

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0066.12
(163 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
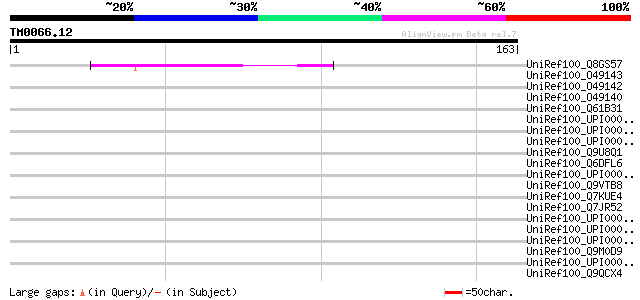
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GS57 Hypothetical protein OJ1477_F01.105 [Oryza sativa] 62 4e-09
UniRef100_O49143 Polyprotein [Arabidopsis thaliana] 42 0.006
UniRef100_O49142 Polyprotein [Arabidopsis thaliana] 42 0.006
UniRef100_O49140 Polyprotein [Arabidopsis thaliana] 42 0.006
UniRef100_Q61B31 Hypothetical protein CBG13518 [Caenorhabditis b... 36 0.31
UniRef100_UPI000029BDC8 UPI000029BDC8 UniRef100 entry 36 0.40
UniRef100_UPI00004577E4 UPI00004577E4 UniRef100 entry 34 1.2
UniRef100_UPI00002B222E UPI00002B222E UniRef100 entry 34 1.5
UniRef100_Q9U8Q1 LagC2 [Dictyostelium discoideum] 34 1.5
UniRef100_Q6DFL6 LOC398539 protein [Xenopus laevis] 33 2.0
UniRef100_UPI000042D98E UPI000042D98E UniRef100 entry 33 2.6
UniRef100_Q9VTB8 CG7958-PA, isoform A [Drosophila melanogaster] 33 2.6
UniRef100_Q7KUE4 CG7958-PB, isoform B [Drosophila melanogaster] 33 2.6
UniRef100_Q7JR52 LD16921p [Drosophila melanogaster] 33 2.6
UniRef100_UPI000035F843 UPI000035F843 UniRef100 entry 33 3.4
UniRef100_UPI000035F83A UPI000035F83A UniRef100 entry 33 3.4
UniRef100_UPI00003C1ED4 UPI00003C1ED4 UniRef100 entry 33 3.4
UniRef100_Q9M0D9 Hypothetical protein AT4g29440 [Arabidopsis tha... 33 3.4
UniRef100_UPI00004307EC UPI00004307EC UniRef100 entry 32 4.4
UniRef100_Q9QCX4 Overlapping protein [Chayote mosaic tymovirus] 32 4.4
>UniRef100_Q8GS57 Hypothetical protein OJ1477_F01.105 [Oryza sativa]
Length = 501
Score = 62.4 bits (150), Expect = 4e-09
Identities = 37/79 (46%), Positives = 42/79 (52%), Gaps = 18/79 (22%)
Query: 27 DMELPLPTTTSSE-TEPFRLDQAVCSHGLFMMAPNSWDPLSNTLTRPLRLHDQDTDPSSS 85
++ELPL PF L+ AVCSHGLFMMAPN WDP S L RPLRL
Sbjct: 20 ELELPLGGAPPYPGAAPFDLEAAVCSHGLFMMAPNRWDPASRALVRPLRL---------- 69
Query: 86 SPSFIVTVSQRSESIAVRV 104
S R+ S+AVRV
Sbjct: 70 -------ASDRAASVAVRV 81
>UniRef100_O49143 Polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 8/91 (8%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLP-------TTTSSETEPFRLDQAVCSHGLFM 56
L PP +P+CS P SPS + ++ P P + TS T P ++++ H
Sbjct: 784 LTTAPPSSPSCSAPHRSPSQSE-NLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSET 842
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSP 87
S PLS RP Q T P SSSP
Sbjct: 843 GPTGSSPPLSPQPQRPQPQSPQSTSPHSSSP 873
>UniRef100_O49142 Polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 8/91 (8%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLP-------TTTSSETEPFRLDQAVCSHGLFM 56
