

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0046a.5
(327 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
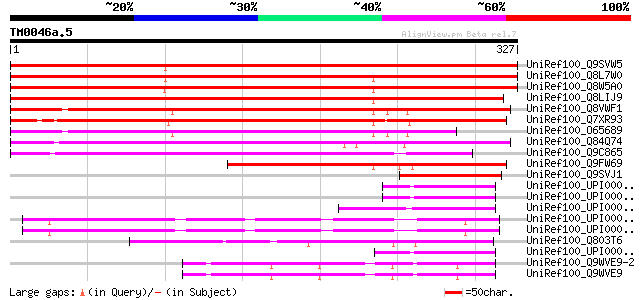
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SVW5 Hypothetical protein F15J5.30 [Arabidopsis thal... 476 e-133
UniRef100_Q8L7W0 AT4g18060/F15J5_30 [Arabidopsis thaliana] 464 e-129
UniRef100_Q8W5A0 SH3 domain-containing protein 3 [Arabidopsis th... 457 e-127
UniRef100_Q8LIJ9 Putative SH3(Src homology) domain-containing pr... 394 e-108
UniRef100_Q8VWF1 Hypothetical protein At4g34660 [Arabidopsis tha... 308 1e-82
UniRef100_Q7XR93 OSJNBa0011L07.8 protein [Oryza sativa] 294 3e-78
UniRef100_O65689 Hypothetical protein T4L20.240 [Arabidopsis tha... 253 5e-66
UniRef100_Q84Q74 Hypothetical protein OJ1364E02.9 [Oryza sativa] 252 1e-65
UniRef100_Q9C865 Hypothetical protein T8E3.10 [Arabidopsis thali... 199 9e-50
UniRef100_Q9FW69 Hypothetical protein OSJNBa0026L12.24 [Oryza sa... 189 9e-47
UniRef100_Q9SVJ1 Hypothetical protein F19H22.120 [Arabidopsis th... 77 8e-13
UniRef100_UPI00002F9C4E UPI00002F9C4E UniRef100 entry 64 7e-09
UniRef100_UPI0000363D0B UPI0000363D0B UniRef100 entry 63 1e-08
UniRef100_UPI0000438E9A UPI0000438E9A UniRef100 entry 62 2e-08
UniRef100_UPI0000438DF1 UPI0000438DF1 UniRef100 entry 62 2e-08
UniRef100_UPI0000438DF0 UPI0000438DF0 UniRef100 entry 62 2e-08
UniRef100_Q803T6 SH3-domain GRB2-like 2 [Brachydanio rerio] 62 2e-08
UniRef100_UPI00004368A9 UPI00004368A9 UniRef100 entry 62 3e-08
UniRef100_Q9WVE9-2 Splice isoform 2 of Q9WVE9 [Rattus norvegicus] 62 3e-08
UniRef100_Q9WVE9 Intersectin 1 [Rattus norvegicus] 62 3e-08
>UniRef100_Q9SVW5 Hypothetical protein F15J5.30 [Arabidopsis thaliana]
Length = 330
Score = 476 bits (1225), Expect = e-133
Identities = 239/330 (72%), Positives = 282/330 (85%), Gaps = 3/330 (0%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA R+QASKLR+QV+KQQ AVIKQFS +GYESSDV+VIDE EMQ H QL+KLY++TR+
Sbjct: 1 MDAFRRQASKLRDQVAKQQLAVIKQFSGTGYESSDVMVIDELEMQRHHQLDKLYRSTRSA 60
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAEN--NID-NILAKAASVLGDARK 117
++FQ++IVKAAE FT IG +HIE GTKLSEDCC+YG EN NID NILAKAA++ GDARK
Sbjct: 61 KEFQRDIVKAAEAFTTIGLRHIEAGTKLSEDCCRYGNENSQNIDENILAKAAAIYGDARK 120
Query: 118 HVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVR 177
HV+KE E+ N+LLASQVLDPLR M+ G PLEDARHLAQRYSRMRQEAET E+S+RQ R
Sbjct: 121 HVDKEQEDFNKLLASQVLDPLRAMVAGSPLEDARHLAQRYSRMRQEAETHATEVSRRQAR 180
Query: 178 VRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSD 237
VRE+P E VAKL AEA+M+ELKANMAVLGKEA AALA+V++QQ RLTFQRLVAMMV++
Sbjct: 181 VREAPIPENVAKLQLAEAKMQELKANMAVLGKEATAALAAVESQQHRLTFQRLVAMMVTE 240
Query: 238 RQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEG 297
+Q KESAPP + +ENGS KT YFLAE HPF SEKEL KGD+IVVRKV+ TGW+EG
Sbjct: 241 KQHKESAPPAIPTENGSEKTSYFLAEVIHPFSAASEKELDLDKGDYIVVRKVSQTGWAEG 300
Query: 298 ECNGKAGWFPSAYVEKRQRVPSSNFSSEVY 327
EC GKAGWFP AY+EKRQR+P++NF++EVY
Sbjct: 301 ECKGKAGWFPMAYIEKRQRLPTTNFAAEVY 330
>UniRef100_Q8L7W0 AT4g18060/F15J5_30 [Arabidopsis thaliana]
Length = 351
Score = 464 bits (1193), Expect = e-129
Identities = 239/351 (68%), Positives = 282/351 (80%), Gaps = 24/351 (6%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA R+QASKLR+QV+KQQ AVIKQFS +GYESSDV+VIDE EMQ H QL+KLY++TR+
Sbjct: 1 MDAFRRQASKLRDQVAKQQLAVIKQFSGTGYESSDVMVIDELEMQRHHQLDKLYRSTRSA 60
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAEN--NID-NILAKAASVLGDARK 117
