

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0067a.13
(212 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
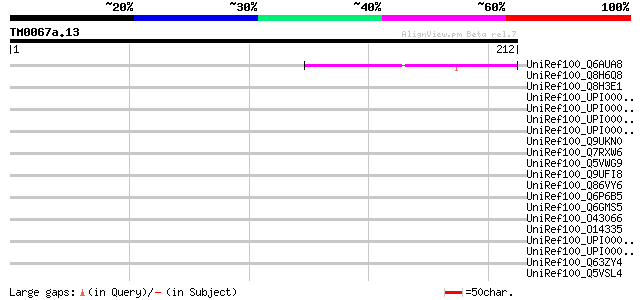
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6AUA8 Hypothetical protein P0017D10.13 [Oryza sativa] 46 6e-04
UniRef100_Q8H6Q8 CTV.20 [Poncirus trifoliata] 45 0.002
UniRef100_Q8H3E1 Hypothetical protein P0018C07.121 [Oryza sativa] 45 0.002
UniRef100_UPI00002BB204 UPI00002BB204 UniRef100 entry 41 0.024
UniRef100_UPI000035F843 UPI000035F843 UniRef100 entry 40 0.040
UniRef100_UPI000035F83A UPI000035F83A UniRef100 entry 40 0.040
UniRef100_UPI0000368EC8 UPI0000368EC8 UniRef100 entry 40 0.053
UniRef100_Q9UKN0 Mucin 11 [Homo sapiens] 39 0.069
UniRef100_Q7RXW6 Hypothetical protein [Neurospora crassa] 39 0.069
UniRef100_Q5VWG9 OTTHUMP00000045008 (TAF3 RNA polymerase II, TAT... 39 0.090
UniRef100_Q9UFI8 Hypothetical protein DKFZp586K1924 [Homo sapiens] 39 0.090
UniRef100_Q86VY6 TAF3 protein [Homo sapiens] 39 0.090
UniRef100_Q6P6B5 TAF3 protein [Homo sapiens] 39 0.090
UniRef100_Q6GMS5 TAF3 protein [Homo sapiens] 39 0.090
UniRef100_O43066 Ark1/Prk1 family protein kinase; serine/threoni... 39 0.090
UniRef100_O14335 DNA-binding protein scr1 [Schizosaccharomyces p... 39 0.090
UniRef100_UPI000036E5BC UPI000036E5BC UniRef100 entry 39 0.12
UniRef100_UPI0000234A11 UPI0000234A11 UniRef100 entry 39 0.12
UniRef100_Q63ZY4 ATXN2L protein [Homo sapiens] 39 0.12
UniRef100_Q5VSL4 OTTHUMP00000021779 [Homo sapiens] 39 0.12
>UniRef100_Q6AUA8 Hypothetical protein P0017D10.13 [Oryza sativa]
Length = 950
Score = 46.2 bits (108), Expect = 6e-04
Identities = 25/92 (27%), Positives = 49/92 (53%), Gaps = 4/92 (4%)
Query: 124 IAKQVFPPNFHYMSGHPSKGRLFYEFILVDTDSIEVTHNKDKNGEIICSKVKILKVLNMQ 183
I + FP + +++ +P K + +YE IL +T S ++ H ++ E+ +K+ I K++++
Sbjct: 675 IMGKYFPQHQYFIPEYPGKDQNYYETILCETRSAQIFHTRN-GDELGFTKLLIQKIISID 733
Query: 184 DW---KHPLFQSKQFSREFTPTGHNYFDYMDA 212
DW +P +S +NY+DY A
Sbjct: 734 DWDKSSNPYVARTIYSTSCANKRYNYWDYQKA 765
>UniRef100_Q8H6Q8 CTV.20 [Poncirus trifoliata]
Length = 3148
Score = 44.7 bits (104), Expect = 0.002
Identities = 40/192 (20%), Positives = 83/192 (42%), Gaps = 12/192 (6%)
Query: 28 SSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSPLVPLSNKYSILDYQNTI 87
+ +P + L+ I + ++P +I + ++S++ ++ + S Q +
Sbjct: 1483 TKSPELMFALQGISKSKALSQIPEEEKPISKEITKLSSQSQNVVISGESSSS----QIVL 1538
Query: 88 SSKTPGKTEQYEYQEKPECLPSVMIEKAWLNCNPRDIAKQVFPPNFHYMSGHPSKGRLFY 147
S TP K ++ +K + +E + + +P + FP ++ + +K + +Y
Sbjct: 1539 SQPTPSKKTS-DWFDKSHFQNVLTMEHGFYHSDPFQAISKFFPQSWFFNPWDLTKPQSYY 1597
Query: 148 EFILVDTDSIEVTH--NKDKNGEIICSKVKILKVLNMQDWKHPLFQSKQFSREFTP---- 201
+ IL T+S++ H + + E S ILKVL+ W L + K F F
Sbjct: 1598 QSILEATESVKFKHFFLSETHSEPAYSTATILKVLSPNQWGDQLHKYKSFPPNFQMRLPH 1657
Query: 202 -TGHNYFDYMDA 212
++Y+DY A
Sbjct: 1658 CLVYSYWDYQQA 1669
>UniRef100_Q8H3E1 Hypothetical protein P0018C07.121 [Oryza sativa]
Length = 376
