

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0062.9
(271 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
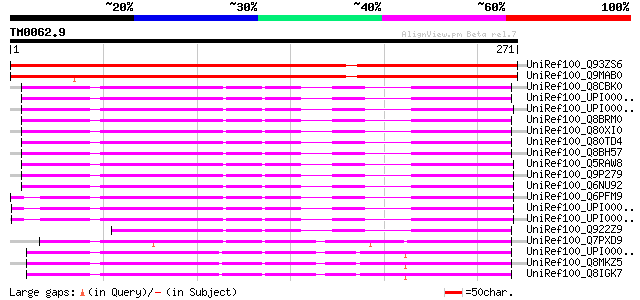
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q93ZS6 Hypothetical protein At3g05090 [Arabidopsis tha... 407 e-112
UniRef100_Q9MAB0 T12H1.5 protein [Arabidopsis thaliana] 402 e-111
UniRef100_Q8CBK0 Mus musculus 16 days neonate cerebellum cDNA, R... 162 1e-38
UniRef100_UPI00001D0125 UPI00001D0125 UniRef100 entry 161 2e-38
UniRef100_UPI00003AB862 UPI00003AB862 UniRef100 entry 161 2e-38
UniRef100_Q8BRM0 Mus musculus 10 days neonate cortex cDNA, RIKEN... 161 2e-38
UniRef100_Q80XI0 8430408H12Rik protein [Mus musculus] 161 2e-38
UniRef100_Q80TD4 MKIAA1449 protein [Mus musculus] 161 2e-38
UniRef100_Q8BH57 Mus musculus adult male testis cDNA, RIKEN full... 161 2e-38
UniRef100_Q5RAW8 Hypothetical protein DKFZp459F151 [Pongo pygmaeus] 161 2e-38
UniRef100_Q9P279 KIAA1449 protein [Homo sapiens] 161 2e-38
UniRef100_Q6NU92 LOC414700 protein [Xenopus laevis] 159 6e-38
UniRef100_Q6PFM9 Zgc:66229 [Brachydanio rerio] 159 7e-38
UniRef100_UPI0000360CB9 UPI0000360CB9 UniRef100 entry 157 2e-37
UniRef100_UPI0000337947 UPI0000337947 UniRef100 entry 157 2e-37
UniRef100_Q922Z9 8430408H12Rik protein [Mus musculus] 151 2e-35
UniRef100_Q7PXD9 ENSANGP00000009742 [Anopheles gambiae str. PEST] 117 3e-25
UniRef100_UPI000007A4E1 UPI000007A4E1 UniRef100 entry 110 4e-23
UniRef100_Q8MKZ5 CG9062-PB [Drosophila melanogaster] 110 4e-23
UniRef100_Q8IGK7 RE72568p [Drosophila melanogaster] 110 4e-23
>UniRef100_Q93ZS6 Hypothetical protein At3g05090 [Arabidopsis thaliana]
Length = 753
Score = 407 bits (1046), Expect = e-112
Identities = 196/271 (72%), Positives = 226/271 (83%), Gaps = 5/271 (1%)
Query: 1 MHRVGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSR 60
MHRVGSAG+ S R RKEK+LTYVLND++DTKHCAGINCL VLKS+VS+ YLFTGSR
Sbjct: 1 MHRVGSAGSNGGSVRTRKEKKLTYVLNDANDTKHCAGINCLDVLKSSVSNDQSYLFTGSR 60
Query: 61 DGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTCTR 120
DG LKRWA EDA CSATFESHVDWVNDA L G+STLVSCSSDTT+KTWD S G CTR
Sbjct: 61 DGTLKRWAFDEDATFCSATFESHVDWVNDAALAGESTLVSCSSDTTVKTWDGLSDGVCTR 120
Query: 121 TLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGING 180
TLRQHSDYVTCLA + KN+N+VASGGLGGEVF+WD+EAA + K DA +D +SNG NG
Sbjct: 121 TLRQHSDYVTCLAVAAKNNNVVASGGLGGEVFIWDIEAALSPVTKPNDANEDSSSNGANG 180
Query: 181 SGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEK 240
P+TSLR++ SSN+ISV ++ + GY P AKGHKESVYALAMN+ GT+LVSGGTEK
Sbjct: 181 -----PVTSLRTVGSSNNISVQSSPSHGYTPTIAKGHKESVYALAMNDTGTMLVSGGTEK 235
Query: 241 VVRVWDPRSGSKTMKLKGHTDNIRALLLDST 271
V+RVWDPR+GSK+MKL+GHTDN+R LLLDST
Sbjct: 236 VLRVWDPRTGSKSMKLRGHTDNVRVLLLDST 266
>UniRef100_Q9MAB0 T12H1.5 protein [Arabidopsis thaliana]
Length = 743
Score = 402 bits (1033), Expect = e-111
Identities = 196/273 (71%), Positives = 226/273 (81%), Gaps = 7/273 (2%)
Query: 1 MHRVGSAGNTNSSTRPRKEKRLTYVLNDSDDTK--HCAGINCLAVLKSAVSDGSDYLFTG 58
MHRVGSAG+ S R RKEK+LTYVLND++DTK HCAGINCL VLKS+VS+ YLFTG
Sbjct: 1 MHRVGSAGSNGGSVRTRKEKKLTYVLNDANDTKLQHCAGINCLDVLKSSVSNDQSYLFTG 60
Query: 59 SRDGRLKRWALGEDAATCSATFESHVDWVNDAVLVGDSTLVSCSSDTTLKTWDAFSTGTC 118
SRDG LKRWA EDA CSATFESHVDWVNDA L G+STLVSCSSDTT+KTWD S G C
