

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
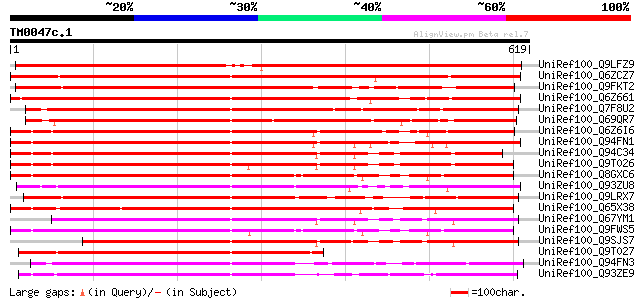
Query= TM0047c.1
(619 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LFZ9 F20N2.11 [Arabidopsis thaliana] 629 e-179
UniRef100_Q6ZCZ7 Phosphatidylinositol transfer-like [Oryza sativa] 618 e-175
UniRef100_Q9FKT2 Similarity to phosphatidylinositol/phosphatidyl... 563 e-159
UniRef100_Q6Z661 Putative phosphatidylinositol transfer [Oryza s... 563 e-159
UniRef100_Q7F8U2 Putative phosphatidylinositol-phosphatidylcholi... 532 e-149
UniRef100_Q69QR7 Putative phosphatidylinositol transfer-like pro... 524 e-147
UniRef100_Q6Z6I6 Putative phosphatidylinositol/ phophatidylcholi... 504 e-141
UniRef100_Q94FN1 Phosphatidylinositol transfer-like protein III ... 499 e-140
UniRef100_Q94C34 AT4g39170/T22F8_70 [Arabidopsis thaliana] 496 e-139
UniRef100_Q9T026 SEC14-like protein [Arabidopsis thaliana] 481 e-134
UniRef100_Q8GXC6 Putative sec14 cytosolic factor [Arabidopsis th... 469 e-131
UniRef100_Q93ZU8 Putative sec14 cytosolic factor [Arabidopsis th... 461 e-128
UniRef100_Q9LRX7 Phosphatidylinositol/phosphatidylcholine transf... 455 e-126
UniRef100_Q65X38 Hypothetical protein OJ1206_C08.12 [Oryza sativa] 450 e-125
UniRef100_Q67YM1 Putative phosphatidylinositol/ phosphatidylchol... 449 e-124
UniRef100_Q9FWS5 F1B16.10 protein [Arabidopsis thaliana] 444 e-123
UniRef100_Q9SJS7 Putative phosphatidylinositol/phosphatidylcholi... 417 e-115
UniRef100_Q9T027 SEC14-like protein [Arabidopsis thaliana] 405 e-111
UniRef100_Q94FN3 Phosphatidylinositol transfer-like protein II [... 405 e-111
UniRef100_Q93ZE9 At2g21540/F2G1.19 [Arabidopsis thaliana] 405 e-111
>UniRef100_Q9LFZ9 F20N2.11 [Arabidopsis thaliana]
Length = 636
Score = 629 bits (1622), Expect = e-179
Identities = 344/616 (55%), Positives = 428/616 (68%), Gaps = 32/616 (5%)
Query: 8 DEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDV 67
DE RERRSD E SEDERR SKIG L+KKA+NAS+KFTHSLKKRGKRKIDYRVPAVSIEDV
Sbjct: 38 DEFRERRSDFEISEDERRRSKIGNLKKKAINASTKFTHSLKKRGKRKIDYRVPAVSIEDV 97
Query: 68 RDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEY 127
RDE EE+VV E R++L+ER LLPPRHD+Y+TLLRFLKARD NIEKT Q+WEEML WRKEY
Sbjct: 98 RDEKEESVVLEFRRKLLERDLLPPRHDEYHTLLRFLKARDLNIEKTTQLWEEMLRWRKEY 157
Query: 128 GTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKY 187
GTDTILE+F+FEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPS+LMRITTIDRYLKY
Sbjct: 158 GTDTILEDFDFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSKLMRITTIDRYLKY 217
Query: 188 HVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYY 247
HVQEFERALQEKFPACSIAAKRRI STTTILDVQGLG+KNFT TAANL+AAM+KID +YY
Sbjct: 218 HVQEFERALQEKFPACSIAAKRRICSTTTILDVQGLGIKNFTPTAANLVAAMSKIDNSYY 277
Query: 248 PETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVID-------S 300
PE L + + M + C P + KS Y +V +
Sbjct: 278 PEVLDFLNFTS------HMFFT-----CIPSCL--------KSSYYFADVAQNVHCKCWN 318
Query: 301 SQLPDFLGGSCTCPTE-GGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDS 359
SQLP+FLGGSC+C + GGCL+++KGPWNDP+I+KL++ E++L RQ +R + S
Sbjct: 319 SQLPEFLGGSCSCFGDGGGCLRSNKGPWNDPEIMKLIYHGESSLFRQSTRKLTDPHYSSS 378
Query: 360 F-QVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDD 418
+ +H K +++S AES S + SSPT + S + EE R D+NGYYSCDD
Sbjct: 379 YISIHPSKAIQAETSAAESISCSDVPSSPTGRLCSASSHVNSAYEEARASDVNGYYSCDD 438
Query: 419 SAPTTEKVV-ENGQFHLSQKQSLQTNDME-NVTHRTISEGTSVNNLFRIIKEIVEKTNHL 476
+K GQ SQ Q + N + T S G + ++ +++K
Sbjct: 439 KFAIPDKATNRKGQERQSQYQMRELNATTIGLKCETSSPGAPIIRWLHDLRVMIDKIKCE 498
Query: 477 YVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHN--INNHSAAVEAAFERDHIL 534
+ + L S M +L R E R+Q V PS+ TE + + S +D IL
Sbjct: 499 NLAKRLLSLMLKLAAVFRYTPLELLRSQTTVSPSSLTEDDSRCSLISPPPREPTMKDRIL 558
Query: 535 PCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQL 594
PC +R+Q+LEK +E++ NKP +P+EKE+MLMDS+DRIKSVEFDL+KTKR+LH+ V +Q+
Sbjct: 559 PCLERIQKLEKSYEDIRNKPVAIPVEKERMLMDSLDRIKSVEFDLDKTKRLLHATVMKQM 618
Query: 595 EIADLVENLQKSNCRV 610
EI ++++N++ S V
Sbjct: 619 EITEMLQNIRDSQLHV 634
>UniRef100_Q6ZCZ7 Phosphatidylinositol transfer-like [Oryza sativa]
Length = 637
Score = 618 bits (1594), Expect = e-175
Identities = 334/625 (53%), Positives = 426/625 (67%), Gaps = 21/625 (3%)
Query: 2 EGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPA 61
EG S DE ++RRSDVENSEDERR IG+L+KKA+NAS+K THSLKKRGKRK++ R P+
Sbjct: 11 EGLFSFDERKDRRSDVENSEDERRRLSIGSLKKKALNASNKLTHSLKKRGKRKVENR-PS 69
Query: 62 VSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEML 121
+IEDVRDE EE V +Q L R LLP +H+DY+ LLRFLKAR F+ EK IQMW EML
Sbjct: 70 FTIEDVRDEEEERAVFSFQQELFSRNLLPDKHNDYHMLLRFLKARKFDTEKAIQMWAEML 129
Query: 122 LWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTI 181
WRKE+G DTILE+F FEEL+EVL YYPQGYHGVD++GRPVYIERLGK P++LM ITT+
Sbjct: 130 QWRKEFGADTILEDFNFEELDEVLVYYPQGYHGVDRQGRPVYIERLGKVEPNKLMHITTV 189
Query: 182 DRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTK 241
DRY+KYHVQEFERA EKFPACSIAAKR I STTTILDV G+G+KNF++TA ++L M K
Sbjct: 190 DRYMKYHVQEFERAFHEKFPACSIAAKRHIDSTTTILDVDGVGLKNFSKTARDMLGRMQK 249
Query: 242 IDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSS 301
ID++YYPETLHQM++VNAG+GF K+LW + F DPKT +KI +L +K KLLEVID+S
Sbjct: 250 IDSDYYPETLHQMFVVNAGNGF-KLLWNTVKGFLDPKTASKIHVLGTKFHGKLLEVIDAS 308
Query: 302 QLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF- 360
QLP+FLGG+CTC EGGCLK++KGPWNDP+I+KL H+ EA R R S +Q SF
Sbjct: 309 QLPEFLGGACTCAAEGGCLKSNKGPWNDPNIMKLAHNKEAKFTRHTRRLSEIEQRRGSFA 368
Query: 361 QVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEE--VRVPDLNGYYSCDD 418
++H LKGR SD+ST ESGSD++D SSP +R RLAPV EE +R D YYSCDD
Sbjct: 369 RLHLLKGRSSDTSTVESGSDVDDLSSPMMRRPVECSRLAPVREEMQIRARDSAAYYSCDD 428
Query: 419 SAPTTEKVVENGQFHL-----------SQKQSLQTNDMENVTHRTISE-GTSVNNLFRII 466
+K V+ G+ +Q + + T + + G S +N R +
Sbjct: 429 HFVVVDKTVDYGRGGAMPDKTSAPEVRAQARPFGGSTTSYATGSSSNRGGISSSNRSRTV 488
