

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0050.15
(95 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
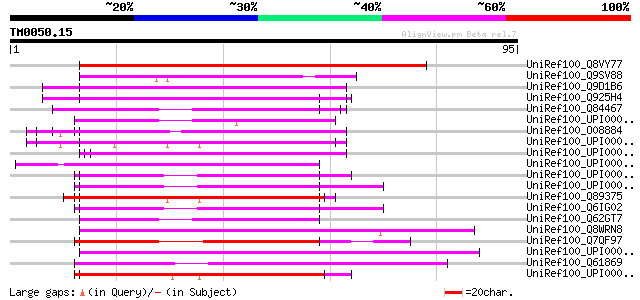
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8VY77 Putative lipase [Arabidopsis thaliana] 121 4e-27
UniRef100_Q9SV88 Hypothetical protein F24G24.130 [Arabidopsis th... 53 2e-06
UniRef100_Q9D1B6 Mus musculus 18-day embryo whole body cDNA, RIK... 52 3e-06
UniRef100_Q925H4 Keratin-associated protein 16.7 [Mus musculus] 52 3e-06
UniRef100_Q84467 A147L protein [Paramecium bursaria chlorella vi... 52 3e-06
UniRef100_UPI000036C6AD UPI000036C6AD UniRef100 entry 51 8e-06
UniRef100_O08884 High-glycine tyrosine keratin type II.4 [Mus mu... 50 1e-05
UniRef100_UPI000021EC11 UPI000021EC11 UniRef100 entry 50 1e-05
UniRef100_UPI000021DFA0 UPI000021DFA0 UniRef100 entry 49 2e-05
UniRef100_UPI000036892C UPI000036892C UniRef100 entry 49 2e-05
UniRef100_UPI000021E808 UPI000021E808 UniRef100 entry 49 3e-05
UniRef100_UPI000021DC2A UPI000021DC2A UniRef100 entry 49 3e-05
UniRef100_Q89375 A40L protein [Paramecium bursaria chlorella vir... 49 3e-05
UniRef100_Q6IG02 Type II keratin Kb2 [Rattus norvegicus] 49 3e-05
UniRef100_Q62GT7 Hypothetical protein [Burkholderia mallei] 49 3e-05
UniRef100_Q8WRN8 Putative antimicrobial peptide [Litopenaeus set... 49 3e-05
UniRef100_Q7QF97 ENSANGP00000007494 [Anopheles gambiae str. PEST] 49 4e-05
UniRef100_UPI0000452EF3 UPI0000452EF3 UniRef100 entry 48 5e-05
UniRef100_Q61869 Keratin 2 epidermis [Mus musculus] 48 5e-05
UniRef100_UPI000043799E UPI000043799E UniRef100 entry 48 7e-05
>UniRef100_Q8VY77 Putative lipase [Arabidopsis thaliana]
Length = 91
Score = 121 bits (304), Expect = 4e-27
Identities = 49/65 (75%), Positives = 60/65 (91%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
GFGVGCGFGVGWGFGGMP+N+LG+G GGGCGVG+GLGWGFG+AFGS YRSS++ FQG+E
Sbjct: 14 GFGVGCGFGVGWGFGGMPMNILGVGAGGGCGVGLGLGWGFGTAFGSHYRSSRLTFQGIEL 73
Query: 74 DSKEK 78
++ +K
Sbjct: 74 ETADK 78
>UniRef100_Q9SV88 Hypothetical protein F24G24.130 [Arabidopsis thaliana]
Length = 130
Score = 53.1 bits (126), Expect = 2e-06
Identities = 30/59 (50%), Positives = 35/59 (58%), Gaps = 9/59 (15%)
Query: 14 GFGVGCGFGVGWG------FG-GMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQ 65
G GVGCGFG G G FG G+P GLG G GCG+GVG G+G G G+ Y S+
Sbjct: 34 GLGVGCGFGFGAGLIGGVGFGPGVPGLQFGLGFGAGCGIGVGFGYGVGR--GAAYDHSR 90
>UniRef100_Q9D1B6 Mus musculus 18-day embryo whole body cDNA, RIKEN full-length
enriched library, clone:1110017B05
product:KERATIN-ASSOCIATED PROTEIN 16.7 homolog [Mus
musculus]
Length = 128
Score = 52.4 bits (124), Expect = 3e-06
Identities = 26/57 (45%), Positives = 32/57 (55%)
Query: 7 GFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G C + S +G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 11 GGCGYGSRYGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 67
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 50 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 99
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 58 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 107
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 26 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 75
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 66 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSRYGCGYGS 115
Score = 37.7 bits (86), Expect = 0.068
Identities = 23/53 (43%), Positives = 26/53 (48%), Gaps = 2/53 (3%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVG--VGLGWGFGSAFGSKYRSS 64
G G GCG+G G+G G G G G GCG G G G+G G K SS
Sbjct: 74 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSRYGCGYGSGCCSYRKCYSS 126
Score = 36.6 bits (83), Expect = 0.15
Identities = 23/53 (43%), Positives = 28/53 (52%), Gaps = 7/53 (13%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGG--GCGVGVGLGWGFGS 55
SG+ C + SG+G G G G G G+G G G G GCG G G G+GS
Sbjct: 67 SGYGCGYGSGYGCGYGSGYGCGYGSG----YGCGYGSGYGCGYGSRYGCGYGS 115