Sbjct: 61 SRDGTLKRWAFDEDATFCSATFESHVDWVNDAALAGESTLVSCSSDTTVKTWDGLSDGVC 120
Query: 119 TRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGI 178
TRTLRQHSDYVTCLA + KN+N+VASGGLGGEVF+WD+EAA + K DA +D +SNG
Sbjct: 121 TRTLRQHSDYVTCLAVAAKNNNVVASGGLGGEVFIWDIEAALSPVTKPNDANEDSSSNGA 180
Query: 179 NGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGT 238
NG P+TSLR++ SSN+ISV ++ + GY P AKGHKESVYALAMN+ GT+LVSGGT
Sbjct: 181 NG-----PVTSLRTVGSSNNISVQSSPSHGYTPTIAKGHKESVYALAMNDTGTMLVSGGT 235
Query: 239 EKVVRVWDPRSGSKTMKLKGHTDNIRALLLDST 271
EKV+RVWDPR+GSK+MKL+GHTDN+R LLLDST
Sbjct: 236 EKVLRVWDPRTGSKSMKLRGHTDNVRVLLLDST 268
>UniRef100_Q8CBK0 Mus musculus 16 days neonate cerebellum cDNA, RIKEN full-length
enriched library, clone:9630012C05 product:hypothetical
Trp-Asp (WD) repeats profile/Trp-Asp (WD) repeats
circular profile/G-protein beta WD-40 repeats containing
protein, full insert sequence [Mus musculus]
Length = 576
Score = 162 bits (409), Expect = 1e-38
Identities = 98/264 (37%), Positives = 145/264 (54%), Gaps = 49/264 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 2 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 56
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLVGDS-TLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL + TL+S SSDTT+K W+A G C TLR
Sbjct: 57 WSVNQHKQDPYIASMEHHTDWVNDVVLCSNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 115
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 116 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 158
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 159 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 194
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WDPR+ +K MKLKGHTDN++ALLL
Sbjct: 195 WDPRTCAKLMKLKGHTDNVKALLL 218
>UniRef100_UPI00001D0125 UPI00001D0125 UniRef100 entry
Length = 753
Score = 161 bits (407), Expect = 2e-38
Identities = 98/264 (37%), Positives = 144/264 (54%), Gaps = 49/264 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 40 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 94
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 95 WSVNQHKQDPYIASMEHHTDWVNDVVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 153
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 154 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 196
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 197 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 232
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WDPR+ +K MKLKGHTDN++ALLL
Sbjct: 233 WDPRTCAKLMKLKGHTDNVKALLL 256
>UniRef100_UPI00003AB862 UPI00003AB862 UniRef100 entry
Length = 680
Score = 161 bits (407), Expect = 2e-38
Identities = 98/265 (36%), Positives = 145/265 (53%), Gaps = 49/265 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 5 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 59
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 60 WSVNQHKQDPYIASMEHHTDWVNDIVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 118
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 119 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 161
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 162 VTTSSL------------------------SGNKDSIYSLAMNQMGTVIVSGSTEKVLRV 197
