

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0055.11
(605 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
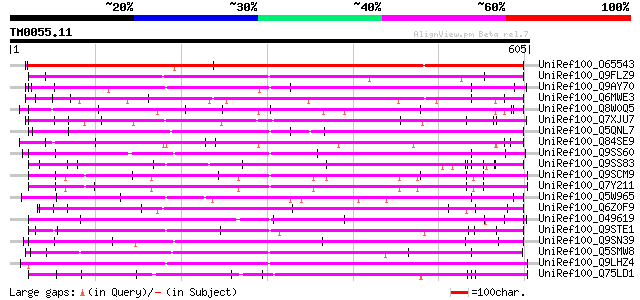
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O65543 Hypothetical protein F6I18.20 [Arabidopsis thal... 568 e-160
UniRef100_Q9FLZ9 Selenium-binding protein-like [Arabidopsis thal... 371 e-101
UniRef100_Q9AY70 Hypothetical protein OSJNBa0091J19.16 [Oryza sa... 367 e-100
UniRef100_Q6MWE3 B1358B12.23 protein [Oryza sativa] 365 3e-99
UniRef100_Q8W0Q5 Putative vegetative storage protein [Sorghum bi... 361 4e-98
UniRef100_Q7XJU7 OSJNBa0016O02.23 protein [Oryza sativa] 361 4e-98
UniRef100_Q5QNL7 Pentatricopeptide repeat protein-like [Oryza sa... 358 3e-97
UniRef100_Q84SE9 P0020E09.15 protein [Oryza sativa] 356 1e-96
UniRef100_Q9SS60 T12J13.14 protein [Arabidopsis thaliana] 352 1e-95
UniRef100_Q9SS83 MZB10.7 protein [Arabidopsis thaliana] 351 3e-95
UniRef100_Q9SCM9 Hypothetical protein T8H10.30 [Arabidopsis thal... 351 3e-95
UniRef100_Q7Y211 Hypothetical protein At3g57430 [Arabidopsis tha... 351 3e-95
UniRef100_Q5W965 PPR868-14 [Physcomitrella patens] 348 2e-94
UniRef100_Q6Z0F9 PPR-repeat protein-like [Oryza sativa] 348 3e-94
UniRef100_O49619 Hypothetical protein M4E13.180 [Arabidopsis tha... 347 4e-94
UniRef100_Q9STE1 Hypothetical protein T6K22.30 [Arabidopsis thal... 347 6e-94
UniRef100_Q9SN39 Hypothetical protein F28A21.160 [Arabidopsis th... 347 8e-94
UniRef100_Q5SMW8 Pentatricopeptide (PPR) repeat-containing prote... 344 5e-93
UniRef100_Q9LHZ4 Pentatricopeptide (PPR) repeat-containing prote... 344 5e-93
UniRef100_Q75LD1 Putative pentatricopeptide repeat domain contia... 340 9e-92
>UniRef100_O65543 Hypothetical protein F6I18.20 [Arabidopsis thaliana]
Length = 613
Score = 568 bits (1465), Expect = e-160
Identities = 291/584 (49%), Positives = 404/584 (68%), Gaps = 7/584 (1%)
Query: 21 QIKSLISQGLYHQTLELFT-QLHFSAPHSISSVLPSVIKACS-SLHSHTLGTLLHGLALT 78
++K L+S Y + L L+ ++H + +++LPSVIKAC+ LG LH L L
Sbjct: 16 KLKGLVSDQFYDEALRLYKLKIHSLGTNGFTAILPSVIKACAFQQEPFLLGAQLHCLCLK 75
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G+ D VVSNSLIS+YAKFS + R+VFD M HRD +S+ S+IN+ Q+G L EA+++
Sbjct: 76 AGADCDTVVSNSLISMYAKFSRKYAVRKVFDEMLHRDTVSYCSIINSCCQDGLLYEAMKL 135
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRK-MGSRIGRQIHALVVVDGRVEHSVVLSTALVDFY 197
+K +Y G +PK EL+AS+++LC R S++ R HALV+VD R++ SV+LSTALVD Y
Sbjct: 136 IKEMYFYGFIPKSELVASLLALCTRMGSSSKVARMFHALVLVDERMQESVLLSTALVDMY 195
Query: 198 FRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLI 257
+ DD A VFD MEVKNEVSWTAMISGC+AN +Y+ FRAMQ E + PNRVTL+
Sbjct: 196 LKFDDHAAAFHVFDQMEVKNEVSWTAMISGCVANQNYEMGVDLFRAMQRENLRPNRVTLL 255
Query: 258 ALLPACAKPGFVKH-GKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSS 316
++LPAC + + KEIHG++FRHG + ++A + MYC+ G ++ L+ +FE S
Sbjct: 256 SVLPACVELNYGSSLVKEIHGFSFRHGCHADERLTAAFMTMYCRCG-NVSLSRVLFETSK 314
Query: 317 FRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCG 376
RDVV+WSS+I Y++ GD + + L N+M E E N VTLLA++SACTN + L
Sbjct: 315 VRDVVMWSSMISGYAETGDCSEVMNLLNQMRKEGIEANSVTLLAIVSACTNSTLLSFAST 374
Query: 377 LHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGH 436
+H ILK G I L NAL++MYAKCG L + ++F E+ +D +SWS+MI+AYGLHGH
Sbjct: 375 VHSQILKCGFMSHILLGNALIDMYAKCGSLSAAREVFYELTEKDLVSWSSMINAYGLHGH 434
Query: 437 GEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYA 496
G +AL++F M + G + D + FLA+LSACNHAGLV E Q +F Q Y +P+T+EHYA
Sbjct: 435 GSEALEIFKGMIKGGHEVDDMAFLAILSACNHAGLVEEAQTIFTQA-GKYHMPVTLEHYA 493
Query: 497 CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIA-EMLGPQLIRSEP 555
C ++LLGR GK++DA E+ MPMKPS RIWSSL+SAC+ HGRLD+A +++ +L++SEP
Sbjct: 494 CYINLLGRFGKIDDAFEVTINMPMKPSARIWSSLLSACETHGRLDVAGKIIANELMKSEP 553
Query: 556 DNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
DN ANY LL+ I+ E G++ E+V M+ ++L KCYGFS+IE
Sbjct: 554 DNPANYVLLSKIHTESGNYHAAEEVRRVMQRRKLNKCYGFSKIE 597
Score = 72.4 bits (176), Expect = 4e-11
Identities = 57/225 (25%), Positives = 100/225 (44%), Gaps = 12/225 (5%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
S I G + + L Q+ + S L +++ AC++ + + +H L
Sbjct: 322 SSMISGYAETGDCSEVMNLLNQMRKEGIEANSVTLLAIVSACTNSTLLSFASTVHSQILK 381
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G S ++ N+LI +YAK + +AR+VF + +D +SW+SMINAY +G EALE+
Sbjct: 382 CGFMSHILLGNALIDMYAKCGSLSAAREVFYELTEKDLVSWSSMINAYGLHGHGSEALEI 441
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLS-----TAL 193
KG+ G +++S C + + + + G+ V L L
Sbjct: 442 FKGMIKGGHEVDDMAFLAILSACNH---AGLVEEAQTIFTQAGKYHMPVTLEHYACYINL 498
Query: 194 VDFYFRCDDALMASRVFDGMEVKNEVS-WTAMISGCIANLDYDAA 237
+ + + DDA V M +K W++++S C + D A
Sbjct: 499 LGRFGKIDDAF---EVTINMPMKPSARIWSSLLSACETHGRLDVA 540
>UniRef100_Q9FLZ9 Selenium-binding protein-like [Arabidopsis thaliana]
Length = 677
Score = 371 bits (953), Expect = e-101
Identities = 205/584 (35%), Positives = 323/584 (55%), Gaps = 8/584 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISS--VLPSVIKACSSLHSHTLGTLLHGLALTT 79
I+ + +GLYH + +F ++ + P V KA L S LG ++HG L +
Sbjct: 87 IRMYVREGLYHDAISVFIRMVSEGVKCVPDGYTYPFVAKAAGELKSMKLGLVVHGRILRS 146
Query: 80 GSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
D V N+L+++Y F ++ AR VFD M +RD ISWN+MI+ Y +NG + +AL M
Sbjct: 147 WFGRDKYVQNALLAMYMNFGKVEMARDVFDVMKNRDVISWNTMISGYYRNGYMNDALMMF 206
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ + + SM+ +CG +GR +H LV + R+ + + ALV+ Y +
Sbjct: 207 DWMVNESVDLDHATIVSMLPVCGHLKDLEMGRNVHKLVE-EKRLGDKIEVKNALVNMYLK 265
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C A VFD ME ++ ++WT MI+G + D + A R MQ EGV PN VT+ +L
Sbjct: 266 CGRMDEARFVFDRMERRDVITWTCMINGYTEDGDVENALELCRLMQFEGVRPNAVTIASL 325
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
+ C V GK +HG+A R + S I ++LI+MY + + + L ++F +S
Sbjct: 326 VSVCGDALKVNDGKCLHGWAVRQQVYSDIIIETSLISMYAKC-KRVDLCFRVFSGASKYH 384
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
WS+II Q AL LF +M E+ EPN TL +++ A L+ L+ +H
Sbjct: 385 TGPWSAIIAGCVQNELVSDALGLFKRMRREDVEPNIATLNSLLPAYAALADLRQAMNIHC 444
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQ----NRDSISWSTMISAYGLHG 435
++ K G S+ + L+++Y+KCG LE + +IF +Q ++D + W +IS YG+HG
Sbjct: 445 YLTKTGFMSSLDAATGLVHVYSKCGTLESAHKIFNGIQEKHKSKDVVLWGALISGYGMHG 504
Query: 436 HGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHY 495
G AL++F EM GV P+ +TF + L+AC+H+GLV EG +F+ + Y+ HY
Sbjct: 505 DGHNALQVFMEMVRSGVTPNEITFTSALNACSHSGLVEEGLTLFRFMLEHYKTLARSNHY 564
Query: 496 ACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEP 555
C+VDLLGR+G++++A ++ T+P +P++ +W +L++AC H + + EM +L EP
Sbjct: 565 TCIVDLLGRAGRLDEAYNLITTIPFEPTSTVWGALLAACVTHENVQLGEMAANKLFELEP 624
Query: 556 DNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
+N NY LL IYA G W E+V M+ L+K G S IE
Sbjct: 625 ENTGNYVLLANIYAALGRWKDMEKVRSMMENVGLRKKPGHSTIE 668
Score = 168 bits (426), Expect = 4e-40
Identities = 130/519 (25%), Positives = 234/519 (45%), Gaps = 30/519 (5%)
Query: 42 HFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDI 101
HF+A SIS +LH H + TG + ++L YA I
Sbjct: 24 HFAATQSISKT--------KALHCHVI----------TGGRVSGHILSTLSVTYALCGHI 65
Query: 102 QSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGL--VPKPELLASMVS 159
AR++F+ MP +S+N +I Y++ G +A+ + + G+ VP +
Sbjct: 66 TYARKLFEEMPQSSLLSYNIVIRMYVREGLYHDAISVFIRMVSEGVKCVPDGYTYPFVAK 125
Query: 160 LCGRKMGSRIGRQIHALVVVD--GRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKN 217
G ++G +H ++ GR ++ + AL+ Y MA VFD M+ ++
Sbjct: 126 AAGELKSMKLGLVVHGRILRSWFGRDKY---VQNALLAMYMNFGKVEMARDVFDVMKNRD 182
Query: 218 EVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHG 277
+SW MISG N + A F M E VD + T++++LP C ++ G+ +H
Sbjct: 183 VISWNTMISGYYRNGYMNDALMMFDWMVNESVDLDHATIVSMLPVCGHLKDLEMGRNVHK 242
Query: 278 YAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSY 337
+ +AL+NMY + G + A +F+R RDV+ W+ +I Y++ GD
Sbjct: 243 LVEEKRLGDKIEVKNALVNMYLKCGR-MDEARFVFDRMERRDVITWTCMINGYTEDGDVE 301
Query: 338 KALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALM 397
AL L M E PN VT+ +++S C + + G LHG+ ++ + I + +L+
Sbjct: 302 NALELCRLMQFEGVRPNAVTIASLVSVCGDALKVNDGKCLHGWAVRQQVYSDIIIETSLI 361
Query: 398 NMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDAL 457
+MYAKC ++ ++F + WS +I+ + AL LF M+ V+P+
Sbjct: 362 SMYAKCKRVDLCFRVFSGASKYHTGPWSAIIAGCVQNELVSDALGLFKRMRREDVEPNIA 421
Query: 458 TFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRT 517
T ++L A + + + + +++ LV + + G +E A +I
Sbjct: 422 TLNSLLPAYAALADLRQAMNIHCYLTKT-GFMSSLDAATGLVHVYSKCGTLESAHKIFNG 480
Query: 518 MPMKPSTR---IWSSLVSACKLHGRLDIAEMLGPQLIRS 553
+ K ++ +W +L+S +HG A + +++RS
Sbjct: 481 IQEKHKSKDVVLWGALISGYGMHGDGHNALQVFMEMVRS 519
>UniRef100_Q9AY70 Hypothetical protein OSJNBa0091J19.16 [Oryza sativa]
Length = 843
Score = 367 bits (941), Expect = e-100
Identities = 195/577 (33%), Positives = 332/577 (56%), Gaps = 2/577 (0%)
Query: 26 ISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDP 85
+ G +ELF + S + L + ++ G LH LA+ G S+
Sbjct: 223 VKAGSVSSAVELFGDMRASGCEPNFATLACFLSVSATESDLFFGVQLHTLAVKYGLESEV 282
Query: 86 VVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLL 145
V+N+L+S+YAK + ++F MP D ++WN MI+ +QNG +++AL + +
Sbjct: 283 AVANTLVSMYAKCKCLDDGWKLFGLMPRDDLVTWNGMISGCVQNGFVDQALLLFCDMQKS 342
Query: 146 GLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALM 205
G+ P L S++ G G+++H +V + V V L +ALVD YF+C M
Sbjct: 343 GIRPDSVTLVSLLPALTDLNGFNQGKELHGYIVRNC-VHMDVFLVSALVDIYFKCRAVRM 401
Query: 206 ASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAK 265
A V+D + + V + MISG + N A FR + +G+ PN V + ++LPACA
Sbjct: 402 AQSVYDSSKAIDVVIGSTMISGYVLNGMSQEAVKMFRYLLEQGIRPNAVAIASVLPACAS 461
Query: 266 PGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSS 325
+K G+E+H YA ++ E SAL++MY + G L L+ IF + S +D V W+S
Sbjct: 462 MAAMKLGQELHSYALKNAYEGRCYVESALMDMYAKCGR-LDLSHYIFSKISAKDEVTWNS 520
Query: 326 IIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFG 385
+I S++Q G+ +AL LF +M E + + VT+ +V+SAC +L ++ +G +HG ++K
Sbjct: 521 MISSFAQNGEPEEALNLFREMCMEGVKYSNVTISSVLSACASLPAIYYGKEIHGVVIKGP 580
Query: 386 LSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFS 445
+ + +AL++MY KCG LE + ++F M ++ +SW+++I++YG +G ++++ L
Sbjct: 581 IRADLFAESALIDMYGKCGNLEWAHRVFESMPEKNEVSWNSIIASYGAYGLVKESVSLLR 640
Query: 446 EMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRS 505
M+E G K D +TFLA++SAC HAG V EG ++F+ + +Y+I +EH+AC+VDL R+
Sbjct: 641 HMQEEGFKADHVTFLALVSACAHAGQVQEGLRLFRCMTEEYQIAPRMEHFACMVDLYSRA 700
Query: 506 GKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLN 565
GK++ A+E++ MP KP IW +L+ AC++H +++AE+ +L + +P N+ Y L++
Sbjct: 701 GKLDKAMELIVDMPFKPDAGIWGALLHACRVHRNVELAEIASQELFKLDPHNSGYYVLMS 760
Query: 566 MIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGN 602
I A G W +V MK +++K G+S ++ N
Sbjct: 761 NINAVAGRWDGVSKVRRLMKDTKVQKIPGYSWVDVNN 797
Score = 284 bits (726), Expect = 6e-75
Identities = 184/571 (32%), Positives = 302/571 (52%), Gaps = 15/571 (2%)
Query: 22 IKSLISQGLYHQTLELFTQL--HFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTT 79
I+ L G Y L + ++ H SAP S P V+K+C++L + LG L+H A T
Sbjct: 116 IRGLTMAGDYRSALLFYLKMWAHPSAPLPDSHTFPYVVKSCAALGAIALGRLVHRTARTL 175
Query: 80 GSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
G D V ++LI +YA + ARQVFD M RD + WN M++ Y++ G + A+E+
Sbjct: 176 GLDGDMFVGSALIKMYANGGLLWDARQVFDGMAERDCVLWNVMMDGYVKAGSVSSAVELF 235
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ G P LA +S+ + G Q+H L V G +E V ++ LV Y +
Sbjct: 236 GDMRASGCEPNFATLACFLSVSATESDLFFGVQLHTLAVKYG-LESEVAVANTLVSMYAK 294
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C ++F M + V+W MISGC+ N D A F MQ G+ P+ VTL++L
Sbjct: 295 CKCLDDGWKLFGLMPRDDLVTWNGMISGCVQNGFVDQALLLFCDMQKSGIRPDSVTLVSL 354
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
LPA GKE+HGY R+ + SAL+++Y + ++ +A+ +++ S D
Sbjct: 355 LPALTDLNGFNQGKELHGYIVRNCVHMDVFLVSALVDIYFKC-RAVRMAQSVYDSSKAID 413
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
VV+ S++I Y G S +A+ +F +L + PN V + +V+ AC +++++K G LH
Sbjct: 414 VVIGSTMISGYVLNGMSQEAVKMFRYLLEQGIRPNAVAIASVLPACASMAAMKLGQELHS 473
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
+ LK + +ALM+MYAKCG L+ S IF ++ +D ++W++MIS++ +G E+
Sbjct: 474 YALKNAYEGRCYVESALMDMYAKCGRLDLSHYIFSKISAKDEVTWNSMISSFAQNGEPEE 533
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYA--C 497
AL LF EM GVK +T +VLSAC + G+++ V + P+ + +A
Sbjct: 534 ALNLFREMCMEGVKYSNVTISSVLSACASLPAIYYGKEIHGVV---IKGPIRADLFAESA 590
Query: 498 LVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDN 557
L+D+ G+ G +E A + +MP K W+S++++ +G + + L + ++ E
Sbjct: 591 LIDMYGKCGNLEWAHRVFESMPEKNEVS-WNSIIASYGAYGLVKESVSL-LRHMQEEGFK 648
Query: 558 AANYTLLNMIYAEHGHWLHKEQVMEAMKLQR 588
A + T L ++ A H QV E ++L R
Sbjct: 649 ADHVTFLALVSA----CAHAGQVQEGLRLFR 675
Score = 179 bits (455), Expect = 2e-43
Identities = 130/493 (26%), Positives = 236/493 (47%), Gaps = 12/493 (2%)
Query: 53 LPSVIKACSSLHSHTLGTLLHGLALTTGSH-SDPVVSNSLISLYAKFSDIQSARQVFDTM 111
L +V++ C S +LG +HG A+T G H +D + L+ +Y + A VF ++
Sbjct: 42 LLAVLRGCVSPSHLSLGLQVHGRAVTAGLHATDTALQTRLVGMYVLARRFRDAVAVFSSL 101
Query: 112 PH---RDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE--LLASMVSLCGRKMG 166
P + WN +I G AL ++ P P+ +V C
Sbjct: 102 PRGAAACALPWNWLIRGLTMAGDYRSALLFYLKMWAHPSAPLPDSHTFPYVVKSCAALGA 161
Query: 167 SRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMIS 226
+GR +H G ++ + + +AL+ Y A +VFDGM ++ V W M+
Sbjct: 162 IALGRLVHRTARTLG-LDGDMFVGSALIKMYANGGLLWDARQVFDGMAERDCVLWNVMMD 220
Query: 227 GCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIES 286
G + +A F M+ G +PN TL L A + G ++H A ++G+ES
Sbjct: 221 GYVKAGSVSSAVELFGDMRASGCEPNFATLACFLSVSATESDLFFGVQLHTLAVKYGLES 280
Query: 287 SHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKM 346
++ L++MY + + L ++F D+V W+ +I Q G +AL LF M
Sbjct: 281 EVAVANTLVSMYAKC-KCLDDGWKLFGLMPRDDLVTWNGMISGCVQNGFVDQALLLFCDM 339
Query: 347 LTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCL 406
P+ VTL++++ A T+L+ G LHG+I++ + + L +AL+++Y KC +
Sbjct: 340 QKSGIRPDSVTLVSLLPALTDLNGFNQGKELHGYIVRNCVHMDVFLVSALVDIYFKCRAV 399
Query: 407 EGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSAC 466
+ ++ + D + STMIS Y L+G ++A+K+F + E+G++P+A+ +VL AC
Sbjct: 400 RMAQSVYDSSKAIDVVIGSTMISGYVLNGMSQEAVKMFRYLLEQGIRPNAVAIASVLPAC 459
Query: 467 NHAGLVTEGQQVFK-QVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTR 525
+ GQ++ + YE +E + L+D+ + G+++ + I + K
Sbjct: 460 ASMAAMKLGQELHSYALKNAYEGRCYVE--SALMDMYAKCGRLDLSHYIFSKISAKDEV- 516
Query: 526 IWSSLVSACKLHG 538
W+S++S+ +G
Sbjct: 517 TWNSMISSFAQNG 529
Score = 138 bits (347), Expect = 5e-31
Identities = 88/314 (28%), Positives = 156/314 (49%), Gaps = 10/314 (3%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
S I + G+ + +++F L + + SV+ AC+S+ + LG LH AL
Sbjct: 418 STMISGYVLNGMSQEAVKMFRYLLEQGIRPNAVAIASVLPACASMAAMKLGQELHSYALK 477
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
V ++L+ +YAK + + +F + +D ++WNSMI+++ QNG EEAL +
Sbjct: 478 NAYEGRCYVESALMDMYAKCGRLDLSHYIFSKISAKDEVTWNSMISSFAQNGEPEEALNL 537
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+ + + G+ ++S++S C G++IH VV+ G + + +AL+D Y
Sbjct: 538 FREMCMEGVKYSNVTISSVLSACASLPAIYYGKEIHG-VVIKGPIRADLFAESALIDMYG 596
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
+C + A RVF+ M KNEVSW ++I+ A + + R MQ EG + VT +A
Sbjct: 597 KCGNLEWAHRVFESMPEKNEVSWNSIIASYGAYGLVKESVSLLRHMQEEGFKADHVTFLA 656
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHI-----FSSALINMYCQSGESLHLAEQIFE 313
L+ ACA G V+ G + FR E I + ++++Y ++G+ E I +
Sbjct: 657 LVSACAHAGQVQEGLRL----FRCMTEEYQIAPRMEHFACMVDLYSRAGKLDKAMELIVD 712
Query: 314 RSSFRDVVLWSSII 327
D +W +++
Sbjct: 713 MPFKPDAGIWGALL 726
>UniRef100_Q6MWE3 B1358B12.23 protein [Oryza sativa]
Length = 918
Score = 365 bits (936), Expect = 3e-99
Identities = 199/554 (35%), Positives = 328/554 (58%), Gaps = 8/554 (1%)
Query: 50 SSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFD 109
S + S ++AC L ++GT LHG + G P V +SL S+Y K + AR +F
Sbjct: 215 SRTMESGLEACGVLGELSVGTCLHGFGVKAGVGHCPSVVSSLFSMYTKCDSTEDARILFP 274
Query: 110 TMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRI 169