>UniRef100_Q925H4 Keratin-associated protein 16.7 [Mus musculus]
Length = 128
Score = 52.4 bits (124), Expect = 3e-06
Identities = 26/57 (45%), Positives = 32/57 (55%)
Query: 7 GFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G C + S +G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 11 GGCGYGSRYGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 67
Score = 50.8 bits (120), Expect = 8e-06
Identities = 25/51 (49%), Positives = 29/51 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G G GCG+G G+G G G G G GCG G G G G+GS +G Y SS
Sbjct: 26 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSS 76
Score = 49.7 bits (117), Expect = 2e-05
Identities = 24/50 (48%), Positives = 29/50 (58%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS++G Y S
Sbjct: 34 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSSYGCGYGS 83
Score = 48.9 bits (115), Expect = 3e-05
Identities = 23/50 (46%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+G+ +G Y S
Sbjct: 58 GSGYGCGYGSGYGCGYGSSYGCGYGSGYGCGYGSGYGCGYGTGYGCGYGS 107
Score = 45.1 bits (105), Expect = 4e-04
Identities = 21/45 (46%), Positives = 25/45 (54%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G GCG+G +G G G G G GCG G G G G+GS +G
Sbjct: 66 GSGYGCGYGSSYGCGYGSGYGCGYGSGYGCGYGTGYGCGYGSRYG 110
Score = 42.0 bits (97), Expect = 0.004
Identities = 21/48 (43%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G+G G G G G G+G G G GCG G G G G+GS +G Y
Sbjct: 56 GYGSGYGCGYGSGYG------CGYGSSYGCGYGSGYGCGYGSGYGCGY 97
Score = 36.2 bits (82), Expect = 0.20
Identities = 22/55 (40%), Positives = 27/55 (49%), Gaps = 7/55 (12%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG+ C + S +G G G G G G+G G G G G G G G G +G GS
Sbjct: 67 SGYGCGYGSSYGCGYGSGYGCGYGS------GYGCGYGTGYGCGYGSRYGCGCGS 115
>UniRef100_Q84467 A147L protein [Paramecium bursaria chlorella virus 1]
Length = 260
Score = 52.0 bits (123), Expect = 3e-06
Identities = 27/54 (50%), Positives = 28/54 (51%), Gaps = 6/54 (11%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYR 62
C F GFG G G G G GFG G G G GCG G G G GFG FG +R
Sbjct: 68 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFGCGFR 115
Score = 50.8 bits (120), Expect = 8e-06
Identities = 26/50 (52%), Positives = 26/50 (52%), Gaps = 6/50 (12%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
C F GFG G G G G GFG G G G GCG G G G GFG FG
Sbjct: 64 CGFRCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFG 107
Score = 48.9 bits (115), Expect = 3e-05
Identities = 27/55 (49%), Positives = 27/55 (49%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C F GFG G G G G GFG G G G GCG G G G GFG F RS
Sbjct: 72 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFGCGFRCGLRS 120
Score = 47.0 bits (110), Expect = 1e-04
Identities = 26/55 (47%), Positives = 27/55 (48%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C F GFG G G G G GFG G G G GCG G G G GF S +RS
Sbjct: 76 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFGCGFRCGLRSGFRS 124
Score = 45.8 bits (107), Expect = 3e-04
Identities = 26/55 (47%), Positives = 27/55 (48%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C F GFG G G G G GFG G G G GCG G G G S F S +RS
Sbjct: 80 CGFGCGFGCGFGCGFGCGFG------CGFGCGFGCGFGCGFRCGLRSGFRSGFRS 128
Score = 42.7 bits (99), Expect = 0.002
Identities = 31/71 (43%), Positives = 33/71 (45%), Gaps = 9/71 (12%)
Query: 9 CLFLSGFGVGCGFGVGWGFG-GMPLNLL-----GLGVGGGCGVGVGLGWGFGSAFGSKYR 62
C F GFG G G G G GFG G L G G GC G G GFG FGS +
Sbjct: 92 CGFGCGFGCGFGCGFGCGFGCGFRCGLRSGFRSGFRSGFGCRFGCGFRSGFGCRFGSGFG 151
Query: 63 SSQIRFQGVEF 73
S F+GV F
Sbjct: 152 SG---FRGVLF 159
>UniRef100_UPI000036C6AD UPI000036C6AD UniRef100 entry
Length = 89
Score = 50.8 bits (120), Expect = 8e-06
Identities = 25/51 (49%), Positives = 29/51 (56%), Gaps = 8/51 (15%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGG--GCGVGVGLGWGFGSAFGSKY 61
SG G GCG+G G+G G G G G GCG G G G G+G FGS+Y