Query: 245 WDPRSGSKTMKLKGHTDNIRALLLD 269
WDPR+ +K MKLKGHTDN++ALLL+
Sbjct: 198 WDPRTCAKLMKLKGHTDNVKALLLN 222
>UniRef100_Q8BRM0 Mus musculus 10 days neonate cortex cDNA, RIKEN full-length
enriched library, clone:A830061E16 product:hypothetical
Trp-Asp (WD) repeats profile/Trp-Asp (WD) repeats
circular profile/G-protein beta WD-40 repeats containing
protein, full insert sequence [Mus musculus]
Length = 676
Score = 161 bits (407), Expect = 2e-38
Identities = 98/264 (37%), Positives = 144/264 (54%), Gaps = 49/264 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 2 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 56
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 57 WSVNQHKQDPYIASMEHHTDWVNDVVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 115
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 116 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 158
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 159 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 194
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WDPR+ +K MKLKGHTDN++ALLL
Sbjct: 195 WDPRTCAKLMKLKGHTDNVKALLL 218
>UniRef100_Q80XI0 8430408H12Rik protein [Mus musculus]
Length = 662
Score = 161 bits (407), Expect = 2e-38
Identities = 98/264 (37%), Positives = 144/264 (54%), Gaps = 49/264 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 2 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 56
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 57 WSVNQHKQDPYIASMEHHTDWVNDVVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 115
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 116 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 158
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 159 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 194
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WDPR+ +K MKLKGHTDN++ALLL
Sbjct: 195 WDPRTCAKLMKLKGHTDNVKALLL 218
>UniRef100_Q80TD4 MKIAA1449 protein [Mus musculus]
Length = 675
Score = 161 bits (407), Expect = 2e-38
Identities = 98/264 (37%), Positives = 144/264 (54%), Gaps = 49/264 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 1 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 55
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 56 WSVNQHKQDPYIASMEHHTDWVNDVVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 114
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 115 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 157
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 158 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 193
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WDPR+ +K MKLKGHTDN++ALLL
Sbjct: 194 WDPRTCAKLMKLKGHTDNVKALLL 217
>UniRef100_Q8BH57 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:4932416M21 product:hypothetical Trp-Asp
(WD) repeats profile/Trp-Asp (WD) repeats circular
profile/G-protein beta WD-40 repeats containing protein,
full insert sequence (Mus musculus adult male cecum
cDNA, RIKEN full-length enriched library,
clone:9130204A17 product:hypothetical Trp-Asp (WD)
repeats profile/Trp-Asp (WD) repeats circular
profile/G-protein beta WD-40 repeats containing protein,
full insert sequence) (Mus musculus adult male urinary
