

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0051b.7
(189 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
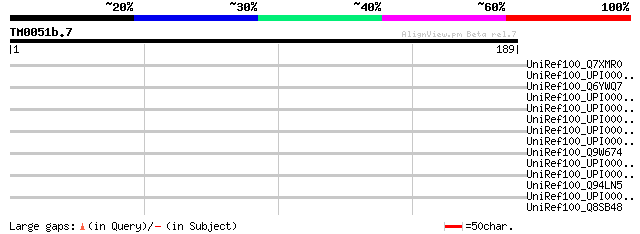
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XMR0 OSJNBa0094O15.8 protein [Oryza sativa] 35 1.0
UniRef100_UPI00003C1477 UPI00003C1477 UniRef100 entry 35 1.3
UniRef100_Q6YWQ7 Hypothetical protein OSJNBa0072H09.31 [Oryza sa... 35 1.3
UniRef100_UPI0000288C52 UPI0000288C52 UniRef100 entry 34 2.2
UniRef100_UPI00002F9FDB UPI00002F9FDB UniRef100 entry 33 2.9
UniRef100_UPI00002215A6 UPI00002215A6 UniRef100 entry 33 2.9
UniRef100_UPI000030F575 UPI000030F575 UniRef100 entry 33 3.8
UniRef100_UPI00002AAEF7 UPI00002AAEF7 UniRef100 entry 33 5.0
UniRef100_Q9W674 CCCH zinc finger protein C3H-4 [Xenopus laevis] 33 5.0
UniRef100_UPI00001D0D41 UPI00001D0D41 UniRef100 entry 32 6.5
UniRef100_UPI00001D0D3B UPI00001D0D3B UniRef100 entry 32 6.5
UniRef100_Q94LN5 Putative retroelement pol polyprotein [Oryza sa... 32 6.5
UniRef100_UPI000028A8B2 UPI000028A8B2 UniRef100 entry 32 8.5
UniRef100_Q8SB48 Putative polyprotein [Oryza sativa] 32 8.5
>UniRef100_Q7XMR0 OSJNBa0094O15.8 protein [Oryza sativa]
Length = 716
Score = 35.0 bits (79), Expect = 1.0
Identities = 16/38 (42%), Positives = 23/38 (60%)
Query: 114 FGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWIVIT 151
F G VG+ Q MV+E++H F GA Y+W+V+T
Sbjct: 110 FMDGNVGYNQIFMVEEDIHNTAFRCPGAISLYEWVVMT 147
>UniRef100_UPI00003C1477 UPI00003C1477 UniRef100 entry
Length = 967
Score = 34.7 bits (78), Expect = 1.3
Identities = 27/87 (31%), Positives = 39/87 (44%), Gaps = 6/87 (6%)
Query: 26 LRQNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVV---AVVRSDWA 82
L +DEQA++ R+ EE K RR GV +A +A +R V A S+W+
Sbjct: 127 LLDEDDEQALAGGRAREEWIKSARRARYGVGGSLARGSLAPSDIRNSTVGFEACTMSNWS 186
Query: 83 PNRNRPIQLIWSRFFPSLNPPSDPSEP 109
+ N S PS +P + S P
Sbjct: 187 SSSNNSPSTSSS---PSTSPSASTSSP 210
>UniRef100_Q6YWQ7 Hypothetical protein OSJNBa0072H09.31 [Oryza sativa]
Length = 158
Score = 34.7 bits (78), Expect = 1.3
Identities = 31/98 (31%), Positives = 41/98 (41%), Gaps = 10/98 (10%)
Query: 12 GRVTGGSGRFHRNRLRQNEDEQAISD---QRSMEEEGKKLRRCSGGVARVVAAQRMAAQV 68
G G G R+R RQ E+ + D +R + G +GGVA+ V R+
Sbjct: 27 GGEAAGGGVVERDRHRQVGGERGVGDAVAERRGLDSGCGGGGVAGGVAKRVVQARLHVHG 86
Query: 69 MRRRVVAVVRSDWAPNRNRPIQLIWSRFFPSLNPPSDP 106
+RR A VR A R P + P L PPS P
Sbjct: 87 CQRRARARVRGSAAMARRCP-------WHPPLPPPSPP 117
>UniRef100_UPI0000288C52 UPI0000288C52 UniRef100 entry
Length = 243