Sbjct: 12 SGSGCGCGYGTGYGCG------YGCGFGSRYGCGYGTGYGCGYGCGFGSRY 56
Score = 42.4 bits (98), Expect = 0.003
Identities = 24/54 (44%), Positives = 28/54 (51%), Gaps = 10/54 (18%)
Query: 9 CLFLSGFG--VGCGFGVGWGFGGMPLNLLGLGVGG--GCGVGVGLGWGFGSAFG 58
C + GFG GCG+G G+G G G G G GCG G G G G+GS G
Sbjct: 26 CGYGCGFGSRYGCGYGTGYGCG------YGCGFGSRYGCGYGTGYGCGYGSGSG 73
Score = 40.4 bits (93), Expect = 0.011
Identities = 20/53 (37%), Positives = 27/53 (50%), Gaps = 6/53 (11%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
C + +G+G G G G G +G G G G GCG G G G +G +G+ Y
Sbjct: 18 CGYGTGYGCGYGCGFGSRYG------CGYGTGYGCGYGCGFGSRYGCGYGTGY 64
Score = 38.5 bits (88), Expect = 0.040
Identities = 16/29 (55%), Positives = 18/29 (61%)
Query: 33 NLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
N G G G GCG G G G G+G FGS+Y
Sbjct: 8 NSCGSGSGCGCGYGTGYGCGYGCGFGSRY 36
Score = 32.3 bits (72), Expect = 2.9
Identities = 17/41 (41%), Positives = 20/41 (48%)
Query: 36 GLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEFDSK 76
G G G G G G G G GFGS +G Y + G F S+
Sbjct: 15 GCGCGYGTGYGCGYGCGFGSRYGCGYGTGYGCGYGCGFGSR 55
>UniRef100_O08884 High-glycine tyrosine keratin type II.4 [Mus musculus]
Length = 159
Score = 50.4 bits (119), Expect = 1e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 37 GSGYGCGYGSGYGCGSGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 86
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 69 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 118
Score = 49.3 bits (116), Expect = 2e-05
Identities = 28/59 (47%), Positives = 34/59 (57%), Gaps = 3/59 (5%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG+ C + SG+G G G G G G+G G G G GCG G G G G+GS +GS Y S
Sbjct: 70 SGYGCGYGSGYGCGYGSGYGCGYGSG--YGCGYGSGYGCGYGSGYGCGYGSGYGSGYGS 126
Score = 48.5 bits (114), Expect = 4e-05
Identities = 24/51 (47%), Positives = 28/51 (54%), Gaps = 8/51 (15%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG G GCG+G G+G G G G GCG G G G G+GS +G Y S
Sbjct: 52 SGSGYGCGYGSGYG--------CGYGSGYGCGYGSGYGCGYGSGYGCGYGS 94
Score = 48.1 bits (113), Expect = 5e-05
Identities = 28/61 (45%), Positives = 34/61 (54%), Gaps = 3/61 (4%)
Query: 4 SHSGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYR 62
S SG+ C + SG+G G G G G G+G G G G GCG G G G G+GS +G Y
Sbjct: 52 SGSGYGCGYGSGYGCGYGSGYGCGYGSG--YGCGYGSGYGCGYGSGYGCGYGSGYGCGYG 109
Query: 63 S 63
S
Sbjct: 110 S 110
Score = 45.8 bits (107), Expect = 3e-04
Identities = 25/55 (45%), Positives = 29/55 (52%), Gaps = 6/55 (10%)
Query: 9 CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
C + SG+G G G G G G+G G G G GCG G G G G GS +G Y S
Sbjct: 14 CGYGSGYGSGYGCGSGSGYG------CGYGSGYGCGYGSGYGCGSGSGYGCGYGS 62
Score = 44.7 bits (104), Expect = 6e-04
Identities = 23/50 (46%), Positives = 27/50 (54%), Gaps = 6/50 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G+G G G G G G+G G G G GCG G G G G+GS +G Y S
Sbjct: 35 GYGSGYGCGYGSGYG------CGSGSGYGCGYGSGYGCGYGSGYGCGYGS 78
Score = 43.5 bits (101), Expect = 0.001
Identities = 20/44 (45%), Positives = 25/44 (56%)
Query: 15 FGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G CG+G G+G G G G G GCG G G G G+GS +G
Sbjct: 6 YGNSCGYGCGYGSGYGSGYGCGSGSGYGCGYGSGYGCGYGSGYG 49
Score = 38.5 bits (88), Expect = 0.040
Identities = 24/59 (40%), Positives = 30/59 (50%), Gaps = 11/59 (18%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
SG+ C + SG+G G G G G G+G + G G G GCG G +GS YRS
Sbjct: 94 SGYGCGYGSGYGCGYGSGYGCGYGSGYGSGYGSGYGSGCGCG----------YGSYYRS 142
Score = 31.2 bits (69), Expect = 6.4
Identities = 15/32 (46%), Positives = 16/32 (49%)
Query: 30 MPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
M N G G GCG G G G G+G GS Y
Sbjct: 1 MCCNYYGNSCGYGCGYGSGYGSGYGCGSGSGY 32
Score = 30.8 bits (68), Expect = 8.3
Identities = 15/31 (48%), Positives = 16/31 (51%)
Query: 33 NLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
N G G G G G G G G G GS +G Y S
Sbjct: 8 NSCGYGCGYGSGYGSGYGCGSGSGYGCGYGS 38
>UniRef100_UPI000021EC11 UPI000021EC11 UniRef100 entry