Query: 467 --KEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAV 524
KE ++ R+L + + ++ F + +N P E ++H A
Sbjct: 489 VPKENTDEGFFRRFFRLLLALIIKVFAFFHIAYGQQEMRVDNPLPPAEPEPTSDDHPAV- 547
Query: 525 EAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKR 584
F D I P +RLQRLE +EL NKP +PLEKE+ L++S DRIK +E DLE+TK+
Sbjct: 548 -ETFSVDRISPVIERLQRLEGKVDELGNKPPEIPLEKERSLLESWDRIKCIESDLERTKK 606
Query: 585 VLHSAVKQQLEIADLVENLQKSNCR 609
VL + V +QLEIA+ +E + + R
Sbjct: 607 VLQATVMKQLEIAESIEEVIRRQLR 631
>UniRef100_Q9FKT2 Similarity to phosphatidylinositol/phosphatidylcholine transfer
protein [Arabidopsis thaliana]
Length = 592
Score = 563 bits (1451), Expect = e-159
Identities = 294/597 (49%), Positives = 399/597 (66%), Gaps = 37/597 (6%)
Query: 8 DEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKR-GKRKIDYRVPAVSIED 66
++ E+ SD E E+E R S+IG L+KKA + S+K TH LK R GKRKID+++P IED
Sbjct: 20 EQTGEKLSDSEYIEEEPRRSRIGNLKKKAFSCSTKLTHPLKMRKGKRKIDFQIPL--IED 77
Query: 67 VRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKE 126
VRDE EE +V +LRQ+L+++ LLPP HDDY+ LLRFLK +F IEKT+ WEEML WRKE
Sbjct: 78 VRDEKEEKLVSKLRQQLLQKDLLPPVHDDYHMLLRFLKTMEFKIEKTVTAWEEMLKWRKE 137
Query: 127 YGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLK 186
+GTD I+++F F+EL+EV ++YPQGYHGVDK+GRP+YIERLGKAHP +LM +TTI+RYLK
Sbjct: 138 FGTDRIIQDFNFKELDEVTRHYPQGYHGVDKDGRPIYIERLGKAHPGKLMEVTTIERYLK 197
Query: 187 YHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNY 246
YHVQEFER LQEK PACS+AAKRR+ +TTTILDV+GLGMKNFT TAANLLA + K+D NY
Sbjct: 198 YHVQEFERTLQEKLPACSVAAKRRVTTTTTILDVEGLGMKNFTPTAANLLATIAKVDCNY 257
Query: 247 YPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDF 306
YPETLH+M+IVNAG GF+ LWPAAQK DP TIAKIQ+L+ +SL KLLE IDSSQLP+F
Sbjct: 258 YPETLHRMFIVNAGIGFRSFLWPAAQKLLDPMTIAKIQVLEPRSLSKLLEAIDSSQLPEF 317
Query: 307 LGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLK 366
LGG C CP EGGCL+++KGPWNDP+I++LVH E V Q + A +++DS
Sbjct: 318 LGGLCKCPNEGGCLRSNKGPWNDPEIVELVHHMEVNNVPQTTTAPLHVRDYDSTTC---- 373
Query: 367 GRCSDSSTAESGSDINDYSSPTRQRSSTHLRLA-PVDEEVRVPDLNGYYSCDDSAPTTEK 425
S T + + +Y S T RSS H + P+ ++ D D TT +
Sbjct: 374 -TISPKETLKEEPEPEEYYSSTGSRSSMHTCIVPPLSDKASTSD-------GDKFITTVE 425
Query: 426 VVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASF 485
+E+ +Q Q L + + ++ EG + F ++E + N ++ ++L F
Sbjct: 426 SIES-----AQSQLLDADTENTFANTSVREGGQILR-FGALREKINSENIFHLVKILLVF 479
Query: 486 MERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEK 545
+L L +W+ QN V +++ +N +L C RL+++EK
Sbjct: 480 PLKLFVLFGFLLPGYWQRQNTVVVPDSSTNN---------------KVLECFDRLKKMEK 524
Query: 546 VFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVEN 602
F E++ K +P EK+L +S++RIKS+E DL+KTK VLH + +QL+I + +E+
Sbjct: 525 EFTEISRKQVKIPEANEKLLAESLERIKSLELDLDKTKSVLHITLTKQLQITEQLES 581
>UniRef100_Q6Z661 Putative phosphatidylinositol transfer [Oryza sativa]
Length = 612
Score = 563 bits (1450), Expect = e-159
Identities = 302/623 (48%), Positives = 418/623 (66%), Gaps = 41/623 (6%)
Query: 2 EGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPA 61
E S DE R+R D E SEDE R ++I +L+KKA++AS++ THSLKKRGKRK+ RVP
Sbjct: 10 EISASNDERRDR-GDAEISEDEPRQTRIRSLKKKALHASTRLTHSLKKRGKRKVGCRVPK 68
Query: 62 VSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEML 121
++IEDVRD EE V R+ L R +LP RHDDY+T+LRFLKAR F++EK MW +ML
Sbjct: 69 ITIEDVRDAEEEQAVSSFREVLFARDMLPERHDDYHTMLRFLKARKFDVEKAAHMWADML 128
Query: 122 LWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTI 181
WRK++GTDTILE+FEF ELEEVLQYYP GYHGVDKEGRPVYIE LGK PS+L++ITT+
Sbjct: 129 HWRKDFGTDTILEDFEFHELEEVLQYYPHGYHGVDKEGRPVYIELLGKVEPSKLVQITTV 188
Query: 182 DRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTK 241
+RY+KYHVQEFERA +EKFPACSIAAK+ I +TTTILDV G+G KNF++ A +L+ M K
Sbjct: 189 ERYIKYHVQEFERAFREKFPACSIAAKKHIDTTTTILDVHGVGWKNFSKIARDLVRCMQK 248
Query: 242 IDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSS 301
ID +YYPETLHQM+IVNAG GF K++W + DPKT +KI +L +K ++LLE IDSS
Sbjct: 249 IDGDYYPETLHQMFIVNAGPGF-KLIWSTVKGLLDPKTSSKIHVLGTKYQHRLLEAIDSS 307
Query: 302 QLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASN-EQQNFDSF 360
QLP+FLGGSCTC ++GGCL+++KGPW+DP I+KLVH E++ ++ + + S+ E+ S
Sbjct: 308 QLPEFLGGSCTCSSQGGCLRSNKGPWSDPLIMKLVHCMESSALKDIGQVSDIEEAITGSV 367
Query: 361 QVHSLK--GRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDD 418
++ +LK R S +S AESGSD++D SP Q + LAPV EE R S
Sbjct: 368 RLRALKLPERISYTSNAESGSDVDDLGSPIGQEDFEYHSLAPVHEEAR-------ESGST 420
Query: 419 SAPTTEKVVE-----------NGQFHLSQKQSLQTNDMENVTH-RTISEGTSVNNLFRII 466
+ + +KVVE +GQ+ Q S+ E H EG + + + +
Sbjct: 421 CSGSDDKVVETNTRYNPPGNGSGQYSARQNPSINRVSPEPAGHVPNDGEGNADHGILK-- 478
Query: 467 KEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEA 526
Y+++ + + +++F+R F R Q P +TT + N +
Sbjct: 479 ----------YISKKVLGVILEVLSFLRI--FIRHRQQLENVPQHTTTVHSNQADLQI-- 524
Query: 527 AFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVL 586
+ D + PC +RL+RLE +F +L+ KP +P +K++ + DS DRIK +EFDLEKTK+VL
Sbjct: 525 -IKEDRVNPCLERLERLETMFNQLSRKPPEIPQDKDRAIQDSFDRIKCIEFDLEKTKKVL 583
Query: 587 HSAVKQQLEIADLVENLQKSNCR 609
H+ V +Q+++A+ +E +++S+ R
Sbjct: 584 HATVIRQMQMAETLEAVKESDLR 606
>UniRef100_Q7F8U2 Putative phosphatidylinositol-phosphatidylcholine transfer protein
[Oryza sativa]
Length = 604
Score = 532 bits (1371), Expect = e-149
Identities = 287/594 (48%), Positives = 395/594 (66%), Gaps = 20/594 (3%)
Query: 19 NSEDERRSSKIGT-LRKKAMNASSKFTHSLKKRGKRKIDYRVP-AVSIEDVRDEGEETVV 76
NSEDERR +IG+ LR+KA+ H++KKRG+R++D R P A+SIEDVRD EE V
Sbjct: 17 NSEDERRRRRIGSNLRRKAI-------HAIKKRGRRRVDCRFPPAISIEDVRDAEEERAV 69
Query: 77 HELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEF 136
RL GLLP +HDDY+ +LRFLKAR F+I++ +QMW +ML WR+E+G DTIL++F
Sbjct: 70 AAFHDRLAAHGLLPDKHDDYHMMLRFLKARKFDIDRAMQMWADMLKWREEFGADTILQDF 129
Query: 137 EFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERAL 196
+F EL+EVL+YYPQGYHGVD+EGRPVYIERLGK P++LM+IT++DRY+KYHVQEFERA