Score = 33.9 bits (76), Expect = 2.2
Identities = 19/48 (39%), Positives = 28/48 (57%)
Query: 129 ENVHINIFDIQGAPLFYQWIVITSVVLGIIDYDISLEYISFYVVIMLN 176
EN INIFDI + L Y I I +++ I Y + L YI+ ++I+ N
Sbjct: 128 ENNEINIFDIGISFLSYLVIFIILIIISFILYRLKLPYINKAILIITN 175
>UniRef100_UPI00002F9FDB UPI00002F9FDB UniRef100 entry
Length = 565
Score = 33.5 bits (75), Expect = 2.9
Identities = 27/90 (30%), Positives = 45/90 (50%), Gaps = 4/90 (4%)
Query: 24 NRLRQNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQR---MAAQVMRRRVVAVVRSD 80
NRL++ S++R ++EE K +R +AR+ AQ + RR+++A +
Sbjct: 414 NRLKEALKAARQSNERRLQEEQKAVRSAGVIMARLTRAQSHHCSDEERKRRKILARKVAP 473
Query: 81 WAPNRNRPIQLIWSRFFPSLNPPSDPSEPS 110
AP+R P S S +PP+ PS P+
Sbjct: 474 LAPSRKPPAPASGSGSGAS-SPPTCPSSPT 502
>UniRef100_UPI00002215A6 UPI00002215A6 UniRef100 entry
Length = 545
Score = 33.5 bits (75), Expect = 2.9
Identities = 16/47 (34%), Positives = 29/47 (61%)
Query: 129 ENVHINIFDIQGAPLFYQWIVITSVVLGIIDYDISLEYISFYVVIML 175
++V IF + GAP FYQWI + + + +S+ Y+S+Y V+++
Sbjct: 97 QHVTAIIFCVCGAPEFYQWIRMDNEGTVSLGSALSVSYLSWYAVLVV 143
>UniRef100_UPI000030F575 UPI000030F575 UniRef100 entry
Length = 202
Score = 33.1 bits (74), Expect = 3.8
Identities = 20/46 (43%), Positives = 31/46 (66%), Gaps = 1/46 (2%)
Query: 23 RNRLRQNEDEQ-AISDQRSMEEEGKKLRRCSGGVARVVAAQRMAAQ 67
+NRLR+ E EQ AI +++ MEE+ + L++ S VAR VA + + Q
Sbjct: 43 QNRLRELEAEQKAIEEKQKMEEQMRVLKQHSEDVARNVAKELVKGQ 88
>UniRef100_UPI00002AAEF7 UPI00002AAEF7 UniRef100 entry
Length = 217
Score = 32.7 bits (73), Expect = 5.0
Identities = 25/87 (28%), Positives = 43/87 (48%), Gaps = 25/87 (28%)
Query: 121 FVQEVMVKENVHINIFDIQGAPLFYQWI--VITS-----------VVLGIIDYD---ISL 164
FV+E ++ NVH +F AP FY W+ ++ + +V+ D D +L
Sbjct: 129 FVEERLIPTNVHFELF---FAPKFYSWVNKIVPNKFSSLEKDHPRLVIPFFDEDNRMFAL 185
Query: 165 EYISF------YVVIMLNPDIKKLIGL 185
+ +F Y+ I+LNP+ +K+ GL
Sbjct: 186 QGRAFGLEEPKYITIILNPEKEKIYGL 212
>UniRef100_Q9W674 CCCH zinc finger protein C3H-4 [Xenopus laevis]
Length = 276
Score = 32.7 bits (73), Expect = 5.0
Identities = 13/27 (48%), Positives = 17/27 (62%)
Query: 92 IWSRFFPSLNPPSDPSEPSGPVFGRGP 118
++S FFP L+PP+DP P P F P
Sbjct: 10 LFSSFFPQLSPPADPETPLLPSFSAPP 36
>UniRef100_UPI00001D0D41 UPI00001D0D41 UniRef100 entry
Length = 325
Score = 32.3 bits (72), Expect = 6.5
Identities = 26/110 (23%), Positives = 46/110 (41%), Gaps = 6/110 (5%)
Query: 28 QNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVVAVVRSDWAPNRNR 87
+NE+ + + DQ ++EE K+ R G + + M++ V +S +