Length = 134
Score = 50.1 bits (118), Expect = 1e-05
Identities = 23/48 (47%), Positives = 28/48 (57%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G GCG+G G+G G G G G GCG G G G G+GS +G +Y
Sbjct: 40 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCRY 87
Score = 49.7 bits (117), Expect = 2e-05
Identities = 27/60 (45%), Positives = 32/60 (53%), Gaps = 2/60 (3%)
Query: 4 SHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
S G C + SG+G G G G G G+G G G G GCG G G G G+GS +G Y S
Sbjct: 8 SSCGGCGYGSGYGCGYGSGYGCGYGSR--YGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 65
Score = 48.1 bits (113), Expect = 5e-05
Identities = 28/65 (43%), Positives = 35/65 (53%), Gaps = 7/65 (10%)
Query: 6 SGF-CLFLSGFGVG------CGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SG+ C + SG+G G CG+G G+G G G G G GCG G G G G+GS +G
Sbjct: 17 SGYGCGYGSGYGCGYGSRYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYG 76
Query: 59 SKYRS 63
Y S
Sbjct: 77 CGYGS 81
Score = 47.0 bits (110), Expect = 1e-04
Identities = 29/65 (44%), Positives = 35/65 (53%), Gaps = 7/65 (10%)
Query: 6 SGF-CLFLSGFGVGCGFGVGWGFG---GMPLNL---LGLGVGGGCGVGVGLGWGFGSAFG 58
SG+ C + SG+G G G G G G+G G L G G G GCG G G G G+GS +G
Sbjct: 57 SGYGCGYGSGYGCGYGSGYGCGYGSGYGCRYGLGYGCGYGSGYGCGYGSGYGCGYGSRYG 116
Query: 59 SKYRS 63
Y S
Sbjct: 117 CGYGS 121
Score = 37.0 bits (84), Expect = 0.12
Identities = 22/53 (41%), Positives = 25/53 (46%), Gaps = 10/53 (18%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVG--VGLGWGFGSAFGSKYRSS 64
G G GCG+G G+G G G G GCG G G G+G G K SS
Sbjct: 88 GLGYGCGYGSGYG--------CGYGSGYGCGYGSRYGCGYGSGCCSYRKCHSS 132
>UniRef100_UPI000021DFA0 UPI000021DFA0 UniRef100 entry
Length = 123
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 17 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 66
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 33 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 82
Score = 49.3 bits (116), Expect = 2e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 25 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 74
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/50 (48%), Positives = 28/50 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 41 GSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSRYGCGYGS 90
Score = 47.8 bits (112), Expect = 7e-05
Identities = 23/48 (47%), Positives = 27/48 (55%)
Query: 16 GVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G GCG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 11 GYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 58
Score = 44.7 bits (104), Expect = 6e-04
Identities = 22/49 (44%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Query: 15 FGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
+G CG+G G+G G G G G GCG G G G G+GS +G Y S
Sbjct: 6 YGNSCGYGCGYGSG----YGCGYGSGYGCGYGSGYGCGYGSGYGCGYGS 50
Score = 42.0 bits (97), Expect = 0.004
Identities = 23/50 (46%), Positives = 26/50 (52%), Gaps = 6/50 (12%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
G+G G G G G G+G G G GCG G G G G G +GS YRS
Sbjct: 63 GYGSGYGCGYGSGYG------CGYGSRYGCGYGSGYGNGCGCGYGSYYRS 106
Score = 34.3 bits (77), Expect = 0.75
Identities = 16/34 (47%), Positives = 18/34 (52%)
Query: 30 MPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRS 63
M N G G GCG G G G G+GS +G Y S
Sbjct: 1 MCCNYYGNSCGYGCGYGSGYGCGYGSGYGCGYGS 34
>UniRef100_UPI000036892C UPI000036892C UniRef100 entry
Length = 493
Score = 49.3 bits (116), Expect = 2e-05
Identities = 28/57 (49%), Positives = 33/57 (57%), Gaps = 1/57 (1%)
Query: 2 SLSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
S+S SG F + FG G G G G+GFGG + G G G G G G+G G GFG FG
Sbjct: 75 SISTSGGS-FRNRFGAGAGAGGGYGFGGGAGSGFGFGGGAGGGFGLGGGAGFGGGFG 130
>UniRef100_UPI000021E808 UPI000021E808 UniRef100 entry
Length = 597
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/45 (57%), Positives = 27/45 (59%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 517 GFGGGSGFGGGRGFGG------GSGFGGGSGFGGGSGFGGGSGFG 555