++FQ++IVKAAE FT IG +HIE GTKLSEDCC+YG EN NID NILAKAA++ GDARK
Sbjct: 61 KEFQRDIVKAAEAFTTIGLRHIEAGTKLSEDCCRYGNENSQNIDENILAKAAAIYGDARK 120
Query: 118 HVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVR 177
HV+KE E+ N+LLASQVLDPLR M+ G PLEDARHLAQRYSRMRQEAET E+S+RQ R
Sbjct: 121 HVDKEQEDFNKLLASQVLDPLRAMVAGSPLEDARHLAQRYSRMRQEAETHATEVSRRQAR 180
Query: 178 VRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
VRE+P E VAKL AEA+M+ELKANMAVLGKEA AALA+V++QQ RLTFQRLVAM
Sbjct: 181 VREAPIPENVAKLQLAEAKMQELKANMAVLGKEATAALAAVESQQHRLTFQRLVAMVEGE 240
Query: 234 -----------------MVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKEL 276
MV+++Q KESAPP + +ENGS KT YFLAE HPF SEKEL
Sbjct: 241 KNYHLRIAAILSDIEAEMVTEKQHKESAPPAIPTENGSEKTSYFLAEVIHPFSAASEKEL 300
Query: 277 SFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSSNFSSEVY 327
KGD+IVVRKV+ TGW+EGEC GKAGWFP AY+EKRQR+P++NF++EVY
Sbjct: 301 DLDKGDYIVVRKVSQTGWAEGECKGKAGWFPMAYIEKRQRLPTTNFAAEVY 351
>UniRef100_Q8W5A0 SH3 domain-containing protein 3 [Arabidopsis thaliana]
Length = 351
Score = 457 bits (1176), Expect = e-127
Identities = 237/351 (67%), Positives = 280/351 (79%), Gaps = 24/351 (6%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA R+QASKLR+QV+KQQ AVIKQFS +GYESSDV+VIDE EMQ H QL+KLY++TR+
Sbjct: 1 MDAFRRQASKLRDQVAKQQLAVIKQFSGTGYESSDVMVIDELEMQRHHQLDKLYRSTRSA 60
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAE--NNID-NILAKAASVLGDARK 117
++FQ++IVKAAE FT IG +HIE GTKLSEDCC+YG E NID NILAKAA++ GDARK
Sbjct: 61 KEFQRDIVKAAEAFTTIGLRHIEAGTKLSEDCCRYGNEIVRNIDENILAKAAAIYGDARK 120
Query: 118 HVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVR 177
HV+KE E+ N+LLASQVLDPLR M+ G PLEDARHLAQRYSRMRQEAET E+S+RQ R
Sbjct: 121 HVDKEQEDFNKLLASQVLDPLRAMVAGSPLEDARHLAQRYSRMRQEAETHATEVSRRQAR 180
Query: 178 VRESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
VRE+P E VAKL AEA+M+ELKANMAVLGKEA AALA+V++QQ RLTFQRLVAM
Sbjct: 181 VREAPIPENVAKLQLAEAKMQELKANMAVLGKEATAALAAVESQQHRLTFQRLVAMVEGE 240
Query: 234 -----------------MVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKEL 276
MV+++Q KESA P + +ENGS KT YFLAE HPF SEKEL
Sbjct: 241 KNYHLRIAAILSDIEAEMVTEKQHKESALPAIPTENGSEKTSYFLAEVIHPFSAVSEKEL 300
Query: 277 SFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSSNFSSEVY 327
KGD+IVVRKV+ TGW+EGEC GKAGWFP AY+EKRQR+P++NF++EVY
Sbjct: 301 DLDKGDYIVVRKVSQTGWAEGECKGKAGWFPMAYIEKRQRLPTTNFAAEVY 351
>UniRef100_Q8LIJ9 Putative SH3(Src homology) domain-containing protein [Oryza sativa]
Length = 348
Score = 394 bits (1013), Expect = e-108
Identities = 204/340 (60%), Positives = 253/340 (74%), Gaps = 22/340 (6%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MD LRKQASK +EQV+KQQQAVIKQFST+GYE SD VVIDE E+Q HQQLEKLY +TR+
Sbjct: 1 MDVLRKQASKFKEQVAKQQQAVIKQFSTTGYEHSDAVVIDEVELQRHQQLEKLYTSTRSG 60
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNI-DNILAKAASVLGDARKHV 119
RDFQK+IV+AAE +IG +H+E GTK SEDC +YG E++ D LAKAAS+ G A ++V
Sbjct: 61 RDFQKDIVRAAEGLVSIGIRHVEVGTKFSEDCYRYGGESSASDEALAKAASLYGGALRNV 120
Query: 120 EKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVR 179
EKE+EE NR+L+SQ +DPLR M G PLEDAR LAQRYSRMR EAE L EI++R+ RVR
Sbjct: 121 EKEYEEFNRILSSQTIDPLRAMAAGAPLEDARGLAQRYSRMRHEAEILSAEIARRKQRVR 180
Query: 180 ESPTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM------ 233
E+P +E KL +E++M E KA+MAVLGKEAAAALA+V++QQQR+T QRLV M
Sbjct: 181 EAPLAEHTTKLQQSESKMIEHKASMAVLGKEAAAALAAVESQQQRITLQRLVGMVEAEKL 240
