

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0045.6
(214 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
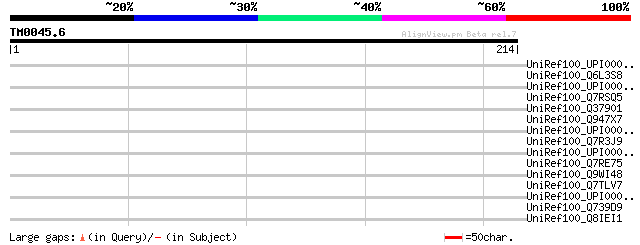
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_UPI000023510D UPI000023510D UniRef100 entry 39 0.092
UniRef100_Q6L3S8 Putative F-Box protein [Solanum demissum] 38 0.16
UniRef100_UPI0000290CF4 UPI0000290CF4 UniRef100 entry 37 0.35
UniRef100_Q7RSQ5 Putative yir4 protein [Plasmodium yoelii yoelii] 37 0.35
UniRef100_Q37901 Tail protein [Bacteriophage BF23] 35 1.0
UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sa... 35 1.3
UniRef100_UPI000034B9A2 UPI000034B9A2 UniRef100 entry 34 2.9
UniRef100_Q7R3J9 GLP_158_8532_13532 [Giardia lamblia ATCC 50803] 34 2.9
UniRef100_UPI00002EC7A5 UPI00002EC7A5 UniRef100 entry 33 5.0
UniRef100_Q7RE75 Hypothetical protein [Plasmodium yoelii yoelii] 33 5.0
UniRef100_Q9WI48 Structural protein [Hepatitis E virus] 33 6.6
UniRef100_Q7TLV7 Enhancin-like [Choristoneura fumiferana nuclear... 33 6.6
UniRef100_UPI00002F670B UPI00002F670B UniRef100 entry 32 8.6
UniRef100_Q739D9 Membrane-bound protease, putative [Bacillus cer... 32 8.6
UniRef100_Q8IEI1 Methionyl-tRNA formyltransferase, putative [Pla... 32 8.6
>UniRef100_UPI000023510D UPI000023510D UniRef100 entry
Length = 569
Score = 38.9 bits (89), Expect = 0.092
Identities = 22/79 (27%), Positives = 36/79 (44%), Gaps = 4/79 (5%)
Query: 27 HGYQNFQNMWRNIGAYEDG----SVRDMITLAKYNGGDLDTYFEHPDFEPLVEGPEALAQ 82
HG+ ++ +I Y+D ++R +I+ K N G D++ F P+ G L
Sbjct: 434 HGHGRGPEVYADITVYDDNDHLSNIRPVISCCKVNLGPNDSFLRAARFAPIFSGSHGLRI 493
Query: 83 VLGFSHVWHFAAPSNVKAF 101
+H W A P NV +F
Sbjct: 494 DYSDTHYWLIAWPVNVVSF 512
>UniRef100_Q6L3S8 Putative F-Box protein [Solanum demissum]
Length = 327
Score = 38.1 bits (87), Expect = 0.16
Identities = 34/112 (30%), Positives = 51/112 (45%), Gaps = 11/112 (9%)
Query: 99 KAFAWRFLLVK---IP-SRLNLWKLGATRGLISFGYNRSKDEFLVVLIKLCNLLHLAKNK 154
+A+ W + K +P SR NL G G FGY+ S+D++ VV I H +
Sbjct: 127 EAYIWNPTITKSKELPKSRSNLCSDGIKCG---FGYDESRDDYKVVFIDYPIHRHNHRTV 183
Query: 155 ITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQFLCDSLYWLKTKKATNVLV 206
+ I SLR + + + D F+ NL G+F+ LYW + N V
Sbjct: 184 VNIYSLR--TKSWTTLHDQLQGFFLLNLH--GRFVNGKLYWTSSSCINNYKV 231
>UniRef100_UPI0000290CF4 UPI0000290CF4 UniRef100 entry
Length = 201
Score = 37.0 bits (84), Expect = 0.35
Identities = 28/102 (27%), Positives = 47/102 (45%), Gaps = 10/102 (9%)
Query: 107 LVKIPSRLNLWKLGATRG-----LISFGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLR 161
+ K +++NL K+G T L+ F N+ E + L+H NKI I ++
Sbjct: 54 MTKESTKINLSKMGTTEKIKKIILVRFKINQENKELIKKTFFF--LMHPKNNKILIKTIY 111
Query: 162 NYVRDYFDIA---DTEYYFYKFNLDYIGQFLCDSLYWLKTKK 200
N V + IA T++ FY L ++ ++WLKT +
Sbjct: 112 NIVDQIWFIAKDESTDFNFYSKRLILSAIYIRTMIFWLKTDR 153
>UniRef100_Q7RSQ5 Putative yir4 protein [Plasmodium yoelii yoelii]
Length = 305
Score = 37.0 bits (84), Expect = 0.35
Identities = 32/121 (26%), Positives = 53/121 (43%), Gaps = 17/121 (14%)