+P +D +SW S+I AY + G E+A+E+ G+ GL P +++ +++ G R
Sbjct: 275 ELPEKDLVSWTSLIGAYCRAGHAEKAVELFLGMEESGLQPDEVVISCLLAGLGNDAKVRG 334
Query: 170 GRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISG-C 228
G+ HA +V SV++ AL+ Y +C +A+ VF + ++ SW++M+ C
Sbjct: 335 GKTFHA-AIVRRNFGDSVLIGNALISMYAKCKQVDIAATVFRMLHQRDTDSWSSMVVAYC 393
Query: 229 IANLDYDAAFARFRAMQVEGVDP---NRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIE 285
A LD +R MQ D + +LI+++ +C++ G ++ G+ H Y+ +H
Sbjct: 394 KAGLDLKC-LELYREMQFRDKDEFEYDTNSLISIISSCSRLGRLRLGQSAHCYSIKHLAG 452
Query: 286 SSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNK 345
+ ++ALI+MY + G + +A +IF +DVV WS++I SYS G S AL L+++
Sbjct: 453 ENSSVANALISMYGRCG-NFDVARKIFGMVKTKDVVTWSALISSYSHLGHSKDALLLYDQ 511
Query: 346 MLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGC 405
MLTE +PN TL++VIS+C NL++L+HG +H + GL +S+ AL++MY KCG
Sbjct: 512 MLTEGVKPNSATLVSVISSCANLAALEHGELIHSHVKDVGLECDLSICTALVDMYMKCGQ 571
Query: 406 LEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSA 465
L + ++F M RD ++W+ MIS YG+HG QALKLFS M+ VKP++LTFLA+LSA
Sbjct: 572 LGIARKMFDSMLERDVVTWNVMISGYGMHGEAIQALKLFSMMERGNVKPNSLTFLAILSA 631
Query: 466 CNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTR 525
C HAGLV +G+++F ++ +Y + ++HYAC+VDLLG+SG +++A +++ MP++P
Sbjct: 632 CCHAGLVDKGRELFTRME-EYSLEPNLKHYACMVDLLGKSGHLQEAEDVVSAMPIEPDGG 690
Query: 526 IWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMK 585
IW +L+ ACK+H ++ + + S+P+N Y L++ Y W E++ + MK
Sbjct: 691 IWGTLLGACKMHDNFEMGLRVAKKAFASDPENDGYYILMSNSYGSAEKWNEIEKLRDMMK 750
Query: 586 LQRLKKCYGFSRIE 599
++K G+S I+
Sbjct: 751 NHGVEKSIGWSTID 764
Score = 209 bits (532), Expect = 2e-52
Identities = 138/518 (26%), Positives = 252/518 (48%), Gaps = 14/518 (2%)
Query: 31 YHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTG---SHSDPVV 87
+ TL ++ S P V A + L + +G +H ++ G V
Sbjct: 88 FASTLSAHRRMRASGARPSRFTAPLVASAAAELGALPVGAAVHAYSVRFGLLEGDGSVAV 147
Query: 88 SNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEAL----EMLKGVY 143
++SL+ +YA+ ++ A ++FD MP RD ++W ++I+ + NG E L M++
Sbjct: 148 ASSLVYMYARCGSVRDAVRLFDEMPERDVVAWTAVISGCVCNGQCGEGLSYLVRMVRSAG 207
Query: 144 LLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDA 203
G P + S + CG +G +H V G V H + ++L Y +CD
Sbjct: 208 DGGARPNSRTMESGLEACGVLGELSVGTCLHGFGVKAG-VGHCPSVVSSLFSMYTKCDST 266
Query: 204 LMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPAC 263
A +F + K+ VSWT++I + A F M+ G+ P+ V + LL
Sbjct: 267 EDARILFPELPEKDLVSWTSLIGAYCRAGHAEKAVELFLGMEESGLQPDEVVISCLLAGL 326
Query: 264 AKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLW 323
V+ GK H R S + +ALI+MY + + + +A +F RD W
Sbjct: 327 GNDAKVRGGKTFHAAIVRRNFGDSVLIGNALISMYAKC-KQVDIAATVFRMLHQRDTDSW 385
Query: 324 SSIIGSYSQRGDSYKALTLFNKML---TEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
SS++ +Y + G K L L+ +M +E E + +L+++IS+C+ L L+ G H +
Sbjct: 386 SSMVVAYCKAGLDLKCLELYREMQFRDKDEFEYDTNSLISIISSCSRLGRLRLGQSAHCY 445
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
+K + S++NAL++MY +CG + + +IF ++ +D ++WS +IS+Y GH + A
Sbjct: 446 SIKHLAGENSSVANALISMYGRCGNFDVARKIFGMVKTKDVVTWSALISSYSHLGHSKDA 505
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
L L+ +M GVKP++ T ++V+S+C + + G+ + V D + + LVD
Sbjct: 506 LLLYDQMLTEGVKPNSATLVSVISSCANLAALEHGELIHSHVK-DVGLECDLSICTALVD 564
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
+ + G++ A ++ +M ++ W+ ++S +HG
Sbjct: 565 MYMKCGQLGIARKMFDSM-LERDVVTWNVMISGYGMHG 601
Score = 183 bits (464), Expect = 1e-44
Identities = 148/533 (27%), Positives = 253/533 (46%), Gaps = 44/533 (8%)
Query: 72 LHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGC 131
LH LA+T+G P + L+S Y+ A F P D WNS++ + +
Sbjct: 28 LHALAVTSGLSPRPDFAAKLVSAYSSAGLPALAALAFAASPCPDAFLWNSLLRSRHRASD 87
Query: 132 LEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGS-RIGRQIHALVVVDGRVE--HSVV 188
L + + G P A +V+ ++G+ +G +HA V G +E SV
Sbjct: 88 FASTLSAHRRMRASGARPS-RFTAPLVASAAAELGALPVGAAVHAYSVRFGLLEGDGSVA 146
Query: 189 LSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIAN------LDYDAAFARFR 242
++++LV Y RC A R+FD M ++ V+WTA+ISGC+ N L Y R
Sbjct: 147 VASSLVYMYARCGSVRDAVRLFDEMPERDVVAWTAVISGCVCNGQCGEGLSY--LVRMVR 204
Query: 243 AMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMY--CQ 300
+ G PN T+ + L AC G + G +HG+ + G+ S+L +MY C
Sbjct: 205 SAGDGGARPNSRTMESGLEACGVLGELSVGTCLHGFGVKAGVGHCPSVVSSLFSMYTKCD 264
Query: 301 SGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLA 360
S E A +F +D+V W+S+IG+Y + G + KA+ LF M +P+ V +
Sbjct: 265 STED---ARILFPELPEKDLVSWTSLIGAYCRAGHAEKAVELFLGMEESGLQPDEVVISC 321
Query: 361 VISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRD 420
+++ N + ++ G H I++ S+ + NAL++MYAKC ++ + +F + RD
Sbjct: 322 LLAGLGNDAKVRGGKTFHAAIVRRNFGDSVLIGNALISMYAKCKQVDIAATVFRMLHQRD 381
Query: 421 SISWSTMISAYGLHGHGEQALKLFSEMKERG---VKPDALTFLAVLSACNHAGLVTEGQQ 477
+ SWS+M+ AY G + L+L+ EM+ R + D + ++++S+C+ G + GQ
Sbjct: 382 TDSWSSMVVAYCKAGLDLKCLELYREMQFRDKDEFEYDTNSLISIISSCSRLGRLRLGQS 441
Query: 478 VFKQVNADYEIPLTIEHYA--------CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSS 529
+I+H A L+ + GR G + A +I + K WS+
Sbjct: 442 AH---------CYSIKHLAGENSSVANALISMYGRCGNFDVARKIFGMVKTK-DVVTWSA 491
Query: 530 LVSACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLL----NMIYAEHGHWLH 576
L+S+ G A +L Q++ +P++A +++ N+ EHG +H
Sbjct: 492 LISSYSHLGHSKDALLLYDQMLTEGVKPNSATLVSVISSCANLAALEHGELIH 544
Score = 63.2 bits (152), Expect = 2e-08
Identities = 40/143 (27%), Positives = 68/143 (46%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
S I S G L L+ Q+ S+ L SVI +C++L + G L+H
Sbjct: 490 SALISSYSHLGHSKDALLLYDQMLTEGVKPNSATLVSVISSCANLAALEHGELIHSHVKD 549
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G D + +L+ +Y K + AR++FD+M RD ++WN MI+ Y +G +AL++
Sbjct: 550 VGLECDLSICTALVDMYMKCGQLGIARKMFDSMLERDVVTWNVMISGYGMHGEAIQALKL 609
Query: 139 LKGVYLLGLVPKPELLASMVSLC 161
+ + P +++S C
Sbjct: 610 FSMMERGNVKPNSLTFLAILSAC 632
>UniRef100_Q8W0Q5 Putative vegetative storage protein [Sorghum bicolor]
Length = 779
Score = 361 bits (926), Expect = 4e-98
Identities = 197/583 (33%), Positives = 334/583 (56%), Gaps = 7/583 (1%)
Query: 22 IKSLISQGLYHQTLELF-TQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTG 80
I++ +G +H ++L+ + L+F P + P V+KACS+L G +H A G
Sbjct: 71 IRAYSWRGPFHAAIDLYRSMLYFRVPPN-KYTFPFVLKACSALADLCAGRTIHAHAAAVG 129
Query: 81 SHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLK 140
H+D VS +LI LY + + A VF MP RD ++WN+M+ Y +G A+ L
Sbjct: 130 LHTDLFVSTALIDLYIRCARFGPAANVFAKMPMRDVVAWNAMLAGYANHGMYHHAIAHLL 189
Query: 141 GVYLLG-LVPKPELLASMVSLCGRKMGSRIGRQIHA--LVVVDGRVEHSVVLSTALVDFY 197
+ G L P L S++ L + G +HA L + E V++ TAL+D Y
Sbjct: 190 DMQDRGGLRPNASTLVSLLPLLAQHGALFQGTSVHAYCLRAYLDQNEEQVLIGTALLDMY 249
Query: 198 FRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLI 257
+C + A RVF GM V+NEV+W+A+I G + AF F+ M VEG+ T +
Sbjct: 250 AKCKHLVYACRVFHGMTVRNEVTWSALIGGFVLCDRMTEAFNLFKDMLVEGMCFLSATSV 309
Query: 258 A-LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSS 316
A L CA ++ G ++H + GI + ++L++MY ++G ++ A +F+ +
Sbjct: 310 ASALRVCASLADLRMGTQLHALLAKSGIHADLTAGNSLLSMYAKAG-LINEATMLFDEIA 368
Query: 317 FRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCG 376
+D + + +++ Y Q G + +A +F KM +P+ T++++I AC++L++L+HG
Sbjct: 369 IKDTISYGALLSGYVQNGKAEEAFLVFKKMQACNVQPDIATMVSLIPACSHLAALQHGRC 428
Query: 377 LHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGH 436
HG ++ GL+ S+ N+L++MYAKCG ++ S Q+F +M RD +SW+TMI+ YG+HG
Sbjct: 429 SHGSVIIRGLALETSICNSLIDMYAKCGRIDLSRQVFDKMPARDIVSWNTMIAGYGIHGL 488
Query: 437 GEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYA 496
G++A LF MK +G +PD +TF+ +++AC+H+GLVTEG+ F + Y I +EHY
Sbjct: 489 GKEATTLFLSMKNQGFEPDDVTFICLIAACSHSGLVTEGKHWFDTMTHKYGILPRMEHYI 548
Query: 497 CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPD 556
C+VDLL R G +++A + +++MP+K R+W +L+ AC++H +D+ + + + + P+
Sbjct: 549 CMVDLLARGGFLDEAYQFIQSMPLKADVRVWGALLGACRIHKNIDLGKQVSRMIQKLGPE 608
Query: 557 NAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
N+ LL+ I++ G + +V K++ KK G S IE
Sbjct: 609 GTGNFVLLSNIFSAAGRFDEAAEVRIIQKVKGFKKSPGCSWIE 651
Score = 197 bits (502), Expect = 6e-49
Identities = 139/502 (27%), Positives = 248/502 (48%), Gaps = 28/502 (5%)
Query: 104 ARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGR 163
ARQVFD +P D ++N++I AY G A+++ + + + P ++ C
Sbjct: 52 ARQVFDRIPAPDARAYNALIRAYSWRGPFHAAIDLYRSMLYFRVPPNKYTFPFVLKACSA 111
Query: 164 KMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTA 223
GR IHA G + + +STAL+D Y RC A+ VF M +++ V+W A
Sbjct: 112 LADLCAGRTIHAHAAAVG-LHTDLFVSTALIDLYIRCARFGPAANVFAKMPMRDVVAWNA 170
Query: 224 MISGCIANLDYDAAFARFRAMQVE-GVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRH 282
M++G + Y A A MQ G+ PN TL++LLP A+ G + G +H Y R
Sbjct: 171 MLAGYANHGMYHHAIAHLLDMQDRGGLRPNASTLVSLLPLLAQHGALFQGTSVHAYCLRA 230
Query: 283 GIESSH---IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKA 339
++ + + +AL++MY + + L A ++F + R+ V WS++IG + +A
Sbjct: 231 YLDQNEEQVLIGTALLDMYAKC-KHLVYACRVFHGMTVRNEVTWSALIGGFVLCDRMTEA 289
Query: 340 LTLFNKMLTEETEPNYVTLLAVISA---CTNLSSLKHGCGLHGFILKFGLSFSISLSNAL 396
LF ML E +++ +V SA C +L+ L+ G LH + K G+ ++ N+L
Sbjct: 290 FNLFKDMLVEGM--CFLSATSVASALRVCASLADLRMGTQLHALLAKSGIHADLTAGNSL 347
Query: 397 MNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDA 456
++MYAK G + + +F E+ +D+IS+ ++S Y +G E+A +F +M+ V+PD
Sbjct: 348 LSMYAKAGLINEATMLFDEIAIKDTISYGALLSGYVQNGKAEEAFLVFKKMQACNVQPDI 407
Query: 457 LTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYAC--LVDLLGRSGKVEDALEI 514
T ++++ AC+H + G+ V L +E C L+D+ + G+++ + ++
Sbjct: 408 ATMVSLIPACSHLAALQHGRCSHGSVIIR---GLALETSICNSLIDMYAKCGRIDLSRQV 464
Query: 515 MRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLL-----NMI 567
MP + W+++++ +HG A L + EPD+ L+ + +
Sbjct: 465 FDKMPARDIVS-WNTMIAGYGIHGLGKEATTLFLSMKNQGFEPDDVTFICLIAACSHSGL 523
Query: 568 YAEHGHWL----HKEQVMEAMK 585
E HW HK ++ M+
Sbjct: 524 VTEGKHWFDTMTHKYGILPRME 545
Score = 107 bits (268), Expect = 8e-22
Identities = 92/394 (23%), Positives = 181/394 (45%), Gaps = 14/394 (3%)
Query: 205 MASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACA 264
+A +VFD + + ++ A+I + AA +R+M V PN+ T +L AC+
Sbjct: 51 LARQVFDRIPAPDARAYNALIRAYSWRGPFHAAIDLYRSMLYFRVPPNKYTFPFVLKACS 110
Query: 265 KPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWS 324
+ G+ IH +A G+ + S+ALI++Y + A +F + RDVV W+
Sbjct: 111 ALADLCAGRTIHAHAAAVGLHTDLFVSTALIDLYIRCAR-FGPAANVFAKMPMRDVVAWN 169
Query: 325 SIIGSYSQRGDSYKALT-LFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILK 383
+++ Y+ G + A+ L + PN TL++++ +L G +H + L+
Sbjct: 170 AMLAGYANHGMYHHAIAHLLDMQDRGGLRPNASTLVSLLPLLAQHGALFQGTSVHAYCLR 229
Query: 384 FGLSFS---ISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
L + + + AL++MYAKC L + ++F M R+ ++WS +I + L +A
Sbjct: 230 AYLDQNEEQVLIGTALLDMYAKCKHLVYACRVFHGMTVRNEVTWSALIGGFVLCDRMTEA 289
Query: 441 LKLFSEMKERGV-KPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
LF +M G+ A + + L C + G Q+ + A I + L+
Sbjct: 290 FNLFKDMLVEGMCFLSATSVASALRVCASLADLRMGTQLHALL-AKSGIHADLTAGNSLL 348
Query: 500 DLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEML--GPQLIRSEPDN 557
+ ++G + +A + + +K T + +L+S +G+ + A ++ Q +PD
Sbjct: 349 SMYAKAGLINEATMLFDEIAIK-DTISYGALLSGYVQNGKAEEAFLVFKKMQACNVQPDI 407
Query: 558 AANYTLL----NMIYAEHGHWLHKEQVMEAMKLQ 587
A +L+ ++ +HG H ++ + L+
Sbjct: 408 ATMVSLIPACSHLAALQHGRCSHGSVIIRGLALE 441
Score = 96.7 bits (239), Expect = 2e-18
Identities = 53/175 (30%), Positives = 93/175 (52%), Gaps = 1/175 (0%)
Query: 305 LHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISA 364
L LA Q+F+R D ++++I +YS RG + A+ L+ ML PN T V+ A
Sbjct: 49 LALARQVFDRIPAPDARAYNALIRAYSWRGPFHAAIDLYRSMLYFRVPPNKYTFPFVLKA 108
Query: 365 CTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISW 424
C+ L+ L G +H GL + +S AL+++Y +C + +F +M RD ++W
Sbjct: 109 CSALADLCAGRTIHAHAAAVGLHTDLFVSTALIDLYIRCARFGPAANVFAKMPMRDVVAW 168
Query: 425 STMISAYGLHGHGEQALKLFSEMKER-GVKPDALTFLAVLSACNHAGLVTEGQQV 478
+ M++ Y HG A+ +M++R G++P+A T +++L G + +G V
Sbjct: 169 NAMLAGYANHGMYHHAIAHLLDMQDRGGLRPNASTLVSLLPLLAQHGALFQGTSV 223
Score = 73.6 bits (179), Expect = 2e-11
Identities = 62/243 (25%), Positives = 101/243 (41%), Gaps = 11/243 (4%)
Query: 12 VSPTPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTL 71
+ T S + + G + +F ++ + + S+I ACS L + G
Sbjct: 369 IKDTISYGALLSGYVQNGKAEEAFLVFKKMQACNVQPDIATMVSLIPACSHLAALQHGRC 428
Query: 72 LHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGC 131
HG + G + + NSLI +YAK I +RQVFD MP RD +SWN+MI Y +G
Sbjct: 429 SHGSVIIRGLALETSICNSLIDMYAKCGRIDLSRQVFDKMPARDIVSWNTMIAGYGIHGL 488
Query: 132 LEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGR-----QIHALVVVDGRVEHS 186
+EA + + G P +++ C G+ H ++ R+EH
Sbjct: 489 GKEATTLFLSMKNQGFEPDDVTFICLIAACSHSGLVTEGKHWFDTMTHKYGILP-RMEHY 547
Query: 187 VVLSTALVDFYFRCDDALMASRVFDGMEVKNEVS-WTAMISGCIANLDYDAAFARFRAMQ 245
+ +VD R A + M +K +V W A++ C + + D R +Q
Sbjct: 548 I----CMVDLLARGGFLDEAYQFIQSMPLKADVRVWGALLGACRIHKNIDLGKQVSRMIQ 603
Query: 246 VEG 248
G
Sbjct: 604 KLG 606
Score = 47.0 bits (110), Expect = 0.002
Identities = 43/171 (25%), Positives = 73/171 (42%), Gaps = 4/171 (2%)
Query: 404 GCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVL 463
G L + Q+F + D+ +++ +I AY G A+ L+ M V P+ TF VL
Sbjct: 47 GQLALARQVFDRIPAPDARAYNALIRAYSWRGPFHAAIDLYRSMLYFRVPPNKYTFPFVL 106
Query: 464 SACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPS 523
AC+ + G+ + A + + L+DL R + A + MPM+
Sbjct: 107 KACSALADLCAGRTIHAHA-AAVGLHTDLFVSTALIDLYIRCARFGPAANVFAKMPMR-D 164
Query: 524 TRIWSSLVSACKLHGRLD--IAEMLGPQLIRSEPDNAANYTLLNMIYAEHG 572
W+++++ HG IA +L Q NA+ L + A+HG
Sbjct: 165 VVAWNAMLAGYANHGMYHHAIAHLLDMQDRGGLRPNASTLVSLLPLLAQHG 215
>UniRef100_Q7XJU7 OSJNBa0016O02.23 protein [Oryza sativa]
Length = 939
Score = 361 bits (926), Expect = 4e-98
Identities = 192/581 (33%), Positives = 336/581 (57%), Gaps = 4/581 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I + G++ + L+LF ++ S V++ C+ L G LH L G+
Sbjct: 237 ISGCVQNGMFLEALDLFRRMQSDGFSMNSYTTVGVLQVCAELAQLNHGRELHAALLKCGT 296
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
+ + N+L+ +YA+ + SA +VF + +D ISWNSM++ Y+QN EA++
Sbjct: 297 EFN-IQCNALLVMYARCGWVDSALRVFREIGDKDYISWNSMLSCYVQNRLYAEAIDFFGE 355
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ G P + S++S G GR++HA V R++ + ++ L+D Y +C
Sbjct: 356 MVQNGFNPDHACIVSLLSAVGHLGRLINGREVHAYAVKQ-RLDSDLQIANTLMDMYIKCY 414
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
++RVFD M +K+ VSWT +I+ + Y A +FR Q EG+ + + + ++L
Sbjct: 415 SVECSARVFDRMRIKDHVSWTTIIACYAQSSRYSEAIGKFRTAQKEGIKVDPMMMGSILE 474
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVV 321
AC+ + K++H YA R+G+ I + +I++Y + GE + A IFE +D+V
Sbjct: 475 ACSGLKSISLLKQVHSYAIRNGLLDL-ILKNRIIDIYGECGEVCY-ALNIFEMLDKKDIV 532
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFI 381
W+S++ +++ G ++A+ LF KML +P+ V L+ ++ A LSSL G +HGF+
Sbjct: 533 TWTSMVNCFAENGLLHEAVALFGKMLNAGIQPDSVALVGILGAIAGLSSLTKGKEIHGFL 592
Query: 382 LKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQAL 441
++ ++ ++L++MY+ CG + +L++F E + +D + W+ MI+A G+HGHG+QA+
Sbjct: 593 IRGKFPVEGAVVSSLVDMYSGCGSMNYALKVFDEAKCKDVVLWTAMINATGMHGHGKQAI 652
Query: 442 KLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDL 501
+F M E GV PD ++FLA+L AC+H+ LV EG+ + + Y++ EHYAC+VDL
Sbjct: 653 YIFKRMLETGVSPDHVSFLALLYACSHSKLVDEGKFYLDMMVSKYKLQPWQEHYACVVDL 712
Query: 502 LGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANY 561
LGRSG+ E+A + +++MP++P + +W +L+ AC++H ++A + +L+ EPDN NY
Sbjct: 713 LGRSGQTEEAYKFIKSMPLEPKSVVWCALLGACRIHKNHELAMIATDKLLELEPDNPGNY 772
Query: 562 TLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGN 602
L++ ++AE G W + +++ M Q L+K S IE GN
Sbjct: 773 VLVSNVFAEMGKWNNVKEIRTKMTEQGLRKDPACSWIEIGN 813
Score = 221 bits (563), Expect = 5e-56
Identities = 147/509 (28%), Positives = 262/509 (50%), Gaps = 16/509 (3%)
Query: 69 GTLLHGLALTTGSHSDP---VVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINA 125
G LH A+ TG+ D ++ L+ +Y K + A ++FD MP R SWN++I A