Query: 234 ---------------MVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSF 278
M S++QK+ESAPP + S + K YFLAEA H F G +EKELS
Sbjct: 241 FHLRLAAILDDVEAEMSSEKQKRESAPPTIHSHKRAEKAQYFLAEAVHNFNGTTEKELSL 300
Query: 279 SKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVP 318
GD++VVR++ P GW+EGEC G AGWFP+AYVE+R+ +P
Sbjct: 301 IVGDYVVVRQIAPNGWAEGECKGVAGWFPAAYVERRENIP 340
>UniRef100_Q8VWF1 Hypothetical protein At4g34660 [Arabidopsis thaliana]
Length = 368
Score = 308 bits (790), Expect = 1e-82
Identities = 174/368 (47%), Positives = 232/368 (62%), Gaps = 48/368 (13%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA+RKQAS+LREQV++QQQAV KQF GY S + DE E+ HQ+LEKLY +TR
Sbjct: 1 MDAIRKQASRLREQVARQQQAVFKQFGGGGYGSG---LADEAELNQHQKLEKLYISTRAA 57
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDN--ILAKAASVLGDARKH 118
+ +Q++IV+ E + G K +E GTKLSED KYG+EN N +L +AA G AR
Sbjct: 58 KHYQRDIVRGVEGYIVTGSKQVEIGTKLSEDSRKYGSENTCTNGNVLTRAALNYGRARAQ 117
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE + + L +QV +PLR M+ G PLEDARHLAQRY RMRQEAE E+++RQ +
Sbjct: 118 MEKERGNMLKALGTQVAEPLRAMVLGAPLEDARHLAQRYDRMRQEAEAQATEVARRQAKA 177
Query: 179 RESPTSEQVA-KLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
RES + + KL +AEA++ +LK+NM +LGKEAA+ALASV+ QQQ+LT +RL++M
Sbjct: 178 RESQGNPDILMKLESAEAKLHDLKSNMTILGKEAASALASVEDQQQKLTLERLLSMVESE 237
Query: 234 -----------------MVSDRQKKE---------SAPPVVVSENGSG------------ 255
MVS+RQ+ E S PP E +G
Sbjct: 238 RAYHQRVLQILDQLEGEMVSERQRIEAPSTPSSADSMPPPPSYEEANGVFASQMHDTSTD 297
Query: 256 KTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQ 315
YFL E P++G ++ ELS S G+++VVRKVT +GW+EGEC GKAGWFP Y+E+R+
Sbjct: 298 SMGYFLGEVLFPYHGVTDVELSLSTGEYVVVRKVTGSGWAEGECKGKAGWFPYGYIERRE 357
Query: 316 RVPSSNFS 323
RV +S S
Sbjct: 358 RVLASKVS 365
>UniRef100_Q7XR93 OSJNBa0011L07.8 protein [Oryza sativa]
Length = 369
Score = 294 bits (752), Expect = 3e-78
Identities = 171/367 (46%), Positives = 224/367 (60%), Gaps = 51/367 (13%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
M+AL KQAS+L+EQVS+Q V K F + Y +S+ DE E+ +HQ+LEKLY +TR
Sbjct: 1 MEALWKQASRLKEQVSRQ--GVFKPFGAA-YGNSENAFTDESEVNLHQRLEKLYLSTRAA 57
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNI--DNILAKAASVLGDARKH 118
+ FQ++IV+ E + G K ++ G KLS+D KYG N DN L+KAA G AR
Sbjct: 58 KHFQRDIVRGVEGYIVTGSKQVDIGNKLSDDSQKYGTGNTCTSDNTLSKAAMYYGKARSL 117
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE + R +QV +PLR M+ G PLEDARHLAQRY RMRQEAE E+S+RQ RV
Sbjct: 118 MEKERGNMLRAFGTQVAEPLRAMVMGAPLEDARHLAQRYDRMRQEAEAQAVEVSRRQNRV 177
Query: 179 RES-PTSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
RES P + + KL AAE +++ELK++M LGKEA AA+A+V+AQQQRLT QRL+AM
Sbjct: 178 RESAPNGDVITKLEAAEYKLEELKSSMVGLGKEAVAAMAAVEAQQQRLTLQRLIAMVEAE 237
Query: 234 -----------------MVSDRQKKESAPPVVVSENGSGK-------------------- 256
MVS+RQK E APP +EN +
Sbjct: 238 RAYHQRVLEILDHLEQEMVSERQKIE-APPTPSAENYMAQPPPSYDEVNGMFASSSVDDS 296
Query: 257 ---TMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEK 313
+FL EA F +SE EL+ S GD ++VRK++ GW+EGEC GKAGWFP Y+E+
Sbjct: 297 VTSVDFFLGEALDSFKAESESELNLSAGDIVIVRKISTNGWAEGECKGKAGWFPHGYIER 356
Query: 314 RQRVPSS 320
R+RV +S
Sbjct: 357 RERVLAS 363
>UniRef100_O65689 Hypothetical protein T4L20.240 [Arabidopsis thaliana]
Length = 397
Score = 253 bits (646), Expect = 5e-66
Identities = 152/345 (44%), Positives = 204/345 (59%), Gaps = 60/345 (17%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