Score = 47.4 bits (111), Expect = 9e-05
Identities = 26/46 (56%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 510 SGFGGSGGFGGGSGFGG------GRGFGGGSGFGGGSGFGGGSGFG 549
Score = 46.6 bits (109), Expect = 1e-04
Identities = 26/52 (50%), Positives = 29/52 (55%), Gaps = 6/52 (11%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
SGFG G GFG G GFGG G G GGG G G G G+G G G ++ S
Sbjct: 522 SGFGGGRGFGGGSGFGG------GSGFGGGSGFGGGSGFGGGGFGGGRFGGS 567
Score = 38.5 bits (88), Expect = 0.040
Identities = 26/52 (50%), Positives = 27/52 (51%), Gaps = 12/52 (23%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG-FGSAFGSKYRS 63
SGFG G GFG G GFGG G G GGG G G G FG + G Y S
Sbjct: 534 SGFGGGSGFGGGSGFGG------GSGFGGG-----GFGGGRFGGSGGGGYSS 574
Score = 37.0 bits (84), Expect = 0.12
Identities = 23/45 (51%), Positives = 23/45 (51%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG GFGG G G GGG G G G GFG G
Sbjct: 505 GFGGGSGFGGSGGFGG------GSGFGGGRGFGG--GSGFGGGSG 541
Score = 36.6 bits (83), Expect = 0.15
Identities = 24/48 (50%), Positives = 26/48 (54%), Gaps = 8/48 (16%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSK 60
SGFG G GFG G GFGG G GGG G G G G+ S GS+
Sbjct: 540 SGFGGGSGFGGGSGFGGG-------GFGGGRFGGSG-GGGYSSGGGSR 579
Score = 33.5 bits (75), Expect = 1.3
Identities = 20/45 (44%), Positives = 21/45 (46%), Gaps = 8/45 (17%)
Query: 22 GVGWGFGGMPLNLLG--------LGVGGGCGVGVGLGWGFGSAFG 58
G G GFGG G G GGG G G G G+G GS FG
Sbjct: 493 GYGGGFGGRSYGGFGGGSGFGGSGGFGGGSGFGGGRGFGGGSGFG 537
Score = 32.3 bits (72), Expect = 2.9
Identities = 19/46 (41%), Positives = 22/46 (47%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SG CG G G G+GG + G+G G G G G GFG G
Sbjct: 76 SGGRSSCGSGFGSGYGGSFGSSRGMGGGFGGASGFGGAGGFGRPGG 121
Score = 31.6 bits (70), Expect = 4.9
Identities = 24/61 (39%), Positives = 29/61 (47%), Gaps = 9/61 (14%)
Query: 1 KSLSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSK 60
KS+S S ++G+G G G GFGG G G GG G G G G G G GS
Sbjct: 485 KSISSS-----VAGYGGGFGGRSYGGFGGGS----GFGGSGGFGGGSGFGGGRGFGGGSG 535
Query: 61 Y 61
+
Sbjct: 536 F 536
>UniRef100_UPI000021DC2A UPI000021DC2A UniRef100 entry
Length = 491
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/45 (57%), Positives = 27/45 (59%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 30 GFGGGSGFGGGRGFGG------GSGFGGGSGFGGGSGFGGGSGFG 68
Score = 47.4 bits (111), Expect = 9e-05
Identities = 26/46 (56%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 23 SGFGGSGGFGGGSGFGG------GRGFGGGSGFGGGSGFGGGSGFG 62
Score = 46.2 bits (108), Expect = 2e-04
Identities = 27/58 (46%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQG 70
SGFG G GFG G GFGG G G GGG G G G G+G G G ++ F G
Sbjct: 35 SGFGGGRGFGGGSGFGG------GSGFGGGSGFGGGSGFGGGGFGGGRFGGGPGGFGG 86
Score = 37.0 bits (84), Expect = 0.12
Identities = 23/45 (51%), Positives = 23/45 (51%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG GFGG G G GGG G G G GFG G
Sbjct: 18 GFGGGSGFGGSGGFGG------GSGFGGGRGFGG--GSGFGGGSG 54
Score = 33.5 bits (75), Expect = 1.3
Identities = 20/45 (44%), Positives = 21/45 (46%), Gaps = 8/45 (17%)
Query: 22 GVGWGFGGMPLNLLG--------LGVGGGCGVGVGLGWGFGSAFG 58
G G GFGG G G GGG G G G G+G GS FG
Sbjct: 6 GYGGGFGGRSYGGFGGGSGFGGSGGFGGGSGFGGGRGFGGGSGFG 50
Score = 33.1 bits (74), Expect = 1.7
Identities = 24/46 (52%), Positives = 24/46 (52%), Gaps = 9/46 (19%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG G GFG G GFGG G G GG G G G GFG G
Sbjct: 53 SGFGGGSGFGGGSGFGGG-----GFG-GGRFGGGPG---GFGGPGG 89
Score = 30.8 bits (68), Expect = 8.3
Identities = 20/50 (40%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
++G+G G G GFGG G G GG G G G G G G GS +
Sbjct: 4 VAGYGGGFGGRSYGGFGGGS----GFGGSGGFGGGSGFGGGRGFGGGSGF 49
>UniRef100_Q89375 A40L protein [Paramecium bursaria chlorella virus 1]
Length = 129
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/51 (50%), Positives = 32/51 (61%), Gaps = 2/51 (3%)