L PP +P+CS P SPS + ++ P P + TS T P ++++ H
Sbjct: 784 LTTAPPSSPSCSAPHRSPSQSE-NLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSET 842
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSP 87
S PLS RP Q T P SSSP
Sbjct: 843 GPTGSSPPLSPQPQRPQPQSPQSTSPHSSSP 873
>UniRef100_O49140 Polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 42.0 bits (97), Expect = 0.006
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 8/91 (8%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLP-------TTTSSETEPFRLDQAVCSHGLFM 56
L PP +P+CS P SPS + ++ P P + TS T P ++++ H
Sbjct: 784 LTTAPPSSPSCSAPHRSPSQSE-NLSPPAPLQPSLSLSPTSPITSPSLSEESLVGHNSET 842
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSP 87
S PLS RP Q T P SSSP
Sbjct: 843 GPTGSSPPLSPQPQRPQPQSPQSTSPHSSSP 873
>UniRef100_Q61B31 Hypothetical protein CBG13518 [Caenorhabditis briggsae]
Length = 1032
Score = 36.2 bits (82), Expect = 0.31
Identities = 40/141 (28%), Positives = 58/141 (40%), Gaps = 13/141 (9%)
Query: 3 ELEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSET-EPFRLDQAVCSHGLFMMAPNS 61
+L+K C P S P +S ST P P+ TSS T +L + V HG+ + S
Sbjct: 506 QLDKKAMC-PTSSSPSTSRGSTTTSQASPAPSHTSSSTSSSVQLRENVARHGVSLKETMS 564
Query: 62 WDPLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQ-----RSESIAVRVHH----GTHLLS 112
+ + L + + P + +PS + SQ RS S HH G S
Sbjct: 565 LRDVGPLSRDAISLRETVSPPITRAPSQPPSYSQPRPPPRSVSSVNNHHHSLPNGDQFSS 624
Query: 113 PHEVRALMVPTS--LTASLNS 131
R + PT+ LT+S S
Sbjct: 625 LDRTRGSVTPTAPPLTSSAAS 645
>UniRef100_UPI000029BDC8 UPI000029BDC8 UniRef100 entry
Length = 828
Score = 35.8 bits (81), Expect = 0.40
Identities = 36/133 (27%), Positives = 51/133 (38%), Gaps = 16/133 (12%)
Query: 15 SKPESSPSSTWFDMELPLPTTTSSETEPFRLDQA---------VCSHGLFMMAPNSWDPL 65
S P+S+ S DM+L + + EP++LD A H F P S D
Sbjct: 544 SSPDSNASYAVEDMDL---NGSYYQKEPYQLDVARERALTISSPFRHSPFSYKPKSLDQN 600
Query: 66 SNTLTRPLRL----HDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSPHEVRALMV 121
NT RPL L H P SS S ++ + S H ++LL
Sbjct: 601 QNTRPRPLSLLQHSHSNSLPPKLSSLSLSLSPTPPSSPSCASPHSSSYLLPRPNTPGSTS 660
Query: 122 PTSLTASLNSWVF 134
P+ S +S+ F
Sbjct: 661 PSPSVRSSSSFNF 673
>UniRef100_UPI00004577E4 UPI00004577E4 UniRef100 entry
Length = 141
Score = 34.3 bits (77), Expect = 1.2
Identities = 16/36 (44%), Positives = 19/36 (52%), Gaps = 8/36 (22%)
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEP 42
P P +P C P+ SP PLPTTTSS +P
Sbjct: 9 PSPAHPTCPSPKPSPQ--------PLPTTTSSHRDP 36
>UniRef100_UPI00002B222E UPI00002B222E UniRef100 entry
Length = 1146