Score = 44.7 bits (104), Expect = 0.002
Identities = 25/92 (27%), Positives = 49/92 (53%), Gaps = 4/92 (4%)
Query: 124 IAKQVFPPNFHYMSGHPSKGRLFYEFILVDTDSIEVTHNKDKNGEIICSKVKILKVLNMQ 183
I + FP + +++ +P K + +YE IL +T S ++ H ++ ++ +K+ I K +++
Sbjct: 69 IMGKYFPQHQYFIPEYPGKDQNYYETILCETRSAQIFHTRN-GDDLGFTKLLIQKNISID 127
Query: 184 DW---KHPLFQSKQFSREFTPTGHNYFDYMDA 212
DW +P +S T +NY+DY A
Sbjct: 128 DWDKSSNPYVARTIYSTSCTNKRYNYWDYQKA 159
>UniRef100_UPI00002BB204 UPI00002BB204 UniRef100 entry
Length = 2314
Score = 40.8 bits (94), Expect = 0.024
Identities = 38/139 (27%), Positives = 52/139 (37%), Gaps = 17/139 (12%)
Query: 13 PPLGSSRKLPSSITGSSAPVAVTPLKTI----KPPGPSTPTKSSQRPTVSQIVQSPAKSK 68
P +S S T SSA VA+ P + + +PP + T+SS PT P K K
Sbjct: 1760 PARPNSAHADHSSTSSSAHVAIAPQQPVGTISQPPIQPSKTESSAAPT-------PGKDK 1812
Query: 69 SPLVPLSNKYSILDYQNTISSKTPGKTEQ------YEYQEKPECLPSVMIEKAWLNCNPR 122
L + S+ D N++ P T + P LPS A LN
Sbjct: 1813 PALAAENQPASVTDGMNSVGFSAPAMTMSAKAEPLQQLPPPPASLPSSEAPPALLNPQIS 1872
Query: 123 DIAKQVFPPNFHYMSGHPS 141
V PP + HP+
Sbjct: 1873 SHLPTVPPPVLSHSVAHPN 1891
>UniRef100_UPI000035F843 UPI000035F843 UniRef100 entry
Length = 331
Score = 40.0 bits (92), Expect = 0.040
Identities = 27/90 (30%), Positives = 39/90 (43%), Gaps = 3/90 (3%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S + +S+P +P + P++ + +S PT S
Sbjct: 198 SSSPTSSSPTSSPTSSS---PTSSSPTSSPTTSSPTSSSPTSSPTSSSPTSSSPTSSSTT 254
Query: 62 QSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
S + S SP S S +T SS T
Sbjct: 255 SSSSTSSSPTSSSSTSSSSTSSSSTRSSST 284
Score = 37.4 bits (85), Expect = 0.26
Identities = 28/90 (31%), Positives = 39/90 (43%), Gaps = 4/90 (4%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S + +S+P + +P + P+T + +S PT S
Sbjct: 184 SSSPTSSSPTSSPTSSS---PTSSSPTSSPTSSSPTSSSPTSSPTTSSPTSSSPTSSPTS 240
Query: 62 QSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
SP S SP + S T SS T
Sbjct: 241 SSPT-SSSPTSSSTTSSSSTSSSPTSSSST 269
Score = 37.0 bits (84), Expect = 0.34
Identities = 25/90 (27%), Positives = 41/90 (44%), Gaps = 3/90 (3%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S SS+P + +P + P++ + +S PT S
Sbjct: 14 SSSPTSSHTSSSPTSSS---PTSSHTSSSPTSSSPTSSSPTSSPTSSSPTSSSPTSSHTS 70
Query: 62 QSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
SP+ S + P S+ + ++ SS +
Sbjct: 71 SSPSSSPTSSSPTSSPTTSSPTSSSTSSSS 100
Score = 33.5 bits (75), Expect = 3.8
Identities = 27/75 (36%), Positives = 34/75 (45%), Gaps = 8/75 (10%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S SS+P + +P + P S+PT SS PT S
Sbjct: 175 SISPTSSHTSSSPTSSS---PTSSPTSSSPTSSSPTSS---PTSSSPTSSS--PTSSPTT 226
Query: 62 QSPAKSKSPLVPLSN 76
SP S P S+
Sbjct: 227 SSPTSSSPTSSPTSS 241
Score = 33.5 bits (75), Expect = 3.8
Identities = 31/94 (32%), Positives = 37/94 (38%), Gaps = 7/94 (7%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLK--TIKPPGPSTPTKS--SQRPTV 57
S PTS P SS P+S SS+P + +P T P S+PT S S PT
Sbjct: 189 SSSPTSSPTSSSPTSSS---PTSSPTSSSPTSSSPTSSPTTSSPTSSSPTSSPTSSSPTS 245