Query: 69 DFEPLVEGPEAL-AQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLIS 127
DFE + G L Q+ G S ++ A SN+ ++L+ + LNL + G +
Sbjct: 45 DFERIDSGCLYLFKQIFGTSQLFKSVANSNINIVD--YILIWLSYMLNLKEDGKNVNNLQ 102
Query: 128 FGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQ 187
F YN + D K K +I + NY ++Y D+ D + YF+ + + I
Sbjct: 103 FFYNTTIDN--------------DKYKNSITGVENYYKNYKDLIDKKTYFFGMDRNIIFN 148
Query: 188 F 188
F
Sbjct: 149 F 149
>UniRef100_Q37901 Tail protein [Bacteriophage BF23]
Length = 595
Score = 35.4 bits (80), Expect = 1.0
Identities = 42/163 (25%), Positives = 64/163 (38%), Gaps = 43/163 (26%)
Query: 34 NMWRNIGAYEDGSVRDMITLAKY---------NGGDLDTYFEHPDFE-PLVEGPEALAQV 83
N++R AY ++ TLA + N G++D F HP+++ P+ G
Sbjct: 167 NLYRPGRAYAAKAMGRAWTLAGFPAGASKVSINSGNVD--FWHPNWQAPIGAGKRG---- 220
Query: 84 LGFSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRS--KDEFLVVL 141
H W + SN++ +A K G T +S YN S D+ V
Sbjct: 221 ----HTWFYVCNSNIRGYA--------------GKKGTTPSNVSVLYNSSTYPDKVYVCR 262
Query: 142 IKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDY 184
NL A K YV+D ++I T+ +Y NL Y
Sbjct: 263 GSTSNLASQAAKK-------QYVQDSYNITPTKVIWYVLNLRY 298
>UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sativa]
Length = 387
Score = 35.0 bits (79), Expect = 1.3
Identities = 15/33 (45%), Positives = 21/33 (63%)
Query: 86 FSHVWHFAAPSNVKAFAWRFLLVKIPSRLNLWK 118
+S +W AAP VK FAW ++P+R+NL K
Sbjct: 226 WSAIWRSAAPPRVKFFAWLMSKNRLPTRVNLHK 258
>UniRef100_UPI000034B9A2 UPI000034B9A2 UniRef100 entry
Length = 286
Score = 33.9 bits (76), Expect = 2.9
Identities = 20/48 (41%), Positives = 26/48 (53%), Gaps = 1/48 (2%)
Query: 157 IISLRNYVRDYFDIADTEYYFYKFNLDYIGQFLCDSLYWLKT-KKATN 203
IIS YV+ + +DT Y K NL G +CDS Y +T KK +N
Sbjct: 97 IISENQYVKSVVEESDTNYEQKKINLCNSGILICDSKYLFETIKKISN 144
>UniRef100_Q7R3J9 GLP_158_8532_13532 [Giardia lamblia ATCC 50803]
Length = 1666
Score = 33.9 bits (76), Expect = 2.9
Identities = 15/36 (41%), Positives = 25/36 (68%), Gaps = 1/36 (2%)
Query: 137 FLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIAD 172
F +V++KLCN L +N+++ S+ ++RDY IAD
Sbjct: 598 FFLVILKLCNF-RLVQNQVSYNSVLTFLRDYVKIAD 632
>UniRef100_UPI00002EC7A5 UPI00002EC7A5 UniRef100 entry
Length = 259
Score = 33.1 bits (74), Expect = 5.0
Identities = 32/122 (26%), Positives = 56/122 (45%), Gaps = 7/122 (5%)
Query: 54 AKYNGGDLDTYFEHPDFEPLVEGPEALAQVLGFSHVWHFAAPSNVKAFAWRFLLVKIPSR 113
++++GG L + PDFE + + + L + F + VK F + +
Sbjct: 133 SRFSGG-LVVDIQKPDFELRKKIVQKKSDELDNLYSGQFKISNEVKDFISNEITASVREL 191
Query: 114 LNLWKLGATRGLISFGYNRSKDEFLV-VLIKLCNLLHLAKNKITIISLRNYVRDYFDIAD 172
+ GA ++SF K LV + L +LL+LA+NK+TI ++ V +F I+
Sbjct: 192 V-----GAVNRIVSFSRIYKKMPTLVETKVVLKDLLNLAENKVTIDLIQTLVCKFFKISK 246
Query: 173 TE 174
E
Sbjct: 247 NE 248
>UniRef100_Q7RE75 Hypothetical protein [Plasmodium yoelii yoelii]
Length = 677
Score = 33.1 bits (74), Expect = 5.0
Identities = 18/55 (32%), Positives = 32/55 (57%), Gaps = 3/55 (5%)
Query: 160 LRNYVRDYFDIADTEYYFYKFNLDYIGQFLCD---SLYWLKTKKATNVLVVVRKL 211