Query: 11 FLSGFGVGCGFGVGWGFG-GMPLNL-LGLGVGGGCGVGVGLGWGFGSAFGS 59
F +G G G G G+G GFG G+ L G GVG G G+G G G GFG+ FG+
Sbjct: 20 FGAGLGTGFGAGLGTGFGTGLGAGLGAGFGVGFGAGLGTGFGAGFGTGFGA 70
Score = 48.5 bits (114), Expect = 4e-05
Identities = 24/49 (48%), Positives = 28/49 (56%), Gaps = 6/49 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
+GFG G G G G G G GLG G G G G GLG GFG+ FG+ +
Sbjct: 26 TGFGAGLGTGFGTGLGA------GLGAGFGVGFGAGLGTGFGAGFGTGF 68
Score = 47.4 bits (111), Expect = 9e-05
Identities = 24/46 (52%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
G G G G G+G GFG GLG G G G+G GLG GFG FG+
Sbjct: 15 GLGTGFGAGLGTGFGA------GLGTGFGTGLGAGLGAGFGVGFGA 54
Score = 45.8 bits (107), Expect = 3e-04
Identities = 22/49 (44%), Positives = 28/49 (56%), Gaps = 6/49 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
+GFG G G G+G GFG +G G G G G G G G GFG+ G+ +
Sbjct: 34 TGFGTGLGAGLGAGFG------VGFGAGLGTGFGAGFGTGFGAGLGAGF 76
Score = 44.3 bits (103), Expect = 7e-04
Identities = 23/48 (47%), Positives = 25/48 (51%), Gaps = 6/48 (12%)
Query: 11 FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
F +G G G G G G GFG GLG G G G G G G G G+ FG
Sbjct: 36 FGTGLGAGLGAGFGVGFGA------GLGTGFGAGFGTGFGAGLGAGFG 77
Score = 40.8 bits (94), Expect = 0.008
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G GFGVG+G G LG G G G G G G G G G G
Sbjct: 41 GAGLGAGFGVGFGAG------LGTGFGAGFGTGFGAGLGAGFGDG 79
Score = 39.7 bits (91), Expect = 0.018
Identities = 20/40 (50%), Positives = 23/40 (57%), Gaps = 6/40 (15%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWG 52
+GFGVG G G+G GFG G G G G G+G G G G
Sbjct: 46 AGFGVGFGAGLGTGFGA------GFGTGFGAGLGAGFGDG 79
Score = 37.7 bits (86), Expect = 0.068
Identities = 18/42 (42%), Positives = 23/42 (53%), Gaps = 6/42 (14%)
Query: 20 GFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKY 61
G G+G GFG GLG G G G+G G G G G+ G+ +
Sbjct: 13 GVGLGTGFGA------GLGTGFGAGLGTGFGTGLGAGLGAGF 48
>UniRef100_Q6IG02 Type II keratin Kb2 [Rattus norvegicus]
Length = 685
Score = 48.9 bits (115), Expect = 3e-05
Identities = 26/45 (57%), Positives = 27/45 (59%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 104 GFGGGSGFGGGRGFGG------GSGFGGGSGFGGGSGFGGGSGFG 142
Score = 47.4 bits (111), Expect = 9e-05
Identities = 26/46 (56%), Positives = 27/46 (58%), Gaps = 6/46 (13%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG GFG G GFGG G G GGG G G G G+G GS FG
Sbjct: 97 SGFGGSGGFGGGSGFGG------GRGFGGGSGFGGGSGFGGGSGFG 136
Score = 46.2 bits (108), Expect = 2e-04
Identities = 27/58 (46%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQG 70
SGFG G GFG G GFGG G G GGG G G G G+G G G ++ F G
Sbjct: 109 SGFGGGRGFGGGSGFGG------GSGFGGGSGFGGGSGFGGGGFGGGRFGGGPGGFGG 160
Score = 37.0 bits (84), Expect = 0.12
Identities = 23/45 (51%), Positives = 23/45 (51%), Gaps = 8/45 (17%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG GFGG G G GGG G G G GFG G
Sbjct: 92 GFGGGSGFGGSGGFGG------GSGFGGGRGFGG--GSGFGGGSG 128
Score = 33.5 bits (75), Expect = 1.3
Identities = 20/45 (44%), Positives = 21/45 (46%), Gaps = 8/45 (17%)
Query: 22 GVGWGFGGMPLNLLG--------LGVGGGCGVGVGLGWGFGSAFG 58
G G GFGG G G GGG G G G G+G GS FG
Sbjct: 80 GYGGGFGGRSYGGFGGGSGFGGSGGFGGGSGFGGGRGFGGGSGFG 124
Score = 31.6 bits (70), Expect = 4.9
Identities = 24/61 (39%), Positives = 29/61 (47%), Gaps = 9/61 (14%)
Query: 1 KSLSHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSK 60
KS+S S ++G+G G G GFGG G G GG G G G G G G GS
Sbjct: 72 KSISSS-----VAGYGGGFGGRSYGGFGGGS----GFGGSGGFGGGSGFGGGRGFGGGSG 122
Query: 61 Y 61
+
Sbjct: 123 F 123
>UniRef100_Q62GT7 Hypothetical protein [Burkholderia mallei]
Length = 188
Score = 48.9 bits (115), Expect = 3e-05
Identities = 24/45 (53%), Positives = 26/45 (57%), Gaps = 6/45 (13%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
GFG G GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 77 GFGFGFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 115