MDA+RKQAS+LREQV++QQQAV KQF GY S + DE E+ HQ+LEKLY +TR
Sbjct: 1 MDAIRKQASRLREQVARQQQAVFKQFGGGGYGSG---LADEAELNQHQKLEKLYISTRAA 57
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDN--ILAKAASVLGDARKH 118
+ +Q++IV+ E + G K +E GTKLSED KYG+EN N +L +AA G AR
Sbjct: 58 KHYQRDIVRGVEGYIVTGSKQVEIGTKLSEDSRKYGSENTCTNGNVLTRAALNYGRARAQ 117
Query: 119 VEKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRV 178
+EKE + + L +QV +PLR M+ G PLEDARHLAQRY RMRQEAE E+++RQ +
Sbjct: 118 MEKERGNMLKALGTQVAEPLRAMVLGAPLEDARHLAQRYDRMRQEAEAQATEVARRQAKA 177
Query: 179 RESPTSEQV-AKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---- 233
RES + + KL +AEA++ +LK+NM +LGKEAA+ALASV+ QQQ+LT +RL++M
Sbjct: 178 RESQGNPDILMKLESAEAKLHDLKSNMTILGKEAASALASVEDQQQKLTLERLLSMVVRL 237
Query: 234 -----------------------------MVSDRQKKE---------SAPPVVVSENGSG 255
MVS+RQ+ E S PP E +G
Sbjct: 238 VTRSISRLNLNAPTIKESSKYLISSKERLMVSERQRIEAPSTPSSADSMPPPPSYEEANG 297
Query: 256 ------------KTMYFLAEATHPFYGDSEKELSFSKGDFIVVRK 288
YFL E P++G ++ ELS S G+++VVRK
Sbjct: 298 VFASQMHDTSTDSMGYFLGEVLFPYHGVTDVELSLSTGEYVVVRK 342
>UniRef100_Q84Q74 Hypothetical protein OJ1364E02.9 [Oryza sativa]
Length = 441
Score = 252 bits (643), Expect = 1e-65
Identities = 147/389 (37%), Positives = 215/389 (54%), Gaps = 68/389 (17%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
M+ LRKQASKLRE V+KQQQAV KQFS + D ++DE E++ HQ L++LY +TR
Sbjct: 1 METLRKQASKLREHVAKQQQAVRKQFSAR--YNQDPSLVDEAELECHQNLQRLYNSTRAA 58
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAEN-NIDNILAKAASVLGDARKHV 119
+ FQ+ IV+ E F A+ K +E KL+EDCC+YG +N N ILA+A+ G++ +
Sbjct: 59 KHFQRSIVRGVEGFIAVSTKQMEIVKKLAEDCCRYGNDNQNFGFILARASVEFGNSHSQM 118
Query: 120 EKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVR 179
EKE E L + L QV +PLR+MI PLEDAR L RY R+RQ+ E+ ++ ++Q++ +
Sbjct: 119 EKERENLLKFLGEQVFEPLREMIMSAPLEDARLLTYRYQRIRQDMESQIADVMRKQLKSK 178
Query: 180 ESP-TSEQVAKLHAAEARMKELKANMAVLGKEAAAA----------------LASVDAQQ 222
ES ++ KL AE+++ EL+ ++ LG+EA AA LA VDA++
Sbjct: 179 ESSGNADNSVKLQHAESKLSELRTTLSALGREATAAMEAVEVQQQQVTFDRLLAMVDAER 238
Query: 223 ---------------------------------------------QRLTFQRLVAMMVSD 237
R T + + S+
Sbjct: 239 AYHQNAADILNKLHDEMVQAKHHDEPENHYDETSSDPKTAATHEHSRSTSEDHIFTNTSE 298
Query: 238 RQKKESAPPVVVSENG---SGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGW 294
+ E++ P + +G+ ++++ E HPF ++ ELS S GD++VVR+V P GW
Sbjct: 299 PTRTETSEPTRTETSEPTRNGQEVHYVGEVIHPFDAQADGELSISVGDYVVVRQVAPNGW 358
Query: 295 SEGECNGKAGWFPSAYVEKRQRVPSSNFS 323
SEGEC GKAGWFPSAYVE+R + P+S S
Sbjct: 359 SEGECKGKAGWFPSAYVEQRDKAPASKQS 387
>UniRef100_Q9C865 Hypothetical protein T8E3.10 [Arabidopsis thaliana]
Length = 439
Score = 199 bits (506), Expect = 9e-50
Identities = 121/300 (40%), Positives = 175/300 (58%), Gaps = 12/300 (4%)
Query: 1 MDALRKQASKLREQVSKQQQAVIKQFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTM 60
M+A+RKQA+KLREQV++QQQAV+K G+ ++D VV+DE E+ HQ+L+ LY +T+
Sbjct: 1 MEAIRKQAAKLREQVARQQQAVLKHL---GHVNADAVVVDEEELHCHQKLQDLYSSTKAA 57
Query: 61 RDFQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNI-LAKAASVLGDARKHV 119
+ Q+ IV+ E F A G K +E G K +ED KYG EN N L++ A G + K V
Sbjct: 58 KRLQRNIVRGLEGFIATGTKVVEIGLKFAEDFKKYGDENPDANTPLSRVAHHFGTSYKSV 117
Query: 120 EKEHEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVR 179
E E E L +L+ QV +P+R MI PLEDARHL Y R+RQE E ++ +R+ +++
Sbjct: 118 EGERETLLGVLSEQVCEPIRTMIYSAPLEDARHLVNHYDRLRQEVEAQATDVLRRRSKLK 177
