
Sbjct: 61 KDAMMAMNGKVRIRVLSRSLVSRMGHPPANPSRPLSPVCRWTADPSR----PGRRRGPRL 116
Query: 132 W 132
W
Sbjct: 117 W 117
>UniRef100_Q9HEQ9 Single-stranded TG1-3 binding protein [Schizosaccharomyces pombe]
Length = 348
Score = 47.8 bits (112), Expect = 5e-04
Identities = 36/108 (33%), Positives = 53/108 (48%), Gaps = 14/108 (12%)
Query: 25 ENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRR------EKFGFVLFQSK 78
++ R+FV L TK S++R LFE GTV V +P +RVRR FV F ++
Sbjct: 42 DDFRVFVGRLSTSTKKSEIRSLFETVGTVRKVTIP--FRRVRRGTRLVPSGIAFVTFNNQ 99
Query: 79 RDGIAAMRSLHGTRLNSFFLSINPARFV------DRKYKWNPPAKQHE 120
D A+ +L+G L+ + + AR V DRK N ++ E
Sbjct: 100 EDVDKAIETLNGKTLDDREIVVQKARPVQEQPIKDRKKSKNKNGEEPE 147
>UniRef100_Q62093 Splicing factor, arginine/serine-rich 2 [Mus musculus]
Length = 220
Score = 47.8 bits (112), Expect = 5e-04
Identities = 39/139 (28%), Positives = 58/139 (41%), Gaps = 24/139 (17%)
Query: 29 LFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREKFGFVLFQSKRDGIAAMRSL 88
L VD L T +R++FE++G V V++P+ F FV F KRD AM ++
Sbjct: 15 LKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDAM 74
Query: 89 HGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIRQEWRIKHRVGAP--FSGTG 146
G L+ L + AR+ PP H R G P G G
Sbjct: 75 DGAVLDGRELRVQMARY------GRPPDSHHS----------------RRGPPPRRYGGG 112
Query: 147 GFSQESQKVWRVKKSTRKS 165
G+ + S+ R ++S +S
Sbjct: 113 GYGRRSRSPRRRRRSRSRS 131
>UniRef100_UPI0000432ABB UPI0000432ABB UniRef100 entry
Length = 175
Score = 47.4 bits (111), Expect = 6e-04
Identities = 45/185 (24%), Positives = 74/185 (39%), Gaps = 34/185 (18%)
Query: 10 PPPRWSCHERSEVAEENIRLFVDGLDRDTKHSQVRKLFERFGTVVSVFVPKTIKRVRREK 69
PPPR + + L VD L T +R++FER G V +++P+
Sbjct: 6 PPPR---------IDGMVSLKVDNLTYRTTPEDLRRVFERCGEVGDIYIPRDRFTRESRG 56
Query: 70 FGFVLFQSKRDGIAAMRSLHGTRLNSFFLSINPARFVDRKYKWNPPAKQHEELPGRKKIR 129
F FV F KRD A+ ++ G L+ L + AR+ P H R++ R
Sbjct: 57 FAFVRFYDKRDAEDALDAMDGRLLDGRELRVQMARY-------GRPTSPHRSRGSRRRGR 109
Query: 130 QEWRIKHRVGA-----------------PFSGTGGFSQESQKVWRVKKSTR-KSPEKKVE 171
R + R + +S + S+ K R K +R KSP+++ +
Sbjct: 110 SRSRSRDRRRSRSRSRSRSRSRDRDRRRSYSRSRSRSRSDSKSSRGKSRSRSKSPDRQKD 169
Query: 172 EQERS 176
+ +S
Sbjct: 170 SRSKS 174
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 585,191,029
Number of Sequences: 2790947
Number of extensions: 23426604
Number of successful extensions: 60491
Number of sequences better than 10.0: 1811
Number of HSP's better than 10.0 without gapping: 1069
Number of HSP's successfully gapped in prelim test: 743
Number of HSP's that attempted gapping in prelim test: 58183
Number of HSP's gapped (non-prelim): 2926
length of query: 351
length of database: 848,049,833
effective HSP length: 128
effective length of query: 223
effective length of database: 490,808,617
effective search space: 109450321591
effective search space used: 109450321591
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0074.9