Sbjct: 130 DFHELDEVLRYYPQGYHGVDREGRPVYIERLGKVDPNKLMQITSVDRYIKYHVQEFERAF 189
Query: 197 QEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYI 256
+E+FPAC++AAKR I STTTILDVQG+G KNF++TA L+ M KID++YYPETLHQM++
Sbjct: 190 RERFPACTLAAKRHIDSTTTILDVQGVGFKNFSKTARELINRMQKIDSDYYPETLHQMFV 249
Query: 257 VNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTE 316
VNAGSGF K++W + + F DPKT +KI +L S +LLEVIDSS+LPDFLGGSC+C +
Sbjct: 250 VNAGSGF-KLIWNSVKGFLDPKTSSKIHVLGSNYQSRLLEVIDSSELPDFLGGSCSCSDK 308
Query: 317 GGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASN-EQQNFDSFQVHSLK--GRCSDSS 373
GGCL ++KGPWNDP I+KL+H+ EA VR++ S+ E+++ S ++ K G SD S
Sbjct: 309 GGCLGSNKGPWNDPFILKLIHNLEAGCVREIKPVSDGEERSSSSLRLEQPKWQGMISDIS 368
Query: 374 TAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFH 433
AESGSD++D+ S Q+ + L PV EEVR D YYSCDD T +
Sbjct: 369 NAESGSDVDDFGS-FFQKGVDYGYLTPVHEEVRGTDSLTYYSCDDQ--TRRDIAPESCKG 425
Query: 434 LSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFI 493
+ +Q +N T N + ++ Y+ V+ +F+ +L++F+
Sbjct: 426 VQATGMVQNQLPDNRQPSTNRNPHDSGNNGHLDGAFARRSLQNYIQVVVTTFI-KLLSFL 484
Query: 494 RSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNK 553
R R NVH + + + D + C QRL LE + L ++
Sbjct: 485 RLFISRPVRRLENVHSCTVP---VPSEEKPEPRSIRDDDMTMCLQRLDSLESLCNHLASR 541
Query: 554 PYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSN 607
P +P EKE ML++S +RIK +E DLE+TKRVLH+ V +Q + + +E +Q+S+
Sbjct: 542 PPEIPREKEHMLLNSFERIKCIEADLERTKRVLHATVVKQKALVETLEAVQESS 595
>UniRef100_Q69QR7 Putative phosphatidylinositol transfer-like protein II| [Oryza
sativa]
Length = 611
Score = 524 bits (1350), Expect = e-147
Identities = 293/600 (48%), Positives = 403/600 (66%), Gaps = 34/600 (5%)
Query: 19 NSEDERRSSKIGT-LRKKAMNASSKFTHSLKKRG----KRKIDYRVPA-VSIEDVRDEGE 72
NSED+RR +G+ LR+KA+ A L+KRG +R++D+R PA +SIEDVRD E
Sbjct: 17 NSEDDRRRRGMGSSLRRKAIRA-------LRKRGGRRRRRRVDFRYPAAMSIEDVRDAEE 69
Query: 73 ETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTI 132
E V R RL LLP +HDDY+ +LRFLKAR F+ EK +QMW EML WRKE+G DTI
Sbjct: 70 ELAVAAFRDRLAVHALLPDKHDDYHMMLRFLKARKFDSEKAMQMWAEMLRWRKEFGADTI 129
Query: 133 LEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEF 192
LEEFEF+EL++VL+YYPQGYHGVD+EGRPVYIERLGK +P++LM+IT++DRY+KYHVQEF
Sbjct: 130 LEEFEFDELDDVLRYYPQGYHGVDREGRPVYIERLGKVYPNKLMQITSVDRYIKYHVQEF 189
Query: 193 ERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLH 252
ERA +E+FPAC++AAKR I STTTILDV G+G+KNF++TA L+ M KID++YYPETLH
Sbjct: 190 ERAFRERFPACTLAAKRHIDSTTTILDVHGVGLKNFSKTARELVHRMQKIDSDYYPETLH 249
Query: 253 QMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCT 312
QMY+VNAGSGF K++W + + F DPKT +KI +L + +LLEVID S+LP+FLGGSCT
Sbjct: 250 QMYVVNAGSGF-KLIWNSVKGFLDPKTSSKIHVLGTNYQSRLLEVIDKSELPEFLGGSCT 308
Query: 313 CPTEGGCLKTSKGPWNDPDIIKLVHS-AEATLVRQLSRAS-NEQQNFDSFQVHSLKGRCS 370
C +EGGCL ++KGPWND I+KL+HS ++ +R++ + S +E ++ S + LKG S
Sbjct: 309 C-SEGGCLGSNKGPWNDHVILKLIHSMRSSSSMREIKQVSDSEDRSGSSLRAEKLKGMMS 367
Query: 371 DSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCD--DSAPTTEKVVE 428
D S AES SD++++S RS+ + L PV EEV+ D + + SC+ D + E
Sbjct: 368 DISNAESESDVDEFSLSAVLRSTDYSFLTPVSEEVKGSDSSTFCSCESCDRKGLPDVTPE 427
Query: 429 NGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRII----KEIVEKTNHLYVTRVLAS 484
+ Q S +QS + + V+H S +NNL + + +T +V RV+ +
Sbjct: 428 SSQ---SVQQSSEMVPNQLVSHEHSSTTRWMNNLGNMAISFHGTLTGRTLSNFV-RVVGT 483
Query: 485 FMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLE 544
M +++ NVHPSN E SA D++ C QRL++LE
Sbjct: 484 LMIKILAVFSLFVSRRGNMLENVHPSN-VEDEPQPRSAT------EDNMSACLQRLEKLE 536
Query: 545 KVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQ 604
+ L +KP MP EKE +L+ S DRIK++E DLE+TKRVLH + +Q+E+ + +E +Q
Sbjct: 537 SLCNHLMSKPPDMPKEKECLLLQSFDRIKTIESDLERTKRVLHMTLVKQMEMMETLEAMQ 596
>UniRef100_Q6Z6I6 Putative phosphatidylinositol/ phophatidylcholine transfer protein
[Oryza sativa]
Length = 624
Score = 504 bits (1298), Expect = e-141
Identities = 287/619 (46%), Positives = 405/619 (65%), Gaps = 40/619 (6%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVP 60
+EG S D+ R+ +SDVENSED++R+ ++G+L+KKA++AS+K HSLKK+ KR+ RV
Sbjct: 13 FEGSSSNDDKRDHKSDVENSEDDKRT-RMGSLKKKAIDASTKIRHSLKKK-KRRSGSRVL 70
Query: 61 AVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEM 120
+VSIEDVRD E V RQ LI LLP RHDDY+ +LRFLKAR F+IEK QMW +M
Sbjct: 71 SVSIEDVRDLEELQAVEAFRQALILDELLPARHDDYHMMLRFLKARRFDIEKAKQMWTDM 130
Query: 121 LLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITT 180
L WRKEYGTDTI+E+F++ EL+ VLQYYP GYHGVDK+GRPVYIERLGK P++LM +TT
Sbjct: 131 LKWRKEYGTDTIVEDFDYNELDAVLQYYPHGYHGVDKDGRPVYIERLGKVDPNKLMHVTT 190
Query: 181 IDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
+DRY++YHV+EFER+ KFPACS+AAKR I S+TTILDVQG+G+KNF++TA L+ +
Sbjct: 191 MDRYVRYHVKEFERSFLIKFPACSLAAKRHIDSSTTILDVQGVGLKNFSKTARELIVRLQ 250
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDS 300
KID + YPETL+QM+IVNAG GF ++LW + F DPKT +KI +L +K KLLEVID+
Sbjct: 251 KIDNDNYPETLYQMFIVNAGPGF-RLLWNTVKSFLDPKTTSKIHVLGNKYQSKLLEVIDA 309
Query: 301 SQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF 360
S+LP+FLGG+CTCP GGCLK KGPW D +I+ +V S EA RQ+ SN ++ S+
Sbjct: 310 SELPEFLGGACTCPEYGGCLKAEKGPWKDQNILNIVLSGEAQCARQIVTVSNGEEKIISY 369
Query: 361 ---QVHSLKGRCSDSSTAESGSDINDYSSPTRQRS-STHLRLAPVDEEVRVPDLNGYYS- 415
+ H+++G SD+STAESGS+ D +SP RS +H +L PV EEV++ + +
Sbjct: 370 AKSKHHTIRG--SDTSTAESGSEAEDVTSPKVLRSYISHPKLTPVREEVKMVRATSFSTR 427
Query: 416 -CDDSAPTTEKVVE-NGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKT 473
+ P +K V+ + +++K + + D + T +E +S +L RI
Sbjct: 428 MPEYDVPVVDKAVDATWKREVTRKTAFSSKD----SSLTSTESSSNGSLDRI-------- 475
Query: 474 NHLYVTRVLASFMERLITFIRSL----------RFEFWRTQNNVHPSNTTEHNINNHSAA 523
V +LA FM +IT +RS+ + E + + ++P + + S
Sbjct: 476 ----VAVLLAVFM-AIITLVRSVKDLAAKRLPDKNESEQKYSTLYPDSMPKEEFRPPS-P 529
Query: 524 VEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTK 583
E + QRL LE+ F L +KP MP EKE++L ++ R+ ++E +L TK