Query: 180 ESPTSEQV-AKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDR 238
ES SE+ KL +E+R+ ELK++M LGKEA A+ VD QQQ +T QRL A++ ++R
Sbjct: 178 ESDISEEAYIKLKNSESRLAELKSSMKTLGKEATKAMLEVDDQQQNVTSQRLRALVEAER 237
Query: 239 QKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGE 298
+A ++ ++ A G S K L D + + P S GE
Sbjct: 238 SYHRNALDIL-------DKLHSEMIAEEEAIGSSPKSLPLHIEDSASLPQQEPNSNSSGE 290
>UniRef100_Q9FW69 Hypothetical protein OSJNBa0026L12.24 [Oryza sativa]
Length = 230
Score = 189 bits (480), Expect = 9e-47
Identities = 108/224 (48%), Positives = 140/224 (62%), Gaps = 44/224 (19%)
Query: 141 MINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRE-SPTSEQVAKLHAAEARMKE 199
M+ G PLEDARHLAQRY RMRQEAE E+SKRQ+++RE S + +++L AAE++++E
Sbjct: 1 MVMGAPLEDARHLAQRYDRMRQEAEAQAIEVSKRQMKLRETSGNGDMISRLEAAESKLQE 60
Query: 200 LKANMAVLGKEAAAALASVDAQQQRLTFQRLVAM---------------------MVSDR 238
LK+NM VLGKEA A++ +V+AQQQRLT QRL+AM MVS+R
Sbjct: 61 LKSNMGVLGKEAVASMTAVEAQQQRLTLQRLIAMVESERSYHQRVLQILDQLEREMVSER 120
Query: 239 QKKESAPPVVVS---------ENGSGKTM-------------YFLAEATHPFYGDSEKEL 276
Q+ E APP V E +G M +FLAEA + +SE EL
Sbjct: 121 QRIEGAPPPAVESSMPPPPSYEEINGVFMRNPTVAELVETVEFFLAEAIQSYRAESETEL 180
Query: 277 SFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSS 320
+ + GD+IVVRKV+ GW+EGEC GKAGWFP Y+EKR RV +S
Sbjct: 181 NLAAGDYIVVRKVSNNGWAEGECRGKAGWFPYDYIEKRDRVLAS 224
>UniRef100_Q9SVJ1 Hypothetical protein F19H22.120 [Arabidopsis thaliana]
Length = 169
Score = 76.6 bits (187), Expect = 8e-13
Identities = 34/67 (50%), Positives = 45/67 (66%), Gaps = 1/67 (1%)
Query: 252 NGSGKTM-YFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAY 310
NG+ M YFL E P+ DS+ ELS S GD++V+R+V + W+EGEC G AGWF Y
Sbjct: 94 NGTSDAMGYFLGEVMFPYQADSDFELSLSVGDYVVIREVVSSVWAEGECKGNAGWFTYIY 153
Query: 311 VEKRQRV 317
+E+R RV
Sbjct: 154 IERRDRV 160
>UniRef100_UPI00002F9C4E UPI00002F9C4E UniRef100 entry
Length = 736
Score = 63.5 bits (153), Expect = 7e-09
Identities = 33/73 (45%), Positives = 43/73 (58%), Gaps = 2/73 (2%)
Query: 241 KESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECN 300
+ + V +SENG G + +A F ++E ELSFSKGD IVV + GW EG N
Sbjct: 146 RRQSKSVEMSENGGGGQQ--MVKARFNFKQNNEDELSFSKGDVIVVTRQEEGGWWEGTLN 203
Query: 301 GKAGWFPSAYVEK 313
G+ GWFPS YV +
Sbjct: 204 GRTGWFPSNYVRE 216
>UniRef100_UPI0000363D0B UPI0000363D0B UniRef100 entry
Length = 744
Score = 63.2 bits (152), Expect = 1e-08
Identities = 33/73 (45%), Positives = 43/73 (58%), Gaps = 2/73 (2%)
Query: 241 KESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECN 300
+ + V ++ENG G + +A F ++E ELSFSKGD IVV + GW EG N
Sbjct: 146 RRQSKSVEMAENGGGGQL--TVKARFNFKQNNEDELSFSKGDVIVVTRQEDGGWWEGTLN 203
Query: 301 GKAGWFPSAYVEK 313
GK GWFPS YV +
Sbjct: 204 GKTGWFPSNYVRE 216
>UniRef100_UPI0000438E9A UPI0000438E9A UniRef100 entry
Length = 733
Score = 62.4 bits (150), Expect = 2e-08
Identities = 37/101 (36%), Positives = 55/101 (53%), Gaps = 3/101 (2%)
Query: 213 AALASVDAQQQRLTFQRLVAMMVSDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDS 272
AALA+ + QQ+R +R A + + P +++N + + + +A F +
Sbjct: 98 AALANENNQQKRSAGERGDAALHDVSEIMPCTSPQDMTDNSNAQ---LVVKARFNFQQTN 154
Query: 273 EKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEK 313
E EL+FSKGD I V + GW EG NGK GWFPS YV++
Sbjct: 155 EDELTFSKGDLISVTRTEEGGWWEGILNGKTGWFPSNYVKE 195
>UniRef100_UPI0000438DF1 UPI0000438DF1 UniRef100 entry
Length = 1668
Score = 62.0 bits (149), Expect = 2e-08