Score = 41.6 bits (96), Expect = 0.005
Identities = 22/47 (46%), Positives = 25/47 (52%), Gaps = 6/47 (12%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+ G V GFG G+GFG G G G G G G G G+GFG FG
Sbjct: 67 IRGRKVRFGFGFGFGFG------FGFGFGFGFGFGFGFGFGFGFGFG 107
>UniRef100_Q8WRN8 Putative antimicrobial peptide [Litopenaeus setiferus]
Length = 188
Score = 48.9 bits (115), Expect = 3e-05
Identities = 30/81 (37%), Positives = 41/81 (50%), Gaps = 7/81 (8%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRF----- 68
G G G G G+G G GG+ L GLG G G G+G GLG G G G + +S R+
Sbjct: 53 GLGGGLGGGLGGGLGGLGGGLGGLGGGLGGGLGGGLGGGLGGGLGGSHGTSDCRYWCKTP 112
Query: 69 --QGVEFDSKEKGDSPPLSKP 87
Q +S + ++P +KP
Sbjct: 113 EGQAYCCESAHEPETPVGTKP 133
Score = 40.8 bits (94), Expect = 0.008
Identities = 28/58 (48%), Positives = 31/58 (53%), Gaps = 4/58 (6%)
Query: 5 HSGFCLFLSGFGV---GCGFGVGWGFGGMPLNLLGLGVGGGCGVGV-GLGWGFGSAFG 58
+ GF L G GV G G GVG G GG LG G+GGG G G+ GLG G G G
Sbjct: 21 YRGFGQPLGGLGVPGGGVGVGVGGGLGGGLGGGLGGGLGGGLGGGLGGLGGGLGGLGG 78
Score = 39.7 bits (91), Expect = 0.018
Identities = 24/53 (45%), Positives = 27/53 (50%), Gaps = 8/53 (15%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCG--------VGVGLGWGFGSAFG 58
G GVG G G+G G GG LG G+GGG G +G GLG G G G
Sbjct: 37 GVGVGVGGGLGGGLGGGLGGGLGGGLGGGLGGLGGGLGGLGGGLGGGLGGGLG 89
Score = 39.3 bits (90), Expect = 0.023
Identities = 24/46 (52%), Positives = 27/46 (58%), Gaps = 2/46 (4%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGG-GCGVGVGLGWGFGSAFG 58
G G G G G+G G GG L LG G+GG G G+G GLG G G G
Sbjct: 49 GLGGGLGGGLGGGLGG-GLGGLGGGLGGLGGGLGGGLGGGLGGGLG 93
>UniRef100_Q7QF97 ENSANGP00000007494 [Anopheles gambiae str. PEST]
Length = 171
Score = 48.5 bits (114), Expect = 4e-05
Identities = 21/46 (45%), Positives = 30/46 (64%), Gaps = 8/46 (17%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
SGFG+G GF +G GFG +G+G G G+G G+ +GFG ++G
Sbjct: 1 SGFGIGSGFEIGSGFG--------IGIGPGIGIGFGISFGFGVSYG 38
Score = 44.7 bits (104), Expect = 6e-04
Identities = 25/62 (40%), Positives = 30/62 (48%), Gaps = 4/62 (6%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
GFG+ GFGV +G G G+G G G G G G+G GF GS S G+E
Sbjct: 26 GFGISFGFGVSYGIGSGIRIGSGIGTGSGIGTGSGIGTGFRIEIGSGIEIS----SGIEI 81
Query: 74 DS 75
S
Sbjct: 82 SS 83
Score = 40.8 bits (94), Expect = 0.008
Identities = 24/63 (38%), Positives = 31/63 (49%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVE 72
SG G+G G G+G GFG G V G G G+ +G G G+ G + S I G+E
Sbjct: 83 SGIGIGSGIGIGSGFGIGCGIRFGFRVSYGIGSGIIIGSGIGTGSGIENGSGIIVSSGIE 142
Query: 73 FDS 75
S
Sbjct: 143 IGS 145
Score = 39.3 bits (90), Expect = 0.023
Identities = 21/57 (36%), Positives = 30/57 (51%), Gaps = 12/57 (21%)
Query: 13 SGFGVGCGFGVGWGF-----------GGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G+G G G+G GF G+ ++ G+G+G G G+G G G G G FG
Sbjct: 51 TGSGIGTGSGIGTGFRIEIGSGIEISSGIEISS-GIGIGSGIGIGSGFGIGCGIRFG 106
>UniRef100_UPI0000452EF3 UPI0000452EF3 UniRef100 entry
Length = 920
Score = 48.1 bits (113), Expect = 5e-05
Identities = 29/75 (38%), Positives = 36/75 (47%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVEF 73
G G+G G G+G G G P +G+G+G G G+GVGLG G GS GS
Sbjct: 698 GPGIGLGIGMGPGLGMGPGLGIGMGMGIGMGMGVGLGLGLGSGSGSDSGLEMKMGSETSL 757
Query: 74 DSKEKGDSPPLSKPT 88
DS S S P+
Sbjct: 758 DSNSNPISKSSSSPS 772
Score = 43.9 bits (102), Expect = 0.001
Identities = 22/45 (48%), Positives = 26/45 (56%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
G G+G G G G G G P LG+G+G G G+G GLG G G G
Sbjct: 682 GIGMGAGIGTGIGAGIGPGIGLGIGMGPGLGMGPGLGIGMGMGIG 726
Score = 42.4 bits (98), Expect = 0.003
Identities = 20/46 (43%), Positives = 27/46 (58%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
+G G G G G+G G G P +G G+G G G+G+G+G G G G
Sbjct: 691 TGIGAGIGPGIGLGIGMGPGLGMGPGLGIGMGMGIGMGMGVGLGLG 736
Score = 42.0 bits (97), Expect = 0.004
Identities = 24/60 (40%), Positives = 31/60 (51%), Gaps = 9/60 (15%)