Sbjct: 530 TPGFVEAELFSSVLQRLGDLEEKFLMLQDKPSEMPCEKEELLNAAVRRVDALEAELIVTK 589
Query: 584 RVLHSAVKQQLEIADLVEN 602
+ LH A+ +Q E+ +++
Sbjct: 590 KALHEALIRQEELLAYIDS 608
>UniRef100_Q94FN1 Phosphatidylinositol transfer-like protein III [Lotus japonicus]
Length = 625
Score = 499 bits (1285), Expect = e-140
Identities = 279/629 (44%), Positives = 400/629 (63%), Gaps = 42/629 (6%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRG-KRKIDYRV 59
+EG +DE RERRSD ENSEDERR+ +IG+L+KKA++AS+KF HSL+K+ +RK D RV
Sbjct: 13 FEGFSGSDEKRERRSDFENSEDERRT-RIGSLKKKALSASTKFKHSLRKKSSRRKSDGRV 71
Query: 60 PAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEE 119
+VSIEDVRD E V RQ LI LLP DDY+ +LRFLKAR F+IEK MW E
Sbjct: 72 SSVSIEDVRDVEELQAVDAFRQSLIMDELLPQAFDDYHMMLRFLKARKFDIEKAKHMWAE 131
Query: 120 MLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRIT 179
ML WRKE+G DTI+++FEF+EL+EV++YYP G+HGVDKEGRPVYIERLGK P++LM++T
Sbjct: 132 MLQWRKEFGADTIMQDFEFQELDEVVRYYPHGHHGVDKEGRPVYIERLGKVDPNKLMQVT 191
Query: 180 TIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAM 239
T+DRY++YHVQEFE++ KFPAC+IAAKR I S+TTILDVQG+G+KNFT++A L+ +
Sbjct: 192 TMDRYVRYHVQEFEKSFAIKFPACTIAAKRHIDSSTTILDVQGVGLKNFTKSARELITRL 251
Query: 240 TKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVID 299
K+D + YPETL QM+I+NAG GF ++LW + F DPKT +KI +L +K KLLEVID
Sbjct: 252 QKVDGDNYPETLCQMFIINAGPGF-RLLWNTVKSFLDPKTTSKIHVLGNKYHSKLLEVID 310
Query: 300 SSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDS 359
+S+LP+FLGG+CTC +GGCL++ KGPW +P+I+K+V + E RQ+ + N + +
Sbjct: 311 ASELPEFLGGACTCEDQGGCLRSDKGPWKNPEILKMVLNGEPRRARQVVKVLNSEGKVIA 370
Query: 360 F---QVHSLKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPD----LNG 412
+ + ++KG SD+STAESGS+ D +SP ++ +HLRL PV EE ++ N
Sbjct: 371 YAKPRYPTVKG--SDTSTAESGSEAEDIASPKAMKNYSHLRLTPVREEAKIVGKSSYTNN 428
Query: 413 YYSCDDSAPTTEKVVE---NGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEI 469
+ D+ P +K V+ Q L + + Q T RT EG L I
Sbjct: 429 FSGYDEYVPMVDKPVDAVLKKQASLQRSYTSQGAPSRPATQRT-PEGIQARILVAI---- 483
Query: 470 VEKTNHLYVTRVLASFMERLITFIRSLRFEFWR---TQNNVHPSNTTEHNINN------H 520
+F+ + T R + + ++ H +T+E +
Sbjct: 484 -------------TAFLLTIFTIFRQVACRVTKKLPAISSNHDQSTSEPTFDTTVVEVIP 530
Query: 521 SAAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLE 580
S++ A E + + +RL LE+ + L +KP MP EKE++L ++ R+ ++E +L
Sbjct: 531 SSSTPAHTEENLLPSMLKRLGELEEKVDTLQSKPSEMPYEKEELLNAAVCRVDALEAELI 590
Query: 581 KTKRVLHSAVKQQLEIADLVENLQKSNCR 609
TK+ L+ A+ +Q E+ ++ ++ R
Sbjct: 591 ATKKALYEALMRQEELLAYIDRQAEAKLR 619
>UniRef100_Q94C34 AT4g39170/T22F8_70 [Arabidopsis thaliana]
Length = 583
Score = 496 bits (1278), Expect = e-139
Identities = 275/596 (46%), Positives = 382/596 (63%), Gaps = 36/596 (6%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVP 60
+EG S+DE +ER+SD ENSEDERR+ +IG+L+KKA+NAS+KF HSLKK+ +RK D RV
Sbjct: 13 FEGILSSDEKKERKSDFENSEDERRT-RIGSLKKKAINASTKFKHSLKKK-RRKSDVRVS 70
Query: 61 AVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEM 120
+VSIEDVRD E V E RQ L+ LLP +HDDY+ +LRFLKAR F+IEK MW +M
Sbjct: 71 SVSIEDVRDVEELQAVDEFRQALVMEELLPHKHDDYHMMLRFLKARKFDIEKAKHMWADM 130
Query: 121 LLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITT 180
+ WRKE+GTDTI+++F+FEE++EVL+YYP GYH VDKEGRPVYIERLGK P++LM++TT
Sbjct: 131 IQWRKEFGTDTIIQDFQFEEIDEVLKYYPHGYHSVDKEGRPVYIERLGKVDPNKLMQVTT 190
Query: 181 IDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
+DRY++YHV+EFER+ KFPAC+IAAK+ I S+TTILDVQG+G+KNFT++A L+ +
Sbjct: 191 LDRYIRYHVKEFERSFMLKFPACTIAAKKYIDSSTTILDVQGVGLKNFTKSARELITRLQ 250
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDS 300
KID + YPETLHQM+I+NAG GF ++LW + F DPKT +KI +L K KLLE+IDS
Sbjct: 251 KIDGDNYPETLHQMFIINAGPGF-RLLWSTVKSFLDPKTTSKIHVLGCKYQSKLLEIIDS 309
Query: 301 SQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF 360
S+LP+FLGG+CTC +GGC+ + KGPW +P+I+K+V A + + + N ++
Sbjct: 310 SELPEFLGGACTCADQGGCMLSDKGPWKNPEIVKMVLHGGAHRAKHVVKVLNSDGKVIAY 369
Query: 361 QVHS---LKGRCSDSSTAESGSDIND-YSSPTRQRSSTHLRLAPVDEEVRVPD-----LN 411
S +KG SD+STAESGS+ D SP +S +HLRL PV EE +V
Sbjct: 370 AKPSYPWIKG--SDTSTAESGSEAEDIVVSPKAVKSYSHLRLTPVREEAKVGSGETSFAG 427
Query: 412 GYYSCDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVE 471
+ D+ P +K V+ +R S+G + V
Sbjct: 428 SFAGYDEYVPMVDKAVD-----------ATWKVKPTAINRAPSKGAH-------MPPNVP 469
Query: 472 KTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERD 531
K + + RVL +FM ++ + R R P ++ I +AA EA D
Sbjct: 470 KDHESFSARVLVTFMAFVMAILTFFRTVSNRVVTKQLPPPPSQPQIEGSAAAEEA----D 525
Query: 532 HILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLH 587
+ ++L LE+ L +KP MP EKE++L ++ R+ ++E +L TK+ L+
Sbjct: 526 LLNSVLKKLTELEEKIGALQSKPSEMPYEKEELLNAAVCRVDALEAELIATKKALY 581
>UniRef100_Q9T026 SEC14-like protein [Arabidopsis thaliana]
Length = 617
Score = 481 bits (1239), Expect = e-134
Identities = 273/613 (44%), Positives = 385/613 (62%), Gaps = 39/613 (6%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVP 60
+EG S+DE +ER+SD ENSEDERR+ +IG+L+KKA+NAS+KF HSLKK+ +RK D RV
Sbjct: 13 FEGILSSDEKKERKSDFENSEDERRT-RIGSLKKKAINASTKFKHSLKKK-RRKSDVRVS 70
Query: 61 AVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEM 120
+VSIEDVRD E V E RQ L+ LLP +HDDY+ +LRFLKAR F+IEK MW +M
Sbjct: 71 SVSIEDVRDVEELQAVDEFRQALVMEELLPHKHDDYHMMLRFLKARKFDIEKAKHMWADM 130
Query: 121 LLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITT 180
+ WRKE+GTDTI+++F+FEE++EVL+YYP GYH VDKEGRPVYIERLGK P++LM++TT
Sbjct: 131 IQWRKEFGTDTIIQDFQFEEIDEVLKYYPHGYHSVDKEGRPVYIERLGKVDPNKLMQVTT 190
Query: 181 IDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
+DRY++YHV+EFER+ KFPAC+IAAK+ I S+TTILDVQG+G+KNFT++A L+ +
Sbjct: 191 LDRYIRYHVKEFERSFMLKFPACTIAAKKYIDSSTTILDVQGVGLKNFTKSARELITRLQ 250
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKI---QILDSKSLYKLLEV 297
KID + YPETLHQM+I+NAG GF ++LW + F DPKT +KI IL + +
Sbjct: 251 KIDGDNYPETLHQMFIINAGPGF-RLLWSTVKSFLDPKTTSKIHNYSILLCFAYISDVSF 309