bladder cDNA, RIKEN full-length enriched library,
clone:9530008B17 product:hypothetical Trp-Asp (WD)
repeats profile/Trp-Asp (WD) repeats circular profile/G-
protein beta WD-40 repeats containing protein, full
insert sequence) [Mus musculus]
Length = 676
Score = 161 bits (407), Expect = 2e-38
Identities = 98/264 (37%), Positives = 144/264 (54%), Gaps = 49/264 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 2 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 56
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 57 WSVNQHKQDPYIASMEHHTDWVNDVVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 115
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 116 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 158
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 159 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 194
Query: 245 WDPRSGSKTMKLKGHTDNIRALLL 268
WDPR+ +K MKLKGHTDN++ALLL
Sbjct: 195 WDPRTCAKLMKLKGHTDNVKALLL 218
>UniRef100_Q5RAW8 Hypothetical protein DKFZp459F151 [Pongo pygmaeus]
Length = 677
Score = 161 bits (407), Expect = 2e-38
Identities = 98/265 (36%), Positives = 145/265 (53%), Gaps = 49/265 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 2 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 56
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 57 WSVNQHKQDPYIASMEHHTDWVNDIVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 115
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 116 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 158
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 159 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 194
Query: 245 WDPRSGSKTMKLKGHTDNIRALLLD 269
WDPR+ +K MKLKGHTDN++ALLL+
Sbjct: 195 WDPRTCAKLMKLKGHTDNVKALLLN 219
>UniRef100_Q9P279 KIAA1449 protein [Homo sapiens]
Length = 680
Score = 161 bits (407), Expect = 2e-38
Identities = 98/265 (36%), Positives = 145/265 (53%), Gaps = 49/265 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 5 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 59
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 60 WSVNQHKQDPYIASMEHHTDWVNDIVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 118
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 119 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 161
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 162 VTTSSL------------------------SGNKDSIYSLAMNQLGTIIVSGSTEKVLRV 197
Query: 245 WDPRSGSKTMKLKGHTDNIRALLLD 269
WDPR+ +K MKLKGHTDN++ALLL+
Sbjct: 198 WDPRTCAKLMKLKGHTDNVKALLLN 222
>UniRef100_Q6NU92 LOC414700 protein [Xenopus laevis]
Length = 410
Score = 159 bits (403), Expect = 6e-38
Identities = 98/265 (36%), Positives = 143/265 (52%), Gaps = 49/265 (18%)
Query: 7 AGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKR 66
A + +T R++ +++YV+ D + + G+N L + + LFT RD ++
Sbjct: 2 AAHHRQNTAGRRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRDSIIRI 56
Query: 67 WALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQ 124
W + + A+ E H DWVND VL TL+S SSDTT+K W+A G C TLR
Sbjct: 57 WNVNQHKQDPYIASMEHHTDWVNDIVLCCNGKTLISASSDTTVKVWNAHK-GFCMSTLRT 115
Query: 125 HSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNS 184
H DYV LA + K+ +VAS GL ++F+WD+ +T + S N+