Sbjct: 74 GRQLHAHAVATGALGDDDAGFLATKLLFMYGKCGRLPDAHRLFDGMPARTVFSWNALIGA 133
Query: 126 YLQNGCLEEALEMLKGVY----LLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDG 181
L +G EA+ + + + + G P LAS++ CG + R G ++H L V G
Sbjct: 134 CLSSGGAGEAVGVYRAMRASEPVAGAAPDGCTLASVLKACGAEGDGRCGSEVHGLAVKSG 193
Query: 182 RVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEV-SWTAMISGCIANLDYDAAFAR 240
++ S +++ ALV Y +C A RVF+ M +V SW + ISGC+ N + A
Sbjct: 194 -LDRSTLVANALVGMYAKCGLLDSALRVFEWMRDGRDVASWNSAISGCVQNGMFLEALDL 252
Query: 241 FRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQ 300
FR MQ +G N T + +L CA+ + HG+E+H + G E +I +AL+ MY +
Sbjct: 253 FRRMQSDGFSMNSYTTVGVLQVCAELAQLNHGRELHAALLKCGTE-FNIQCNALLVMYAR 311
Query: 301 SGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLA 360
G + A ++F +D + W+S++ Y Q +A+ F +M+ P++ +++
Sbjct: 312 CG-WVDSALRVFREIGDKDYISWNSMLSCYVQNRLYAEAIDFFGEMVQNGFNPDHACIVS 370
Query: 361 VISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRD 420
++SA +L L +G +H + +K L + ++N LM+MY KC +E S ++F M+ +D
Sbjct: 371 LLSAVGHLGRLINGREVHAYAVKQRLDSDLQIANTLMDMYIKCYSVECSARVFDRMRIKD 430
Query: 421 SISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFK 480
+SW+T+I+ Y +A+ F ++ G+K D + ++L AC+ ++ +QV
Sbjct: 431 HVSWTTIIACYAQSSRYSEAIGKFRTAQKEGIKVDPMMMGSILEACSGLKSISLLKQVHS 490
Query: 481 QVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRL 540
+ + L +++ ++D+ G G+V AL I + K W+S+V+ +G L
Sbjct: 491 YAIRNGLLDLILKNR--IIDIYGECGEVCYALNIFEMLDKKDIV-TWTSMVNCFAENGLL 547
Query: 541 DIAEMLGPQLIRS--EPDNAANYTLLNMI 567
A L +++ + +PD+ A +L I
Sbjct: 548 HEAVALFGKMLNAGIQPDSVALVGILGAI 576
Score = 213 bits (542), Expect = 1e-53
Identities = 138/515 (26%), Positives = 269/515 (51%), Gaps = 9/515 (1%)
Query: 53 LPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTM- 111
L SV+KAC + G+ +HGLA+ +G +V+N+L+ +YAK + SA +VF+ M
Sbjct: 166 LASVLKACGAEGDGRCGSEVHGLAVKSGLDRSTLVANALVGMYAKCGLLDSALRVFEWMR 225
Query: 112 PHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGR 171
RD SWNS I+ +QNG EAL++ + + G ++ +C GR
Sbjct: 226 DGRDVASWNSAISGCVQNGMFLEALDLFRRMQSDGFSMNSYTTVGVLQVCAELAQLNHGR 285
Query: 172 QIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIAN 231
++HA ++ G E ++ + LV Y RC A RVF + K+ +SW +M+S + N
Sbjct: 286 ELHAALLKCG-TEFNIQCNALLV-MYARCGWVDSALRVFREIGDKDYISWNSMLSCYVQN 343
Query: 232 LDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFS 291
Y A F M G +P+ +++LL A G + +G+E+H YA + ++S +
Sbjct: 344 RLYAEAIDFFGEMVQNGFNPDHACIVSLLSAVGHLGRLINGREVHAYAVKQRLDSDLQIA 403
Query: 292 SALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEET 351
+ L++MY + S+ + ++F+R +D V W++II Y+Q +A+ F E
Sbjct: 404 NTLMDMYIKC-YSVECSARVFDRMRIKDHVSWTTIIACYAQSSRYSEAIGKFRTAQKEGI 462
Query: 352 EPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQ 411
+ + + + +++ AC+ L S+ +H + ++ GL + L N ++++Y +CG + +L
Sbjct: 463 KVDPMMMGSILEACSGLKSISLLKQVHSYAIRNGL-LDLILKNRIIDIYGECGEVCYALN 521
Query: 412 IFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGL 471
IF + +D ++W++M++ + +G +A+ LF +M G++PD++ + +L A
Sbjct: 522 IFEMLDKKDIVTWTSMVNCFAENGLLHEAVALFGKMLNAGIQPDSVALVGILGAIAGLSS 581
Query: 472 VTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
+T+G+++ + + P+ + LVD+ G + AL++ K +W++++
Sbjct: 582 LTKGKEIHGFLIRG-KFPVEGAVVSSLVDMYSGCGSMNYALKVFDEAKCK-DVVLWTAMI 639
Query: 532 SACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLL 564
+A +HG A + +++ + PD+ + LL
Sbjct: 640 NATGMHGHGKQAIYIFKRMLETGVSPDHVSFLALL 674
Score = 201 bits (511), Expect = 5e-50
Identities = 137/440 (31%), Positives = 225/440 (51%), Gaps = 24/440 (5%)
Query: 108 FDTMPHRD--PISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKP-ELLASMVSLCGRK 164
F P R P S + + ++G L EAL L G P P + ++ L +
Sbjct: 9 FHPTPRRKLPPASAGASLRQLCKDGDLREALRQLAARSARGRAPPPTDHYGWVLDLVAVR 68
Query: 165 MGSRIGRQIHALVVVDGRV--EHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWT 222
GRQ+HA V G + + + L+T L+ Y +C A R+FDGM + SW
Sbjct: 69 RAVSEGRQLHAHAVATGALGDDDAGFLATKLLFMYGKCGRLPDAHRLFDGMPARTVFSWN 128
Query: 223 AMISGCIANLDYDAAFARFRAMQ----VEGVDPNRVTLIALLPACAKPGFVKHGKEIHGY 278
A+I C+++ A +RAM+ V G P+ TL ++L AC G + G E+HG
Sbjct: 129 ALIGACLSSGGAGEAVGVYRAMRASEPVAGAAPDGCTLASVLKACGAEGDGRCGSEVHGL 188
Query: 279 AFRHGIESSHIFSSALINMYCQSGESLHLAEQIFE-RSSFRDVVLWSSIIGSYSQRGDSY 337
A + G++ S + ++AL+ MY + G L A ++FE RDV W+S I Q G
Sbjct: 189 AVKSGLDRSTLVANALVGMYAKCG-LLDSALRVFEWMRDGRDVASWNSAISGCVQNGMFL 247
Query: 338 KALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALM 397
+AL LF +M ++ N T + V+ C L+ L HG LH +LK G F+I NAL+
Sbjct: 248 EALDLFRRMQSDGFSMNSYTTVGVLQVCAELAQLNHGRELHAALLKCGTEFNIQ-CNALL 306
Query: 398 NMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDAL 457
MYA+CG ++ +L++F E+ ++D ISW++M+S Y + +A+ F EM + G PD
Sbjct: 307 VMYARCGWVDSALRVFREIGDKDYISWNSMLSCYVQNRLYAEAIDFFGEMVQNGFNPDHA 366
Query: 458 TFLAVLSACNHAGLVTEGQQVF-----KQVNADYEIPLTIEHYACLVDLLGRSGKVEDAL 512
+++LSA H G + G++V +++++D +I T L+D+ + VE +
Sbjct: 367 CIVSLLSAVGHLGRLINGREVHAYAVKQRLDSDLQIANT------LMDMYIKCYSVECSA 420
Query: 513 EIMRTMPMKPSTRIWSSLVS 532
+ M +K W+++++
Sbjct: 421 RVFDRMRIKDHVS-WTTIIA 439
Score = 164 bits (415), Expect = 7e-39
Identities = 105/439 (23%), Positives = 215/439 (48%), Gaps = 7/439 (1%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
+ + + LY + ++ F ++ + + + + S++ A L G +H A+
Sbjct: 334 NSMLSCYVQNRLYAEAIDFFGEMVQNGFNPDHACIVSLLSAVGHLGRLINGREVHAYAVK 393
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
SD ++N+L+ +Y K ++ + +VFD M +D +SW ++I Y Q+ EA+
Sbjct: 394 QRLDSDLQIANTLMDMYIKCYSVECSARVFDRMRIKDHVSWTTIIACYAQSSRYSEAIGK 453
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+ G+ P ++ S++ C + +Q+H+ + +G ++ ++L ++D Y
Sbjct: 454 FRTAQKEGIKVDPMMMGSILEACSGLKSISLLKQVHSYAIRNGLLD--LILKNRIIDIYG 511
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
C + A +F+ ++ K+ V+WT+M++ N A A F M G+ P+ V L+
Sbjct: 512 ECGEVCYALNIFEMLDKKDIVTWTSMVNCFAENGLLHEAVALFGKMLNAGIQPDSVALVG 571
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
+L A A + GKEIHG+ R S+L++MY G S++ A ++F+ + +
Sbjct: 572 ILGAIAGLSSLTKGKEIHGFLIRGKFPVEGAVVSSLVDMYSGCG-SMNYALKVFDEAKCK 630
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHG-CGL 377
DVVLW+++I + G +A+ +F +ML P++V+ LA++ AC++ + G L
Sbjct: 631 DVVLWTAMINATGMHGHGKQAIYIFKRMLETGVSPDHVSFLALLYACSHSKLVDEGKFYL 690
Query: 378 HGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQ-NRDSISWSTMISAYGLHGH 436
+ K+ L ++++ + G E + + M S+ W ++ A +H +
Sbjct: 691 DMMVSKYKLQPWQEHYACVVDLLGRSGQTEEAYKFIKSMPLEPKSVVWCALLGACRIHKN 750
Query: 437 GEQALKLFSEMKERGVKPD 455
E A+ ++ E ++PD
Sbjct: 751 HELAMIATDKLLE--LEPD 767
>UniRef100_Q5QNL7 Pentatricopeptide repeat protein-like [Oryza sativa]
Length = 658
Score = 358 bits (918), Expect = 3e-97
Identities = 196/531 (36%), Positives = 307/531 (56%), Gaps = 3/531 (0%)
Query: 69 GTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQ 128
G ++HG + G + V N+LIS YAK + I+ A VFD MP RD ISWNS+I
Sbjct: 3 GLVVHGYLVKYGFGAQCAVCNALISFYAKSNRIEDALMVFDEMPQRDIISWNSIIGGCAS 62
Query: 129 NGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVV 188
NG ++A+E+ ++L G L S++ C + S IG +H V G + +
Sbjct: 63 NGLYDKAVELFVRMWLEGQELDSTTLLSVMPACVQSHYSFIGGVVHGYSVRTGLISETS- 121
Query: 189 LSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEG 248
L AL+D Y C D +++F ME KN VSWTAMI+ +D F+ M +EG
Sbjct: 122 LGNALLDMYSNCSDWRSTNKIFRNMEQKNVVSWTAMITSYTRAGHFDKVAGLFQEMGLEG 181
Query: 249 VDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLA 308
+ P+ + + L A A +KHGK +HGYA R+GIE ++AL+ MY + G + A
Sbjct: 182 IRPDVFAITSALDAFAGNESLKHGKSVHGYAIRNGIEEVLPVANALMEMYVKCGY-MEEA 240
Query: 309 EQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNL 368
IF+ + +D + W+++IG YS+ + +A TLFN+ML + PN VT+ ++ A +L
Sbjct: 241 RFIFDHVTKKDTISWNTLIGGYSRSNLANEAFTLFNEMLLQ-LRPNAVTMACILPAAASL 299
Query: 369 SSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMI 428
SSL+ G +H + ++ G ++NAL++MY KCG L + ++F + N++ ISW+ MI
Sbjct: 300 SSLERGREMHAYAVRRGYLEDNFVANALVDMYVKCGALLLARRLFDMLTNKNLISWTIMI 359
Query: 429 SAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEI 488
+ YG+HG G A+ LF +MK G++PDA +F A+L AC+H+GL EG + F + ++ I
Sbjct: 360 AGYGMHGRGRDAIALFEQMKGSGIQPDAGSFSAILYACSHSGLRDEGWRFFNAMRNEHRI 419
Query: 489 PLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGP 548
++HYAC+VDLL +G +++A E + TMP++P + IW SL+ C++H + +AE +
Sbjct: 420 EPKLKHYACMVDLLCHTGNLKEAYEFIETMPIEPDSSIWVSLLRGCRIHRNVKLAEKVAE 479
Query: 549 QLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
+ EP+N Y LL IYAE W ++ + + L++ G S IE
Sbjct: 480 MVFELEPENTGYYVLLANIYAEAERWEAVRKLKNKVGGRGLRENTGCSWIE 530
Score = 171 bits (434), Expect = 4e-41
Identities = 94/341 (27%), Positives = 182/341 (52%), Gaps = 3/341 (0%)
Query: 27 SQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPV 86
S GLY + +ELF ++ S+ L SV+ AC H +G ++HG ++ TG S+
Sbjct: 62 SNGLYDKAVELFVRMWLEGQELDSTTLLSVMPACVQSHYSFIGGVVHGYSVRTGLISETS 121
Query: 87 VSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLG 146
+ N+L+ +Y+ SD +S ++F M ++ +SW +MI +Y + G ++ + + + L G
Sbjct: 122 LGNALLDMYSNCSDWRSTNKIFRNMEQKNVVSWTAMITSYTRAGHFDKVAGLFQEMGLEG 181
Query: 147 LVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMA 206
+ P + S + + G+ +H + +G +E + ++ AL++ Y +C A
Sbjct: 182 IRPDVFAITSALDAFAGNESLKHGKSVHGYAIRNG-IEEVLPVANALMEMYVKCGYMEEA 240
Query: 207 SRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKP 266
+FD + K+ +SW +I G + + AF F M ++ + PN VT+ +LPA A
Sbjct: 241 RFIFDHVTKKDTISWNTLIGGYSRSNLANEAFTLFNEMLLQ-LRPNAVTMACILPAAASL 299
Query: 267 GFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSI 326
++ G+E+H YA R G + ++AL++MY + G +L LA ++F+ + ++++ W+ +
Sbjct: 300 SSLERGREMHAYAVRRGYLEDNFVANALVDMYVKCG-ALLLARRLFDMLTNKNLISWTIM 358
Query: 327 IGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTN 367
I Y G A+ LF +M +P+ + A++ AC++
Sbjct: 359 IAGYGMHGRGRDAIALFEQMKGSGIQPDAGSFSAILYACSH 399
Score = 119 bits (299), Expect = 2e-25
Identities = 85/307 (27%), Positives = 141/307 (45%), Gaps = 3/307 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I S G + + LF ++ + S + A + S G +HG A+ G
Sbjct: 158 ITSYTRAGHFDKVAGLFQEMGLEGIRPDVFAITSALDAFAGNESLKHGKSVHGYAIRNGI 217
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
V+N+L+ +Y K ++ AR +FD + +D ISWN++I Y ++ EA +
Sbjct: 218 EEVLPVANALMEMYVKCGYMEEARFIFDHVTKKDTISWNTLIGGYSRSNLANEAFTLFNE 277
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ LL L P +A ++ GR++HA V G +E + V + ALVD Y +C
Sbjct: 278 M-LLQLRPNAVTMACILPAAASLSSLERGREMHAYAVRRGYLEDNFV-ANALVDMYVKCG 335
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
L+A R+FD + KN +SWT MI+G + A A F M+ G+ P+ + A+L
Sbjct: 336 ALLLARRLFDMLTNKNLISWTIMIAGYGMHGRGRDAIALFEQMKGSGIQPDAGSFSAILY 395
Query: 262 ACAKPGFVKHG-KEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
AC+ G G + + H IE + ++++ C +G E I D
Sbjct: 396 ACSHSGLRDEGWRFFNAMRNEHRIEPKLKHYACMVDLLCHTGNLKEAYEFIETMPIEPDS 455
Query: 321 VLWSSII 327
+W S++
Sbjct: 456 SIWVSLL 462
>UniRef100_Q84SE9 P0020E09.15 protein [Oryza sativa]
Length = 813
Score = 356 bits (913), Expect = 1e-96
Identities = 193/570 (33%), Positives = 323/570 (55%), Gaps = 15/570 (2%)
Query: 42 HFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDI 101
H AP++ + P +KACS+L H G +H A+ G +D VS +L+ +Y K + +
Sbjct: 119 HRVAPNNYT--FPFALKACSALADHHCGRAIHRHAIHAGLQADLFVSTALLDMYVKCACL 176
Query: 102 QSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLL--GLVPKPELLASMVS 159
A +F TMP RD ++WN+M+ Y +G A+ L + + L P L +++
Sbjct: 177 PDAAHIFATMPARDLVAWNAMLAGYAHHGMYHHAVAHLLSMQMQMHRLRPNASTLVALLP 236
Query: 160 LCGRKMGSRIGRQIHALVV---------VDGRVEHSVVLSTALVDFYFRCDDALMASRVF 210
L ++ G +HA + ++ V+L TAL+D Y +C L A RVF
Sbjct: 237 LLAQQGALAQGTSVHAYCIRACLHPNRNSKSKLTDGVLLGTALLDMYAKCGSLLYARRVF 296
Query: 211 DGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA-LLPACAKPGFV 269
D M +NEV+W+A+I G + AF F+AM +G+ T IA L ACA +
Sbjct: 297 DAMPARNEVTWSALIGGFVLCSRMTQAFLLFKAMLAQGLCFLSPTSIASALRACASLDHL 356
Query: 270 KHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGS 329
+ G+++H + G+ + ++L++MY ++G + A +F+ + +D V +S+++
Sbjct: 357 RMGEQLHALLAKSGVHADLTAGNSLLSMYAKAG-LIDQAIALFDEMAVKDTVSYSALVSG 415
Query: 330 YSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFS 389
Y Q G + +A +F KM EP+ T++++I AC++L++L+HG HG ++ GL+
Sbjct: 416 YVQNGRAEEAFLVFKKMQACNVEPDAATMVSLIPACSHLAALQHGRCSHGSVIIRGLASE 475
Query: 390 ISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKE 449
S+ NAL++MYAKCG ++ S Q+F M +RD +SW+TMI+ YG+HG G++A LF EM
Sbjct: 476 TSICNALIDMYAKCGRIDLSRQVFNMMPSRDIVSWNTMIAGYGIHGLGKEATALFLEMNN 535
Query: 450 RGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVE 509
G PD +TF+ +LSAC+H+GLV EG+ F + Y + +EHY C+VDLL R G ++
Sbjct: 536 LGFPPDGVTFICLLSACSHSGLVIEGKHWFHVMGHGYGLTPRMEHYICMVDLLSRGGFLD 595
Query: 510 DALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYA 569
+A E +++MP++ R+W +L+ AC+++ +D+ + + + P+ N+ LL+ IY+
Sbjct: 596 EAYEFIQSMPLRADVRVWVALLGACRVYKNIDLGKKVSRMIQELGPEGTGNFVLLSNIYS 655
Query: 570 EHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
G + +V K+Q KK G S IE
Sbjct: 656 AAGRFDEAAEVRIIQKVQGFKKSPGCSWIE 685
Score = 190 bits (482), Expect = 1e-46
Identities = 129/503 (25%), Positives = 243/503 (47%), Gaps = 36/503 (7%)
Query: 101 IQSARQVFDTMPHRDPISWNSMINAYLQNG--CLEEALEMLKGVYLLGLVPKPELLASMV 158
+ A +FD +P D ++N +I AY + + L + + + + P +
Sbjct: 73 LSRAHHLFDQIPSPDVRTYNDLIRAYSSSSPTAAADGLHLYRRMLRHRVAPNNYTFPFAL 132
Query: 159 SLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNE 218
C GR IH + G ++ + +STAL+D Y +C A+ +F M ++
Sbjct: 133 KACSALADHHCGRAIHRHAIHAG-LQADLFVSTALLDMYVKCACLPDAAHIFATMPARDL 191
Query: 219 VSWTAMISGCIANLDYDAAFARFRAMQVE--GVDPNRVTLIALLPACAKPGFVKHGKEIH 276
V+W AM++G + Y A A +MQ++ + PN TL+ALLP A+ G + G +H
Sbjct: 192 VAWNAMLAGYAHHGMYHHAVAHLLSMQMQMHRLRPNASTLVALLPLLAQQGALAQGTSVH 251
Query: 277 GYAFRHGIESSH----------IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSI 326
Y R + + + +AL++MY + G SL A ++F+ R+ V WS++
Sbjct: 252 AYCIRACLHPNRNSKSKLTDGVLLGTALLDMYAKCG-SLLYARRVFDAMPARNEVTWSAL 310
Query: 327 IGSYSQRGDSYKALTLFNKMLTEE-TEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFG 385
IG + +A LF ML + + ++ + + AC +L L+ G LH + K G
Sbjct: 311 IGGFVLCSRMTQAFLLFKAMLAQGLCFLSPTSIASALRACASLDHLRMGEQLHALLAKSG 370
Query: 386 LSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFS 445
+ ++ N+L++MYAK G ++ ++ +F EM +D++S+S ++S Y +G E+A +F
Sbjct: 371 VHADLTAGNSLLSMYAKAGLIDQAIALFDEMAVKDTVSYSALVSGYVQNGRAEEAFLVFK 430
Query: 446 EMKERGVKPDALTFLAVLSACNHA-----GLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
+M+ V+PDA T ++++ AC+H G + G + + + ++ I L+D
Sbjct: 431 KMQACNVEPDAATMVSLIPACSHLAALQHGRCSHGSVIIRGLASETSI------CNALID 484
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLH--GRLDIAEMLGPQLIRSEPDNA 558
+ + G+++ + ++ MP + W+++++ +H G+ A L + PD
Sbjct: 485 MYAKCGRIDLSRQVFNMMPSRDIVS-WNTMIAGYGIHGLGKEATALFLEMNNLGFPPDGV 543
Query: 559 ANYTLLNM-----IYAEHGHWLH 576
LL+ + E HW H
Sbjct: 544 TFICLLSACSHSGLVIEGKHWFH 566
Score = 111 bits (278), Expect = 5e-23
Identities = 95/363 (26%), Positives = 167/363 (45%), Gaps = 22/363 (6%)
Query: 241 FRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQ 300
+R M V PN T L AC+ G+ IH +A G+++ S+AL++MY +
Sbjct: 113 YRRMLRHRVAPNNYTFPFALKACSALADHHCGRAIHRHAIHAGLQADLFVSTALLDMYVK 172
Query: 301 SGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALT--LFNKMLTEETEPNYVTL 358
L A IF RD+V W++++ Y+ G + A+ L +M PN TL
Sbjct: 173 CA-CLPDAAHIFATMPARDLVAWNAMLAGYAHHGMYHHAVAHLLSMQMQMHRLRPNASTL 231
Query: 359 LAVISACTNLSSLKHGCGLHGFIL----------KFGLSFSISLSNALMNMYAKCGCLEG 408
+A++ +L G +H + + K L+ + L AL++MYAKCG L
Sbjct: 232 VALLPLLAQQGALAQGTSVHAYCIRACLHPNRNSKSKLTDGVLLGTALLDMYAKCGSLLY 291
Query: 409 SLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLA-VLSACN 467
+ ++F M R+ ++WS +I + L QA LF M +G+ + T +A L AC
Sbjct: 292 ARRVFDAMPARNEVTWSALIGGFVLCSRMTQAFLLFKAMLAQGLCFLSPTSIASALRACA 351
Query: 468 HAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIW 527
+ G+Q+ + A + + L+ + ++G ++ A+ + M +K T +
Sbjct: 352 SLDHLRMGEQLHALL-AKSGVHADLTAGNSLLSMYAKAGLIDQAIALFDEMAVK-DTVSY 409
Query: 528 SSLVSACKLHGRLDIAEML--GPQLIRSEPDNAANYTLL----NMIYAEHGHWLHKEQVM 581