Identities = 71/317 (22%), Positives = 129/317 (40%), Gaps = 42/317 (13%)
Query: 9 SKLREQVSKQQQAVIK------QFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRD 62
+KLRE+ +++QA ++ + E + +E E + Q+ +L + R RD
Sbjct: 491 AKLREEEERERQARLQAAKEQAERERKAREEEERRRREEEEERREQERRRLEEEDRRRRD 550
Query: 63 FQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKE 122
++ ++ + +E K E+ + A+ + R+ E+E
Sbjct: 551 EERRKLEEERRKLEEERRKLEDERKRQEEIRLKEEREREERRRAE------EERRRKEEE 604
Query: 123 HEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESP 182
E R + + R + DA R R E E R+E +R+ +
Sbjct: 605 RREEERKREREEEERRRHRAQEAAMRDAEE------RKRIEEERKRKEEERRKQEEEDRR 658
Query: 183 TSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQR-LVAMMVSDRQKK 241
E+ + E + K+ + +EAAAA D Q ++ T + LV+ ++ +++
Sbjct: 659 RREEEQRKQDEERKRKQQQ-------EEAAAARLRDDEQNKKNTVKSDLVSALLRGLEER 711
Query: 242 ESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPT--GWSEGEC 299
++A +A +PF +E+ELSF D I V + GW G
Sbjct: 712 KAALTTF--------------KALYPFTARNEEELSFESDDLIEVDESVEREQGWLYGSW 757
Query: 300 NGKAGWFPSAYVEKRQR 316
GK GWFP +YVEK+ +
Sbjct: 758 QGKMGWFPESYVEKQTK 774
Score = 36.6 bits (83), Expect = 0.96
Identities = 19/49 (38%), Positives = 26/49 (52%)
Query: 264 ATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVE 312
A + + +E ELSFSK I V + W +GE NG G P+ YV+
Sbjct: 1111 AIYDYTAANEDELSFSKSQVINVLDKSNPDWWKGELNGVTGLIPTNYVK 1159
Score = 36.2 bits (82), Expect = 1.3
Identities = 19/50 (38%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Query: 262 AEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYV 311
A+A + ++ L FSK D I V + W GE +G GWFP YV
Sbjct: 877 AQALCSWSAKTDNHLDFSKDDVITVLEQQENWWL-GELDGVQGWFPKTYV 925
>UniRef100_UPI0000438DF0 UPI0000438DF0 UniRef100 entry
Length = 1670
Score = 62.0 bits (149), Expect = 2e-08
Identities = 71/317 (22%), Positives = 129/317 (40%), Gaps = 42/317 (13%)
Query: 9 SKLREQVSKQQQAVIK------QFSTSGYESSDVVVIDEGEMQIHQQLEKLYKATRTMRD 62
+KLRE+ +++QA ++ + E + +E E + Q+ +L + R RD
Sbjct: 506 AKLREEEERERQARLQAAKEQAERERKAREEEERRRREEEEERREQERRRLEEEDRRRRD 565
Query: 63 FQKEIVKAAETFTAIGYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKE 122
++ ++ + +E K E+ + A+ + R+ E+E
Sbjct: 566 EERRKLEEERRKLEEERRKLEDERKRQEEIRLKEEREREERRRAE------EERRRKEEE 619
Query: 123 HEELNRLLASQVLDPLRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESP 182
E R + + R + DA R R E E R+E +R+ +
Sbjct: 620 RREEERKREREEEERRRHRAQEAAMRDAEE------RKRIEEERKRKEEERRKQEEEDRR 673
Query: 183 TSEQVAKLHAAEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQR-LVAMMVSDRQKK 241
E+ + E + K+ + +EAAAA D Q ++ T + LV+ ++ +++
Sbjct: 674 RREEEQRKQDEERKRKQQQ-------EEAAAARLRDDEQNKKNTVKSDLVSALLRGLEER 726
Query: 242 ESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPT--GWSEGEC 299
++A +A +PF +E+ELSF D I V + GW G
Sbjct: 727 KAALTTF--------------KALYPFTARNEEELSFESDDLIEVDESVEREQGWLYGSW 772
Query: 300 NGKAGWFPSAYVEKRQR 316
GK GWFP +YVEK+ +
Sbjct: 773 QGKMGWFPESYVEKQTK 789
Score = 36.6 bits (83), Expect = 0.96
Identities = 19/49 (38%), Positives = 26/49 (52%)
Query: 264 ATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVE 312
A + + +E ELSFSK I V + W +GE NG G P+ YV+
Sbjct: 1113 AIYDYTAANEDELSFSKSQVINVLDKSNPDWWKGELNGVTGLIPTNYVK 1161
Score = 36.2 bits (82), Expect = 1.3
Identities = 19/50 (38%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Query: 262 AEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYV 311