Query: 58 SQIVQSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
S S S S S +T SS T
Sbjct: 246 SSPTSSSTTSSSSTSSSPTSSSSTSSSSTSSSST 279
>UniRef100_UPI000035F83A UPI000035F83A UniRef100 entry
Length = 464
Score = 40.0 bits (92), Expect = 0.040
Identities = 27/90 (30%), Positives = 39/90 (43%), Gaps = 3/90 (3%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S + +S+P +P + P++ + +S PT S
Sbjct: 323 SSSPTSSSPTSSPTSSS---PTSSSPTSSPTTSSPTSSSPTSSPTSSSPTSSSPTSSSTT 379
Query: 62 QSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
S + S SP S S +T SS T
Sbjct: 380 SSSSTSSSPTSSSSTSSSSTSSSSTRSSST 409
Score = 37.4 bits (85), Expect = 0.26
Identities = 28/90 (31%), Positives = 39/90 (43%), Gaps = 4/90 (4%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S + +S+P + +P + P+T + +S PT S
Sbjct: 309 SSSPTSSSPTSSPTSSS---PTSSSPTSSPTSSSPTSSSPTSSPTTSSPTSSSPTSSPTS 365
Query: 62 QSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
SP S SP + S T SS T
Sbjct: 366 SSPT-SSSPTSSSTTSSSSTSSSPTSSSST 394
Score = 33.5 bits (75), Expect = 3.8
Identities = 27/75 (36%), Positives = 34/75 (45%), Gaps = 8/75 (10%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S PTS P SS P+S SS+P + +P + P S+PT SS PT S
Sbjct: 300 SISPTSSHTSSSPTSSS---PTSSPTSSSPTSSSPTSS---PTSSSPTSSS--PTSSPTT 351
Query: 62 QSPAKSKSPLVPLSN 76
SP S P S+
Sbjct: 352 SSPTSSSPTSSPTSS 366
Score = 33.5 bits (75), Expect = 3.8
Identities = 31/94 (32%), Positives = 37/94 (38%), Gaps = 7/94 (7%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLK--TIKPPGPSTPTKS--SQRPTV 57
S PTS P SS P+S SS+P + +P T P S+PT S S PT
Sbjct: 314 SSSPTSSPTSSSPTSSS---PTSSPTSSSPTSSSPTSSPTTSSPTSSSPTSSPTSSSPTS 370
Query: 58 SQIVQSPAKSKSPLVPLSNKYSILDYQNTISSKT 91
S S S S S +T SS T
Sbjct: 371 SSPTSSSTTSSSSTSSSPTSSSSTSSSSTSSSST 404
>UniRef100_UPI0000368EC8 UPI0000368EC8 UniRef100 entry
Length = 341
Score = 39.7 bits (91), Expect = 0.053
Identities = 26/61 (42%), Positives = 36/61 (58%), Gaps = 3/61 (4%)
Query: 1 MSEKPTSRDQ-YGPPLGSS-RKLPSSITGSSAPVAVTPLKTIK-PPGPSTPTKSSQRPTV 57
+SE+P+ R GP GSS +LP+S G SAPV L+T K PP ++ T S PT+
Sbjct: 148 LSEQPSHRSSPVGPAPGSSPSELPASPAGGSAPVGKQKLETSKRPPSGTSTTSKSTSPTL 207
Query: 58 S 58
+
Sbjct: 208 T 208
>UniRef100_Q9UKN0 Mucin 11 [Homo sapiens]
Length = 957
Score = 39.3 bits (90), Expect = 0.069
Identities = 24/91 (26%), Positives = 37/91 (40%)
Query: 5 PTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSP 64
P+S G L +R S + G S P ++P T P +PT S + SP
Sbjct: 746 PSSSGSTGTTLSPARSTTSGLVGESTPSRLSPSSTETTTLPGSPTTPSLSEKSTTFYTSP 805
Query: 65 AKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
+ L P + S + +++ S PG T
Sbjct: 806 RSPDATLSPATTTSSGVSEESSTSHSQPGST 836
>UniRef100_Q7RXW6 Hypothetical protein [Neurospora crassa]
Length = 1448
Score = 39.3 bits (90), Expect = 0.069
Identities = 27/91 (29%), Positives = 40/91 (43%), Gaps = 2/91 (2%)
Query: 20 KLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSS--QRPTVSQIVQSPAKSKSPLVPLSNK 77