S+LVS +GR + A ++ Q EPD A +L+ ++ +HG H ++
Sbjct: 410 SALVSGYVQNGRAEEAFLVFKKMQACNVEPDAATMVSLIPACSHLAALQHGRCSHGSVII 469
Query: 582 EAM 584
+
Sbjct: 470 RGL 472
Score = 77.4 bits (189), Expect = 1e-12
Identities = 61/233 (26%), Positives = 99/233 (42%), Gaps = 31/233 (13%)
Query: 12 VSPTPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTL 71
V T S S + + G + +F ++ ++ + S+I ACS L + G
Sbjct: 403 VKDTVSYSALVSGYVQNGRAEEAFLVFKKMQACNVEPDAATMVSLIPACSHLAALQHGRC 462
Query: 72 LHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGC 131
HG + G S+ + N+LI +YAK I +RQVF+ MP RD +SWN+MI Y +G
Sbjct: 463 SHGSVIIRGLASETSICNALIDMYAKCGRIDLSRQVFNMMPSRDIVSWNTMIAGYGIHGL 522
Query: 132 LEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDG---------- 181
+EA + + LG P ++S C H+ +V++G
Sbjct: 523 GKEATALFLEMNNLGFPPDGVTFICLLSACS-----------HSGLVIEGKHWFHVMGHG 571
Query: 182 -----RVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVS-WTAMISGC 228
R+EH + +VD R A M ++ +V W A++ C
Sbjct: 572 YGLTPRMEHYI----CMVDLLSRGGFLDEAYEFIQSMPLRADVRVWVALLGAC 620
>UniRef100_Q9SS60 T12J13.14 protein [Arabidopsis thaliana]
Length = 882
Score = 352 bits (904), Expect = 1e-95
Identities = 204/583 (34%), Positives = 328/583 (55%), Gaps = 9/583 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I S G Y + LE++ +L S S + SV+ A +L G LHG AL +G
Sbjct: 179 ISGYSSHGYYEEALEIYHELKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSGV 238
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
+S VV+N L+++Y KF AR+VFD M RD +S+N+MI YL+ +EE++ M
Sbjct: 239 NSVVVVNNGLVAMYLKFRRPTDARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRM--- 295
Query: 142 VYLLGLVP-KPELL--ASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+L L KP+LL +S++ CG + + I+ ++ G V S V + L+D Y
Sbjct: 296 -FLENLDQFKPDLLTVSSVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNI-LIDVYA 353
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
+C D + A VF+ ME K+ VSW ++ISG I + D A F+ M + + +T +
Sbjct: 354 KCGDMITARDVFNSMECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLM 413
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
L+ + +K GK +H + GI S+ALI+MY + GE + + +IF
Sbjct: 414 LISVSTRLADLKFGKGLHSNGIKSGICIDLSVSNALIDMYAKCGE-VGDSLKIFSSMGTG 472
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLH 378
D V W+++I + + GD L + +M E P+ T L + C +L++ + G +H
Sbjct: 473 DTVTWNTVISACVRFGDFATGLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIH 532
Query: 379 GFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGE 438
+L+FG + + NAL+ MY+KCGCLE S ++F M RD ++W+ MI AYG++G GE
Sbjct: 533 CCLLRFGYESELQIGNALIEMYSKCGCLENSSRVFERMSRRDVVTWTGMIYAYGMYGEGE 592
Query: 439 QALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACL 498
+AL+ F++M++ G+ PD++ F+A++ AC+H+GLV EG F+++ Y+I IEHYAC+
Sbjct: 593 KALETFADMEKSGIVPDSVVFIAIIYACSHSGLVDEGLACFEKMKTHYKIDPMIEHYACV 652
Query: 499 VDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNA 558
VDLL RS K+ A E ++ MP+KP IW+S++ AC+ G ++ AE + ++I PD+
Sbjct: 653 VDLLSRSQKISKAEEFIQAMPIKPDASIWASVLRACRTSGDMETAERVSRRIIELNPDDP 712
Query: 559 ANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETG 601
L + YA W + +++K + + K G+S IE G
Sbjct: 713 GYSILASNAYAALRKWDKVSLIRKSLKDKHITKNPGYSWIEVG 755
Score = 258 bits (660), Expect = 3e-67
Identities = 163/518 (31%), Positives = 270/518 (51%), Gaps = 7/518 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I++ GL+ + LE + +L S PSVIKAC+ L +G L++ L G
Sbjct: 78 IRAFSKNGLFPEALEFYGKLRESKVSPDKYTFPSVIKACAGLFDAEMGDLVYEQILDMGF 137
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
SD V N+L+ +Y++ + ARQVFD MP RD +SWNS+I+ Y +G EEALE+
Sbjct: 138 ESDLFVGNALVDMYSRMGLLTRARQVFDEMPVRDLVSWNSLISGYSSHGYYEEALEIYHE 197
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ +VP ++S++ G + + G+ +H + G V VV++ LV Y +
Sbjct: 198 LKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSG-VNSVVVVNNGLVAMYLKFR 256
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
A RVFD M+V++ VS+ MI G + L+ R ++ P+ +T+ ++L
Sbjct: 257 RPTDARRVFDEMDVRDSVSYNTMICGYL-KLEMVEESVRMFLENLDQFKPDLLTVSSVLR 315
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVV 321
AC + K I+ Y + G + LI++Y + G+ + A +F +D V
Sbjct: 316 ACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYAKCGDMI-TARDVFNSMECKDTV 374
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFI 381
W+SII Y Q GD +A+ LF M+ E + +++T L +IS T L+ LK G GLH
Sbjct: 375 SWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADLKFGKGLHSNG 434
Query: 382 LKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQAL 441
+K G+ +S+SNAL++MYAKCG + SL+IF M D+++W+T+ISA G L
Sbjct: 435 IKSGICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGTGDTVTWNTVISACVRFGDFATGL 494
Query: 442 KLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVF-KQVNADYEIPLTIEHYACLVD 500
++ ++M++ V PD TFL L C G+++ + YE L I + L++
Sbjct: 495 QVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIHCCLLRFGYESELQIGN--ALIE 552
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
+ + G +E++ + M + W+ ++ A ++G
Sbjct: 553 MYSKCGCLENSSRVFERMSRR-DVVTWTGMIYAYGMYG 589
Score = 198 bits (504), Expect = 3e-49
Identities = 142/529 (26%), Positives = 266/529 (49%), Gaps = 10/529 (1%)
Query: 54 PSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTM-P 112
P + +A SS + +H L ++ G S S LI Y+ F + S+ VF + P
Sbjct: 8 PFISRALSSSSNLNELRRIHALVISLGLDSSDFFSGKLIDKYSHFREPASSLSVFRRVSP 67
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQ 172
++ WNS+I A+ +NG EALE + + P S++ C + +G
Sbjct: 68 AKNVYLWNSIIRAFSKNGLFPEALEFYGKLRESKVSPDKYTFPSVIKACAGLFDAEMGDL 127
Query: 173 IHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANL 232
++ ++D E + + ALVD Y R A +VFD M V++ VSW ++ISG ++
Sbjct: 128 VYE-QILDMGFESDLFVGNALVDMYSRMGLLTRARQVFDEMPVRDLVSWNSLISGYSSHG 186
Query: 233 DYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSS 292
Y+ A + ++ + P+ T+ ++LPA VK G+ +HG+A + G+ S + ++
Sbjct: 187 YYEEALEIYHELKNSWIVPDSFTVSSVLPAFGNLLVVKQGQGLHGFALKSGVNSVVVVNN 246
Query: 293 ALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE 352
L+ MY + A ++F+ RD V ++++I Y + +++ +F + L ++ +
Sbjct: 247 GLVAMYLKFRRPTD-ARRVFDEMDVRDSVSYNTMICGYLKLEMVEESVRMFLENL-DQFK 304
Query: 353 PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQI 412
P+ +T+ +V+ AC +L L ++ ++LK G ++ N L+++YAKCG + + +
Sbjct: 305 PDLLTVSSVLRACGHLRDLSLAKYIYNYMLKAGFVLESTVRNILIDVYAKCGDMITARDV 364
Query: 413 FLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLV 472
F M+ +D++SW+++IS Y G +A+KLF M + D +T+L ++S +
Sbjct: 365 FNSMECKDTVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADL 424
Query: 473 TEGQQVFKQ-VNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
G+ + + + I L++ + L+D+ + G+V D+L+I +M T W++++
Sbjct: 425 KFGKGLHSNGIKSGICIDLSVSN--ALIDMYAKCGEVGDSLKIFSSMG-TGDTVTWNTVI 481
Query: 532 SACKLHGRLDIAEMLGPQLIRSE--PDNAANYTLLNMIYAEHGHWLHKE 578
SAC G + Q+ +SE PD A L M + L KE
Sbjct: 482 SACVRFGDFATGLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKE 530
Score = 108 bits (269), Expect = 6e-22
Identities = 81/346 (23%), Positives = 151/346 (43%), Gaps = 3/346 (0%)
Query: 15 TPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHG 74
T S + I I G + ++LF + + +I + L G LH
Sbjct: 373 TVSWNSIISGYIQSGDLMEAMKLFKMMMIMEEQADHITYLMLISVSTRLADLKFGKGLHS 432
Query: 75 LALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEE 134
+ +G D VSN+LI +YAK ++ + ++F +M D ++WN++I+A ++ G
Sbjct: 433 NGIKSGICIDLSVSNALIDMYAKCGEVGDSLKIFSSMGTGDTVTWNTVISACVRFGDFAT 492
Query: 135 ALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALV 194
L++ + +VP + +C R+G++IH ++ G E + + AL+
Sbjct: 493 GLQVTTQMRKSEVVPDMATFLVTLPMCASLAAKRLGKEIHCCLLRFG-YESELQIGNALI 551
Query: 195 DFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRV 254
+ Y +C +SRVF+ M ++ V+WT MI + + A F M+ G+ P+ V
Sbjct: 552 EMYSKCGCLENSSRVFERMSRRDVVTWTGMIYAYGMYGEGEKALETFADMEKSGIVPDSV 611
Query: 255 TLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFER 314
IA++ AC+ G V G H I A + + + AE+ +
Sbjct: 612 VFIAIIYACSHSGLVDEGLACFEKMKTHYKIDPMIEHYACVVDLLSRSQKISKAEEFIQA 671
Query: 315 SSFR-DVVLWSSIIGSYSQRGDSYKALTLFNKML-TEETEPNYVTL 358
+ D +W+S++ + GD A + +++ +P Y L
Sbjct: 672 MPIKPDASIWASVLRACRTSGDMETAERVSRRIIELNPDDPGYSIL 717
>UniRef100_Q9SS83 MZB10.7 protein [Arabidopsis thaliana]
Length = 1028
Score = 351 bits (901), Expect = 3e-95
Identities = 189/580 (32%), Positives = 318/580 (54%), Gaps = 4/580 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I+ G H+ +ELF + S + S++ C++ H +G+ H + +
Sbjct: 400 IRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSLLSTCAASHDLEMGSQFHSIIIKKKL 459
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
+ V N+L+ +YAK ++ ARQ+F+ M RD ++WN++I +Y+Q+ EA ++ K
Sbjct: 460 AKNLFVGNALVDMYAKCGALEDARQIFERMCDRDNVTWNTIIGSYVQDENESEAFDLFKR 519
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ L G+V LAS + C G G+Q+H L V G ++ + ++L+D Y +C
Sbjct: 520 MNLCGIVSDGACLASTLKACTHVHGLYQGKQVHCLSVKCG-LDRDLHTGSSLIDMYSKCG 578
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
A +VF + + VS A+I+G N + + A F+ M GV+P+ +T ++
Sbjct: 579 IIKDARKVFSSLPEWSVVSMNALIAGYSQN-NLEEAVVLFQEMLTRGVNPSEITFATIVE 637
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESS-HIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
AC KP + G + HG + G S +L+ MY S E SS + +
Sbjct: 638 ACHKPESLTLGTQFHGQITKRGFSSEGEYLGISLLGMYMNSRGMTEACALFSELSSPKSI 697
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
VLW+ ++ +SQ G +AL + +M + P+ T + V+ C+ LSSL+ G +H
Sbjct: 698 VLWTGMMSGHSQNGFYEEALKFYKEMRHDGVLPDQATFVTVLRVCSVLSSLREGRAIHSL 757
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSI-SWSTMISAYGLHGHGEQ 439
I SN L++MYAKCG ++GS Q+F EM+ R ++ SW+++I+ Y +G+ E
Sbjct: 758 IFHLAHDLDELTSNTLIDMYAKCGDMKGSSQVFDEMRRRSNVVSWNSLINGYAKNGYAED 817
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
ALK+F M++ + PD +TFL VL+AC+HAG V++G+++F+ + Y I ++H AC+V
Sbjct: 818 ALKIFDSMRQSHIMPDEITFLGVLTACSHAGKVSDGRKIFEMMIGQYGIEARVDHVACMV 877
Query: 500 DLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAA 559
DLLGR G +++A + + +KP R+WSSL+ AC++HG E+ +LI EP N++
Sbjct: 878 DLLGRWGYLQEADDFIEAQNLKPDARLWSSLLGACRIHGDDIRGEISAEKLIELEPQNSS 937
Query: 560 NYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
Y LL+ IYA G W + + M+ + +KK G+S I+
Sbjct: 938 AYVLLSNIYASQGCWEKANALRKVMRDRGVKKVPGYSWID 977
Score = 207 bits (526), Expect = 9e-52
Identities = 134/508 (26%), Positives = 252/508 (49%), Gaps = 10/508 (1%)
Query: 35 LELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISL 94
+E F + S+ S S L SV+ A + + LG ++H A+ G S+ V +SL+S+
Sbjct: 312 IEYFFNMRKSSVKSTRSTLGSVLSAIGIVANLDLGLVVHAEAIKLGLASNIYVGSSLVSM 371
Query: 95 YAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELL 154
Y+K +++A +VF+ + ++ + WN+MI Y NG + +E+ + G
Sbjct: 372 YSKCEKMEAAAKVFEALEEKNDVFWNAMIRGYAHNGESHKVMELFMDMKSSGYNIDDFTF 431
Query: 155 ASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGME 214
S++S C +G Q H+ +++ ++ ++ + ALVD Y +C A ++F+ M
Sbjct: 432 TSLLSTCAASHDLEMGSQFHS-IIIKKKLAKNLFVGNALVDMYAKCGALEDARQIFERMC 490
Query: 215 VKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKE 274
++ V+W +I + + + AF F+ M + G+ + L + L AC + GK+
Sbjct: 491 DRDNVTWNTIIGSYVQDENESEAFDLFKRMNLCGIVSDGACLASTLKACTHVHGLYQGKQ 550
Query: 275 IHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRG 334
+H + + G++ S+LI+MY + G + A ++F VV +++I YSQ
Sbjct: 551 VHCLSVKCGLDRDLHTGSSLIDMYSKCG-IIKDARKVFSSLPEWSVVSMNALIAGYSQ-N 608
Query: 335 DSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSIS-LS 393
+ +A+ LF +MLT P+ +T ++ AC SL G HG I K G S L
Sbjct: 609 NLEEAVVLFQEMLTRGVNPSEITFATIVEACHKPESLTLGTQFHGQITKRGFSSEGEYLG 668
Query: 394 NALMNMYAKCGCLEGSLQIFLEMQNRDSI-SWSTMISAYGLHGHGEQALKLFSEMKERGV 452
+L+ MY + + +F E+ + SI W+ M+S + +G E+ALK + EM+ GV
Sbjct: 669 ISLLGMYMNSRGMTEACALFSELSSPKSIVLWTGMMSGHSQNGFYEEALKFYKEMRHDGV 728
Query: 453 KPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYA--CLVDLLGRSGKVED 510
PD TF+ VL C+ + EG+ + + + + ++ L+D+ + G ++
Sbjct: 729 LPDQATFVTVLRVCSVLSSLREGRAIHSLI---FHLAHDLDELTSNTLIDMYAKCGDMKG 785
Query: 511 ALEIMRTMPMKPSTRIWSSLVSACKLHG 538
+ ++ M + + W+SL++ +G
Sbjct: 786 SSQVFDEMRRRSNVVSWNSLINGYAKNG 813
Score = 192 bits (489), Expect = 2e-47
Identities = 132/492 (26%), Positives = 236/492 (47%), Gaps = 11/492 (2%)
Query: 80 GSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEML 139
G D + ++I+ Y + ++ AR +F M D ++WN MI+ + + GC A+E
Sbjct: 256 GHRPDHLAFVTVINTYIRLGKLKDARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYF 315
Query: 140 KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFR 199
+ + L S++S G +G +HA + G + ++ + ++LV Y +
Sbjct: 316 FNMRKSSVKSTRSTLGSVLSAIGIVANLDLGLVVHAEAIKLG-LASNIYVGSSLVSMYSK 374
Query: 200 CDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIAL 259
C+ A++VF+ +E KN+V W AMI G N + F M+ G + + T +L
Sbjct: 375 CEKMEAAAKVFEALEEKNDVFWNAMIRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSL 434
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
L CA ++ G + H + + + +AL++MY + G +L A QIFER RD
Sbjct: 435 LSTCAASHDLEMGSQFHSIIIKKKLAKNLFVGNALVDMYAKCG-ALEDARQIFERMCDRD 493
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
V W++IIGSY Q + +A LF +M + L + + ACT++ L G +H
Sbjct: 494 NVTWNTIIGSYVQDENESEAFDLFKRMNLCGIVSDGACLASTLKACTHVHGLYQGKQVHC 553
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
+K GL + ++L++MY+KCG ++ + ++F + +S + +I+ Y + E+
Sbjct: 554 LSVKCGLDRDLHTGSSLIDMYSKCGIIKDARKVFSSLPEWSVVSMNALIAGYS-QNNLEE 612
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
A+ LF EM RGV P +TF ++ AC+ +T G Q Q+ + + E +
Sbjct: 613 AVVLFQEMLTRGVNPSEITFATIVEACHKPESLTLGTQFHGQIT---KRGFSSEGEYLGI 669
Query: 500 DLLG---RSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSE-- 554
LLG S + +A + + S +W+ ++S +G + A ++
Sbjct: 670 SLLGMYMNSRGMTEACALFSELSSPKSIVLWTGMMSGHSQNGFYEEALKFYKEMRHDGVL 729
Query: 555 PDNAANYTLLNM 566
PD A T+L +
Sbjct: 730 PDQATFVTVLRV 741
Score = 175 bits (444), Expect = 3e-42
Identities = 124/467 (26%), Positives = 220/467 (46%), Gaps = 42/467 (8%)
Query: 68 LGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYL 127
+G +H +L G S+ + N+++ LYAK + + A + FD + +D +WNSM++ Y
Sbjct: 78 IGKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQVSYAEKQFDFL-EKDVTAWNSMLSMYS 136
Query: 128 QNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSV 187
G + L ++ + P + ++S C R+ GRQIH ++ G +E +
Sbjct: 137 SIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSMIKMG-LERNS 195
Query: 188 VLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCI-ANLDYDAAFARFRAMQV 246
ALVD Y +CD A RVF+ + N V WT + SG + A L +A F M+
Sbjct: 196 YCGGALVDMYAKCDRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLV-FERMRD 254
Query: 247 EGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLH 306
EG P+ H+ +IN Y + G+ L
Sbjct: 255 EGHRPD-----------------------------------HLAFVTVINTYIRLGK-LK 278
Query: 307 LAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACT 366
A +F S DVV W+ +I + +RG A+ F M + TL +V+SA
Sbjct: 279 DARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYFFNMRKSSVKSTRSTLGSVLSAIG 338
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
+++L G +H +K GL+ +I + ++L++MY+KC +E + ++F ++ ++ + W+
Sbjct: 339 IVANLDLGLVVHAEAIKLGLASNIYVGSSLVSMYSKCEKMEAAAKVFEALEEKNDVFWNA 398
Query: 427 MISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY 486
MI Y +G + ++LF +MK G D TF ++LS C + + G Q F +
Sbjct: 399 MIRGYAHNGESHKVMELFMDMKSSGYNIDDFTFTSLLSTCAASHDLEMGSQ-FHSIIIKK 457
Query: 487 EIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSA 533
++ + LVD+ + G +EDA +I M + + W++++ +
Sbjct: 458 KLAKNLFVGNALVDMYAKCGALEDARQIFERMCDRDNV-TWNTIIGS 503
Score = 107 bits (267), Expect = 1e-21
Identities = 69/284 (24%), Positives = 143/284 (50%), Gaps = 10/284 (3%)
Query: 272 GKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYS 331
GK +H + GI+S +A++++Y + + + AE+ F+ +DV W+S++ YS
Sbjct: 79 GKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQ-VSYAEKQFDFLE-KDVTAWNSMLSMYS 136
Query: 332 QRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSIS 391
G K L F + + PN T V+S C ++++ G +H ++K GL +
Sbjct: 137 SIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSMIKMGLERNSY 196