Sbjct: 116 HKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALTASNNT 158
Query: 185 LPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRV 244
+ +SL G+K+S+Y+LAMN+ GT++VSG TEKV+RV
Sbjct: 159 VTTSSL------------------------SGNKDSIYSLAMNQMGTVIVSGSTEKVLRV 194
Query: 245 WDPRSGSKTMKLKGHTDNIRALLLD 269
WDPR+ K MKLKGHTDN++ALLL+
Sbjct: 195 WDPRTCQKLMKLKGHTDNVKALLLN 219
>UniRef100_Q6PFM9 Zgc:66229 [Brachydanio rerio]
Length = 677
Score = 159 bits (402), Expect = 7e-38
Identities = 97/271 (35%), Positives = 147/271 (53%), Gaps = 57/271 (21%)
Query: 1 MHRVGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSR 60
+HR +AG R++ +++YV+ D + + G+N L + + LFT R
Sbjct: 4 LHRQNAAG--------RRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGR 50
Query: 61 DGRLKRWALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTC 118
D ++ W++ + A+ E H DWVND +L TL+S SSDTT+K W+A G C
Sbjct: 51 DSIIRIWSVNQHKQDPYIASMEHHTDWVNDIILCCNGKTLISASSDTTVKVWNAHK-GFC 109
Query: 119 TRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGI 178
TLR H DYV LA + K+ +VAS GL ++F+WD+ +T +
Sbjct: 110 MSTLRTHKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTAL 152
Query: 179 NGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGT 238
S N++ +SL G+K+S+Y+LAMN+ GT+++SG T
Sbjct: 153 TASNNTVTTSSL------------------------SGNKDSIYSLAMNQTGTVIISGST 188
Query: 239 EKVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
EKV+RVWDPR+ +K MKLKGHTDN+++LLL+
Sbjct: 189 EKVLRVWDPRTCAKLMKLKGHTDNVKSLLLN 219
>UniRef100_UPI0000360CB9 UPI0000360CB9 UniRef100 entry
Length = 680
Score = 157 bits (398), Expect = 2e-37
Identities = 99/270 (36%), Positives = 146/270 (53%), Gaps = 57/270 (21%)
Query: 2 HRVGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRD 61
HR +AG R++ +++YV+ D + + G+N L + + LFT RD
Sbjct: 8 HRQNAAG--------RRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRD 54
Query: 62 GRLKRWALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCT 119
++ W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C
Sbjct: 55 SIIRIWSVYQHKQDPYIASMEHHTDWVNDIVLCCNGKTLISASSDTTVKVWNAHK-GFCM 113
Query: 120 RTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGIN 179
TLR H DYV LA + K+ +VAS GL ++F+WD+ +T +
Sbjct: 114 STLRTHKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALT 156
Query: 180 GSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTE 239
S N++ +SL G+K+S+Y+LAMN+ GT++VSG TE
Sbjct: 157 ASNNTVTTSSL------------------------SGNKDSIYSLAMNQMGTVIVSGSTE 192
Query: 240 KVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
KV+RVWDPR+ +K MKLKGHTDN+++LLL+
Sbjct: 193 KVLRVWDPRTCAKLMKLKGHTDNVKSLLLN 222
>UniRef100_UPI0000337947 UPI0000337947 UniRef100 entry
Length = 707
Score = 157 bits (398), Expect = 2e-37
Identities = 99/270 (36%), Positives = 146/270 (53%), Gaps = 57/270 (21%)
Query: 2 HRVGSAGNTNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRD 61
HR +AG R++ +++YV+ D + + G+N L + + LFT RD
Sbjct: 8 HRQNAAG--------RRKVQVSYVIRDEVEKYNRNGVNALQL-----DPALNRLFTAGRD 54
Query: 62 GRLKRWALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCT 119
++ W++ + A+ E H DWVND VL TL+S SSDTT+K W+A G C
Sbjct: 55 SIIRIWSVYQHKQDPYIASMEHHTDWVNDIVLCCNGKTLISASSDTTVKVWNAHK-GFCM 113
Query: 120 RTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGIN 179