++P S S VTP + P PT SS +PT + S + S P S +
Sbjct: 54 EVPISSVSPSPVTPVTPATPVMPTVDLEPTMSSTAPQPTGDPVDHSLSPSVPAETPKSKR 113
Query: 78 YSILDYQNTISSKTPGKTEQYEYQEKPECLP 108
+S+L ++N S+ K + EKP LP
Sbjct: 114 FSMLRFRNASDSQLAAKAKLQAASEKPPPLP 144
>UniRef100_Q5VWG9 OTTHUMP00000045008 (TAF3 RNA polymerase II, TATA box binding
protein (TBP)-associated factor, 140kDa
(TAF3)(TAF140,TAFII140)) [Homo sapiens]
Length = 929
Score = 38.9 bits (89), Expect = 0.090
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 255 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 307
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 308 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 351
>UniRef100_Q9UFI8 Hypothetical protein DKFZp586K1924 [Homo sapiens]
Length = 681
Score = 38.9 bits (89), Expect = 0.090
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 233 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 285
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 286 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 329
>UniRef100_Q86VY6 TAF3 protein [Homo sapiens]
Length = 539
Score = 38.9 bits (89), Expect = 0.090
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 255 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 307
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 308 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 351
>UniRef100_Q6P6B5 TAF3 protein [Homo sapiens]
Length = 771
Score = 38.9 bits (89), Expect = 0.090
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 312 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 364
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 365 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 408
>UniRef100_Q6GMS5 TAF3 protein [Homo sapiens]
Length = 601
Score = 38.9 bits (89), Expect = 0.090
Identities = 32/104 (30%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + TVS+
Sbjct: 320 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TVSK 372
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 373 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 416
>UniRef100_O43066 Ark1/Prk1 family protein kinase; serine/threonine protein kinase
(Predicted); similar to S. cerevisiae YNL020C and
YIL095W and YBR059C; similar to S. pombe SPBC557.04 and
SPCP1E11.02; actin cortical patch component (Predicted);
consensus phosphorylation target site
[Schizosaccharomyces pombe]
Length = 953
Score = 38.9 bits (89), Expect = 0.090
Identities = 29/77 (37%), Positives = 38/77 (48%), Gaps = 8/77 (10%)
Query: 11 YGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAKSKSP 70
Y PP ++ PS GS P++ P T PP P+ T SS P V+ + KSKSP
Sbjct: 346 YNPPRAPLQRTPS---GSLTPLSSRPAHTSLPPIPTVQTTSSNVPPVN---RPSLKSKSP 399
Query: 71 LVP--LSNKYSILDYQN 85
V LSN+ S + N
Sbjct: 400 SVSNILSNQLSPISSAN 416
>UniRef100_O14335 DNA-binding protein scr1 [Schizosaccharomyces pombe]
Length = 565
Score = 38.9 bits (89), Expect = 0.090