Sbjct: 175 RNENRELLWDQIALEESIKETTRFCGEASMKICITSSLRAAMKQNYPEVHQS---ASEKD 231
Query: 88 PIQLIWSRFFPSLNPPSDPSEPSGPVFGRGPVGFVQEVMVKENVHINIFD 137
+ ++W+ FP DPS P P+ G +G V V +N D
Sbjct: 232 ELLILWNGIFPETR-DKDPSLP--PLLCFGTLGLSGVAQVLRGVPVNDID 278
>UniRef100_UPI00001D0D3B UPI00001D0D3B UniRef100 entry
Length = 355
Score = 32.3 bits (72), Expect = 6.5
Identities = 26/110 (23%), Positives = 46/110 (41%), Gaps = 6/110 (5%)
Query: 28 QNEDEQAISDQRSMEEEGKKLRRCSGGVARVVAAQRMAAQVMRRRVVAVVRSDWAPNRNR 87
+NE+ + + DQ ++EE K+ R G + + M++ V +S +
Sbjct: 205 RNENRELLWDQIALEESIKETTRFCGEASMKICITSSLRAAMKQNYPEVHQS---ASEKD 261
Query: 88 PIQLIWSRFFPSLNPPSDPSEPSGPVFGRGPVGFVQEVMVKENVHINIFD 137
+ ++W+ FP DPS P P+ G +G V V +N D
Sbjct: 262 ELLILWNGIFPETR-DKDPSLP--PLLCFGTLGLSGVAQVLRGVPVNDID 308
>UniRef100_Q94LN5 Putative retroelement pol polyprotein [Oryza sativa]
Length = 1580
Score = 32.3 bits (72), Expect = 6.5
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 1/55 (1%)
Query: 114 FGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWIVITSVVLGI-IDYDISLEYI 167
F G G+ Q M +E++H F GA ++W+VIT V+ Y ++ YI
Sbjct: 673 FMDGNAGYNQIFMAEEDIHKTTFRCPGAIGLFEWVVITFVLKSAGATYQRAMNYI 727
>UniRef100_UPI000028A8B2 UPI000028A8B2 UniRef100 entry
Length = 113
Score = 32.0 bits (71), Expect = 8.5
Identities = 18/60 (30%), Positives = 27/60 (45%), Gaps = 4/60 (6%)
Query: 57 RVVAAQRMAAQVMRRRVVAVVRSDWAPNRNRPIQL----IWSRFFPSLNPPSDPSEPSGP 112
R Q + V+ +V +VR + N P+ L + RFFP LNPP + + P
Sbjct: 8 RAALVQNVQKAVVHIKVEKIVRGSDGRSFNNPMDLYNDEFFRRFFPELNPPKNNQQREQP 67
>UniRef100_Q8SB48 Putative polyprotein [Oryza sativa]
Length = 931
Score = 32.0 bits (71), Expect = 8.5
Identities = 17/55 (30%), Positives = 29/55 (51%), Gaps = 1/55 (1%)
Query: 114 FGRGPVGFVQEVMVKENVHINIFDIQGAPLFYQWIVITSVVLGI-IDYDISLEYI 167
F G VG+ Q M +E++H F GA ++W+V+T + + Y ++ YI
Sbjct: 437 FMDGNVGYNQIFMAEEDIHKTTFTCPGAIGLFEWVVMTFGLKSVGATYQRAMNYI 491
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.143 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 324,649,167
Number of Sequences: 2790947
Number of extensions: 13074180
Number of successful extensions: 48478
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 48470
Number of HSP's gapped (non-prelim): 15
length of query: 189
length of database: 848,049,833
effective HSP length: 120
effective length of query: 69
effective length of database: 513,136,193
effective search space: 35406397317
effective search space used: 35406397317
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0051b.7