TLR H DYV LA + K+ +VAS GL ++F+WD+ +T +
Sbjct: 114 STLRTHKDYVKALAYA-KDKELVASAGLDRQIFLWDV----------------NTLTALT 156
Query: 180 GSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTE 239
S N++ +SL G+K+S+Y+LAMN+ GT++VSG TE
Sbjct: 157 ASNNTVTTSSL------------------------SGNKDSIYSLAMNQMGTVIVSGSTE 192
Query: 240 KVVRVWDPRSGSKTMKLKGHTDNIRALLLD 269
KV+RVWDPR+ +K MKLKGHTDN+++LLL+
Sbjct: 193 KVLRVWDPRTCAKLMKLKGHTDNVKSLLLN 222
>UniRef100_Q922Z9 8430408H12Rik protein [Mus musculus]
Length = 635
Score = 151 bits (382), Expect = 2e-35
Identities = 89/216 (41%), Positives = 122/216 (56%), Gaps = 44/216 (20%)
Query: 55 LFTGSRDGRLKRWALGEDAAT-CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDA 112
LFT RD ++ W++ + A+ E H DWVND VL TL+S SSDTT+K W+A
Sbjct: 4 LFTAGRDSIIRIWSVNQHKQDPYIASMEHHTDWVNDVVLCCNGKTLISASSDTTVKVWNA 63
Query: 113 FSTGTCTRTLRQHSDYVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDD 172
G C TLR H DYV LA + K+ +VAS GL ++F+WD+
Sbjct: 64 HK-GFCMSTLRTHKDYVKALAYA-KDKELVASAGLDRQIFLWDV---------------- 105
Query: 173 DTSNGINGSGNSLPLTSLRSISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTL 232
+T + S N++ +SL G+K+S+Y+LAMN+ GT+
Sbjct: 106 NTLTALTASNNTVTTSSL------------------------SGNKDSIYSLAMNQLGTI 141
Query: 233 LVSGGTEKVVRVWDPRSGSKTMKLKGHTDNIRALLL 268
+VSG TEKV+RVWDPR+ +K MKLKGHTDN++ALLL
Sbjct: 142 IVSGSTEKVLRVWDPRTCAKLMKLKGHTDNVKALLL 177
>UniRef100_Q7PXD9 ENSANGP00000009742 [Anopheles gambiae str. PEST]
Length = 676
Score = 117 bits (293), Expect = 3e-25
Identities = 80/262 (30%), Positives = 137/262 (51%), Gaps = 22/262 (8%)
Query: 17 RKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWALGEDAAT- 75
RK+ ++ +V+ D+++ +H G+N L + + L++ RDG ++ W + +++
Sbjct: 11 RKKMQVAFVIRDAEEKRHRNGVNALQL-----DSINGRLYSAGRDGIIRLWNSTQTSSSE 65
Query: 76 -CSATFESHVDWVNDAVLV-GDSTLVSCSSDTTLKTWDAFSTGTCTRTLRQHSDYVTCLA 133
+ E H DWVND VL G L+S S DTT+K W+A G C TLR H DYV LA
Sbjct: 66 PYIQSMEHHNDWVNDIVLCCGGRNLISASCDTTVKVWNAHK-GFCMSTLRTHRDYVQALA 124
Query: 134 ASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPLTSLR-- 191
+ K+ VAS GL +F+WD+ A A + + T++ I+GS +S+ ++
Sbjct: 125 YA-KDREQVASAGLDKAIFLWDVNTLTALTA----SNNTVTTSSISGSKDSIYSLAMNPS 179
Query: 192 -----SISSSNSISVHTTQNQGYIPIAAKGHKESVYALAMNEGGTLLVSGGTEKVVRVWD 246
S S+ N++ + + I KGH E+V AL ++E GT +VSG ++ +++W
Sbjct: 180 GTIIVSGSTENTLRIWDPRTCNKIA-KLKGHTENVKALIVSEDGTQVVSGSSDGKIKLWS 238
Query: 247 PRSGSKTMKLKGHTDNIRALLL 268
+ H++ + LL+
Sbjct: 239 IGQQRCIQTISVHSEGVWCLLM 260
>UniRef100_UPI000007A4E1 UPI000007A4E1 UniRef100 entry
Length = 680
Score = 110 bits (275), Expect = 4e-23
Identities = 79/269 (29%), Positives = 136/269 (50%), Gaps = 23/269 (8%)
Query: 10 TNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWAL 69
T+ + + RK+ ++++V+ D+++ +H G+N L + + L++ RD ++ W
Sbjct: 3 THKTCQARKKMQVSFVIRDAEEKQHRNGVNALQL-----DANNGKLYSAGRDAIIRVWNT 57
Query: 70 GEDAAT-CSATFESHVDWVNDAVLVGDS-TLVSCSSDTTLKTWDAFSTGTCTRTLRQHSD 127
D++ + E H DWVND VL + L+S S DTT+K W+A G C TLR H D
Sbjct: 58 RTDSSEKYIQSMEHHNDWVNDIVLCCNGRNLISASCDTTVKVWNA-QKGFCMSTLRTHRD 116
Query: 128 YVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPL 187
YV LA + K+ VAS GL +F+WD+ A A + + T++ + GS +S+
Sbjct: 117 YVQALAYA-KDREQVASAGLDKAIFLWDVNTLTALTA----SNNTVTTSSLTGSKDSI-- 169