Query: 392 LSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERG 451
AL++MYAKC + + ++F + + +++ W+ + S Y G E+A+ +F M++ G
Sbjct: 197 CGGALVDMYAKCDRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLVFERMRDEG 256
Query: 452 VKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDA 511
+PD L F+ V++ G + + + +F ++++ + + ++ G+ G A
Sbjct: 257 HRPDHLAFVTVINTYIRLGKLKDARLLFGEMSSPDVVAWNV-----MISGHGKRGCETVA 311
Query: 512 LEI---MRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIR 552
+E MR +K + S++SA + LD+ ++ + I+
Sbjct: 312 IEYFFNMRKSSVKSTRSTLGSVLSAIGIVANLDLGLVVHAEAIK 355
Score = 104 bits (260), Expect = 7e-21
Identities = 85/366 (23%), Positives = 161/366 (43%), Gaps = 44/366 (12%)
Query: 168 RIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISG 227
RIG+ +H+ ++ G ++ L A+VD Y +C A + FD +E K+ +W +M+S
Sbjct: 77 RIGKAVHSKSLILG-IDSEGRLGNAIVDLYAKCAQVSYAEKQFDFLE-KDVTAWNSMLSM 134
Query: 228 CIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESS 287
+ F ++ + PN+ T +L CA+ V+ G++IH + G+E +
Sbjct: 135 YSSIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSMIKMGLERN 194
Query: 288 HIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKML 347
AL++MY + + + A ++FE + V W+ + Y + G +A+ +F +M
Sbjct: 195 SYCGGALVDMYAKC-DRISDARRVFEWIVDPNTVCWTCLFSGYVKAGLPEEAVLVFERMR 253
Query: 348 TEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLE 407
E P+++ + VI N Y + G L+
Sbjct: 254 DEGHRPDHLAFVTVI-----------------------------------NTYIRLGKLK 278
Query: 408 GSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACN 467
+ +F EM + D ++W+ MIS +G G A++ F M++ VK T +VLSA
Sbjct: 279 DARLLFGEMSSPDVVAWNVMISGHGKRGCETVAIEYFFNMRKSSVKSTRSTLGSVLSAIG 338
Query: 468 HAGLVTEGQQVFKQVNADYEIPLTIEHY--ACLVDLLGRSGKVEDALEIMRTMPMKPSTR 525
+ G V + ++ L Y + LV + + K+E A ++ + K
Sbjct: 339 IVANLDLGLVVHAEA---IKLGLASNIYVGSSLVSMYSKCEKMEAAAKVFEALEEKNDV- 394
Query: 526 IWSSLV 531
W++++
Sbjct: 395 FWNAMI 400
Score = 64.7 bits (156), Expect = 8e-09
Identities = 53/201 (26%), Positives = 97/201 (47%), Gaps = 11/201 (5%)
Query: 370 SLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMIS 429
+L+ G +H L G+ L NA++++YAKC + + + F + +D +W++M+S
Sbjct: 75 ALRIGKAVHSKSLILGIDSEGRLGNAIVDLYAKCAQVSYAEKQF-DFLEKDVTAWNSMLS 133
Query: 430 AYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIP 489
Y G + L+ F + E + P+ TF VLS C V G+Q+ + ++
Sbjct: 134 MYSSIGKPGKVLRSFVSLFENQIFPNKFTFSIVLSTCARETNVEFGRQIHCSM---IKMG 190
Query: 490 LTIEHY--ACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLG 547
L Y LVD+ + ++ DA + + + P+T W+ L S G + A ++
Sbjct: 191 LERNSYCGGALVDMYAKCDRISDARRVFEWI-VDPNTVCWTCLFSGYVKAGLPEEAVLVF 249
Query: 548 PQLIRSE---PDNAANYTLLN 565
++ R E PD+ A T++N
Sbjct: 250 ERM-RDEGHRPDHLAFVTVIN 269
>UniRef100_Q9SCM9 Hypothetical protein T8H10.30 [Arabidopsis thaliana]
Length = 803
Score = 351 bits (901), Expect = 3e-95
Identities = 197/598 (32%), Positives = 336/598 (55%), Gaps = 18/598 (3%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSL---HSHTLGTLLHGLALT 78
I SL S + LE F + S L SV+ ACS+L +G +H L
Sbjct: 84 ISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMMGKQVHAYGLR 143
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G + ++ N+L+++Y K + S++ + + RD ++WN+++++ QN L EALE
Sbjct: 144 KGELNSFII-NTLVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLCQNEQLLEALEY 202
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
L+ + L G+ P ++S++ C R G+++HA + +G ++ + + +ALVD Y
Sbjct: 203 LREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENSFVGSALVDMYC 262
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVE-GVDPNRVTLI 257
C L RVFDGM + W AMI+G N A F M+ G+ N T+
Sbjct: 263 NCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGMEESAGLLANSTTMA 322
Query: 258 ALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSF 317
++PAC + G + IHG+ + G++ + L++MY + G+ + +A +IF +
Sbjct: 323 GVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGK-IDIAMRIFGKMED 381
Query: 318 RDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE-----------PNYVTLLAVISACT 366
RD+V W+++I Y AL L +KM E + PN +TL+ ++ +C
Sbjct: 382 RDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKPNSITLMTILPSCA 441
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
LS+L G +H + +K L+ +++ +AL++MYAKCGCL+ S ++F ++ ++ I+W+
Sbjct: 442 ALSALAKGKEIHAYAIKNNLATDVAVGSALVDMYAKCGCLQMSRKVFDQIPQKNVITWNV 501
Query: 427 MISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY 486
+I AYG+HG+G++A+ L M +GVKP+ +TF++V +AC+H+G+V EG ++F + DY
Sbjct: 502 IIMAYGMHGNGQEAIDLLRMMMVQGVKPNEVTFISVFAACSHSGMVDEGLRIFYVMKPDY 561
Query: 487 EIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMK-PSTRIWSSLVSACKLHGRLDIAEM 545
+ + +HYAC+VDLLGR+G++++A ++M MP WSSL+ A ++H L+I E+
Sbjct: 562 GVEPSSDHYACVVDLLGRAGRIKEAYQLMNMMPRDFNKAGAWSSLLGASRIHNNLEIGEI 621
Query: 546 LGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
LI+ EP+ A++Y LL IY+ G W +V MK Q ++K G S IE G++
Sbjct: 622 AAQNLIQLEPNVASHYVLLANIYSSAGLWDKATEVRRNMKEQGVRKEPGCSWIEHGDE 679
Score = 216 bits (550), Expect = 2e-54
Identities = 146/522 (27%), Positives = 269/522 (50%), Gaps = 30/522 (5%)
Query: 54 PSVIKACSSLHSHTLGTLLHGLALTTGSHSDPV-VSNSLISLYAKFSDIQSARQVFDTMP 112
P+++KA + L LG +H G D V V+N+L++LY K D + +VFD +
Sbjct: 14 PALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDFGAVYKVFDRIS 73
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGR---KMGSRI 169
R+ +SWNS+I++ E ALE + + + P L S+V+ C G +
Sbjct: 74 ERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMM 133
Query: 170 GRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCI 229
G+Q+HA + G + +S +++T LV Y + + + ++ V+W ++S
Sbjct: 134 GKQVHAYGLRKGEL-NSFIINT-LVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLC 191
Query: 230 ANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHG-IESSH 288
N A R M +EGV+P+ T+ ++LPAC+ ++ GKE+H YA ++G ++ +
Sbjct: 192 QNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENS 251
Query: 289 IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLT 348
SAL++MYC + L ++F+ R + LW+++I YSQ +AL LF M
Sbjct: 252 FVGSALVDMYCNCKQVLS-GRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGM-- 308
Query: 349 EETE---PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGC 405
EE+ N T+ V+ AC + +HGF++K GL + N LM+MY++ G
Sbjct: 309 EESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGK 368
Query: 406 LEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK--ERGV---------KP 454
++ +++IF +M++RD ++W+TMI+ Y H E AL L +M+ ER V KP
Sbjct: 369 IDIAMRIFGKMEDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKP 428
Query: 455 DALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEI 514
+++T + +L +C + +G+++ + + + + LVD+ + G ++ + ++
Sbjct: 429 NSITLMTILPSCAALSALAKGKEIHAYAIKN-NLATDVAVGSALVDMYAKCGCLQMSRKV 487
Query: 515 MRTMPMKPSTRIWSSLVSACKLHGR----LDIAEMLGPQLIR 552
+P K + W+ ++ A +HG +D+ M+ Q ++
Sbjct: 488 FDQIPQK-NVITWNVIIMAYGMHGNGQEAIDLLRMMMVQGVK 528
Score = 164 bits (415), Expect = 7e-39
Identities = 114/399 (28%), Positives = 205/399 (50%), Gaps = 19/399 (4%)
Query: 144 LLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDA 203
+LG+ P +++ +G+QIHA V G SV ++ LV+ Y +C D
Sbjct: 3 VLGIKPDNYAFPALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDF 62
Query: 204 LMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPAC 263
+VFD + +N+VSW ++IS + ++ A FR M E V+P+ TL++++ AC
Sbjct: 63 GAVYKVFDRISERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTAC 122
Query: 264 AK---PGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
+ P + GK++H Y R G +S I ++ L+ MY + G+ L ++ + RD+
Sbjct: 123 SNLPMPEGLMMGKQVHAYGLRKGELNSFIINT-LVAMYGKLGK-LASSKVLLGSFGGRDL 180
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
V W++++ S Q +AL +M+ E EP+ T+ +V+ AC++L L+ G LH +
Sbjct: 181 VTWNTVLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAY 240
Query: 381 ILKFG-LSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
LK G L + + +AL++MY C + ++F M +R W+ MI+ Y + H ++
Sbjct: 241 ALKNGSLDENSFVGSALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKE 300
Query: 440 ALKLFSEMKE-RGVKPDALTFLAVLSACNHAGLVT-----EGQQVFKQVNADYEIPLTIE 493
AL LF M+E G+ ++ T V+ AC +G + G V + ++ D + T
Sbjct: 301 ALLLFIGMEESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNT-- 358
Query: 494 HYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVS 532
L+D+ R GK++ A+ I M + W+++++
Sbjct: 359 ----LMDMYSRLGKIDIAMRIFGKMEDRDLV-TWNTMIT 392
Score = 115 bits (289), Expect = 3e-24
Identities = 78/240 (32%), Positives = 129/240 (53%), Gaps = 8/240 (3%)
Query: 244 MQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHG--YAFRHGIESSHIFSSALINMYCQS 301
M V G+ P+ ALL A A ++ GK+IH Y F +G++S + ++ L+N+Y +
Sbjct: 1 MIVLGIKPDNYAFPALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTV-ANTLVNLYRKC 59
Query: 302 GESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAV 361
G+ ++F+R S R+ V W+S+I S AL F ML E EP+ TL++V
Sbjct: 60 GD-FGAVYKVFDRISERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSV 118
Query: 362 ISACTNL---SSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQN 418
++AC+NL L G +H + L+ G + + N L+ MY K G L S +
Sbjct: 119 VTACSNLPMPEGLMMGKQVHAYGLRKG-ELNSFIINTLVAMYGKLGKLASSKVLLGSFGG 177
Query: 419 RDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQV 478
RD ++W+T++S+ + +AL+ EM GV+PD T +VL AC+H ++ G+++
Sbjct: 178 RDLVTWNTVLSSLCQNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKEL 237
>UniRef100_Q7Y211 Hypothetical protein At3g57430 [Arabidopsis thaliana]
Length = 890
Score = 351 bits (901), Expect = 3e-95
Identities = 197/598 (32%), Positives = 336/598 (55%), Gaps = 18/598 (3%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSL---HSHTLGTLLHGLALT 78
I SL S + LE F + S L SV+ ACS+L +G +H L
Sbjct: 171 ISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMMGKQVHAYGLR 230
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G + ++ N+L+++Y K + S++ + + RD ++WN+++++ QN L EALE
Sbjct: 231 KGELNSFII-NTLVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLCQNEQLLEALEY 289
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
L+ + L G+ P ++S++ C R G+++HA + +G ++ + + +ALVD Y
Sbjct: 290 LREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENSFVGSALVDMYC 349
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVE-GVDPNRVTLI 257
C L RVFDGM + W AMI+G N A F M+ G+ N T+
Sbjct: 350 NCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGMEESAGLLANSTTMA 409
Query: 258 ALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSF 317
++PAC + G + IHG+ + G++ + L++MY + G+ + +A +IF +
Sbjct: 410 GVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGK-IDIAMRIFGKMED 468
Query: 318 RDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE-----------PNYVTLLAVISACT 366
RD+V W+++I Y AL L +KM E + PN +TL+ ++ +C
Sbjct: 469 RDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKPNSITLMTILPSCA 528
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
LS+L G +H + +K L+ +++ +AL++MYAKCGCL+ S ++F ++ ++ I+W+
Sbjct: 529 ALSALAKGKEIHAYAIKNNLATDVAVGSALVDMYAKCGCLQMSRKVFDQIPQKNVITWNV 588
Query: 427 MISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY 486
+I AYG+HG+G++A+ L M +GVKP+ +TF++V +AC+H+G+V EG ++F + DY
Sbjct: 589 IIMAYGMHGNGQEAIDLLRMMMVQGVKPNEVTFISVFAACSHSGMVDEGLRIFYVMKPDY 648
Query: 487 EIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMK-PSTRIWSSLVSACKLHGRLDIAEM 545
+ + +HYAC+VDLLGR+G++++A ++M MP WSSL+ A ++H L+I E+
Sbjct: 649 GVEPSSDHYACVVDLLGRAGRIKEAYQLMNMMPRDFNKAGAWSSLLGASRIHNNLEIGEI 708
Query: 546 LGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
LI+ EP+ A++Y LL IY+ G W +V MK Q ++K G S IE G++
Sbjct: 709 AAQNLIQLEPNVASHYVLLANIYSSAGLWDKATEVRRNMKEQGVRKEPGCSWIEHGDE 766
Score = 216 bits (550), Expect = 2e-54
Identities = 146/522 (27%), Positives = 269/522 (50%), Gaps = 30/522 (5%)
Query: 54 PSVIKACSSLHSHTLGTLLHGLALTTGSHSDPV-VSNSLISLYAKFSDIQSARQVFDTMP 112
P+++KA + L LG +H G D V V+N+L++LY K D + +VFD +
Sbjct: 101 PALLKAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDFGAVYKVFDRIS 160
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGR---KMGSRI 169
R+ +SWNS+I++ E ALE + + + P L S+V+ C G +
Sbjct: 161 ERNQVSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMM 220
Query: 170 GRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCI 229
G+Q+HA + G + +S +++T LV Y + + + ++ V+W ++S
Sbjct: 221 GKQVHAYGLRKGEL-NSFIINT-LVAMYGKLGKLASSKVLLGSFGGRDLVTWNTVLSSLC 278
Query: 230 ANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHG-IESSH 288
N A R M +EGV+P+ T+ ++LPAC+ ++ GKE+H YA ++G ++ +
Sbjct: 279 QNEQLLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENS 338
Query: 289 IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLT 348
SAL++MYC + L ++F+ R + LW+++I YSQ +AL LF M
Sbjct: 339 FVGSALVDMYCNCKQVLS-GRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGM-- 395
Query: 349 EETE---PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGC 405
EE+ N T+ V+ AC + +HGF++K GL + N LM+MY++ G
Sbjct: 396 EESAGLLANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNTLMDMYSRLGK 455
Query: 406 LEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK--ERGV---------KP 454
++ +++IF +M++RD ++W+TMI+ Y H E AL L +M+ ER V KP
Sbjct: 456 IDIAMRIFGKMEDRDLVTWNTMITGYVFSEHHEDALLLLHKMQNLERKVSKGASRVSLKP 515
Query: 455 DALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEI 514
+++T + +L +C + +G+++ + + + + LVD+ + G ++ + ++
Sbjct: 516 NSITLMTILPSCAALSALAKGKEIHAYAIKN-NLATDVAVGSALVDMYAKCGCLQMSRKV 574
Query: 515 MRTMPMKPSTRIWSSLVSACKLHGR----LDIAEMLGPQLIR 552
+P K + W+ ++ A +HG +D+ M+ Q ++
Sbjct: 575 FDQIPQK-NVITWNVIIMAYGMHGNGQEAIDLLRMMMVQGVK 615
Score = 174 bits (441), Expect = 7e-42
Identities = 123/444 (27%), Positives = 225/444 (49%), Gaps = 20/444 (4%)
Query: 99 SDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMV 158
S + A +F + R P W ++ + +++ L EA+ + +LG+ P +++
Sbjct: 46 SAVSGAPSIFISQS-RSPEWWIDLLRSKVRSNLLREAVLTYVDMIVLGIKPDNYAFPALL 104
Query: 159 SLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNE 218
+G+QIHA V G SV ++ LV+ Y +C D +VFD + +N+
Sbjct: 105 KAVADLQDMELGKQIHAHVYKFGYGVDSVTVANTLVNLYRKCGDFGAVYKVFDRISERNQ 164
Query: 219 VSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAK---PGFVKHGKEI 275
VSW ++IS + ++ A FR M E V+P+ TL++++ AC+ P + GK++
Sbjct: 165 VSWNSLISSLCSFEKWEMALEAFRCMLDENVEPSSFTLVSVVTACSNLPMPEGLMMGKQV 224
Query: 276 HGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGD 335
H Y R G +S I ++ L+ MY + G+ L ++ + RD+V W++++ S Q
Sbjct: 225 HAYGLRKGELNSFIINT-LVAMYGKLGK-LASSKVLLGSFGGRDLVTWNTVLSSLCQNEQ 282
Query: 336 SYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFG-LSFSISLSN 394
+AL +M+ E EP+ T+ +V+ AC++L L+ G LH + LK G L + + +
Sbjct: 283 LLEALEYLREMVLEGVEPDEFTISSVLPACSHLEMLRTGKELHAYALKNGSLDENSFVGS 342
Query: 395 ALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKE-RGVK 453
AL++MY C + ++F M +R W+ MI+ Y + H ++AL LF M+E G+
Sbjct: 343 ALVDMYCNCKQVLSGRRVFDGMFDRKIGLWNAMIAGYSQNEHDKEALLLFIGMEESAGLL 402
Query: 454 PDALTFLAVLSACNHAGLVT-----EGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKV 508
++ T V+ AC +G + G V + ++ D + T L+D+ R GK+
Sbjct: 403 ANSTTMAGVVPACVRSGAFSRKEAIHGFVVKRGLDRDRFVQNT------LMDMYSRLGKI 456
Query: 509 EDALEIMRTMPMKPSTRIWSSLVS 532
+ A+ I M + W+++++
Sbjct: 457 DIAMRIFGKMEDRDLV-TWNTMIT 479
>UniRef100_Q5W965 PPR868-14 [Physcomitrella patens]
Length = 868
Score = 348 bits (894), Expect = 2e-94
Identities = 188/582 (32%), Positives = 318/582 (54%), Gaps = 40/582 (6%)
Query: 55 SVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHR 114
S++KAC++ G +H + G +D V+ +LI++Y+K +I A +VF M R
Sbjct: 162 SILKACNNYSILEKGRKIHTIVKAMGMETDVAVATALITMYSKCGEISVACEVFHKMTER 221
Query: 115 DPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIH 174
+ +SW ++I A Q+ L EA E+ + + G+ P S+++ C GR+IH
Sbjct: 222 NVVSWTAIIQANAQHRKLNEAFELYEQMLQAGISPNAVTFVSLLNSCNTPEALNRGRRIH 281
Query: 175 ALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDY 234
+ + G +E ++++ AL+ Y +C+ A +FD M ++ +SW+AMI+G A Y
Sbjct: 282 SHISERG-LETDMIVANALITMYCKCNSVQEAREIFDRMSKRDVISWSAMIAG-YAQSGY 339
Query: 235 ------DAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSH 288
D F M+ EGV PN+VT +++L AC G ++ G++IH + G E
Sbjct: 340 KDKESIDEVFQLLERMRREGVFPNKVTFMSILRACTAHGALEQGRQIHAELSKVGFELDR 399