A+A + ++ L FSK D I V + W GE +G GWFP YV
Sbjct: 892 AQALCSWSAKTDNHLDFSKDDVITVLEQQENWWL-GELDGVQGWFPKTYV 940
Score = 33.9 bits (76), Expect = 6.2
Identities = 33/117 (28%), Positives = 51/117 (43%), Gaps = 8/117 (6%)
Query: 204 MAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDRQKKESAPPVVVSENGSGKTMYFLAE 263
+ VL ++ L +D Q + + SD Q E P V + S + E
Sbjct: 914 ITVLEQQENWWLGELDGVQGWFPKTYVTVLSTSDAQA-EFPKPYVTFISSSEYVALYTYE 972
Query: 264 ATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVEKRQRVPSS 320
+ P GD L+F++GD I+V + W G +AG FPS YV+ ++ SS
Sbjct: 973 SPEP--GD----LTFNEGDTILVSEREGEWW-RGSIGDRAGLFPSNYVKPKETDTSS 1022
>UniRef100_Q803T6 SH3-domain GRB2-like 2 [Brachydanio rerio]
Length = 347
Score = 62.0 bits (149), Expect = 2e-08
Identities = 59/263 (22%), Positives = 112/263 (42%), Gaps = 33/263 (12%)
Query: 78 GYKHIETGTKLSEDCCKYGAENNIDNILAKAASVLGDARKHVEKEHEELNRLLASQVLDP 137
G + +T T L E K G E ++ A +G+A + + + + L+ + +DP
Sbjct: 83 GPGYTQTETTLGEAMQKAGRELGEESCFGVALIDVGEAMRELGEVKDALDMEVKQNFIDP 142
Query: 138 LRQMINGVPLEDARHLAQRYSRMRQEAETLREEISKRQVRVRESPTSEQVAKLH-----A 192
L Q I+ L++ +H ++ R + + + KRQ +V E + + K A
Sbjct: 143 L-QNIHDKDLKEIQHHLKKLEGRRLDFDYKK----KRQGKVTEDEIKQALEKFDDSKEIA 197
Query: 193 AEARMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDRQKKESAPP------ 246
++ L+ ++ + + +A A V+ +Q + ++ + DR ++ S P
Sbjct: 198 EQSMFNLLENDIEQVSQLSALVQAQVNYHRQAAEILQQLSSKIEDRIREASCKPKREYVP 257
Query: 247 -----VVVSENGSGKTMYF------------LAEATHPFYGDSEKELSFSKGDFIVVRKV 289
+ SEN +G + A + F ++E EL F +GD I +
Sbjct: 258 KPRTSLDFSENHNGSIGHSGPSRSPAPMDQPCCRALYDFDPENEGELGFKEGDIITLTSK 317
Query: 290 TPTGWSEGECNGKAGWFPSAYVE 312
W EG NG++G+FP YV+
Sbjct: 318 IDDNWYEGMVNGQSGFFPVNYVD 340
>UniRef100_UPI00004368A9 UPI00004368A9 UniRef100 entry
Length = 739
Score = 61.6 bits (148), Expect = 3e-08
Identities = 33/78 (42%), Positives = 44/78 (56%), Gaps = 2/78 (2%)
Query: 236 SDRQKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWS 295
S R + + V +SENG G + + +A F ++E ELSFSKG+ I V + GW
Sbjct: 86 SSRSLRRQSKAVEMSENGGGGQL--VVKARFNFKQNNEDELSFSKGELIHVTRQEEGGWW 143
Query: 296 EGECNGKAGWFPSAYVEK 313
EG N K GWFPS YV +
Sbjct: 144 EGTLNSKTGWFPSNYVRE 161
>UniRef100_Q9WVE9-2 Splice isoform 2 of Q9WVE9 [Rattus norvegicus]
Length = 1146
Score = 61.6 bits (148), Expect = 3e-08
Identities = 60/226 (26%), Positives = 99/226 (43%), Gaps = 34/226 (15%)
Query: 112 LGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHL-AQRYSRMRQEAETL--- 167
L + K + +E++ + + L LR++ + L+ R + A+R + QE ++L
Sbjct: 585 LDEVEKETRSKLQEID--VFNNQLKELREIHSKQQLQKQRSIEAERLKQKEQERKSLELE 642
Query: 168 -REEISKRQVRVRESPTSEQVAKLHAAEARMK----ELKANMAVLGKEAAA-ALASVDAQ 221
++E +R+V+ R+ E V + R +LK +V KEA A V +
Sbjct: 643 KQKEEGQRRVQERDKQWQEHVQQEEQQRPRKPHEEDKLKREDSVKKKEAEERAKPEVQDK 702
Query: 222 QQRLTFQRLVAMMVSDRQKKESAP-------PVVVSENGSGKTMYFLAEATHPFYGDSEK 274
Q RL + K AP P+ +S S K +Y+ A +PF S
Sbjct: 703 QSRLFHPH------QEPAKPAQAPWPTTEKGPLTISAQESAKVVYY--RALYPFESRSHD 754
Query: 275 ELSFSKGDFIVVR-------KVTPTGWSEGECNGKAGWFPSAYVEK 313
E++ GD ++V+ + GW GE GK GWFP+ Y EK
Sbjct: 755 EITIQPGDIVMVKGEWVDESQTGEPGWLGGEPKGKTGWFPANYAEK 800
Score = 41.6 bits (96), Expect = 0.030
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Query: 262 AEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVE 312
A+A +P+ + L+F+K D I V + W GE G+ GWFP +YV+
Sbjct: 915 AQALYPWRAKKDNHLNFNKSDVITVLEQQDMWWF-GEVQGQKGWFPKSYVK 964