Query: 298 IDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNF 357
I S+LP+FLGG+CTC +GGC+ + KGPW +P+I+K+V A +Q+ + N
Sbjct: 310 ICFSELPEFLGGACTCADQGGCMLSDKGPWKNPEIVKMVLHGGAHRAKQVVKVLNSDGKV 369
Query: 358 DSFQVHS---LKGRCSDSSTAESGSDIND-YSSPTRQRSSTHLRLAPVDEEVRVPD---- 409
++ S +KG SD+STAESGS+ D SP +S +HLRL PV EE +V
Sbjct: 370 IAYAKPSYPWIKG--SDTSTAESGSEAEDIVVSPKAVKSYSHLRLTPVREEAKVGSGETS 427
Query: 410 -LNGYYSCDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKE 468
+ D+ P +K V+ +R S+G +
Sbjct: 428 FAGSFAGYDEYVPMVDKAVD-----------ATWKVKPTAINRAPSKGAH-------MPP 469
Query: 469 IVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAF 528
V K + + RVL +FM ++ + R R P ++ I +AA EA
Sbjct: 470 NVPKDHESFSARVLVTFMAFVMAILTFFRTVSNRVVTKQLPPPPSQPQIEGSAAAEEA-- 527
Query: 529 ERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHS 588
D + ++L LE+ L +KP MP EKE++L ++ R+ ++E +L TK+ L+
Sbjct: 528 --DLLNSVLKKLTELEEKIGALQSKPSEMPYEKEELLNAAVCRVDALEAELIATKKALYE 585
Query: 589 AVKQQLEIADLVE 601
A+ +Q E+ ++
Sbjct: 586 ALMRQEELLAYID 598
>UniRef100_Q8GXC6 Putative sec14 cytosolic factor [Arabidopsis thaliana]
Length = 612
Score = 469 bits (1208), Expect = e-131
Identities = 273/614 (44%), Positives = 379/614 (61%), Gaps = 42/614 (6%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTL--RKKAMNASSKFTHSLKKRG--KRKID 56
+EG S DE RERRSD E SEDE+++ +IG +KKA ASSK HSLKK+G +R+
Sbjct: 13 FEGVSSNDERRERRSDFEVSEDEKKT-RIGNFNFKKKAAKASSKLRHSLKKKGSSRRRSS 71
Query: 57 YRVPAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQM 116
R +++IED+ D E V E R L+ LLPP DDY+ +LRFLKAR F+I KT M
Sbjct: 72 DRTFSLTIEDIHDVEELRAVDEFRNLLVSENLLPPTLDDYHIMLRFLKARKFDIGKTKLM 131
Query: 117 WEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLM 176
W M+ WRK++GTDTI E+FEFEE +EVL+YYP GYHGVDKEGRPVYIERLG P++LM
Sbjct: 132 WSNMIKWRKDFGTDTIFEDFEFEEFDEVLKYYPHGYHGVDKEGRPVYIERLGLVDPAKLM 191
Query: 177 RITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLL 236
++TT++R+++YHV+EFE+ + K PAC IAAKR I S+TTILDVQG+G KNF++ A +L+
Sbjct: 192 QVTTVERFIRYHVREFEKTVNIKLPACCIAAKRHIDSSTTILDVQGVGFKNFSKPARDLI 251
Query: 237 AAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLE 296
+ KID + YPETLH+M+I+N GSGF K++W ++F DPKT+ KI ++ +K KLLE
Sbjct: 252 IQLQKIDNDNYPETLHRMFIINGGSGF-KLVWATVKQFLDPKTVTKIHVIGNKYQNKLLE 310
Query: 297 VIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQN 356
+ID+SQLPDFLGG+CTC GGC+++ KGPWNDP+I+K++ S L R S A N
Sbjct: 311 IIDASQLPDFLGGTCTCADRGGCMRSDKGPWNDPEILKMLQSG-GPLCRHNS-ALNSFSR 368
Query: 357 FDSFQVHSLKG-RCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYS 415
S S G + SD+STAESGS++ + +SP R +L PV E++R ++ Y
Sbjct: 369 VSSCDKPSFSGIKASDTSTAESGSEVEEMASPKVNRELRVPKLTPVCEDIRGTAIS--YP 426
Query: 416 CDDS---APTTEKVVENG-QFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVE 471
D S +P +KVV+ H K S SE T + R +
Sbjct: 427 TDSSEYDSPMVDKVVDVAWMAHEKPKASKG------------SEDTPDSGKIRTV----- 469
Query: 472 KTNHLYVTRVLASFMERLITFIRSL----RFEFWRTQNNVHPSNTTEHNINNHSAAVEAA 527
Y+ R L F L T + SL R +++++V N E S A
Sbjct: 470 ----TYIWRWLMMFFVNLFTLLISLALPQREGHSQSESSVDGPNARES--RPPSPAFATI 523
Query: 528 FERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLH 587
ER+ RL LEK E L++K + MP EKE++L ++ R+ ++E +L TK+ LH
Sbjct: 524 AERNVFSSVVNRLGDLEKQVETLHSKRHEMPREKEELLNTAVYRVDALEAELIATKKALH 583
Query: 588 SAVKQQLEIADLVE 601
A+ +Q ++ ++
Sbjct: 584 EALMRQDDLLAYID 597
>UniRef100_Q93ZU8 Putative sec14 cytosolic factor [Arabidopsis thaliana]
Length = 608
Score = 461 bits (1187), Expect = e-128
Identities = 262/613 (42%), Positives = 370/613 (59%), Gaps = 40/613 (6%)
Query: 9 EIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVR 68
E RE++SD E SEDE+++ G L+KK+ + SKF HSLK+RG R ID R +++ ED+
Sbjct: 18 EKREKKSDFEVSEDEKKTRIGGILKKKS--SKSKFRHSLKRRGSRSID-RTLSLTFEDIH 74
Query: 69 DEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYG 128
D E V E RQ LI LLPP DDY+ +LRFL AR F++ K MW M+ WR+++G
Sbjct: 75 DAEELRYVSEFRQSLISDHLLPPNLDDYHIMLRFLFARKFDLGKAKLMWTNMIQWRRDFG 134
Query: 129 TDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYH 188
TDTILE+FEF EL+EVL+YYPQGYHGVDKEGRPVYIERLGK S+LM++TT++RYL+YH
Sbjct: 135 TDTILEDFEFPELDEVLRYYPQGYHGVDKEGRPVYIERLGKVDASKLMQVTTLERYLRYH 194
Query: 189 VQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYP 248
V+EFE+ + KFPAC IAAKR I S+TTILDVQGLG+KNFT+TA +L+ + KID++ YP
Sbjct: 195 VKEFEKTITVKFPACCIAAKRHIDSSTTILDVQGLGLKNFTKTARDLIIQLQKIDSDNYP 254
Query: 249 ETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLG 308
ETLH+M+I+NAGSGF K+LW + F DPKT++KI +L +K KLLE+ID+SQLPDF G
Sbjct: 255 ETLHRMFIINAGSGF-KLLWGTVKSFLDPKTVSKIHVLGNKYQNKLLEMIDASQLPDFFG 313
Query: 309 GSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGR 368
G+CTC +GGC+++ KGPW D +I+K+ S T R S + +
Sbjct: 314 GTCTCADQGGCMRSDKGPWKDSEILKMGRSG-GTFCRHAGAFLTSDSQISSSDKPTYSLK 372
Query: 369 CSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDE----EVRVPDLNGYYSCDDSAPTTE 424
SD+STA+SGS++ + +SP ++ +L PV E + L+ Y C P +
Sbjct: 373 VSDTSTAKSGSELEEMASPKTNTNNHVPKLTPVSEYANGNISPTVLSEYEEC---VPMVD 429
Query: 425 KVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLAS 484
KVV+ + Q +M N SEG + I + ++ L +
Sbjct: 430 KVVD---------VAWQLQEMPNA-----SEGPQYTSSLGKIGSV------RHIWSWLTA 469
Query: 485 FMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNH--------SAAVEAAFERDHILPC 536
F T + SL + + +H S+ + S ER I
Sbjct: 470 FFISFFTLLASLALPQTKEHSQLHSSSVRAELCDERIARESRPPSPPRSTITERVIISSV 529
Query: 537 AQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEI 596
RL LEK E L+++ MP EKE++L ++ R+ ++E +L TK+ LH A+ +Q E+
Sbjct: 530 LSRLGDLEKQIENLHSRKSEMPHEKEELLNAAVYRVDALEAELITTKKALHEALIRQEEL 589
Query: 597 ADLVENLQKSNCR 609
++ +++ CR
Sbjct: 590 LGYIDRQKEAKCR 602
>UniRef100_Q9LRX7 Phosphatidylinositol/phosphatidylcholine transfer protein-like
[Arabidopsis thaliana]
Length = 627
Score = 455 bits (1171), Expect = e-126
Identities = 259/594 (43%), Positives = 364/594 (60%), Gaps = 58/594 (9%)