Query: 289 IFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSS----------------------- 325
+A+ NMY + G S++ AEQ+F + + ++VV W+S
Sbjct: 400 SLQTAIFNMYAKCG-SIYEAEQVFSKMANKNVVAWTSFLSMYIKCGDLSSAEKVFSEMPT 458
Query: 326 --------IIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGL 377
+I Y+Q GD K L + M E +P+ VT++ ++ AC L+ L+ G +
Sbjct: 459 RNVVSWNLMIAGYAQNGDIVKVFELLSSMKAEGFQPDRVTVITILEACGALAGLERGKLV 518
Query: 378 HGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHG 437
H +K GL ++ +L+ MY+KCG + + +F +M NRD+++W+ M++ YG HG G
Sbjct: 519 HAEAVKLGLESDTVVATSLIGMYSKCGQVAEARTVFDKMSNRDTVAWNAMLAGYGQHGDG 578
Query: 438 EQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYAC 497
+A+ LF M + V P+ +T AV+SAC+ AGLV EG+++F+ + D+++ +HY C
Sbjct: 579 LEAVDLFKRMLKERVSPNEITLTAVISACSRAGLVQEGREIFRMMQEDFKMTPRKQHYGC 638
Query: 498 LVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDN 557
+VDLLGR+G++++A E +++MP +P +W +L+ ACK H + +AE ++ EP
Sbjct: 639 MVDLLGRAGRLQEAEEFIQSMPCEPDISVWHALLGACKSHNNVQLAERAAHHILELEPSY 698
Query: 558 AANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
A+ Y L+ IYA+ G W +V M + LKK G S IE
Sbjct: 699 ASVYITLSNIYAQAGRWDDSTKVRRVMDDRGLKKDRGESSIE 740
Score = 249 bits (636), Expect = 2e-64
Identities = 164/562 (29%), Positives = 262/562 (46%), Gaps = 42/562 (7%)
Query: 14 PTPSPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLH 73
PT ++ L G + ++L + S+ VI+ C+ G ++H
Sbjct: 20 PTSVSGGEVWRLCKAGRLREAIQLLGIIKQRGLLVNSNTYGCVIEHCAKARRFEDGKMVH 79
Query: 74 GLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLE 133
G D + NSLI+ Y+KF D+ SA QVF M RD ++W+SMI AY N
Sbjct: 80 KQLDELGVEIDIYLGNSLINFYSKFEDVASAEQVFRRMTLRDVVTWSSMIAAYAGNNHPA 139
Query: 134 EALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTAL 193
+A + + + + P S++ C GR+IH +V G +E V ++TAL
Sbjct: 140 KAFDTFERMTDANIEPNRITFLSILKACNNYSILEKGRKIHTIVKAMG-METDVAVATAL 198
Query: 194 VDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNR 253
+ Y +C + +A VF M +N VSWTA+I + + AF + M G+ PN
Sbjct: 199 ITMYSKCGEISVACEVFHKMTERNVVSWTAIIQANAQHRKLNEAFELYEQMLQAGISPNA 258
Query: 254 VTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFE 313
VT ++LL +C P + G+ IH + G+E+ I ++ALI MYC+ S+ A +IF+
Sbjct: 259 VTFVSLLNSCNTPEALNRGRRIHSHISERGLETDMIVANALITMYCKC-NSVQEAREIFD 317
Query: 314 RSSFRDVVLWSSIIGSYSQRGDSYK-----ALTLFNKMLTEETEPNYVTLLAVISACTNL 368
R S RDV+ WS++I Y+Q G K L +M E PN VT ++++ ACT
Sbjct: 318 RMSKRDVISWSAMIAGYAQSGYKDKESIDEVFQLLERMRREGVFPNKVTFMSILRACTAH 377
Query: 369 SSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGC----------------------- 405
+L+ G +H + K G SL A+ NMYAKCG
Sbjct: 378 GALEQGRQIHAELSKVGFELDRSLQTAIFNMYAKCGSIYEAEQVFSKMANKNVVAWTSFL 437
Query: 406 --------LEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDAL 457
L + ++F EM R+ +SW+ MI+ Y +G + +L S MK G +PD +
Sbjct: 438 SMYIKCGDLSSAEKVFSEMPTRNVVSWNLMIAGYAQNGDIVKVFELLSSMKAEGFQPDRV 497
Query: 458 TFLAVLSACNHAGLVTEGQQVFKQ-VNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMR 516
T + +L AC + G+ V + V E + L+ + + G+V +A +
Sbjct: 498 TVITILEACGALAGLERGKLVHAEAVKLGLESDTVVA--TSLIGMYSKCGQVAEARTVFD 555
Query: 517 TMPMKPSTRIWSSLVSACKLHG 538
M + T W+++++ HG
Sbjct: 556 KMSNR-DTVAWNAMLAGYGQHG 576
Score = 209 bits (531), Expect = 2e-52
Identities = 126/464 (27%), Positives = 245/464 (52%), Gaps = 14/464 (3%)
Query: 130 GCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVL 189
G L EA+++L + GL+ ++ C + G+ +H + G VE + L
Sbjct: 35 GRLREAIQLLGIIKQRGLLVNSNTYGCVIEHCAKARRFEDGKMVHKQLDELG-VEIDIYL 93
Query: 190 STALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGV 249
+L++FY + +D A +VF M +++ V+W++MI+ N AF F M +
Sbjct: 94 GNSLINFYSKFEDVASAEQVFRRMTLRDVVTWSSMIAAYAGNNHPAKAFDTFERMTDANI 153
Query: 250 DPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAE 309
+PNR+T +++L AC ++ G++IH G+E+ ++ALI MY + GE + +A
Sbjct: 154 EPNRITFLSILKACNNYSILEKGRKIHTIVKAMGMETDVAVATALITMYSKCGE-ISVAC 212
Query: 310 QIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLS 369
++F + + R+VV W++II + +Q +A L+ +ML PN VT ++++++C
Sbjct: 213 EVFHKMTERNVVSWTAIIQANAQHRKLNEAFELYEQMLQAGISPNAVTFVSLLNSCNTPE 272
Query: 370 SLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMIS 429
+L G +H I + GL + ++NAL+ MY KC ++ + +IF M RD ISWS MI+
Sbjct: 273 ALNRGRRIHSHISERGLETDMIVANALITMYCKCNSVQEAREIFDRMSKRDVISWSAMIA 332
Query: 430 AYGLHGHG-----EQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVN- 483
Y G+ ++ +L M+ GV P+ +TF+++L AC G + +G+Q+ +++
Sbjct: 333 GYAQSGYKDKESIDEVFQLLERMRREGVFPNKVTFMSILRACTAHGALEQGRQIHAELSK 392
Query: 484 ADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIA 543
+E+ +++ + ++ + G + +A ++ M K + W+S +S G L A
Sbjct: 393 VGFELDRSLQ--TAIFNMYAKCGSIYEAEQVFSKMANK-NVVAWTSFLSMYIKCGDLSSA 449
Query: 544 EMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQ 587
E + ++ N ++ L+ YA++G + +++ +MK +
Sbjct: 450 EKVFSEM---PTRNVVSWNLMIAGYAQNGDIVKVFELLSSMKAE 490
>UniRef100_Q6Z0F9 PPR-repeat protein-like [Oryza sativa]
Length = 819
Score = 348 bits (892), Expect = 3e-94
Identities = 197/565 (34%), Positives = 309/565 (53%), Gaps = 2/565 (0%)
Query: 35 LELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISL 94
LELF ++ VL S + ACS+L G +HG A + + +D V N LI L
Sbjct: 206 LELFDRMGIEGVRPDRFVLASAVSACSALGFLEGGRQIHGYAYRSATETDTSVINVLIDL 265
Query: 95 YAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELL 154
Y K S + +AR++FD M +R+ +SW +MI+ Y+QN EA+ M + G P
Sbjct: 266 YCKCSRLSAARKLFDCMEYRNLVSWTTMISGYMQNSFNAEAITMFWNMTQAGWQPDGFAC 325
Query: 155 ASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGME 214
S+++ CG GRQIHA V+ +E + AL+D Y +C+ A VFD +
Sbjct: 326 TSILNSCGSLAAIWQGRQIHAHVI-KADLEADEYVKNALIDMYAKCEHLTEARAVFDALA 384
Query: 215 VKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKE 274
+ +S+ AMI G N D A F+ M+ + P+ +T ++LL + ++ K+
Sbjct: 385 EDDAISYNAMIEGYSKNRDLAEAVNIFQRMRFFSLRPSLLTFVSLLGVSSSQLAIELSKQ 444
Query: 275 IHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRG 334
IHG + G +SALI++Y + ++ A+ +F ++D+V+W+S+I ++Q
Sbjct: 445 IHGLIIKSGTSLDLYAASALIDVYSKCS-LVNDAKTVFNMLHYKDMVIWNSMIFGHAQNE 503
Query: 335 DSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSN 394
+A+ LFN++L PN T +A+++ + L+S+ HG H +I+K G+ +SN
Sbjct: 504 QGEEAIKLFNQLLLSGMAPNEFTFVALVTVASTLASMFHGQQFHAWIIKAGVDNDPHVSN 563
Query: 395 ALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKP 454
AL++MYAKCG ++ +F D I W++MI+ Y HGH E+AL++F M E V+P
Sbjct: 564 ALIDMYAKCGFIKEGRMLFESTCGEDVICWNSMITTYAQHGHAEEALQVFRLMGEAEVEP 623
Query: 455 DALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEI 514
+ +TF+ VLSAC HAG V EG F + ++Y+I IEHYA +V+L GRSGK+ A E
Sbjct: 624 NYVTFVGVLSACAHAGFVGEGLNHFNSMKSNYDIEPGIEHYASVVNLFGRSGKLHAAKEF 683
Query: 515 MRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHW 574
+ MP+KP+ +W SL+SAC L G +I + ++P ++ Y LL+ IYA G W
Sbjct: 684 IERMPIKPAAAVWRSLLSACHLFGNAEIGRYAAEMALLADPTDSGPYVLLSNIYASKGLW 743
Query: 575 LHKEQVMEAMKLQRLKKCYGFSRIE 599
+ + M K G S IE
Sbjct: 744 ADVHNLRQQMDSSGTVKETGCSWIE 768
Score = 228 bits (580), Expect = 5e-58
Identities = 135/493 (27%), Positives = 255/493 (51%), Gaps = 6/493 (1%)
Query: 52 VLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTM 111
+L SV++AC+ + +LG +HG+A+ ++ V +LI+LYAK + A VF +
Sbjct: 122 LLASVLRACTQSKAVSLGEQVHGIAVKLDLDANVYVGTALINLYAKLGCMDEAMLVFHAL 181
Query: 112 PHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGR 171
P R P++WN++I Y Q GC ALE+ + + G+ P +LAS VS C GR
Sbjct: 182 PVRTPVTWNTVITGYAQIGCGGVALELFDRMGIEGVRPDRFVLASAVSACSALGFLEGGR 241
Query: 172 QIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIAN 231
QIH + V++ L+D Y +C A ++FD ME +N VSWT MISG + N
Sbjct: 242 QIHGYAYRSATETDTSVIN-VLIDLYCKCSRLSAARKLFDCMEYRNLVSWTTMISGYMQN 300
Query: 232 LDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFS 291
A F M G P+ ++L +C + G++IH + + +E+
Sbjct: 301 SFNAEAITMFWNMTQAGWQPDGFACTSILNSCGSLAAIWQGRQIHAHVIKADLEADEYVK 360
Query: 292 SALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEET 351
+ALI+MY + E L A +F+ + D + ++++I YS+ D +A+ +F +M
Sbjct: 361 NALIDMYAKC-EHLTEARAVFDALAEDDAISYNAMIEGYSKNRDLAEAVNIFQRMRFFSL 419
Query: 352 EPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQ 411
P+ +T ++++ ++ +++ +HG I+K G S + ++AL+++Y+KC + +
Sbjct: 420 RPSLLTFVSLLGVSSSQLAIELSKQIHGLIIKSGTSLDLYAASALIDVYSKCSLVNDAKT 479
Query: 412 IFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGL 471
+F + +D + W++MI + + GE+A+KLF+++ G+ P+ TF+A+++ +
Sbjct: 480 VFNMLHYKDMVIWNSMIFGHAQNEQGEEAIKLFNQLLLSGMAPNEFTFVALVTVASTLAS 539
Query: 472 VTEGQQVFKQ-VNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSL 530
+ GQQ + A + + + L+D+ + G +++ + + W+S+
Sbjct: 540 MFHGQQFHAWIIKAGVDNDPHVSN--ALIDMYAKCGFIKEGRMLFES-TCGEDVICWNSM 596
Query: 531 VSACKLHGRLDIA 543
++ HG + A
Sbjct: 597 ITTYAQHGHAEEA 609
Score = 214 bits (544), Expect = 8e-54
Identities = 142/527 (26%), Positives = 263/527 (48%), Gaps = 21/527 (3%)
Query: 68 LGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYL 127
L +H A G D ++N L+ Y+ ++ AR +FD MPHR+ +SW S+I+ Y
Sbjct: 36 LNPAIHARATVAGRLDDLFLTNLLLRGYSNLGRLRDARHLFDRMPHRNLVSWGSVISMYT 95
Query: 128 QNGCLEEALEMLKGVYLLGL-VPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHS 186
Q+G + A+ + VP LLAS++ C + +G Q+H + V ++ +
Sbjct: 96 QHGRDDCAISLFVAFQKASCEVPNEFLLASVLRACTQSKAVSLGEQVHG-IAVKLDLDAN 154
Query: 187 VVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQV 246
V + TAL++ Y + A VF + V+ V+W +I+G A F M +
Sbjct: 155 VYVGTALINLYAKLGCMDEAMLVFHALPVRTPVTWNTVITGYAQIGCGGVALELFDRMGI 214
Query: 247 EGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLH 306
EGV P+R L + + AC+ GF++ G++IHGYA+R E+ + LI++YC+ L
Sbjct: 215 EGVRPDRFVLASAVSACSALGFLEGGRQIHGYAYRSATETDTSVINVLIDLYCKCSR-LS 273
Query: 307 LAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACT 366
A ++F+ +R++V W+++I Y Q + +A+T+F M +P+ +++++C
Sbjct: 274 AARKLFDCMEYRNLVSWTTMISGYMQNSFNAEAITMFWNMTQAGWQPDGFACTSILNSCG 333
Query: 367 NLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWST 426
+L+++ G +H ++K L + NAL++MYAKC L + +F + D+IS++
Sbjct: 334 SLAAIWQGRQIHAHVIKADLEADEYVKNALIDMYAKCEHLTEARAVFDALAEDDAISYNA 393
Query: 427 MISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADY 486
MI Y + +A+ +F M+ ++P LTF+++L + + +Q+ +
Sbjct: 394 MIEGYSKNRDLAEAVNIFQRMRFFSLRPSLLTFVSLLGVSSSQLAIELSKQIHGLI---I 450
Query: 487 EIPLTIEHYA--CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV----------SAC 534
+ +++ YA L+D+ + V DA + + K IW+S++ A
Sbjct: 451 KSGTSLDLYAASALIDVYSKCSLVNDAKTVFNMLHYKDMV-IWNSMIFGHAQNEQGEEAI 509
Query: 535 KLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGH-WLHKEQV 580
KL +L ++ M + A+ TL +M + + H W+ K V
Sbjct: 510 KLFNQLLLSGMAPNEFTFVALVTVAS-TLASMFHGQQFHAWIIKAGV 555
Score = 157 bits (397), Expect = 9e-37
Identities = 106/363 (29%), Positives = 184/363 (50%), Gaps = 9/363 (2%)
Query: 154 LASMVSLCGRKMGSRIGR---QIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVF 210
LA ++ C G R+ R IHA V GR++ + L+ L+ Y A +F
Sbjct: 18 LARVLLSCLPTGGDRLRRLNPAIHARATVAGRLD-DLFLTNLLLRGYSNLGRLRDARHLF 76
Query: 211 DGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVD-PNRVTLIALLPACAKPGFV 269
D M +N VSW ++IS + D A + F A Q + PN L ++L AC + V
Sbjct: 77 DRMPHRNLVSWGSVISMYTQHGRDDCAISLFVAFQKASCEVPNEFLLASVLRACTQSKAV 136
Query: 270 KHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGS 329
G+++HG A + ++++ +ALIN+Y + G + A +F R V W+++I
Sbjct: 137 SLGEQVHGIAVKLDLDANVYVGTALINLYAKLG-CMDEAMLVFHALPVRTPVTWNTVITG 195
Query: 330 YSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFS 389
Y+Q G AL LF++M E P+ L + +SAC+ L L+ G +HG+ +
Sbjct: 196 YAQIGCGGVALELFDRMGIEGVRPDRFVLASAVSACSALGFLEGGRQIHGYAYRSATETD 255
Query: 390 ISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKE 449
S+ N L+++Y KC L + ++F M+ R+ +SW+TMIS Y + +A+ +F M +
Sbjct: 256 TSVINVLIDLYCKCSRLSAARKLFDCMEYRNLVSWTTMISGYMQNSFNAEAITMFWNMTQ 315
Query: 450 RGVKPDALTFLAVLSACNHAGLVTEGQQVFKQV-NADYEIPLTIEHYACLVDLLGRSGKV 508
G +PD ++L++C + +G+Q+ V AD E +++ L+D+ + +
Sbjct: 316 AGWQPDGFACTSILNSCGSLAAIWQGRQIHAHVIKADLEADEYVKN--ALIDMYAKCEHL 373
Query: 509 EDA 511
+A
Sbjct: 374 TEA 376
Score = 126 bits (316), Expect = 2e-27
Identities = 79/299 (26%), Positives = 151/299 (50%), Gaps = 4/299 (1%)
Query: 33 QTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLI 92
+ + +F ++ F + S++ SS + L +HGL + +G+ D +++LI
Sbjct: 406 EAVNIFQRMRFFSLRPSLLTFVSLLGVSSSQLAIELSKQIHGLIIKSGTSLDLYAASALI 465
Query: 93 SLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPE 152
+Y+K S + A+ VF+ + ++D + WNSMI + QN EEA+++ + L G+ P
Sbjct: 466 DVYSKCSLVNDAKTVFNMLHYKDMVIWNSMIFGHAQNEQGEEAIKLFNQLLLSGMAPNEF 525
Query: 153 LLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDG 212
++V++ G+Q HA ++ G V++ +S AL+D Y +C +F+
Sbjct: 526 TFVALVTVASTLASMFHGQQFHAWIIKAG-VDNDPHVSNALIDMYAKCGFIKEGRMLFES 584
Query: 213 MEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHG 272
++ + W +MI+ + + A FR M V+PN VT + +L ACA GFV G
Sbjct: 585 TCGEDVICWNSMITTYAQHGHAEEALQVFRLMGEAEVEPNYVTFVGVLSACAHAGFVGEG 644
Query: 273 -KEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD-VVLWSSIIGS 329
+ + IE ++++N++ +SG+ LH A++ ER + +W S++ +
Sbjct: 645 LNHFNSMKSNYDIEPGIEHYASVVNLFGRSGK-LHAAKEFIERMPIKPAAAVWRSLLSA 702
>UniRef100_O49619 Hypothetical protein M4E13.180 [Arabidopsis thaliana]
Length = 804
Score = 347 bits (891), Expect = 4e-94
Identities = 202/580 (34%), Positives = 318/580 (54%), Gaps = 8/580 (1%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
IK S GLY + ++ ++++ F+ + + P VIK+ + + S G +H + + G
Sbjct: 102 IKGFTSCGLYIEAVQFYSRMVFAGVKADTFTYPFVIKSVAGISSLEEGKKIHAMVIKLGF 161
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
SD V NSLISLY K A +VF+ MP RD +SWNSMI+ YL G +L + K
Sbjct: 162 VSDVYVCNSLISLYMKLGCAWDAEKVFEEMPERDIVSWNSMISGYLALGDGFSSLMLFKE 221
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ G P S + C ++G++IH V V++ T+++D Y +
Sbjct: 222 MLKCGFKPDRFSTMSALGACSHVYSPKMGKEIHCHAVRSRIETGDVMVMTSILDMYSKYG 281
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIA-NLDYDAAFARFRAMQVE-GVDPNRVTLIAL 259
+ A R+F+GM +N V+W MI GC A N AF F+ M + G+ P+ +T I L
Sbjct: 282 EVSYAERIFNGMIQRNIVAWNVMI-GCYARNGRVTDAFLCFQKMSEQNGLQPDVITSINL 340
Query: 260 LPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRD 319
LPA A + G+ IHGYA R G + +ALI+MY + G+ L AE IF+R + ++
Sbjct: 341 LPASA----ILEGRTIHGYAMRRGFLPHMVLETALIDMYGECGQ-LKSAEVIFDRMAEKN 395
Query: 320 VVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHG 379
V+ W+SII +Y Q G +Y AL LF ++ P+ T+ +++ A SL G +H
Sbjct: 396 VISWNSIIAAYVQNGKNYSALELFQELWDSSLVPDSTTIASILPAYAESLSLSEGREIHA 455
Query: 380 FILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQ 439
+I+K + + N+L++MYA CG LE + + F + +D +SW+++I AY +HG G
Sbjct: 456 YIVKSRYWSNTIILNSLVHMYAMCGDLEDARKCFNHILLKDVVSWNSIIMAYAVHGFGRI 515
Query: 440 ALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLV 499
++ LFSEM V P+ TF ++L+AC+ +G+V EG + F+ + +Y I IEHY C++
Sbjct: 516 SVWLFSEMIASRVNPNKSTFASLLAACSISGMVDEGWEYFESMKREYGIDPGIEHYGCML 575
Query: 500 DLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAA 559
DL+GR+G A + MP P+ RIW SL++A + H + IAE Q+ + E DN
Sbjct: 576 DLIGRTGNFSAAKRFLEEMPFVPTARIWGSLLNASRNHKDITIAEFAAEQIFKMEHDNTG 635
Query: 560 NYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
Y LL +YAE G W ++ M+ + + + S +E
Sbjct: 636 CYVLLLNMYAEAGRWEDVNRIKLLMESKGISRTSSRSTVE 675
Score = 187 bits (474), Expect = 1e-45
Identities = 135/473 (28%), Positives = 241/473 (50%), Gaps = 11/473 (2%)
Query: 83 SDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGV 142
+DP ++ +L +A ++ A Q+FD M D WN MI + G EA++ +
Sbjct: 63 NDPALTRALRG-FADSRLMEDALQLFDEMNKADAFLWNVMIKGFTSCGLYIEAVQFYSRM 121
Query: 143 YLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDD 202
G+ ++ G++IHA+V+ G V V + +L+ Y +
Sbjct: 122 VFAGVKADTFTYPFVIKSVAGISSLEEGKKIHAMVIKLGFVS-DVYVCNSLISLYMKLGC 180
Query: 203 ALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPA 262
A A +VF+ M ++ VSW +MISG +A D ++ F+ M G P+R + ++ L A
Sbjct: 181 AWDAEKVFEEMPERDIVSWNSMISGYLALGDGFSSLMLFKEMLKCGFKPDRFSTMSALGA 240
Query: 263 CAKPGFVKHGKEIHGYAFRHGIESSHIF-SSALINMYCQSGESLHLAEQIFERSSFRDVV 321
C+ K GKEIH +A R IE+ + +++++MY + GE + AE+IF R++V