+RN V +A+ YY+ KF +I + + + +L++ TK +NV++VVR L
Sbjct: 175 MRNVVDILISLANVNYYYDKFINFFIYEIIKNEKNNLFFFNTKNMSNVILVVRSL 229
>UniRef100_Q9WI48 Structural protein [Hepatitis E virus]
Length = 436
Score = 32.7 bits (73), Expect = 6.6
Identities = 16/66 (24%), Positives = 36/66 (54%), Gaps = 1/66 (1%)
Query: 76 GPEALAQVLGFSHVWHFAAP-SNVKAFAWRFLLVKIPSRLNLWKLGATRGLISFGYNRSK 134
G +A+A+ L ++ V P S ++ ++ F ++ + +L+ W+ G T+ + YN +
Sbjct: 282 GAQAVARSLDWTKVTLDGRPLSTIQQYSKTFFVLPLRGKLSFWEAGTTKAWYPYNYNTTA 341
Query: 135 DEFLVV 140
+ L+V
Sbjct: 342 SDQLLV 347
>UniRef100_Q7TLV7 Enhancin-like [Choristoneura fumiferana nuclear polyhedrosis virus]
Length = 758
Score = 32.7 bits (73), Expect = 6.6
Identities = 21/66 (31%), Positives = 32/66 (47%), Gaps = 13/66 (19%)
Query: 25 IAHGYQNFQNMWRNIGAYEDGSVRDMITLAKYNGGDLDTYFEH-------PDFEPLVEGP 77
++H +F N+W +D +TL+ ++ D D YFEH P FE L+ G
Sbjct: 45 VSHNDVSF-NLWN-----DDSETEQNLTLSAHDMSDPDDYFEHVVEHDSVPFFECLLNGN 98
Query: 78 EALAQV 83
E AQ+
Sbjct: 99 EYTAQI 104
>UniRef100_UPI00002F670B UPI00002F670B UniRef100 entry
Length = 245
Score = 32.3 bits (72), Expect = 8.6
Identities = 26/78 (33%), Positives = 41/78 (52%), Gaps = 7/78 (8%)
Query: 125 LISFGYNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDY---FDIADTEYYFYKFN 181
LI +S F L+KL +L +L K++I+I++ ++R F IADTE ++ K N
Sbjct: 151 LIGNSDQKSIYTFKEALLKLIDLSNLKKDEISIVTEDKFIRPTNVPFLIADTEKFYKKTN 210
Query: 182 LD---YIGQFLCDSL-YW 195
+ L D+L YW
Sbjct: 211 WSPKINFKEILSDTLNYW 228
>UniRef100_Q739D9 Membrane-bound protease, putative [Bacillus cereus]
Length = 742
Score = 32.3 bits (72), Expect = 8.6
Identities = 21/69 (30%), Positives = 36/69 (51%), Gaps = 5/69 (7%)
Query: 150 LAKNKIT----IISLRNYVRDY-FDIADTEYYFYKFNLDYIGQFLCDSLYWLKTKKATNV 204
L K+K T ++++ NY D+ F T F N DY+ QFL D+ +T++
Sbjct: 428 LTKDKDTRYDKVLAIENYFTDHSFVYETTNVSFPTKNQDYVDQFLFDTKSGYCNNFSTSM 487
Query: 205 LVVVRKLGL 213
+V++R G+
Sbjct: 488 IVLLRSAGI 496
>UniRef100_Q8IEI1 Methionyl-tRNA formyltransferase, putative [Plasmodium falciparum]
Length = 665
Score = 32.3 bits (72), Expect = 8.6
Identities = 21/73 (28%), Positives = 29/73 (38%), Gaps = 7/73 (9%)
Query: 130 YNRSKDEFLVVLIKLCNLLHLAKNKITIISLRNYVRDYFDIADTEYYFYKFNLDYIGQFL 189
Y D+ ++ I L N L + K K NY+R Y YY YK NL +
Sbjct: 131 YKNQLDDINILYILLFNTLMIYKKKYDFFMNENYIRSY-------YYIYKNNLGRDKKIY 183
Query: 190 CDSLYWLKTKKAT 202
Y++ T T
Sbjct: 184 NTKNYFINTFSIT 196
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.144 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 373,221,366
Number of Sequences: 2790947
Number of extensions: 15596264
Number of successful extensions: 40690
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 40682
Number of HSP's gapped (non-prelim): 18
length of query: 214
length of database: 848,049,833
effective HSP length: 122
effective length of query: 92
effective length of database: 507,554,299
effective search space: 46694995508
effective search space used: 46694995508
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0045.6