Query: 17 VENSEDERRS-SKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETV 75
+E SEDE+ + ++ +L+KKA+ AS+K THSL+KRGKR D P V IEDVRDE EE
Sbjct: 27 IEVSEDEKITRTRSRSLKKKAIKASNKLTHSLRKRGKRVADQYAPIV-IEDVRDEEEEKA 85
Query: 76 VHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEE 135
V+ R+ L+ LLPPRHDDY+T+LRFLKAR F++EKT+QMWEEML WRKE G DTI+++
Sbjct: 86 VNVFRKALVSLDLLPPRHDDYHTMLRFLKARRFDLEKTVQMWEEMLKWRKENGVDTIIQD 145
Query: 136 FEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERA 195
F ++E EEV QYYP GYHGVD+EGRPVYIERLGK P +LM++TT++R+L+YHVQ FE+
Sbjct: 146 FVYDEYEEVQQYYPHGYHGVDREGRPVYIERLGKIDPGKLMKVTTLERFLRYHVQGFEKT 205
Query: 196 LQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMY 255
EKFPACSIAAKR I S+TTI+DV G+ +F + A +L+ M KID + YPETL+QMY
Sbjct: 206 FSEKFPACSIAAKRHINSSTTIIDVHGVSWMSFRKLAQDLVMRMQKIDGDNYPETLNQMY 265
Query: 256 IVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPT 315
I+NAG+GF K++W + F DPKT +KI +L +K LLE+ID S+LP+FLGG+C C
Sbjct: 266 IINAGNGF-KLVWNTVKGFLDPKTTSKIHVLGNKYRSHLLEIIDPSELPEFLGGNCKCAH 324
Query: 316 EGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTA 375
EGGC++ +KGPWNDP+I+KLV S +A + +N + ++ SL+ +D S+
Sbjct: 325 EGGCMRFNKGPWNDPEIMKLVRSRDA---MYKPKEMGLLENGEVAKLFSLRHVNTDMSSP 381
Query: 376 ESGSDINDYSSPTRQRSS--THLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFH 433
+ G R+R S H + A + + + D ++P +
Sbjct: 382 DGGH--------VRERESHPEHDKRAQLSNQAEAVGVGRMEQSDSTSPLPN--------N 425
Query: 434 LSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFI 493
L+ ++SL T+ + V LA F+ +L+ +
Sbjct: 426 LAVERSLTTSLQK-------------------------------VASFLARFILQLLGSL 454
Query: 494 RSLRFEFWRTQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNK 553
L F N P N ++ + + + H PC RLQ LE + L +K
Sbjct: 455 -CLMFRILGRLVNKQPENQLRPELSVSVSQQQVPPPQVH--PCWLRLQNLETMVTVLCDK 511
Query: 554 PYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSN 607
P +P EKE +L DS+DRIKS+E DL+KTK+ L +Q+E+A+ ENL++S+
Sbjct: 512 PSSIPQEKEDILRDSLDRIKSIEQDLQKTKKALFLTASKQIELAECFENLKESS 565
>UniRef100_Q65X38 Hypothetical protein OJ1206_C08.12 [Oryza sativa]
Length = 613
Score = 450 bits (1157), Expect = e-125
Identities = 257/611 (42%), Positives = 368/611 (60%), Gaps = 34/611 (5%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVP 60
+EG DE +E RSD +NSE E+++ KIG+ +KKA+NA +KF HSL++R K+K + P
Sbjct: 13 FEGFTHNDEKKEIRSDADNSEGEKKT-KIGSFKKKAINAGNKFRHSLRRRSKKKNE---P 68
Query: 61 AVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEM 120
SIED+RD + V RQ L++ LLP +HDDY+T+LRFLKAR F++EK MW +M
Sbjct: 69 RGSIEDIRDVQDLQAVDAFRQCLVDEDLLPQQHDDYHTMLRFLKARKFDVEKAKSMWSDM 128
Query: 121 LLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITT 180
L WRKE+G D I EEF++ E +EV++YYPQ YHGVDKEGRP+YIE +GK ++LM++TT
Sbjct: 129 LKWRKEFGADNI-EEFDYTEADEVMKYYPQFYHGVDKEGRPIYIELIGKVDANKLMQVTT 187
Query: 181 IDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMT 240
I+RY+KYHV+EFER Q +FPACSIAAKR I S+TTILDVQG+G+KNF++ A +L+ +
Sbjct: 188 IERYVKYHVKEFERCFQMRFPACSIAAKRPIDSSTTILDVQGVGLKNFSKAARDLITRLQ 247
Query: 241 KIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDS 300
KID + YPETL +MYI+NAG GF KMLW + F DPKT +KI +L SK KLLE+ID
Sbjct: 248 KIDNDNYPETLRRMYIINAGQGF-KMLWSTVKSFLDPKTASKIHVLGSKYQNKLLEIIDE 306
Query: 301 SQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSF 360
++LP+F GG C C GGC K+ KGPW DP+IIK V + EA RQ+ S+ +
Sbjct: 307 NELPEFFGGKCKCEAFGGCKKSDKGPWKDPNIIKRVLNGEANYGRQIVTISSTDGKIIRY 366
Query: 361 QVHSLKGRCSDSSTAESGSDINDYSSPTRQRS-STHLRLAPVDEEVRVPDLNGYYSC--- 416
R +AESGS++ D +SP R+ T+ L PV EE ++ +G+ S
Sbjct: 367 AGPQYPTRKGSDGSAESGSEVEDGASPMASRNLITNPLLTPVHEESKLA-AHGFTSASPS 425
Query: 417 --DDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTN 474
++S P +KVV++G S + S + +T + +
Sbjct: 426 IIEESIPVVDKVVDDGWG--SPRASSSPSRSLPITFDGL---------------WTQVIT 468
Query: 475 HLYVTRVLASFMERLITFIRSLRFEFWRTQNN----VHPSNTTEHNINNHSAAVEAAFER 530
L V V M R + + RF T ++ +P + + E+
Sbjct: 469 WLTVLIVSLFAMVRSVPSRMAKRFSSQSTDHDHSYVEYPQEAEYKEEFRPPSPAPSYTEK 528
Query: 531 DHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAV 590
D + +RL LE+ + L KP MP EKE++L ++ R+ ++E +L TK+ L+ A+
Sbjct: 529 DVLSSMVRRLGELEEKVQALETKPSEMPFEKEELLNAAVRRVDALEAELISTKKALYEAL 588
Query: 591 KQQLEIADLVE 601
+Q E+ ++
Sbjct: 589 MRQDELLAYID 599
>UniRef100_Q67YM1 Putative phosphatidylinositol/ phosphatidylcholine transfer protein
[Arabidopsis thaliana]
Length = 572
Score = 449 bits (1154), Expect = e-124
Identities = 250/582 (42%), Positives = 352/582 (59%), Gaps = 53/582 (9%)
Query: 51 GKRKIDYRVPAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNI 110
G+RK D RV +VSIEDVRD E V RQ L+ LLP RHDDY+ +LRFLKAR F++
Sbjct: 2 GRRKSDGRVSSVSIEDVRDVEELQAVDAFRQSLLMDELLPDRHDDYHMMLRFLKARKFDV 61
Query: 111 EKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKA 170
EK QMW +M+ WRKE+GTDTI+++F+FEE+ EVL++YPQ YHGVDKEGRP+YIERLGK
Sbjct: 62 EKAKQMWADMIQWRKEFGTDTIIQDFDFEEINEVLKHYPQCYHGVDKEGRPIYIERLGKV 121
Query: 171 HPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTR 230
P+RLM++T++DRY++YHV+EFER+ KFP+C+I+AKR I S+TTILDVQG+G+KNF +
Sbjct: 122 DPNRLMQVTSMDRYVRYHVKEFERSFMIKFPSCTISAKRHIDSSTTILDVQGVGLKNFNK 181
Query: 231 TAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKS 290
+A +L+ + KID + YPETLHQM+I+NAG GF ++LW + F DPKT AKI +L K
Sbjct: 182 SARDLITRLQKIDGDNYPETLHQMFIINAGPGF-RLLWNTVKSFLDPKTSAKIHVLGYKY 240
Query: 291 LYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRA 350
L KLLEVID ++LP+FLGG+CTC +GGC+ + KGPW +P+I+K+V A RQ+ +
Sbjct: 241 LSKLLEVIDVNELPEFLGGACTCADQGGCMLSDKGPWKNPEIVKMVLHGGAHRARQVVKV 300
Query: 351 SNEQQNFDSFQVHS---LKGRCSDSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRV 407
N + ++ S +KG SD+STAESGSD D SP +S +HLRL PV EE ++
Sbjct: 301 LNSEGKVIAYAKPSYTWIKG--SDTSTAESGSDAEDIGSPKAIKSFSHLRLTPVREEAKI 358
Query: 408 PD----LNGYYSCDDSAPTTEKVVENGQFHLSQKQSLQTNDMENVTHRTISEGTSVNNLF 463
+ D+ P +K V+ T ++ R S G
Sbjct: 359 AGETSLAGSFPGYDEYVPMVDKAVD------------ATWKVKPAIQRVASRGA------ 400