Score = 40.8 bits (94), Expect = 0.051
Identities = 18/57 (31%), Positives = 30/57 (52%), Gaps = 5/57 (8%)
Query: 261 LAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKA-----GWFPSAYVE 312
+A+ + ++L+ + G I++RK P GW EGE + GWFP+ YV+
Sbjct: 1004 IAQVIASYTATGPEQLTLAPGQLILIRKKNPGGWWEGELQARGKKRQIGWFPANYVK 1060
Score = 39.3 bits (90), Expect = 0.15
Identities = 18/47 (38%), Positives = 27/47 (57%)
Query: 266 HPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVE 312
+ + ++ EL+FSKG I V W +GE +G+ G FPS YV+
Sbjct: 1090 YDYTAQNDDELAFSKGQIINVLSKEDPDWWKGEVSGQVGLFPSNYVK 1136
>UniRef100_Q9WVE9 Intersectin 1 [Rattus norvegicus]
Length = 1217
Score = 61.6 bits (148), Expect = 3e-08
Identities = 60/226 (26%), Positives = 99/226 (43%), Gaps = 34/226 (15%)
Query: 112 LGDARKHVEKEHEELNRLLASQVLDPLRQMINGVPLEDARHL-AQRYSRMRQEAETL--- 167
L + K + +E++ + + L LR++ + L+ R + A+R + QE ++L
Sbjct: 585 LDEVEKETRSKLQEID--VFNNQLKELREIHSKQQLQKQRSIEAERLKQKEQERKSLELE 642
Query: 168 -REEISKRQVRVRESPTSEQVAKLHAAEARMK----ELKANMAVLGKEAAA-ALASVDAQ 221
++E +R+V+ R+ E V + R +LK +V KEA A V +
Sbjct: 643 KQKEEGQRRVQERDKQWQEHVQQEEQQRPRKPHEEDKLKREDSVKKKEAEERAKPEVQDK 702
Query: 222 QQRLTFQRLVAMMVSDRQKKESAP-------PVVVSENGSGKTMYFLAEATHPFYGDSEK 274
Q RL + K AP P+ +S S K +Y+ A +PF S
Sbjct: 703 QSRLFHPH------QEPAKPAQAPWPTTEKGPLTISAQESAKVVYY--RALYPFESRSHD 754
Query: 275 ELSFSKGDFIVVR-------KVTPTGWSEGECNGKAGWFPSAYVEK 313
E++ GD ++V+ + GW GE GK GWFP+ Y EK
Sbjct: 755 EITIQPGDIVMVKGEWVDESQTGEPGWLGGEPKGKTGWFPANYAEK 800
Score = 43.5 bits (101), Expect = 0.008
Identities = 33/143 (23%), Positives = 59/143 (41%), Gaps = 26/143 (18%)
Query: 196 RMKELKANMAVLGKEAAAALASVDAQQQRLTFQRLVAMMVSDR----------------- 238
R+ A A+ G+E A ++ LTFQ+ ++V+ +
Sbjct: 989 RVASPAAKPAIPGEEFVAMYTYESSEHGDLTFQQGHVIVVTKKDGDWWTGTVGETSGVFP 1048
Query: 239 ----QKKESAPPVVVSENGSGKTMYFLAEATHPFYGDSEKELSFSKGDFIVVRKVTPTGW 294
+ K+S + GS + +A+ + ++L+ + G I++RK P GW
Sbjct: 1049 SNYVRLKDSEGSGTAGKTGSLEKKPEIAQVIASYTATGPEQLTLAPGQLILIRKKNPGGW 1108
Query: 295 SEGECNGKA-----GWFPSAYVE 312
EGE + GWFP+ YV+
Sbjct: 1109 WEGELQARGKKRQIGWFPANYVK 1131
Score = 41.6 bits (96), Expect = 0.030
Identities = 19/51 (37%), Positives = 29/51 (56%), Gaps = 1/51 (1%)
Query: 262 AEATHPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVE 312
A+A +P+ + L+F+K D I V + W GE G+ GWFP +YV+
Sbjct: 915 AQALYPWRAKKDNHLNFNKSDVITVLEQQDMWWF-GEVQGQKGWFPKSYVK 964
Score = 39.3 bits (90), Expect = 0.15
Identities = 18/47 (38%), Positives = 27/47 (57%)
Query: 266 HPFYGDSEKELSFSKGDFIVVRKVTPTGWSEGECNGKAGWFPSAYVE 312
+ + ++ EL+FSKG I V W +GE +G+ G FPS YV+
Sbjct: 1161 YDYTAQNDDELAFSKGQIINVLSKEDPDWWKGEVSGQVGLFPSNYVK 1207
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.127 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 474,210,129
Number of Sequences: 2790947
Number of extensions: 18315619
Number of successful extensions: 67717
Number of sequences better than 10.0: 2521
Number of HSP's better than 10.0 without gapping: 1125
Number of HSP's successfully gapped in prelim test: 1420
Number of HSP's that attempted gapping in prelim test: 64389
Number of HSP's gapped (non-prelim): 4195
length of query: 327
length of database: 848,049,833
effective HSP length: 127
effective length of query: 200
effective length of database: 493,599,564
effective search space: 98719912800
effective search space used: 98719912800
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0046a.5