Score = 33.9 bits (76), Expect = 1.5
Identities = 27/108 (25%), Positives = 49/108 (45%), Gaps = 12/108 (11%)
Query: 10 CNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSNTL 69
C+ CS+P P S P P+++S+ E S G + P+S +S+ +
Sbjct: 813 CSCCCSEPPQPPPSPGLG---PYPSSSSAIKER-------PSDGSLPVPPSSCSEISSEI 862
Query: 70 TRPLRLHDQDTDPSSSSP--SFIVTVSQRSESIAVRVHHGTHLLSPHE 115
+ + + SS P S +V + +++ + +HH TH LS H+
Sbjct: 863 STSEMSSEVGSTASSDEPPQSSLVLLHEQAHHPHLHLHHQTHPLSTHQ 910
>UniRef100_Q9U8Q1 LagC2 [Dictyostelium discoideum]
Length = 895
Score = 33.9 bits (76), Expect = 1.5
Identities = 26/99 (26%), Positives = 37/99 (37%), Gaps = 6/99 (6%)
Query: 10 CNPNCSKPESSPSSTWF-DMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSNT 68
C S P STW D P P P V GLF+ + L+N+
Sbjct: 97 CTRTVSDPTEDCQSTWITDNAFPEPLNVKFSGYPSTSGGDVVFTGLFLRLAGGPNTLANS 156
Query: 69 LTRP-----LRLHDQDTDPSSSSPSFIVTVSQRSESIAV 102
T P +++H +DP+ + +F T S I V
Sbjct: 157 FTNPSKTFLIKVHGNFSDPNFNCNNFTATFPPSSGYIKV 195
>UniRef100_Q6DFL6 LOC398539 protein [Xenopus laevis]
Length = 2414
Score = 33.5 bits (75), Expect = 2.0
Identities = 31/115 (26%), Positives = 54/115 (46%), Gaps = 19/115 (16%)
Query: 3 ELEKPPPCNPNCSK----PESSPSSTWFDMELPLPTTTSSETEPFR-LDQAVCS-HGLFM 56
E+E+P P +P C + P+++PS+T +P TT E + + VC + +
Sbjct: 2135 EIEEPNPFDPCCPRYRCEPKTTPSTT-------MPITTKQECDNVKCYKDVVCGPNDNEV 2187
Query: 57 MAPNSWDPLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQ--RSESIAVRVHHGTH 109
PN +DP R + T PS+ + S I+T ++ S+ I V++ H
Sbjct: 2188 EEPNPFDP----CCPKYRCEPKSTTPSTITTSTIITTTKIDCSKRICVKISCRLH 2238
>UniRef100_UPI000042D98E UPI000042D98E UniRef100 entry
Length = 438
Score = 33.1 bits (74), Expect = 2.6
Identities = 28/111 (25%), Positives = 44/111 (39%), Gaps = 8/111 (7%)
Query: 4 LEKPPPCNPNCSKPESSPSSTWFDMELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWD 63
L +PP P + P SSP + P +T+S+ T Q+V + M P S
Sbjct: 52 LSRPPNSGPASTAPSSSPRFQPYSRPQPQRSTSSNST------QSVQQNNNMGMPPPSLP 105
Query: 64 PLSNTLTRPLRLH-DQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLSP 113
P SN++ P ++ P S + + S + H HL +P
Sbjct: 106 P-SNSIRPPTQISPSHQPSPLHQSALSFLPPQELSRGYSESTHSPNHLSNP 155
>UniRef100_Q9VTB8 CG7958-PA, isoform A [Drosophila melanogaster]
Length = 1135
Score = 33.1 bits (74), Expect = 2.6
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 13/136 (9%)
Query: 8 PPCNPNCSKPESSPS-------STWFDMELPLPTTTSSETEPFRL--DQAVCSHGLFMMA 58
PP P CS P SP+ +M L P PFRL + +V +H +F +
Sbjct: 533 PPLTPACSVPYVSPNPDIKPPMDNSEEMRLTFPVRDGIILAPFRLLHNLSVSNH-VFHLK 591