Query: 464 RIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQNNVHPSNTTEHNINNHSAA 523
++ V K + RVL FM L+ F F+RT P+ TT A
Sbjct: 401 -LMSPTVPKDHEGIKARVLVMFMAFLMAV-----FTFFRTVTKKLPATTTSSPAETQGNA 454
Query: 524 VEAA-------------------FERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKM 564
+E E D + ++L LE L +KP MP EKE++
Sbjct: 455 IELGSNGEGVKEECRPPSPVPDLTETDLLNCVTKKLTELEGKIGTLQSKPNEMPYEKEEL 514
Query: 565 LMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKS 606
L ++ R+ ++E +L TK+ L+ A+ +Q E+ ++ +++
Sbjct: 515 LNAAVCRVDALEAELIATKKALYEALMRQEELLAYIDRQEEA 556
>UniRef100_Q9FWS5 F1B16.10 protein [Arabidopsis thaliana]
Length = 640
Score = 444 bits (1142), Expect = e-123
Identities = 268/633 (42%), Positives = 373/633 (58%), Gaps = 61/633 (9%)
Query: 1 YEGQCSTDEIRERRSDVENSEDERRSSKIGTL--RKKAMNASSKFTHSLKKRG--KRKID 56
+EG S DE RERRSD E SEDE+++ +IG +KKA ASSK HSLKK+G +R+
Sbjct: 13 FEGVSSNDERRERRSDFEVSEDEKKT-RIGNFNFKKKAAKASSKLRHSLKKKGSSRRRSS 71
Query: 57 YRVPAVSIEDVRDEGEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQM 116
R +++IED+ D E V E R L+ LLPP DDY+ +LRFLKAR F+I KT M
Sbjct: 72 DRTFSLTIEDIHDVEELRAVDEFRNLLVSENLLPPTLDDYHIMLRFLKARKFDIGKTKLM 131
Query: 117 WEEMLLWRKEYGTDTILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLM 176
W M+ WRK++GTDTI E+FEFEE +EVL+YYP GYHGVDKEGRPVYIERLG P++LM
Sbjct: 132 WSNMIKWRKDFGTDTIFEDFEFEEFDEVLKYYPHGYHGVDKEGRPVYIERLGLVDPAKLM 191
Query: 177 RITTIDRYLKYHVQEFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLL 236
++TT++R+++YHV+EFE+ + K PAC IAAKR I S+TTILDVQG+G KNF++ A +L+
Sbjct: 192 QVTTVERFIRYHVREFEKTVNIKLPACCIAAKRHIDSSTTILDVQGVGFKNFSKPARDLI 251
Query: 237 AAMTKIDTNYYPETLHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQ------------ 284
+ KID + YPETLH+M+I+N GSGF K++W ++F DPKT+ KI
Sbjct: 252 IQLQKIDNDNYPETLHRMFIINGGSGF-KLVWATVKQFLDPKTVTKIHVNLPFYAYPSTF 310
Query: 285 -------ILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPWNDPDIIKLVH 337
I + + + + VI QLPDFLGG+CTC GGC+++ KGPWNDP+I+K++
Sbjct: 311 PQVILKIIFNRINYCRCVVVIGHCQLPDFLGGTCTCADRGGCMRSDKGPWNDPEILKMLQ 370
Query: 338 SAEATLVRQLSRASNEQQNFDSFQVHSLKG-RCSDSSTAESGSDINDYSSPTRQRSSTHL 396
S L R S A N S S G + SD+STAESGS++ + +SP R
Sbjct: 371 SG-GPLCRHNS-ALNSFSRVSSCDKPSFSGIKASDTSTAESGSEVEEMASPKVNRELRVP 428
Query: 397 RLAPVDEEVRVPDLNGYYSCDDS---APTTEKVVENG-QFHLSQKQSLQTNDMENVTHRT 452
+L PV E++R ++ Y D S +P +KVV+ H K S
Sbjct: 429 KLTPVCEDIRGTAIS--YPTDSSEYDSPMVDKVVDVAWMAHEKPKASKG----------- 475
Query: 453 ISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSL----RFEFWRTQNNVH 508
SE T + R + Y+ R L F L T + SL R +++++V
Sbjct: 476 -SEDTPDSGKIRTV---------TYIWRWLMMFFVNLFTLLISLALPQREGHSQSESSVD 525
Query: 509 PSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYGMPLEKEKMLMDS 568
N E S A ER+ RL LEK E L++K + MP EKE++L +
Sbjct: 526 GPNARES--RPPSPAFATIAERNVFSSVVNRLGDLEKQVETLHSKRHEMPREKEELLNTA 583
Query: 569 MDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVE 601
+ R+ ++E +L TK+ LH A+ +Q ++ ++
Sbjct: 584 VYRVDALEAELIATKKALHEALMRQDDLLAYID 616
>UniRef100_Q9SJS7 Putative phosphatidylinositol/phosphatidylcholine transfer protein
[Arabidopsis thaliana]
Length = 531
Score = 417 bits (1073), Expect = e-115
Identities = 230/541 (42%), Positives = 328/541 (60%), Gaps = 51/541 (9%)
Query: 88 LLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEVLQY 147
LLP RHDDY+ +LRFLKAR F++EK QMW +M+ WRKE+GTDTI+++F+FEE+ EVL++
Sbjct: 4 LLPDRHDDYHMMLRFLKARKFDVEKAKQMWADMIQWRKEFGTDTIIQDFDFEEINEVLKH 63
Query: 148 YPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACSIAA 207
YPQ YHGVDKEGRP+YIERLGK P+RLM++T++DRY++YHV+EFER+ KFP+C+I+A
Sbjct: 64 YPQCYHGVDKEGRPIYIERLGKVDPNRLMQVTSMDRYVRYHVKEFERSFMIKFPSCTISA 123
Query: 208 KRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFKKML 267
KR I S+TTILDVQG+G+KNF ++A +L+ + KID + YPETLHQM+I+NAG GF ++L
Sbjct: 124 KRHIDSSTTILDVQGVGLKNFNKSARDLITRLQKIDGDNYPETLHQMFIINAGPGF-RLL 182
Query: 268 WPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSKGPW 327
W + F DPKT AKI +L K L KLLEVID ++LP+FLGG+CTC +GGC+ + KGPW
Sbjct: 183 WNTVKSFLDPKTSAKIHVLGYKYLSKLLEVIDVNELPEFLGGACTCADQGGCMLSDKGPW 242
Query: 328 NDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHS---LKGRCSDSSTAESGSDINDY 384
+P+I+K+V A RQ+ + N + ++ S +KG SD+STAESGSD D
Sbjct: 243 KNPEIVKMVLHGGAHRARQVVKVLNSEGKVIAYAKPSYTWIKG--SDTSTAESGSDAEDI 300
Query: 385 SSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENGQFHLSQKQSLQTND 444
SP +S +HLRL PV E + + D+ P +K V+ T
Sbjct: 301 GSPKAIKSFSHLRLTPVPGETSL--AGSFPGYDEYVPMVDKAVD------------ATWK 346
Query: 445 MENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWRTQ 504
++ R S G ++ V K + RVL FM L+ F F+RT
Sbjct: 347 VKPAIQRVASRGA-------LMSPTVPKDHEGIKARVLVMFMAFLMAV-----FTFFRTV 394
Query: 505 NNVHPSNTTEHNINNHSAAVEAA-------------------FERDHILPCAQRLQRLEK 545
P+ TT A+E E D + ++L LE
Sbjct: 395 TKKLPATTTSSPAETQGNAIELGSNGEGVKEECRPPSPVPDLTETDLLNCVTKKLTELEG 454
Query: 546 VFEELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
L +KP MP EKE++L ++ R+ ++E +L TK+ L+ A+ +Q E+ ++ ++
Sbjct: 455 KIGTLQSKPNEMPYEKEELLNAAVCRVDALEAELIATKKALYEALMRQEELLAYIDRQEE 514
Query: 606 S 606
+
Sbjct: 515 A 515
>UniRef100_Q9T027 SEC14-like protein [Arabidopsis thaliana]
Length = 550
Score = 405 bits (1042), Expect = e-111
Identities = 196/364 (53%), Positives = 267/364 (72%), Gaps = 5/364 (1%)
Query: 11 RERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDE 70
R + DVE SED++R +K+ +L+KKA+NA++KF HS+ K+G+R RV VSI D D
Sbjct: 11 RHNKIDVEISEDDKRLTKLCSLKKKAINATNKFKHSMTKKGRRHS--RVACVSIVDEIDT 68
Query: 71 GEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTD 130
E V RQ LI LLP +HDD++ +LRFL+AR F++EK QMW +ML WRKEYG D
Sbjct: 69 EELQAVDAFRQALILDELLPSKHDDHHMMLRFLRARKFDLEKAKQMWSDMLNWRKEYGAD 128
Query: 131 TILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQ 190
TI+E+F+F+E+EEV++YYPQGYHGVDKEGRP+YIERLG+ ++LM++TTIDRY+KYHV+