Sbjct: 241 CSHVYSPKMGKEIHCHAVRSRIETGDVMVMTSILDMYSKYGE-VSYAERIFNGMIQRNIV 299
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEE-TEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
W+ +IG Y++ G A F KM + +P+ +T + ++ A S++ G +HG+
Sbjct: 300 AWNVMIGCYARNGRVTDAFLCFQKMSEQNGLQPDVITSINLLPA----SAILEGRTIHGY 355
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
++ G + L AL++MY +CG L+ + IF M ++ ISW+++I+AY +G A
Sbjct: 356 AMRRGFLPHMVLETALIDMYGECGQLKSAEVIFDRMAEKNVISWNSIIAAYVQNGKNYSA 415
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
L+LF E+ + + PD+ T ++L A + ++EG+++ + TI LV
Sbjct: 416 LELFQELWDSSLVPDSTTIASILPAYAESLSLSEGREIHAYIVKSRYWSNTI-ILNSLVH 474
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS 553
+ G +EDA + + +K W+S++ A +HG I+ L ++I S
Sbjct: 475 MYAMCGDLEDARKCFNHILLKDVVS-WNSIIMAYAVHGFGRISVWLFSEMIAS 526
Score = 108 bits (270), Expect = 5e-22
Identities = 69/272 (25%), Positives = 131/272 (47%), Gaps = 4/272 (1%)
Query: 308 AEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTN 367
A Q+F+ + D LW+ +I ++ G +A+ +++M+ + + T VI +
Sbjct: 83 ALQLFDEMNKADAFLWNVMIKGFTSCGLYIEAVQFYSRMVFAGVKADTFTYPFVIKSVAG 142
Query: 368 LSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTM 427
+SSL+ G +H ++K G + + N+L+++Y K GC + ++F EM RD +SW++M
Sbjct: 143 ISSLEEGKKIHAMVIKLGFVSDVYVCNSLISLYMKLGCAWDAEKVFEEMPERDIVSWNSM 202
Query: 428 ISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYE 487
IS Y G G +L LF EM + G KPD + ++ L AC+H G+++
Sbjct: 203 ISGYLALGDGFSSLMLFKEMLKCGFKPDRFSTMSALGACSHVYSPKMGKEIHCHAVRSRI 262
Query: 488 IPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLG 547
+ ++D+ + G+V A I M ++ + W+ ++ +GR+ A +
Sbjct: 263 ETGDVMVMTSILDMYSKYGEVSYAERIFNGM-IQRNIVAWNVMIGCYARNGRVTDAFLCF 321
Query: 548 PQLIRS---EPDNAANYTLLNMIYAEHGHWLH 576
++ +PD + LL G +H
Sbjct: 322 QKMSEQNGLQPDVITSINLLPASAILEGRTIH 353
Score = 57.4 bits (137), Expect = 1e-06
Identities = 54/216 (25%), Positives = 98/216 (45%), Gaps = 19/216 (8%)
Query: 391 SLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKER 450
+L+ AL +A +E +LQ+F EM D+ W+ MI + G +A++ +S M
Sbjct: 66 ALTRALRG-FADSRLMEDALQLFDEMNKADAFLWNVMIKGFTSCGLYIEAVQFYSRMVFA 124
Query: 451 GVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYAC--LVDLLGRSGKV 508
GVK D T+ V+ + + EG+++ V ++ + Y C L+ L + G
Sbjct: 125 GVKADTFTYPFVIKSVAGISSLEEGKKIHAMV---IKLGFVSDVYVCNSLISLYMKLGCA 181
Query: 509 EDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRS--EPDNAANYTLLNM 566
DA ++ MP + W+S++S G + ML ++++ +PD + + L
Sbjct: 182 WDAEKVFEEMPERDIVS-WNSMISGYLALGDGFSSLMLFKEMLKCGFKPDRFSTMSALGA 240
Query: 567 IYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGN 602
+ + KE A++ SRIETG+
Sbjct: 241 CSHVYSPKMGKEIHCHAVR----------SRIETGD 266
>UniRef100_Q9STE1 Hypothetical protein T6K22.30 [Arabidopsis thaliana]
Length = 857
Score = 347 bits (890), Expect = 6e-94
Identities = 184/545 (33%), Positives = 314/545 (56%), Gaps = 3/545 (0%)
Query: 56 VIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRD 115
V+ C+S LG LHGL + +G + + NSL+S+Y+K A ++F M D
Sbjct: 245 VLSVCASKLLIDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGRFDDASKLFRMMSRAD 304
Query: 116 PISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHA 175
++WN MI+ Y+Q+G +EE+L + G++P +S++ + +QIH
Sbjct: 305 TVTWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKFENLEYCKQIHC 364
Query: 176 LVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYD 235
++ + + L++AL+D YF+C MA +F + V +TAMISG + N Y
Sbjct: 365 YIMRHS-ISLDIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMISGYLHNGLYI 423
Query: 236 AAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALI 295
+ FR + + PN +TL+++LP +K G+E+HG+ + G ++ A+I
Sbjct: 424 DSLEMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFDNRCNIGCAVI 483
Query: 296 NMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNY 355
+MY + G ++LA +IFER S RD+V W+S+I +Q + A+ +F +M +
Sbjct: 484 DMYAKCGR-MNLAYEIFERLSKRDIVSWNSMITRCAQSDNPSAAIDIFRQMGVSGICYDC 542
Query: 356 VTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLE 415
V++ A +SAC NL S G +HGF++K L+ + + L++MYAKCG L+ ++ +F
Sbjct: 543 VSISAALSACANLPSESFGKAIHGFMIKHSLASDVYSESTLIDMYAKCGNLKAAMNVFKT 602
Query: 416 MQNRDSISWSTMISAYGLHGHGEQALKLFSEMKER-GVKPDALTFLAVLSACNHAGLVTE 474
M+ ++ +SW+++I+A G HG + +L LF EM E+ G++PD +TFL ++S+C H G V E
Sbjct: 603 MKEKNIVSWNSIIAACGNHGKLKDSLCLFHEMVEKSGIRPDQITFLEIISSCCHVGDVDE 662
Query: 475 GQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSAC 534
G + F+ + DY I EHYAC+VDL GR+G++ +A E +++MP P +W +L+ AC
Sbjct: 663 GVRFFRSMTEDYGIQPQQEHYACVVDLFGRAGRLTEAYETVKSMPFPPDAGVWGTLLGAC 722
Query: 535 KLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYG 594
+LH +++AE+ +L+ +P N+ Y L++ +A W +V MK + ++K G
Sbjct: 723 RLHKNVELAEVASSKLMDLDPSNSGYYVLISNAHANAREWESVTKVRSLMKEREVQKIPG 782
Query: 595 FSRIE 599
+S IE
Sbjct: 783 YSWIE 787
Score = 234 bits (597), Expect = 5e-60
Identities = 151/547 (27%), Positives = 279/547 (50%), Gaps = 6/547 (1%)
Query: 22 IKSLISQGLYHQTLEL-FTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTG 80
I S + GL +Q L F L F +S+ P ++KAC +L + L + G
Sbjct: 110 ISSFVRNGLLNQALAFYFKMLCFGVSPDVST-FPCLVKACVALKNFKGIDFLSDTVSSLG 168
Query: 81 SHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLK 140
+ V++SLI Y ++ I ++FD + +D + WN M+N Y + G L+ ++
Sbjct: 169 MDCNEFVASSLIKAYLEYGKIDVPSKLFDRVLQKDCVIWNVMLNGYAKCGALDSVIKGFS 228
Query: 141 GVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRC 200
+ + + P ++S+C K+ +G Q+H LVVV G V+ + +L+ Y +C
Sbjct: 229 VMRMDQISPNAVTFDCVLSVCASKLLIDLGVQLHGLVVVSG-VDFEGSIKNSLLSMYSKC 287
Query: 201 DDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALL 260
AS++F M + V+W MISG + + + + F M GV P+ +T +LL
Sbjct: 288 GRFDDASKLFRMMSRADTVTWNCMISGYVQSGLMEESLTFFYEMISSGVLPDAITFSSLL 347
Query: 261 PACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDV 320
P+ +K +++ K+IH Y RH I +SALI+ Y + + +A+ IF + + DV
Sbjct: 348 PSVSKFENLEYCKQIHCYIMRHSISLDIFLTSALIDAYFKC-RGVSMAQNIFSQCNSVDV 406
Query: 321 VLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGF 380
V+++++I Y G +L +F ++ + PN +TL++++ L +LK G LHGF
Sbjct: 407 VVFTAMISGYLHNGLYIDSLEMFRWLVKVKISPNEITLVSILPVIGILLALKLGRELHGF 466
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
I+K G ++ A+++MYAKCG + + +IF + RD +SW++MI+ + A
Sbjct: 467 IIKKGFDNRCNIGCAVIDMYAKCGRMNLAYEIFERLSKRDIVSWNSMITRCAQSDNPSAA 526
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
+ +F +M G+ D ++ A LSAC + + G+ + + + + + + L+D
Sbjct: 527 IDIFRQMGVSGICYDCVSISAALSACANLPSESFGKAIHGFM-IKHSLASDVYSESTLID 585
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAAN 560
+ + G ++ A+ + +TM K + W+S+++AC HG+L + L +++
Sbjct: 586 MYAKCGNLKAAMNVFKTMKEK-NIVSWNSIIAACGNHGKLKDSLCLFHEMVEKSGIRPDQ 644
Query: 561 YTLLNMI 567
T L +I
Sbjct: 645 ITFLEII 651
Score = 150 bits (380), Expect = 8e-35
Identities = 110/507 (21%), Positives = 234/507 (45%), Gaps = 9/507 (1%)
Query: 31 YHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNS 90
Y ++L L F +I L +++ACS+ + G +H + D
Sbjct: 17 YKKSLPLRNSSRF-LEETIPRRLSLLLQACSNPNLLRQGKQVHAFLIVNSISGDSYTDER 75
Query: 91 LISLYAKFSDIQSARQVFDTMPHRDPI--SWNSMINAYLQNGCLEEALEMLKGVYLLGLV 148
++ +YA ++F + R WNS+I+++++NG L +AL + G+
Sbjct: 76 ILGMYAMCGSFSDCGKMFYRLDLRRSSIRPWNSIISSFVRNGLLNQALAFYFKMLCFGVS 135
Query: 149 PKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASR 208
P +V C + G + V ++ + ++++L+ Y + S+
Sbjct: 136 PDVSTFPCLVKACVALKNFK-GIDFLSDTVSSLGMDCNEFVASSLIKAYLEYGKIDVPSK 194
Query: 209 VFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGF 268
+FD + K+ V W M++G D+ F M+++ + PN VT +L CA
Sbjct: 195 LFDRVLQKDCVIWNVMLNGYAKCGALDSVIKGFSVMRMDQISPNAVTFDCVLSVCASKLL 254
Query: 269 VKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIG 328
+ G ++HG G++ ++L++MY + G A ++F S D V W+ +I
Sbjct: 255 IDLGVQLHGLVVVSGVDFEGSIKNSLLSMYSKCGR-FDDASKLFRMMSRADTVTWNCMIS 313
Query: 329 SYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSF 388
Y Q G ++LT F +M++ P+ +T +++ + + +L++ +H +I++ +S
Sbjct: 314 GYVQSGLMEESLTFFYEMISSGVLPDAITFSSLLPSVSKFENLEYCKQIHCYIMRHSISL 373
Query: 389 SISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMK 448
I L++AL++ Y KC + + IF + + D + ++ MIS Y +G +L++F +
Sbjct: 374 DIFLTSALIDAYFKCRGVSMAQNIFSQCNSVDVVVFTAMISGYLHNGLYIDSLEMFRWLV 433
Query: 449 ERGVKPDALTFLAVLSACNHAGLVTEGQQVFK-QVNADYEIPLTIEHYACLVDLLGRSGK 507
+ + P+ +T +++L + G+++ + ++ I ++D+ + G+
Sbjct: 434 KVKISPNEITLVSILPVIGILLALKLGRELHGFIIKKGFDNRCNIG--CAVIDMYAKCGR 491
Query: 508 VEDALEIMRTMPMKPSTRIWSSLVSAC 534
+ A EI + K W+S+++ C
Sbjct: 492 MNLAYEIFERL-SKRDIVSWNSMITRC 517
Score = 108 bits (269), Expect = 6e-22
Identities = 76/303 (25%), Positives = 141/303 (46%), Gaps = 17/303 (5%)
Query: 246 VEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESL 305
+E P R++L LL AC+ P ++ GK++H + + I ++ MY G
Sbjct: 30 LEETIPRRLSL--LLQACSNPNLLRQGKQVHAFLIVNSISGDSYTDERILGMYAMCGSFS 87
Query: 306 HLAEQIFE----RSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAV 361
+ + RSS R W+SII S+ + G +AL + KML P+ T +
Sbjct: 88 DCGKMFYRLDLRRSSIRP---WNSIISSFVRNGLLNQALAFYFKMLCFGVSPDVSTFPCL 144
Query: 362 ISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDS 421
+ AC L + K L + G+ + ++++L+ Y + G ++ ++F + +D
Sbjct: 145 VKACVALKNFKGIDFLSDTVSSLGMDCNEFVASSLIKAYLEYGKIDVPSKLFDRVLQKDC 204
Query: 422 ISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQ 481
+ W+ M++ Y G + +K FS M+ + P+A+TF VLS C L+ G Q+
Sbjct: 205 VIWNVMLNGYAKCGALDSVIKGFSVMRMDQISPNAVTFDCVLSVCASKLLIDLGVQLHGL 264
Query: 482 V---NADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
V D+E + L+ + + G+ +DA ++ R M + T W+ ++S G
Sbjct: 265 VVVSGVDFEGSIK----NSLLSMYSKCGRFDDASKLFRMM-SRADTVTWNCMISGYVQSG 319
Query: 539 RLD 541
++
Sbjct: 320 LME 322
>UniRef100_Q9SN39 Hypothetical protein F28A21.160 [Arabidopsis thaliana]
Length = 871
Score = 347 bits (889), Expect = 8e-94
Identities = 194/579 (33%), Positives = 325/579 (55%), Gaps = 3/579 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
+ L G + ++ LF ++ S S V K+ SSL S G LHG L +G
Sbjct: 167 MNELAKSGDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGF 226
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
V NSL++ Y K + SAR+VFD M RD ISWNS+IN Y+ NG E+ L +
Sbjct: 227 GERNSVGNSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQ 286
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ + G+ + S+ + C +GR +H++ V +T L+D Y +C
Sbjct: 287 MLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNT-LLDMYSKCG 345
Query: 202 DALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLP 261
D A VF M ++ VS+T+MI+G A F M+ EG+ P+ T+ A+L
Sbjct: 346 DLDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLN 405
Query: 262 ACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVV 321
CA+ + GK +H + + + S+AL++MY + G S+ AE +F +D++
Sbjct: 406 CCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCG-SMQEAELVFSEMRVKDII 464
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEET-EPNYVTLLAVISACTNLSSLKHGCGLHGF 380
W++IIG YS+ + +AL+LFN +L E+ P+ T+ V+ AC +LS+ G +HG+
Sbjct: 465 SWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGY 524
Query: 381 ILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQA 440
I++ G ++N+L++MYAKCG L + +F ++ ++D +SW+ MI+ YG+HG G++A
Sbjct: 525 IMRNGYFSDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEA 584
Query: 441 LKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVD 500
+ LF++M++ G++ D ++F+++L AC+H+GLV EG + F + + +I T+EHYAC+VD
Sbjct: 585 IALFNQMRQAGIEADEISFVSLLYACSHSGLVDEGWRFFNIMRHECKIEPTVEHYACIVD 644
Query: 501 LLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAAN 560
+L R+G + A + MP+ P IW +L+ C++H + +AE + ++ EP+N
Sbjct: 645 MLARTGDLIKAYRFIENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGY 704
Query: 561 YTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIE 599
Y L+ IYAE W +++ + + + L+K G S IE
Sbjct: 705 YVLMANIYAEAEKWEQVKRLRKRIGQRGLRKNPGCSWIE 743
Score = 191 bits (484), Expect = 7e-47
Identities = 135/519 (26%), Positives = 249/519 (47%), Gaps = 8/519 (1%)
Query: 53 LPSVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMP 112
L SV++ C+ S G + G D + + L +Y D++ A +VFD +
Sbjct: 97 LCSVLQLCADSKSLKDGKEVDNFIRGNGFVIDSNLGSKLSLMYTNCGDLKEASRVFDEVK 156
Query: 113 HRDPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQ 172
+ WN ++N ++G ++ + K + G+ + + G Q
Sbjct: 157 IEKALFWNILMNELAKSGDFSGSIGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQ 216
Query: 173 IHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANL 232
+H ++ G E + V +LV FY + A +VFD M ++ +SW ++I+G ++N
Sbjct: 217 LHGFILKSGFGERNSV-GNSLVAFYLKNQRVDSARKVFDEMTERDVISWNSIINGYVSNG 275
Query: 233 DYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSS 292
+ + F M V G++ + T++++ CA + G+ +H + F +
Sbjct: 276 LAEKGLSVFVQMLVSGIEIDLATIVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCN 335
Query: 293 ALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE 352
L++MY + G+ L A+ +F S R VV ++S+I Y++ G + +A+ LF +M E
Sbjct: 336 TLLDMYSKCGD-LDSAKAVFREMSDRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGIS 394
Query: 353 PNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQI 412
P+ T+ AV++ C L G +H +I + L F I +SNALM+MYAKCG ++ + +
Sbjct: 395 PDVYTVTAVLNCCARYRLLDEGKRVHEWIKENDLGFDIFVSNALMDMYAKCGSMQEAELV 454
Query: 413 FLEMQNRDSISWSTMISAYGLHGHGEQALKLFS-EMKERGVKPDALTFLAVLSACNHAGL 471
F EM+ +D ISW+T+I Y + + +AL LF+ ++E+ PD T VL AC
Sbjct: 455 FSEMRVKDIISWNTIIGGYSKNCYANEALSLFNLLLEEKRFSPDERTVACVLPACASLSA 514
Query: 472 VTEGQQVFKQVNADYEIPLTIEHYA-CLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSL 530
+G+++ + + + H A LVD+ + G + A + + K W+ +
Sbjct: 515 FDKGREIHGYIMRNGY--FSDRHVANSLVDMYAKCGALLLAHMLFDDIASKDLVS-WTVM 571
Query: 531 VSACKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYA 569
++ +HG A L Q+ R A + ++++YA
Sbjct: 572 IAGYGMHGFGKEAIALFNQM-RQAGIEADEISFVSLLYA 609
Score = 180 bits (457), Expect = 9e-44
Identities = 109/414 (26%), Positives = 214/414 (51%), Gaps = 10/414 (2%)
Query: 120 NSMINAYLQNGCLEEALEML--KGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALV 177
N+ + + ++G LE A+++L G + + P L S++ LC + G+++ +
Sbjct: 65 NTQLRRFCESGNLENAVKLLCVSGKWDID----PRTLCSVLQLCADSKSLKDGKEVDNFI 120
Query: 178 VVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAA 237
+G V S L + L Y C D ASRVFD ++++ + W +++ + D+ +
Sbjct: 121 RGNGFVIDSN-LGSKLSLMYTNCGDLKEASRVFDEVKIEKALFWNILMNELAKSGDFSGS 179
Query: 238 FARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINM 297
F+ M GV+ + T + + + V G+++HG+ + G + ++L+
Sbjct: 180 IGLFKKMMSSGVEMDSYTFSCVSKSFSSLRSVHGGEQLHGFILKSGFGERNSVGNSLVAF 239
Query: 298 YCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVT 357
Y ++ + + A ++F+ + RDV+ W+SII Y G + K L++F +ML E + T
Sbjct: 240 YLKN-QRVDSARKVFDEMTERDVISWNSIINGYVSNGLAEKGLSVFVQMLVSGIEIDLAT 298
Query: 358 LLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQ 417
+++V + C + + G +H +K S N L++MY+KCG L+ + +F EM
Sbjct: 299 IVSVFAGCADSRLISLGRAVHSIGVKACFSREDRFCNTLLDMYSKCGDLDSAKAVFREMS 358
Query: 418 NRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQ 477
+R +S+++MI+ Y G +A+KLF EM+E G+ PD T AVL+ C L+ EG++
Sbjct: 359 DRSVVSYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKR 418
Query: 478 VFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLV 531
V + + + ++ I L+D+ + G +++A + M +K W++++
Sbjct: 419 VHEWIK-ENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIIS-WNTII 470
Score = 110 bits (276), Expect = 9e-23
Identities = 93/354 (26%), Positives = 155/354 (43%), Gaps = 10/354 (2%)
Query: 17 SPSQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLA 76
S + I +GL + ++LF ++ + +V+ C+ G +H
Sbjct: 364 SYTSMIAGYAREGLAGEAVKLFEEMEEEGISPDVYTVTAVLNCCARYRLLDEGKRVHEWI 423
Query: 77 LTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEAL 136
D VSN+L+ +YAK +Q A VF M +D ISWN++I Y +N EAL
Sbjct: 424 KENDLGFDIFVSNALMDMYAKCGSMQEAELVFSEMRVKDIISWNTIIGGYSKNCYANEAL 483
Query: 137 EMLKGVYLL---GLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTAL 193
+ LL P +A ++ C GR+IH ++ +G V + +L
Sbjct: 484 SLFN--LLLEEKRFSPDERTVACVLPACASLSAFDKGREIHGYIMRNGYFSDRHV-ANSL 540
Query: 194 VDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNR 253