Query: 59 PNSWDPL--SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAV-RVHHGTHLLSPHE 115
N ++ L N L L+ QD +++ VTVS + + + R + L P
Sbjct: 592 QNVYNTLMCRNDLELQLKCFHQDDRQMNTNWPHTVTVSANATPLNIERSEKNSTALRPLY 651
Query: 116 VRALMVPTSLTASLNS 131
++A+ P T L +
Sbjct: 652 LKAVCQPGRNTLQLTA 667
>UniRef100_Q7KUE4 CG7958-PB, isoform B [Drosophila melanogaster]
Length = 1149
Score = 33.1 bits (74), Expect = 2.6
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 13/136 (9%)
Query: 8 PPCNPNCSKPESSPS-------STWFDMELPLPTTTSSETEPFRL--DQAVCSHGLFMMA 58
PP P CS P SP+ +M L P PFRL + +V +H +F +
Sbjct: 547 PPLTPACSVPYVSPNPDIKPPMDNSEEMRLTFPVRDGIILAPFRLLHNLSVSNH-VFHLK 605
Query: 59 PNSWDPL--SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAV-RVHHGTHLLSPHE 115
N ++ L N L L+ QD +++ VTVS + + + R + L P
Sbjct: 606 QNVYNTLMCRNDLELQLKCFHQDDRQMNTNWPHTVTVSANATPLNIERSEKNSTALRPLY 665
Query: 116 VRALMVPTSLTASLNS 131
++A+ P T L +
Sbjct: 666 LKAVCQPGRNTLQLTA 681
>UniRef100_Q7JR52 LD16921p [Drosophila melanogaster]
Length = 1109
Score = 33.1 bits (74), Expect = 2.6
Identities = 36/136 (26%), Positives = 57/136 (41%), Gaps = 13/136 (9%)
Query: 8 PPCNPNCSKPESSPS-------STWFDMELPLPTTTSSETEPFRL--DQAVCSHGLFMMA 58
PP P CS P SP+ +M L P PFRL + +V +H +F +
Sbjct: 507 PPLTPACSVPYVSPNPDIKPPMDNSEEMRLTFPVRDGIILAPFRLLHNLSVSNH-VFHLK 565
Query: 59 PNSWDPL--SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAV-RVHHGTHLLSPHE 115
N ++ L N L L+ QD +++ VTVS + + + R + L P
Sbjct: 566 QNVYNTLMCRNDLELQLKCFHQDDRQMNTNWPHTVTVSANATPLNIERSEKNSTALRPLY 625
Query: 116 VRALMVPTSLTASLNS 131
++A+ P T L +
Sbjct: 626 LKAVCQPGRNTLQLTA 641
>UniRef100_UPI000035F843 UPI000035F843 UniRef100 entry
Length = 331
Score = 32.7 bits (73), Expect = 3.4
Identities = 29/94 (30%), Positives = 38/94 (39%), Gaps = 7/94 (7%)
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPT-TTSSETEPFRLDQAVCSHGLFMMAPNSWDPL 65
P +P S P SSP+S+ P + TTSS T S +P S P
Sbjct: 196 PTSSSPTSSSPTSSPTSSSPTSSSPTSSPTTSSPTSSSPTSSPTSS------SPTSSSPT 249
Query: 66 SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSES 99
S++ T T SS+S S + S RS S
Sbjct: 250 SSSTTSSSSTSSSPTSSSSTSSSSTSSSSTRSSS 283
>UniRef100_UPI000035F83A UPI000035F83A UniRef100 entry
Length = 464
Score = 32.7 bits (73), Expect = 3.4
Identities = 29/94 (30%), Positives = 38/94 (39%), Gaps = 7/94 (7%)
Query: 7 PPPCNPNCSKPESSPSSTWFDMELPLPT-TTSSETEPFRLDQAVCSHGLFMMAPNSWDPL 65
P +P S P SSP+S+ P + TTSS T S +P S P
Sbjct: 321 PTSSSPTSSSPTSSPTSSSPTSSSPTSSPTTSSPTSSSPTSSPTSS------SPTSSSPT 374
Query: 66 SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSES 99
S++ T T SS+S S + S RS S