Identities = 27/75 (36%), Positives = 38/75 (50%), Gaps = 4/75 (5%)
Query: 1 MSEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQI 60
M KP S+ GP SS + + SS V+V+ L PP PS+ TKS+ + S
Sbjct: 493 MPHKPASQSNVGPVRISSNRRSRKFSSSSR-VSVSNLLAGSPPSPSSSTKSA---SSSYS 548
Query: 61 VQSPAKSKSPLVPLS 75
+PA S PL P++
Sbjct: 549 TTTPAFSIGPLTPMT 563
>UniRef100_UPI000036E5BC UPI000036E5BC UniRef100 entry
Length = 729
Score = 38.5 bits (88), Expect = 0.12
Identities = 31/104 (29%), Positives = 49/104 (46%), Gaps = 9/104 (8%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVT--PLKTIKPPGPSTPTKSSQRPTVSQ 59
S+ PT++ PL + P + T +S+P T P P +P +S + T+S+
Sbjct: 255 SQMPTAK-----PLETKSFTPKTKTKTSSPGQKTKSPKTAQSPAMVGSPIRSPK--TISK 307
Query: 60 IVQSPAKSKSPLVPLSNKYSILDYQNTISSKTPGKTEQYEYQEK 103
+SP +SKSP P S K + Q + +TP +T EK
Sbjct: 308 EKKSPGRSKSPKSPKSPKVTTHIPQTPVRPETPNRTPSATLSEK 351
>UniRef100_UPI0000234A11 UPI0000234A11 UniRef100 entry
Length = 889
Score = 38.5 bits (88), Expect = 0.12
Identities = 29/91 (31%), Positives = 46/91 (49%), Gaps = 13/91 (14%)
Query: 7 SRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIVQSPAK 66
SR GPPLG++ +LP + S P +V + + P TP SS RPT+
Sbjct: 8 SRTASGPPLGAAAQLPIPKSRKSLPTSVESPRAVSPSKLRTP--SSPRPTL--------- 56
Query: 67 SKSPLVPLSNKYSILDYQNTISSKTPGKTEQ 97
SKSPL ++ +I ++T ++TP ++
Sbjct: 57 SKSPL--SNSATNISAARSTTVARTPSSPDK 85
>UniRef100_Q63ZY4 ATXN2L protein [Homo sapiens]
Length = 1062
Score = 38.5 bits (88), Expect = 0.12
Identities = 28/100 (28%), Positives = 39/100 (39%), Gaps = 14/100 (14%)
Query: 2 SEKPTSRDQYGPPLGSSRKLPSSITGSSAPVAVTPLKTIKPPGPSTPTKSSQRPTVSQIV 61
S +P+ PP R P S+AP ++ P G + PT S+ P S +
Sbjct: 426 SNRPSGETSVPPPPAVGRMYPPRSPKSAAPAPISASCPEPPIGSAVPTSSASIPVTSSVS 485
Query: 62 Q------SPAKSKSPLVPLSNKYSILDYQNTISSKTPGKT 95
SPA K L P K +S+K PG+T
Sbjct: 486 DPGVGSISPASPKISLAPTDVK--------ELSTKEPGRT 517
>UniRef100_Q5VSL4 OTTHUMP00000021779 [Homo sapiens]
Length = 334
Score = 38.5 bits (88), Expect = 0.12
Identities = 25/61 (40%), Positives = 35/61 (56%), Gaps = 3/61 (4%)
Query: 1 MSEKPTSRDQ-YGPPLGSS-RKLPSSITGSSAPVAVTPLKTIK-PPGPSTPTKSSQRPTV 57
+ +KP+ R GP GSS +LP+S G SAPV L+T K PP ++ T S PT+
Sbjct: 141 LEQKPSHRSSPVGPAPGSSPSELPASPAGGSAPVGKQKLETSKRPPSGTSTTSKSTSPTL 200
Query: 58 S 58
+
Sbjct: 201 T 201
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.130 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 404,604,773
Number of Sequences: 2790947
Number of extensions: 17887150
Number of successful extensions: 60570
Number of sequences better than 10.0: 417
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 383
Number of HSP's that attempted gapping in prelim test: 59665
Number of HSP's gapped (non-prelim): 1117
length of query: 212
length of database: 848,049,833
effective HSP length: 122
effective length of query: 90
effective length of database: 507,554,299
effective search space: 45679886910
effective search space used: 45679886910
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0067a.13