VD Y +C L+A +FD + K+ VSWT MI+G + A A F M+ G++ +
Sbjct: 541 VDMYAKCGALLLAHMLFDDIASKDLVSWTVMIAGYGMHGFGKEAIALFNQMRQAGIEADE 600
Query: 254 VTLIALLPACAKPGFVKHGKEIHGYAFRH--GIESSHIFSSALINMYCQSGESLHLAEQI 311
++ ++LL AC+ G V G RH IE + + +++M ++G+ + I
Sbjct: 601 ISFVSLLYACSHSGLVDEGWRFFN-IMRHECKIEPTVEHYACIVDMLARTGDLIKAYRFI 659
Query: 312 FERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETE-PNYVTLLAVISA 364
D +W +++ D A + K+ E E Y L+A I A
Sbjct: 660 ENMPIPPDATIWGALLCGCRIHHDVKLAEKVAEKVFELEPENTGYYVLMANIYA 713
>UniRef100_Q5SMW8 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 673
Score = 344 bits (882), Expect = 5e-93
Identities = 185/576 (32%), Positives = 315/576 (54%), Gaps = 5/576 (0%)
Query: 25 LISQGLYHQTLELFTQLHFSAPHSISS--VLPSVIKACSSLHSHTLGTLLHGLALTTGSH 82
L+ G + + L + P + + VL +KAC G LH + G
Sbjct: 96 LVDAGSHADAVALHRDMRRRCPAAAQADVVLSLALKACVRSADFRYGRRLHCDVVKAGG- 154
Query: 83 SDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGV 142
+D V NSL+ +YAK D+++AR+VFD +P R+ +SW SM++ +QNG EE L + +
Sbjct: 155 ADGFVMNSLVDMYAKAGDLENARKVFDRVPERNVVSWTSMLSGSIQNGIAEEGLVLFNEM 214
Query: 143 YLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDD 202
+ P + S+++ C G GR IH V+ G +S + S +L+D Y +C+
Sbjct: 215 RQDNVHPSEYTMVSVLAACAMLGGLHQGRWIHGSVIKYGLSTNSFI-SASLLDMYAKCEK 273
Query: 203 ALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPA 262
A RVFD +E + V WTAMI G N A F + + PN VT+ ++ A
Sbjct: 274 VEDARRVFDELEFVDIVLWTAMIVGYTQNKRPLDALQLFLHKKFVSIVPNSVTIATVISA 333
Query: 263 CAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVL 322
A+ + G+ +H + G S + +AL++MY + ++L A IF R +DVV
Sbjct: 334 SAQLRHLPLGRSVHAIGVKLGTMESDVVRNALVDMYAKC-QALPEANSIFGRILIKDVVA 392
Query: 323 WSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFIL 382
W+S++ YS+ G + ++L LFN+M + P+ ++++ +SAC L+ L G G H + +
Sbjct: 393 WNSMMAGYSENGMANESLVLFNRMRMQGISPDAISVVNALSACVCLADLHIGKGFHTYAI 452
Query: 383 KFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALK 442
K+ +I ++ AL+N+Y+KC L + ++F +M +R+S++WS MI YG+ G ++
Sbjct: 453 KYAFMSNIYVNTALLNLYSKCADLPSAQRVFNDMTDRNSVTWSAMIGGYGMQGDSAGSID 512
Query: 443 LFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLL 502
LF+EM + + P+ + F ++LSAC+H G+VT G++ F + + I +++HYAC+VD++
Sbjct: 513 LFNEMLKENIHPNEVVFTSILSACSHTGMVTAGKEYFDSMARHFNITPSMKHYACMVDVM 572
Query: 503 GRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIAEMLGPQLIRSEPDNAANYT 562
R+G +E+ALE ++ MP+K +W S + CKLH RL+ E ++ P+ Y
Sbjct: 573 ARAGNLEEALEFIQNMPIKAGISVWGSFLHGCKLHSRLEFGEEAIKKMAALHPETPDFYV 632
Query: 563 LLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRI 598
L++ +Y +G W + + M+ Q L K G S +
Sbjct: 633 LMSNLYTSYGRWDKSQTIRRWMQEQGLVKLPGCSSV 668
Score = 215 bits (547), Expect = 3e-54
Identities = 150/499 (30%), Positives = 246/499 (49%), Gaps = 13/499 (2%)
Query: 44 SAPHSISSVLPSVIKACSSLHSHTLGTLLHG--LALTTGSHSDPVVSNSLISLYAKFSDI 101
SAP +S L ++ AC +L S LHG L LT+G L+S YA D+
Sbjct: 16 SAPRD-ASALVLLLPACGTLRSLRA---LHGRLLLLTSGLLRGIRARTKLLSCYAALGDL 71
Query: 102 QSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGV--YLLGLVPKPELLASMVS 159
SAR V D P D ++ M+ + G +A+ + + + +L+ +
Sbjct: 72 ASARGVLDGTPRPDAYAYRVMLGWLVDAGSHADAVALHRDMRRRCPAAAQADVVLSLALK 131
Query: 160 LCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEV 219
C R R GR++H VV G + V+ +LVD Y + D A +VFD + +N V
Sbjct: 132 ACVRSADFRYGRRLHCDVVKAGGADGFVM--NSLVDMYAKAGDLENARKVFDRVPERNVV 189
Query: 220 SWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYA 279
SWT+M+SG I N + F M+ + V P+ T++++L ACA G + G+ IHG
Sbjct: 190 SWTSMLSGSIQNGIAEEGLVLFNEMRQDNVHPSEYTMVSVLAACAMLGGLHQGRWIHGSV 249
Query: 280 FRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKA 339
++G+ ++ S++L++MY + E + A ++F+ F D+VLW+++I Y+Q A
Sbjct: 250 IKYGLSTNSFISASLLDMYAKC-EKVEDARRVFDELEFVDIVLWTAMIVGYTQNKRPLDA 308
Query: 340 LTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNM 399
L LF PN VT+ VISA L L G +H +K G S + NAL++M
Sbjct: 309 LQLFLHKKFVSIVPNSVTIATVISASAQLRHLPLGRSVHAIGVKLGTMESDVVRNALVDM 368
Query: 400 YAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTF 459
YAKC L + IF + +D ++W++M++ Y +G ++L LF+ M+ +G+ PDA++
Sbjct: 369 YAKCQALPEANSIFGRILIKDVVAWNSMMAGYSENGMANESLVLFNRMRMQGISPDAISV 428
Query: 460 LAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMP 519
+ LSAC + G+ F Y I L++L + + A + M
Sbjct: 429 VNALSACVCLADLHIGKG-FHTYAIKYAFMSNIYVNTALLNLYSKCADLPSAQRVFNDMT 487
Query: 520 MKPSTRIWSSLVSACKLHG 538
+ S WS+++ + G
Sbjct: 488 DRNSV-TWSAMIGGYGMQG 505
Score = 203 bits (516), Expect = 1e-50
Identities = 126/451 (27%), Positives = 231/451 (50%), Gaps = 12/451 (2%)
Query: 19 SQQIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALT 78
+ + I G+ + L LF ++ H + SV+ AC+ L G +HG +
Sbjct: 192 TSMLSGSIQNGIAEEGLVLFNEMRQDNVHPSEYTMVSVLAACAMLGGLHQGRWIHGSVIK 251
Query: 79 TGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEM 138
G ++ +S SL+ +YAK ++ AR+VFD + D + W +MI Y QN +AL++
Sbjct: 252 YGLSTNSFISASLLDMYAKCEKVEDARRVFDELEFVDIVLWTAMIVGYTQNKRPLDALQL 311
Query: 139 LKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYF 198
+ +VP +A+++S + +GR +HA+ V G +E VV ALVD Y
Sbjct: 312 FLHKKFVSIVPNSVTIATVISASAQLRHLPLGRSVHAIGVKLGTMESDVV-RNALVDMYA 370
Query: 199 RCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIA 258
+C A+ +F + +K+ V+W +M++G N + + F M+++G+ P+ ++++
Sbjct: 371 KCQALPEANSIFGRILIKDVVAWNSMMAGYSENGMANESLVLFNRMRMQGISPDAISVVN 430
Query: 259 LLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFR 318
L AC + GK H YA ++ S+ ++AL+N+Y + + L A+++F + R
Sbjct: 431 ALSACVCLADLHIGKGFHTYAIKYAFMSNIYVNTALLNLYSKCAD-LPSAQRVFNDMTDR 489
Query: 319 DVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLH 378
+ V WS++IG Y +GDS ++ LFN+ML E PN V +++SAC++ + G
Sbjct: 490 NSVTWSAMIGGYGMQGDSAGSIDLFNEMLKENIHPNEVVFTSILSACSHTGMVTAGKEYF 549
Query: 379 GFILK-FGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS-WSTMISAYGLHGH 436
+ + F ++ S+ ++++ A+ G LE +L+ M + IS W + + LH
Sbjct: 550 DSMARHFNITPSMKHYACMVDVMARAGNLEEALEFIQNMPIKAGISVWGSFLHGCKLHSR 609
Query: 437 ---GEQALKLFSEMKERGVKPDALTFLAVLS 464
GE+A+K K + P+ F ++S
Sbjct: 610 LEFGEEAIK-----KMAALHPETPDFYVLMS 635
>UniRef100_Q9LHZ4 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 734
Score = 344 bits (882), Expect = 5e-93
Identities = 200/600 (33%), Positives = 318/600 (52%), Gaps = 11/600 (1%)
Query: 13 SPTPSPSQ--------QIKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLH 64
SP P P+ ++++ ++ L F + + + S++K C++
Sbjct: 14 SPPPKPTALAADDHHARLRAAAARSDLPAALAAFVAMSSAGAPPVLRTFTSLLKLCAARG 73
Query: 65 SHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMIN 124
G +H G S+ + + +L ++YAK AR+VFD MP RD ++WN+++
Sbjct: 74 DLATGRAVHAQLAARGIDSEALAATALANMYAKCRRPADARRVFDRMPVRDRVAWNALVA 133
Query: 125 AYLQNGCLEEALEMLKGVYLL-GLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRV 183
Y +NG A+EM+ + G P L S++ C R+ HA + G +
Sbjct: 134 GYARNGLARMAMEMVVRMQEEEGERPDSITLVSVLPACANARALAACREAHAFAIRSG-L 192
Query: 184 EHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRA 243
E V ++TA++D Y +C D A VFD M KN VSW AMI G N D A A F
Sbjct: 193 EELVNVATAILDAYCKCGDIRAARVVFDWMPTKNSVSWNAMIDGYAQNGDSREALALFNR 252
Query: 244 MQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGE 303
M EGVD V+++A L AC + G + G +H R G++S+ +ALI MY + +
Sbjct: 253 MVEEGVDVTDVSVLAALQACGELGCLDEGMRVHELLVRIGLDSNVSVMNALITMYSKC-K 311
Query: 304 SLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVIS 363
+ LA +F+ R V W+++I +Q G S A+ LF +M E +P+ TL++VI
Sbjct: 312 RVDLASHVFDELDRRTQVSWNAMILGCAQNGCSEDAVRLFTRMQLENVKPDSFTLVSVIP 371
Query: 364 ACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSIS 423
A ++S +HG+ ++ L + + AL++MYAKCG + + +F + R I+
Sbjct: 372 ALADISDPLQARWIHGYSIRLHLDQDVYVLTALIDMYAKCGRVNIARILFNSARERHVIT 431
Query: 424 WSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVN 483
W+ MI YG HG G+ A++LF EMK G+ P+ TFL+VLSAC+HAGLV EG++ F +
Sbjct: 432 WNAMIHGYGSHGFGKAAVELFEEMKSIGIVPNETTFLSVLSACSHAGLVDEGREYFTSMK 491
Query: 484 ADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHGRLDIA 543
DY + +EHY +VDLLGR+GK+++A ++ MPM P ++ +++ ACKLH +++A
Sbjct: 492 EDYGLEPGMEHYGTMVDLLGRAGKLDEAWAFIQKMPMDPGLSVYGAMLGACKLHKNVELA 551
Query: 544 EMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCYGFSRIETGNQ 603
E ++ P + LL IYA W +V AM+ L+K G+S I+ N+
Sbjct: 552 EESAQKIFELGPQEGVYHVLLANIYANASMWKDVARVRTAMEKNGLQKTPGWSIIQLKNE 611
>UniRef100_Q75LD1 Putative pentatricopeptide repeat domain contianing protein [Oryza
sativa]
Length = 702
Score = 340 bits (871), Expect = 9e-92
Identities = 191/550 (34%), Positives = 302/550 (54%), Gaps = 9/550 (1%)
Query: 55 SVIKACSSLHSHTLGTLLHGLALTTGSHSDPVVSNSLISLYAKFSDIQSARQVFDTMPHR 114
+ + AC+ L + G +H LA+ G D + + LI +Y++ + +A++VFD M
Sbjct: 119 AALVACADLGALRAGEQVHSLAVRAGFAGDAWIGSCLIEMYSRCGSLPAAKEVFDRMDSP 178
Query: 115 DPISWNSMINAYLQNGCLEEALEMLKGVYLLGLVPKPELLASMVSLCGRKMGSRIGRQIH 174
D + + S+I+A+ +NG E A E L + GL P + ++++ C R +G +QIH
Sbjct: 179 DVVGYTSLISAFCRNGEFELAAEALIQMLKQGLKPNEHTMTTILTACPRVLG----QQIH 234
Query: 175 ALVVVD-GRVEHSVVLSTALVDFYFRCDDALMASRVFDGMEVKNEVSWTAMISGCIANLD 233
++ G SV STAL+DFY R + +A VFD + KN VSW +M+ I +
Sbjct: 235 GYLIKKIGLRSQSVYSSTALIDFYSRNGEFKLAKAVFDSLHCKNVVSWCSMMQLYIRDGR 294
Query: 234 YDAAFARFRAMQVEGVDPNRVTLIALLPACAKPGFVKHGKEIHGYAFRHGIESSHIFSSA 293
+ A F M EGVDPN L +L AC G G+++H A +H + + S+A
Sbjct: 295 LEEALQVFGDMISEGVDPNEFALSIVLGACGSIGL---GRQLHCSAIKHDLITDIRVSNA 351
Query: 294 LINMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEP 353
L++MY ++G L E + + D+V W++ I + Q G KA+ L +M +E P
Sbjct: 352 LLSMYGRTGLVEEL-EAMLNKIENPDLVSWTTAISANFQNGFGEKAIALLCQMHSEGFTP 410
Query: 354 NYVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIF 413
N +V+S+C +++SL G H LK G I NAL+NMY+KCG + + F
Sbjct: 411 NGYAFSSVLSSCADVASLDQGMQFHCLALKLGCDSEICTGNALINMYSKCGQMGSARLAF 470
Query: 414 LEMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVT 473
M D SW+++I + HG +AL++FS+M+ G+KPD TFL VL CNH+G+V
Sbjct: 471 DVMHTHDVTSWNSLIHGHAQHGDANKALEVFSKMRSNGIKPDDSTFLGVLMGCNHSGMVE 530
Query: 474 EGQQVFKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSA 533
EG+ F+ + Y HYAC++D+LGR+G+ ++AL ++ MP +P IW +L+++
Sbjct: 531 EGELFFRLMIDQYSFTPAPSHYACMIDMLGRNGRFDEALRMINDMPFEPDALIWKTLLAS 590
Query: 534 CKLHGRLDIAEMLGPQLIRSEPDNAANYTLLNMIYAEHGHWLHKEQVMEAMKLQRLKKCY 593
CKLH LDI ++ +L+ ++A+Y L++ IYA HG W +V M +KK
Sbjct: 591 CKLHRNLDIGKLAADRLMELSDRDSASYVLMSNIYAMHGEWEDARKVRRRMDETGVKKDA 650
Query: 594 GFSRIETGNQ 603
G S IE N+
Sbjct: 651 GCSWIEINNE 660
Score = 201 bits (510), Expect = 7e-50
Identities = 137/460 (29%), Positives = 228/460 (48%), Gaps = 19/460 (4%)
Query: 84 DPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKGVY 143
D V+ ++ K + A +FD MP ++ ++W S+++ Y +NG E AL M +
Sbjct: 47 DVVLECKRLNRLVKSGRLADALDLFDRMPRKNVVAWTSVMSGYTRNGRPEAALAMFADMV 106
Query: 144 LLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCDDA 203
G+ P + + C R G Q+H+L V G + + L++ Y RC
Sbjct: 107 ESGVAPNDFACNAALVACADLGALRAGEQVHSLAVRAG-FAGDAWIGSCLIEMYSRCGSL 165
Query: 204 LMASRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPAC 263
A VFD M+ + V +T++IS N +++ A M +G+ PN T+ +L AC
Sbjct: 166 PAAKEVFDRMDSPDVVGYTSLISAFCRNGEFELAAEALIQMLKQGLKPNEHTMTTILTAC 225
Query: 264 AKPGFVKHGKEIHGYAFRH-GIESSHIFSS-ALINMYCQSGESLHLAEQIFERSSFRDVV 321
+ G++IHGY + G+ S ++SS ALI+ Y ++GE LA+ +F+ ++VV
Sbjct: 226 PR----VLGQQIHGYLIKKIGLRSQSVYSSTALIDFYSRNGE-FKLAKAVFDSLHCKNVV 280
Query: 322 LWSSIIGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFI 381
W S++ Y + G +AL +F M++E +PN L V+ AC S+ G LH
Sbjct: 281 SWCSMMQLYIRDGRLEEALQVFGDMISEGVDPNEFALSIVLGAC---GSIGLGRQLHCSA 337
Query: 382 LKFGLSFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQAL 441
+K L I +SNAL++MY + G +E + +++N D +SW+T ISA +G GE+A+
Sbjct: 338 IKHDLITDIRVSNALLSMYGRTGLVEELEAMLNKIENPDLVSWTTAISANFQNGFGEKAI 397
Query: 442 KLFSEMKERGVKPDALTFLAVLSACNHAGLVTEGQQ---VFKQVNADYEIPLTIEHYACL 498
L +M G P+ F +VLS+C + +G Q + ++ D EI T +
Sbjct: 398 ALLCQMHSEGFTPNGYAFSSVLSSCADVASLDQGMQFHCLALKLGCDSEI-CTGNALINM 456
Query: 499 VDLLGRSGKVEDALEIMRTMPMKPSTRIWSSLVSACKLHG 538
G+ G A ++M T + W+SL+ HG
Sbjct: 457 YSKCGQMGSARLAFDVMHTHDVTS----WNSLIHGHAQHG 492
Score = 154 bits (388), Expect = 9e-36
Identities = 116/389 (29%), Positives = 192/389 (48%), Gaps = 14/389 (3%)
Query: 149 PKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHS--VVLSTALVDFYFRCDDALMA 206
P+P +LA+ + R+ + + D S VVL ++ + A
Sbjct: 8 PRPRVLAAAPCHVPSRRRRRVCDGLCSAAAADNGCAESPDVVLECKRLNRLVKSGRLADA 67
Query: 207 SRVFDGMEVKNEVSWTAMISGCIANLDYDAAFARFRAMQVEGVDPNRVTLIALLPACAKP 266
+FD M KN V+WT+++SG N +AA A F M GV PN A L ACA
Sbjct: 68 LDLFDRMPRKNVVAWTSVMSGYTRNGRPEAALAMFADMVESGVAPNDFACNAALVACADL 127
Query: 267 GFVKHGKEIHGYAFRHGIESSHIFSSALINMYCQSGESLHLAEQIFERSSFRDVVLWSSI 326
G ++ G+++H A R G S LI MY + G SL A+++F+R DVV ++S+
Sbjct: 128 GALRAGEQVHSLAVRAGFAGDAWIGSCLIEMYSRCG-SLPAAKEVFDRMDSPDVVGYTSL 186
Query: 327 IGSYSQRGDSYKALTLFNKMLTEETEPNYVTLLAVISACTNLSSLKHGCGLHGFIL-KFG 385
I ++ + G+ A +ML + +PN T+ +++AC + G +HG+++ K G
Sbjct: 187 ISAFCRNGEFELAAEALIQMLKQGLKPNEHTMTTILTACPRVL----GQQIHGYLIKKIG 242
Query: 386 L-SFSISLSNALMNMYAKCGCLEGSLQIFLEMQNRDSISWSTMISAYGLHGHGEQALKLF 444
L S S+ S AL++ Y++ G + + +F + ++ +SW +M+ Y G E+AL++F
Sbjct: 243 LRSQSVYSSTALIDFYSRNGEFKLAKAVFDSLHCKNVVSWCSMMQLYIRDGRLEEALQVF 302
Query: 445 SEMKERGVKPDALTFLAVLSACNHAGLVTEGQQVFKQVNADYEIPLTIEHYACLVDLLGR 504
+M GV P+ VL AC GL G+Q+ +++ I L+ + GR
Sbjct: 303 GDMISEGVDPNEFALSIVLGACGSIGL---GRQLHCSA-IKHDLITDIRVSNALLSMYGR 358
Query: 505 SGKVEDALEIMRTMPMKPSTRIWSSLVSA 533
+G VE+ LE M P W++ +SA
Sbjct: 359 TGLVEE-LEAMLNKIENPDLVSWTTAISA 386
Score = 100 bits (250), Expect = 9e-20
Identities = 66/253 (26%), Positives = 128/253 (50%), Gaps = 13/253 (5%)
Query: 295 INMYCQSGESLHLAEQIFERSSFRDVVLWSSIIGSYSQRGDSYKALTLFNKMLTEETEPN 354
+N +SG L A +F+R ++VV W+S++ Y++ G AL +F M+ PN
Sbjct: 55 LNRLVKSGR-LADALDLFDRMPRKNVVAWTSVMSGYTRNGRPEAALAMFADMVESGVAPN 113
Query: 355 YVTLLAVISACTNLSSLKHGCGLHGFILKFGLSFSISLSNALMNMYAKCGCLEGSLQIFL 414
A + AC +L +L+ G +H ++ G + + + L+ MY++CG L + ++F
Sbjct: 114 DFACNAALVACADLGALRAGEQVHSLAVRAGFAGDAWIGSCLIEMYSRCGSLPAAKEVFD 173
Query: 415 EMQNRDSISWSTMISAYGLHGHGEQALKLFSEMKERGVKPDALTFLAVLSACNHAGLVTE 474
M + D + ++++ISA+ +G E A + +M ++G+KP+ T +L+AC
Sbjct: 174 RMDSPDVVGYTSLISAFCRNGEFELAAEALIQMLKQGLKPNEHTMTTILTACPR----VL 229
Query: 475 GQQV----FKQVNADYEIPLTIEHYACLVDLLGRSGKVEDALEIMRTMPMKPSTRIWSSL 530
GQQ+ K++ + ++ L+D R+G+ + A + ++ K + W S+
Sbjct: 230 GQQIHGYLIKKIGLRSQ---SVYSSTALIDFYSRNGEFKLAKAVFDSLHCK-NVVSWCSM 285
Query: 531 VSACKLHGRLDIA 543
+ GRL+ A
Sbjct: 286 MQLYIRDGRLEEA 298
Score = 71.6 bits (174), Expect = 6e-11
Identities = 56/239 (23%), Positives = 99/239 (40%), Gaps = 2/239 (0%)
Query: 22 IKSLISQGLYHQTLELFTQLHFSAPHSISSVLPSVIKACSSLHSHTLGTLLHGLALTTGS 81
I + G + + L Q+H SV+ +C+ + S G H LAL G
Sbjct: 384 ISANFQNGFGEKAIALLCQMHSEGFTPNGYAFSSVLSSCADVASLDQGMQFHCLALKLGC 443
Query: 82 HSDPVVSNSLISLYAKFSDIQSARQVFDTMPHRDPISWNSMINAYLQNGCLEEALEMLKG 141
S+ N+LI++Y+K + SAR FD M D SWNS+I+ + Q+G +ALE+
Sbjct: 444 DSEICTGNALINMYSKCGQMGSARLAFDVMHTHDVTSWNSLIHGHAQHGDANKALEVFSK 503
Query: 142 VYLLGLVPKPELLASMVSLCGRKMGSRIGRQIHALVVVDGRVEHSVVLSTALVDFYFRCD 201
+ G+ P ++ C G L++ + ++D R
Sbjct: 504 MRSNGIKPDDSTFLGVLMGCNHSGMVEEGELFFRLMIDQYSFTPAPSHYACMIDMLGRNG 563
Query: 202 DALMASRVFDGMEVK-NEVSWTAMISGCIANLDYD-AAFARFRAMQVEGVDPNRVTLIA 258
A R+ + M + + + W +++ C + + D A R M++ D L++
Sbjct: 564 RFDEALRMINDMPFEPDALIWKTLLASCKLHRNLDIGKLAADRLMELSDRDSASYVLMS 622
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 952,532,712
Number of Sequences: 2790947
Number of extensions: 37054499
Number of successful extensions: 100573
Number of sequences better than 10.0: 1005
Number of HSP's better than 10.0 without gapping: 813
Number of HSP's successfully gapped in prelim test: 195
Number of HSP's that attempted gapping in prelim test: 83823
Number of HSP's gapped (non-prelim): 4935
length of query: 605
length of database: 848,049,833
effective HSP length: 133
effective length of query: 472
effective length of database: 476,853,882
effective search space: 225075032304
effective search space used: 225075032304
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0055.11