Sbjct: 375 SSSTTSSSSTSSSPTSSSSTSSSSTSSSSTRSSS 408
>UniRef100_UPI00003C1ED4 UPI00003C1ED4 UniRef100 entry
Length = 1444
Score = 32.7 bits (73), Expect = 3.4
Identities = 16/49 (32%), Positives = 24/49 (48%)
Query: 64 PLSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGTHLLS 112
PL+ + TRP LH TDP S+ I T R + + ++ H L+
Sbjct: 889 PLATSSTRPAGLHRSSTDPQLSACKPIATQPPRQQQLHAQIQASEHQLA 937
>UniRef100_Q9M0D9 Hypothetical protein AT4g29440 [Arabidopsis thaliana]
Length = 1071
Score = 32.7 bits (73), Expect = 3.4
Identities = 32/104 (30%), Positives = 43/104 (40%), Gaps = 14/104 (13%)
Query: 12 PNCSKPESSPSSTWFDMELPLPTT-------TSSETEPFRLDQAVCSHGLFMMAPNSWDP 64
P+ P+SS S DMELP + T S T P L V L P
Sbjct: 730 PSSIPPDSSSSDDESDMELPKRVSFRYQEKRTESRTRPTHLHSGVSHKDLEEEIPTR--- 786
Query: 65 LSNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHHGT 108
++T ++ R H T P+S+S S+ T+S E VH T
Sbjct: 787 -ASTRSQDRRTH--KTTPASASASYFHTMSSDDED-EKEVHRDT 826
>UniRef100_UPI00004307EC UPI00004307EC UniRef100 entry
Length = 322
Score = 32.3 bits (72), Expect = 4.4
Identities = 27/80 (33%), Positives = 35/80 (43%), Gaps = 12/80 (15%)
Query: 9 PCNPNCSKPESSPSSTWFDMEL-PLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPLSN 67
P +P S+PSST D P PT+TSS TEP S P+S +P +
Sbjct: 85 PDTTTSPEPTSTPSSTEPDTTTSPEPTSTSSSTEPDTTTSPEPS-----STPSSTEPATT 139
Query: 68 TLTRPLRLHDQDTDPSSSSP 87
T P + PSS+ P
Sbjct: 140 TSPEP------TSTPSSTEP 153
>UniRef100_Q9QCX4 Overlapping protein [Chayote mosaic tymovirus]
Length = 680
Score = 32.3 bits (72), Expect = 4.4
Identities = 30/101 (29%), Positives = 39/101 (37%), Gaps = 21/101 (20%)
Query: 7 PPPCNPNCSKPESSPSSTWFD-MELPLPTTTSSETEPFRLDQAVCSHGLFMMAPNSWDPL 65
P P P S+P S PSS+ LPL + + + P Q C PL
Sbjct: 520 PHPSIPTMSQPPSLPSSSSGPPRHLPLRSPSPPVSAPMEAPQLPCVES---------PPL 570
Query: 66 SNTLTRPLRLHDQDTDPSSSSPSFIVTVSQRSESIAVRVHH 106
S L+ P SSP IV +RS S++ HH
Sbjct: 571 SPPLSHP-----------PSSPELIVYRPRRSASLSPHSHH 600
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.132 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 285,564,720
Number of Sequences: 2790947
Number of extensions: 10714308
Number of successful extensions: 34792
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 34737
Number of HSP's gapped (non-prelim): 72
length of query: 163
length of database: 848,049,833
effective HSP length: 117
effective length of query: 46
effective length of database: 521,509,034
effective search space: 23989415564
effective search space used: 23989415564
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0066.12