Query: 188 TSLRSISSSNSISVHTTQNQGYI--------PIAAKGHKESVYALAMNEGGTLLVSGGTE 239
SL S I +T+N I + +GH E+V L ++ G +VSG ++
Sbjct: 170 YSLAMNPSGTVIVSGSTENILRIWDPRTCMRRMKLRGHTENVRCLVVSPDGNQVVSGSSD 229
Query: 240 KVVRVWDPRSGSKTMKLKGHTDNIRALLL 268
++VW+ + H + + +LL+
Sbjct: 230 GTIKVWNLGQQRCVQTIHVHKEGVWSLLM 258
>UniRef100_Q8MKZ5 CG9062-PB [Drosophila melanogaster]
Length = 668
Score = 110 bits (275), Expect = 4e-23
Identities = 79/269 (29%), Positives = 136/269 (50%), Gaps = 23/269 (8%)
Query: 10 TNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWAL 69
T+ + + RK+ ++++V+ D+++ +H G+N L + + L++ RD ++ W
Sbjct: 3 THKTCQARKKMQVSFVIRDAEEKQHRNGVNALQL-----DANNGKLYSAGRDAIIRVWNT 57
Query: 70 GEDAAT-CSATFESHVDWVNDAVLVGDS-TLVSCSSDTTLKTWDAFSTGTCTRTLRQHSD 127
D++ + E H DWVND VL + L+S S DTT+K W+A G C TLR H D
Sbjct: 58 RTDSSEKYIQSMEHHNDWVNDIVLCCNGRNLISASCDTTVKVWNA-QKGFCMSTLRTHRD 116
Query: 128 YVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPL 187
YV LA + K+ VAS GL +F+WD+ A A + + T++ + GS +S+
Sbjct: 117 YVQALAYA-KDREQVASAGLDKAIFLWDVNTLTALTA----SNNTVTTSSLTGSKDSI-- 169
Query: 188 TSLRSISSSNSISVHTTQNQGYI--------PIAAKGHKESVYALAMNEGGTLLVSGGTE 239
SL S I +T+N I + +GH E+V L ++ G +VSG ++
Sbjct: 170 YSLAMNPSGTVIVSGSTENILRIWDPRTCMRRMKLRGHTENVRCLVVSPDGNQVVSGSSD 229
Query: 240 KVVRVWDPRSGSKTMKLKGHTDNIRALLL 268
++VW+ + H + + +LL+
Sbjct: 230 GTIKVWNLGQQRCVQTIHVHKEGVWSLLM 258
>UniRef100_Q8IGK7 RE72568p [Drosophila melanogaster]
Length = 668
Score = 110 bits (275), Expect = 4e-23
Identities = 79/269 (29%), Positives = 136/269 (50%), Gaps = 23/269 (8%)
Query: 10 TNSSTRPRKEKRLTYVLNDSDDTKHCAGINCLAVLKSAVSDGSDYLFTGSRDGRLKRWAL 69
T+ + + RK+ ++++V+ D+++ +H G+N L + + L++ RD ++ W
Sbjct: 3 THKTCQARKKMQVSFVIRDAEEKQHRNGVNALQL-----DANNGKLYSAGRDAIIRVWNT 57
Query: 70 GEDAAT-CSATFESHVDWVNDAVLVGDS-TLVSCSSDTTLKTWDAFSTGTCTRTLRQHSD 127
D++ + E H DWVND VL + L+S S DTT+K W+A G C TLR H D
Sbjct: 58 RTDSSEKYIQSMEHHNDWVNDIVLCCNGRNLISASCDTTVKVWNA-QKGFCMSTLRTHRD 116
Query: 128 YVTCLAASEKNSNIVASGGLGGEVFVWDLEAAHASAAKCIDATDDDTSNGINGSGNSLPL 187
YV LA + K+ VAS GL +F+WD+ A A + + T++ + GS +S+
Sbjct: 117 YVQALAYA-KDREQVASAGLDKAIFLWDVNTLTALTA----SNNTVTTSSLTGSKDSI-- 169
Query: 188 TSLRSISSSNSISVHTTQNQGYI--------PIAAKGHKESVYALAMNEGGTLLVSGGTE 239
SL S I +T+N I + +GH E+V L ++ G +VSG ++
Sbjct: 170 YSLAMNPSGTVIVSGSTENILRIWDPRTCMRRMKLRGHTENVRCLVVSPDGNQVVSGSSD 229
Query: 240 KVVRVWDPRSGSKTMKLKGHTDNIRALLL 268
++VW+ + H + + +LL+
Sbjct: 230 GTIKVWNLGQQRCVQTIHVHKEGVWSLLM 258
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.127 0.372
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 463,844,862
Number of Sequences: 2790947
Number of extensions: 18644495
Number of successful extensions: 64158
Number of sequences better than 10.0: 3828
Number of HSP's better than 10.0 without gapping: 1896
Number of HSP's successfully gapped in prelim test: 1970
Number of HSP's that attempted gapping in prelim test: 45797
Number of HSP's gapped (non-prelim): 14528
length of query: 271
length of database: 848,049,833
effective HSP length: 125
effective length of query: 146
effective length of database: 499,181,458
effective search space: 72880492868
effective search space used: 72880492868
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0062.9