Sbjct: 129 TIMEDFDFKEIEEVVKYYPQGYHGVDKEGRPIYIERLGQVDATKLMKVTTIDRYVKYHVK 188
Query: 191 EFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPET 250
EFE+ KFPACSIAAKR I +TTILDVQG+G+ NF + A +LL ++ KID + YPET
Sbjct: 189 EFEKTFNVKFPACSIAAKRHIDQSTTILDVQGVGLSNFNKAAKDLLQSIQKIDNDNYPET 248
Query: 251 LHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGS 310
L++M+I+NAG GF ++LW + F DPKT AKI +L +K KLLE+ID+++LP+FLGG
Sbjct: 249 LNRMFIINAGCGF-RLLWNTVKSFLDPKTTAKIHVLGNKYQTKLLEIIDANELPEFLGGK 307
Query: 311 CTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCS 370
CTC +GGC+++ KGPWNDP+I KLV + E +R+ E+ F+ + K +C
Sbjct: 308 CTCADKGGCMRSDKGPWNDPEIFKLVQNGEGRCLRRSLSGIEEKTIFE--YNNETKKKCE 365
Query: 371 DSST 374
T
Sbjct: 366 PEET 369
>UniRef100_Q94FN3 Phosphatidylinositol transfer-like protein II [Lotus japonicus]
Length = 550
Score = 405 bits (1042), Expect = e-111
Identities = 235/593 (39%), Positives = 344/593 (57%), Gaps = 59/593 (9%)
Query: 25 RSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDEGEETVVHELRQRLI 84
+S + G+L+KKAM+ S + R R+ +V +V IEDVRD + V E RQ LI
Sbjct: 13 KSDRAGSLKKKAMSLRSSLS-----RKSRRSSSKVMSVEIEDVRDAEDLKAVDEFRQALI 67
Query: 85 ERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTDTILEEFEFEELEEV 144
LLP +HDDY+ LLRFLKAR F+IEK+ QMW +ML WRKE+G DTI+E+F+F+E++EV
Sbjct: 68 LDELLPEKHDDYHQLLRFLKARKFDIEKSKQMWSDMLQWRKEFGADTIVEDFDFKEIDEV 127
Query: 145 LQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQEFERALQEKFPACS 204
++YYP G+HGVDK+GRPVYIE +G+ ++LM++TT+DRY+KYHV+EFER KF ACS
Sbjct: 128 VKYYPHGHHGVDKDGRPVYIENIGQVDATKLMQVTTMDRYIKYHVKEFERTFDLKFAACS 187
Query: 205 IAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPETLHQMYIVNAGSGFK 264
IAAK+ I +TTILDVQG+G+KNF + A L+ + KID + YPETL++M+I+NAGSGF
Sbjct: 188 IAAKKHIDQSTTILDVQGVGLKNFNKHARELITRLQKIDGDNYPETLNRMFIINAGSGF- 246
Query: 265 KMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGSCTCPTEGGCLKTSK 324
+MLW + F DPKT +KI +L +K KLLEVID+SQLP+FLGG+CTC +GGC+++ K
Sbjct: 247 RMLWSTVKSFLDPKTTSKIHVLGNKYQSKLLEVIDASQLPEFLGGTCTCADQGGCMRSDK 306
Query: 325 GPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCSDSSTAESGSDINDY 384
GPW DP+++++V + E H +C E
Sbjct: 307 GPWKDPELVRMVQNGE----------------------HKCSRKCESPVVEEKKIS---- 340
Query: 385 SSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTT--EKVVENGQFHLSQKQSLQT 442
T+ ++ +L+ V EV + Y +A T +KV EN QF +S K ++T
Sbjct: 341 EETTKMGANFTSQLSSVFGEVPATKVCNYEDFVPAADETVWKKVEENEQFQMS-KAVVET 399
Query: 443 NDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLITFIRSLRFEFWR 502
M + +I EK N T V+A F+ ++T +R R +
Sbjct: 400 FSMVDSC------------------KIHEKVNSQIFTGVMA-FVMGIVTMVRMTRNMPKK 440
Query: 503 TQNNVHPSNTTEHNINNHSAAVEAAFERDHILPCAQRLQRLEKVFEELNNKPYG--MPLE 560
+ SN+ N S + + +R+ LE E++ N Y MP E
Sbjct: 441 LTDANFYSNSVYSGGQNPSDQTNPSISAQEFMTVMKRMAELE---EKMGNMNYNTCMPPE 497
Query: 561 KEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQKSNCRVSWS 613
KE+ML ++ R ++E +L TK+ L ++ +Q E++ +E +K +W+
Sbjct: 498 KEEMLNAAISRADALEQELMSTKKALEDSLAKQEELSAYIEKKKKKKKLFAWA 550
>UniRef100_Q93ZE9 At2g21540/F2G1.19 [Arabidopsis thaliana]
Length = 548
Score = 405 bits (1040), Expect = e-111
Identities = 234/597 (39%), Positives = 344/597 (57%), Gaps = 68/597 (11%)
Query: 11 RERRSDVENSEDERRSSKIGTLRKKAMNASSKFTHSLKKRGKRKIDYRVPAVSIEDVRDE 70
R + D + SEDE+++ K+ +L+KKA+NAS+KF HS KR +R + RV +VSI D D
Sbjct: 11 RHNKLDYDGSEDEKKT-KLCSLKKKAINASNKFKHSFTKRTRR--NSRVMSVSIVDDIDL 67
Query: 71 GEETVVHELRQRLIERGLLPPRHDDYYTLLRFLKARDFNIEKTIQMWEEMLLWRKEYGTD 130
E V RQ LI LLP +HDD++ +LRFL+AR F++EK QMW +M+ WRKE+G D
Sbjct: 68 EELQAVDAFRQALILDELLPSKHDDHHMMLRFLRARKFDLEKAKQMWTDMIHWRKEFGVD 127
Query: 131 TILEEFEFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSRLMRITTIDRYLKYHVQ 190
TI+E+F+F+E++EVL+YYPQGYHGVDK+GRPVYIERLG+ ++LM++TTIDRY+KYHV+
Sbjct: 128 TIMEDFDFKEIDEVLKYYPQGYHGVDKDGRPVYIERLGQVDATKLMQVTTIDRYVKYHVR 187
Query: 191 EFERALQEKFPACSIAAKRRIFSTTTILDVQGLGMKNFTRTAANLLAAMTKIDTNYYPET 250
EFE+ K PACSIAAK+ I +TTILDVQG+G+K+F++ A +LL + KID++ YPET
Sbjct: 188 EFEKTFNIKLPACSIAAKKHIDQSTTILDVQGVGLKSFSKAARDLLQRIQKIDSDNYPET 247
Query: 251 LHQMYIVNAGSGFKKMLWPAAQKFCDPKTIAKIQILDSKSLYKLLEVIDSSQLPDFLGGS 310
L++M+I+NAGSGF ++LW + F DPKT AKI +L +K KLLE+IDS++LP+FLGG+
Sbjct: 248 LNRMFIINAGSGF-RLLWSTVKSFLDPKTTAKIHVLGNKYQSKLLEIIDSNELPEFLGGN 306
Query: 311 CTCPTEGGCLKTSKGPWNDPDIIKLVHSAEATLVRQLSRASNEQQNFDSFQVHSLKGRCS 370
CTC +GGC+++ KGPWNDPDI K+V + E G+C
Sbjct: 307 CTCADKGGCMRSDKGPWNDPDIFKMVQNGE--------------------------GKC- 339
Query: 371 DSSTAESGSDINDYSSPTRQRSSTHLRLAPVDEEVRVPDLNGYYSCDDSAPTTEKVVENG 430
P + S+ + VDE + DS + EN
Sbjct: 340 ----------------PRKTLSNIEEKTISVDENTTMK--------SDSFAKNKFDAENT 375
Query: 431 QFHLSQKQSLQTNDMENVTHRTISEGTSVNNLFRIIKEIVEKTNHLYVTRVLASFMERLI 490
+F +++ + H+ S +L+ +K + Y+ + S + L+
Sbjct: 376 KFIPMIDKTVNASTWPTNLHK--SNYPEPEDLYSAVKPSQRRGGEGYLFGGVMSLVMGLM 433
Query: 491 TFIRSLRFEFWRTQNNVHPSNTTEHNI--NNHSAAVEAAFERDHILPCAQRLQRLEKVFE 548
T +R T+N P TE I A + +R+ LE+
Sbjct: 434 TVVR-------LTKN--MPRKLTEAAIYGGEVDKAETTMVSNQEYMSMVKRMAELEEKCR 484
Query: 549 ELNNKPYGMPLEKEKMLMDSMDRIKSVEFDLEKTKRVLHSAVKQQLEIADLVENLQK 605
L+N+P EKE++L ++ R+ +E L +TK+ L + Q I ++ +K
Sbjct: 485 SLDNQPAAFSPEKEQILTAALSRVDELELQLAQTKKTLEETMATQHVIMAYIDKKKK 541
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,004,212,459
Number of Sequences: 2790947
Number of extensions: 41993888
Number of successful extensions: 131490
Number of sequences better than 10.0: 398
Number of HSP's better than 10.0 without gapping: 151
Number of HSP's successfully gapped in prelim test: 252
Number of HSP's that attempted gapping in prelim test: 130731
Number of HSP's gapped (non-prelim): 526
length of query: 619
length of database: 848,049,833
effective HSP length: 133
effective length of query: 486
effective length of database: 476,853,882
effective search space: 231750986652
effective search space used: 231750986652
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0047c.1