Query: 2 SLSHSGFCL---FLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
S+S SG F G G+G G G G G G +G G+G G G+G+G+G G G G
Sbjct: 663 SISDSGLATNPGFGPGLGIGIGMGAGIGTG------IGAGIGPGIGLGIGMGPGLGMGPG 716
>UniRef100_Q61869 Keratin 2 epidermis [Mus musculus]
Length = 707
Score = 48.1 bits (113), Expect = 5e-05
Identities = 31/70 (44%), Positives = 32/70 (45%), Gaps = 6/70 (8%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSSQIRFQGVE 72
SGFG G GFG G GFGG G GGG G G G G+G GS FG F G
Sbjct: 94 SGFGGGRGFGGGQGFGGSG------GFGGGSGFGGGQGFGGGSRFGGGSGFGGGGFGGGS 147
Query: 73 FDSKEKGDSP 82
F G P
Sbjct: 148 FGGGRFGGGP 157
Score = 36.2 bits (82), Expect = 0.20
Identities = 26/57 (45%), Positives = 27/57 (46%), Gaps = 8/57 (14%)
Query: 4 SHSGFCLFLSGFGVGC--GFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
S G F G G G GFG GFGG G G GGG G G G +G GS FG
Sbjct: 89 SFGGSSGFGGGRGFGGGQGFGGSGGFGG------GSGFGGGQGFGGGSRFGGGSGFG 139
Score = 32.7 bits (73), Expect = 2.2
Identities = 22/47 (46%), Positives = 22/47 (46%), Gaps = 6/47 (12%)
Query: 13 SGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGS 59
SG G G G G G GG G G GGG G G G G GS GS
Sbjct: 596 SGGGYGSGGGSRGGSGG------GYGSGGGSGSGGGYSSGGGSRGGS 636
Score = 32.7 bits (73), Expect = 2.2
Identities = 24/57 (42%), Positives = 28/57 (49%), Gaps = 2/57 (3%)
Query: 4 SHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSK 60
S SG + SG G G G G+G GG G G G G G G G G G+ S GS+
Sbjct: 579 SGSGGGGYSSGGGSRGGSGGGYGSGGGSRG--GSGGGYGSGGGSGSGGGYSSGGGSR 633
>UniRef100_UPI000043799E UPI000043799E UniRef100 entry
Length = 577
Score = 47.8 bits (112), Expect = 7e-05
Identities = 26/51 (50%), Positives = 30/51 (57%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFGSKYRSS 64
G G G G G+G G GG LGLG+GGG G G G +G GS+F S SS
Sbjct: 527 GLGGGLGGGLGGGLGGGLGGGLGLGLGGGLGFGAGSSFGAGSSFMSSSLSS 577
Score = 46.2 bits (108), Expect = 2e-04
Identities = 26/51 (50%), Positives = 33/51 (63%), Gaps = 4/51 (7%)
Query: 13 SGFGVGCGFGVGWGFGG-MPLNL---LGLGVGGGCGVGVGLGWGFGSAFGS 59
+G G+G GFG+G G GG + L LG G+GGG G G+GLG G G FG+
Sbjct: 510 AGLGLGSGFGLGGGLGGGLGGGLGGGLGGGLGGGLGGGLGLGLGGGLGFGA 560
Score = 40.4 bits (93), Expect = 0.011
Identities = 22/43 (51%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Query: 14 GFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSA 56
GFG G G G G G G+ L L +GGG G+G GLG G G A
Sbjct: 49 GFGAGAGAGAGAG-AGLGLGGGSLALGGGLGLGGGLGLGAGGA 90
Score = 38.1 bits (87), Expect = 0.052
Identities = 21/55 (38%), Positives = 29/55 (52%), Gaps = 3/55 (5%)
Query: 4 SHSGFCLFLSGFGVGCGFGVGWGFGGMPLNLLGLGVGGGCGVGVGLGWGFGSAFG 58
S G + +S +G G G G GG+ G G+GGG G+G+G G+G G G
Sbjct: 474 SGGGVTVHMSKTLLGGGAG---GAGGLGAGAGGAGLGGGAGLGLGSGFGLGGGLG 525
Score = 34.7 bits (78), Expect = 0.58
Identities = 23/52 (44%), Positives = 25/52 (47%), Gaps = 5/52 (9%)
Query: 12 LSGFGVGCGFGVGWGFGGMPLNL-----LGLGVGGGCGVGVGLGWGFGSAFG 58
L G G G G+G G GG L LG G G G G+G GLG G G G
Sbjct: 486 LLGGGAGGAGGLGAGAGGAGLGGGAGLGLGSGFGLGGGLGGGLGGGLGGGLG 537
Score = 30.8 bits (68), Expect = 8.3
Identities = 14/25 (56%), Positives = 14/25 (56%)
Query: 35 LGLGVGGGCGVGVGLGWGFGSAFGS 59
LGLG G G G G G G G G GS
Sbjct: 46 LGLGFGAGAGAGAGAGAGLGLGGGS 70
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.147 0.481
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 190,780,556
Number of Sequences: 2790947
Number of extensions: 8905729
Number of successful extensions: 78118
Number of sequences better than 10.0: 1253
Number of HSP's better than 10.0 without gapping: 714
Number of HSP's successfully gapped in prelim test: 574
Number of HSP's that attempted gapping in prelim test: 46244
Number of HSP's gapped (non-prelim): 13712
length of query: 95
length of database: 848,049,833
effective HSP length: 71
effective length of query: 24
effective length of database: 649,892,596
effective search space: 15597422304
effective search space used: 15597422304
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0050.15


